Abstract
Animal models for cystic fibrosis (CF) have enhanced our understanding of the pathology and contributed to the development of new treatments. In the field of CF, many animal models have been developed and described. To our knowledge, thus far, none of the reviews of CF animal models has used a systematic methodology. A systematic approach to creating model overviews can lead to an objective, evidence-based choice of an animal model for new research questions. We searched Pubmed and Embase for the currently available animal models for CF. Two independent reviewers screened the results. We included all primary studies describing an animal model for CF. After duplicate removal, 12,304 publications were left. Because of the large number of models, in the current paper, only the genetic models are presented. A total of 636 publications were identified describing genetic animal models for CF in mice, pigs, ferrets, rats and zebrafish. Most of these models have an altered Cftr gene. An overview of basic model characteristics and outcome measures for these genetic models is provided, together with advice on using these data. As far as the authors are aware, this is one of the largest systematic mapping reviews on genetic animal models for CF. It can aid in selecting a suitable model and outcome measures. In general, the reporting quality of the included publications was poor. Further systematic reviews are warranted to determine the quality and translational value of these models further.
Introduction
Cystic fibrosis (CF) can be caused by several mutations in the CF transmembrane conductance regulator (Cftr) gene. This gene was identified in 1989, 1 after the mapping of its location to chromosome 7q31 by restriction fragment length polymorphism analysis in the early 1980s. Development of the first CF mouse was reported three years later in 1992. 2 Currently, a multitude of animal models are available to the preclinical researcher studying CF. When choosing the most appropriate model to answer a specific research question, all known models should be considered, and the selection should be based on scientific evidence on model validity.
Animal models for CF are thought to have enhanced our understanding of the pathology of this debilitating disease.3,4 In addition, these models have aided the development of new treatments. In theory, genetic models have the highest construct validity possible (i.e. the highest degree of resemblance in aetiology and pathophysiology between experimental animals and humans 5 ). In practice, among other issues, genetic background can substantially alter the model pathology. 6
A complete and structured overview of the literature on the available models would be very helpful for scientists choosing an animal model. More and more types of reviews (i.e. papers summarising the results of several primary studies) are being published. 7 Most review types do not clearly describe their methodology, and a reader cannot easily estimate the completeness of and the possible bias in their findings. Systematised review types are in this respect preferred over narrative reviews 8 because they transparently describe the review methodology and limitations, and therefore make it easier to assess potential bias in the conclusions. Several systematised review types exist, which are suitable for different purposes: full systematic reviews (SRs), mapping reviews, scoping reviews and rapid reviews. Mapping reviews comprise a systematic search of a broad field that presents results in a user-friendly format. 9 James et al. 10 have outlined the differences between SRs and systematic maps.
Broadly speaking, the objective of a mapping review ranges wider than answering a specific research question as in an SR. In addition, data extraction is limited, and critical appraisal is optional. Hence, mapping reviews usually do not synthesise data from the included studies. Otherwise, mapping reviews mainly follow the same methods as SRs. In the environmental sciences, 11 health care and social sciences, 12 mapping reviews are increasingly encouraged. As far as we know, this is one of the first mapping reviews of animal studies. It describes animal models for CF.
While several informative reviews on animal models have been published, none of these has used systematic approaches as in an SR. Narrative reviews of these models focus on, for example, genetic mouse models 4 or on specific disease aspects. 3 While informative, narrative reviews are at risk of bias based on subjective preferences or awareness of the included literature. We thus performed a systematic mapping review describing all published animal models for CF. The central question in this review is: ‘What are the currently available animal models for CF (to perform, e.g., a proof-of-principle/preclinical efficacy study for a new compound)?’. Due to the extensive volume of published studies retrieved by our searches (n = 12,304 references) and the number of included publications (n = 799), we had to limit the intended coverage of our review. We list models published up to our search date, and provide data on species, sex, mutations, survival and outcomes measured.
This systematic map is one of the largest preclinical literature summaries to date. To improve legibility, we have split the presentation of our results into two papers. We describe genetic and non-genetic models separately, grouping the included models by type of model induction (and thereby by construct validity). The current paper focuses on genetic models.
This mapping review should aid preclinical scientists in making an evidence-based choice for the most appropriate model for their studies. Moreover, it can be used to identify the relevant background literature for the techniques to measure relevant outcome parameters. We provide guidance for using our systematic map. With this comprehensive overview of the available genetic animal models for CF, scientists can carefully weigh the usefulness and translatability of the considered animal models, taking animal welfare and alternatives into consideration.
Materials and methods
The original protocol for this review was developed in collaboration with an information specialist (focussing on the search strings), a clinician (providing feedback on a draft version of the protocol and search strategy) and an anonymous patient participant (providing feedback on a draft version of the protocol, search strategy and, at a later stage, this publication). It was posted on the SYRCLE website (www.SYRCLE.nl) on 23 December 2015.
Search strings for CF.

CF: cystic fibrosis.
We included primary studies describing a mutation or intervention inducing CF-like symptoms in live animals, with the authors' intention to study CF. All references were independently screened by two reviewers using Early Review Organising Software (EROS; Institute of Clinical Effectiveness and Health Policy, Buenos Aires, Argentina). Discrepancies were resolved by discussion between the reviewers. During the screening of the titles and abstracts, papers that were not about CF and those that did not describe an animal model for CF were excluded. During screening of the full texts, reviews not containing new data and other non-primary studies were furthermore excluded. Studies with and without a control group for the CF model were included.
The following data were extracted from the full-text papers by one reviewer: language, animal (species), strain/stock, sex, type of model induction and mutation, percentage survival, outcome measures and, if available, changes of the model animal compared to the control animals (either wild types or heterozygous mice). We extracted data for each group of model animals separately. Correct data extraction was verified by a second reviewer for a sample of 10% of the publications included in this paper, which were used as convenience samples for projects on lung disease, cellular immunity in inflammation, development of symptoms over time and failure to thrive.
Protocol deviations
Our search retrieved more references than anticipated. During the early phase of data extraction, we therefore decided that our original plans were not feasible. We thus made two adaptations to the protocol to provide this map of the literature. First, we did not screen the reference lists of relevant reviews to find further publications. Second, we restricted our data extraction to language, species, strain/stock, sex, type of model induction (genetic for the papers included in this publication), mutations, percentage survival, outcome measures and (when a control group was provided) changes in these outcome measures of the model animals compared to control animals.
Data analysis
Mutations: abbreviations used in our output with the synonyms used in the literature.
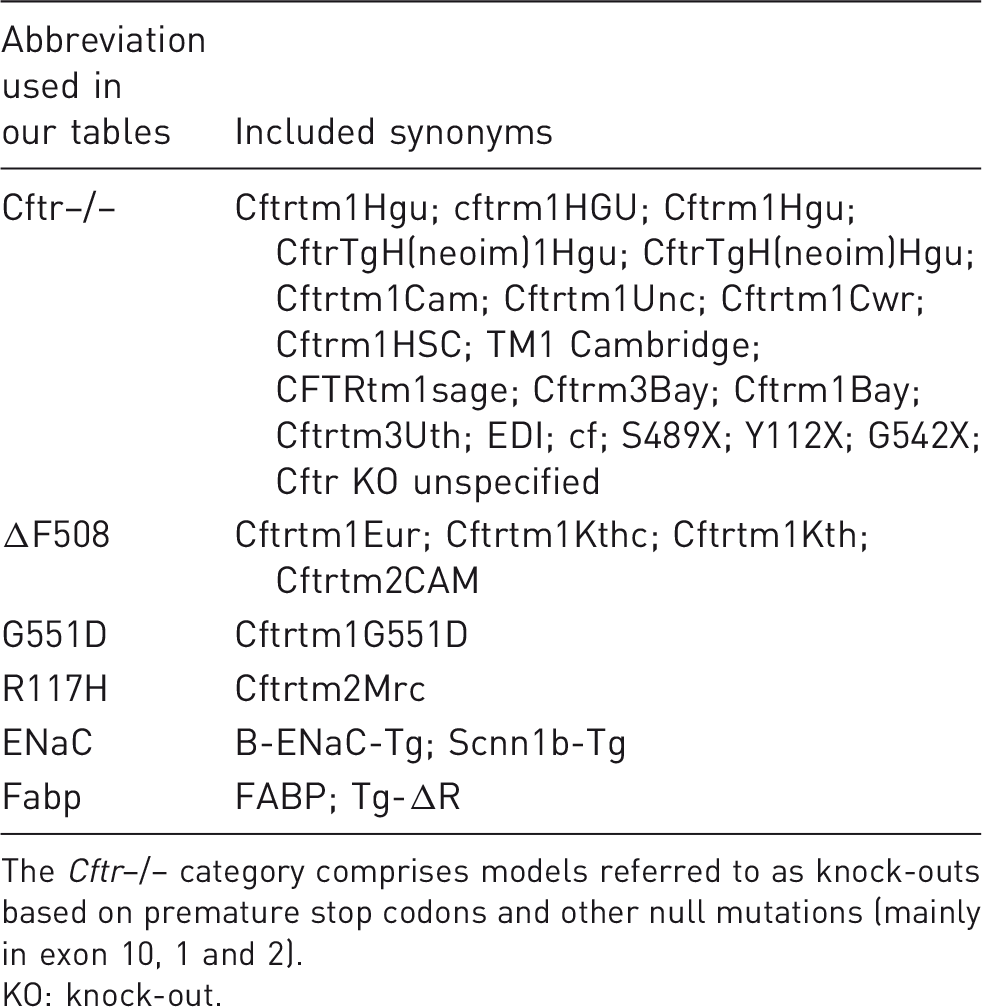
The Cftr–/– category comprises models referred to as knock-outs based on premature stop codons and other null mutations (mainly in exon 10, 1 and 2).
KO: knock-out.
In this paper, we only present the genetic animal models, sorted by species and mutation. To provide an indication of the relevance of the animal model for CF and the aspects of the human disease that are modelled, we tabulated the outcome measures from all studies, with the direction of any potential difference between model animals and controls. We compared the outcome measures reported for the animal models to those used in clinical trials.
The Core Outcome Measures in Effectiveness Trials Initiative (COMET; http://comet-initiative.org/; accessed 4 August 2018) encourages the use of agreed standardised sets of outcomes to improve the comparison, contrast and combination of results of trials. We searched the COMET database to check for preclinical versus clinical alignment.
Before we decided to restrict our data extraction, additional study design characteristics were extracted for a non-random sample of 166 papers (the same articles on which we based our quality control) included in this review. We calculated the percentages of reporting based on this sample to provide a crude indication of study quality.
Results
Search and selection
From Pubmed, 7976 references were retrieved, and from Embase, 9403 were retrieved. After duplicate removal, 12,304 references were imported into EROS for screening of the title and abstract. We excluded 9700 references during screening of the titles and abstracts, and another 1805 during screening of the full texts. Of the remaining 799 references, 636 described genetic animal models and are presented in this paper. The non-genetic models will be described in a following paper. The flow of the retrieved references is provided in Figure 1.
Study selection.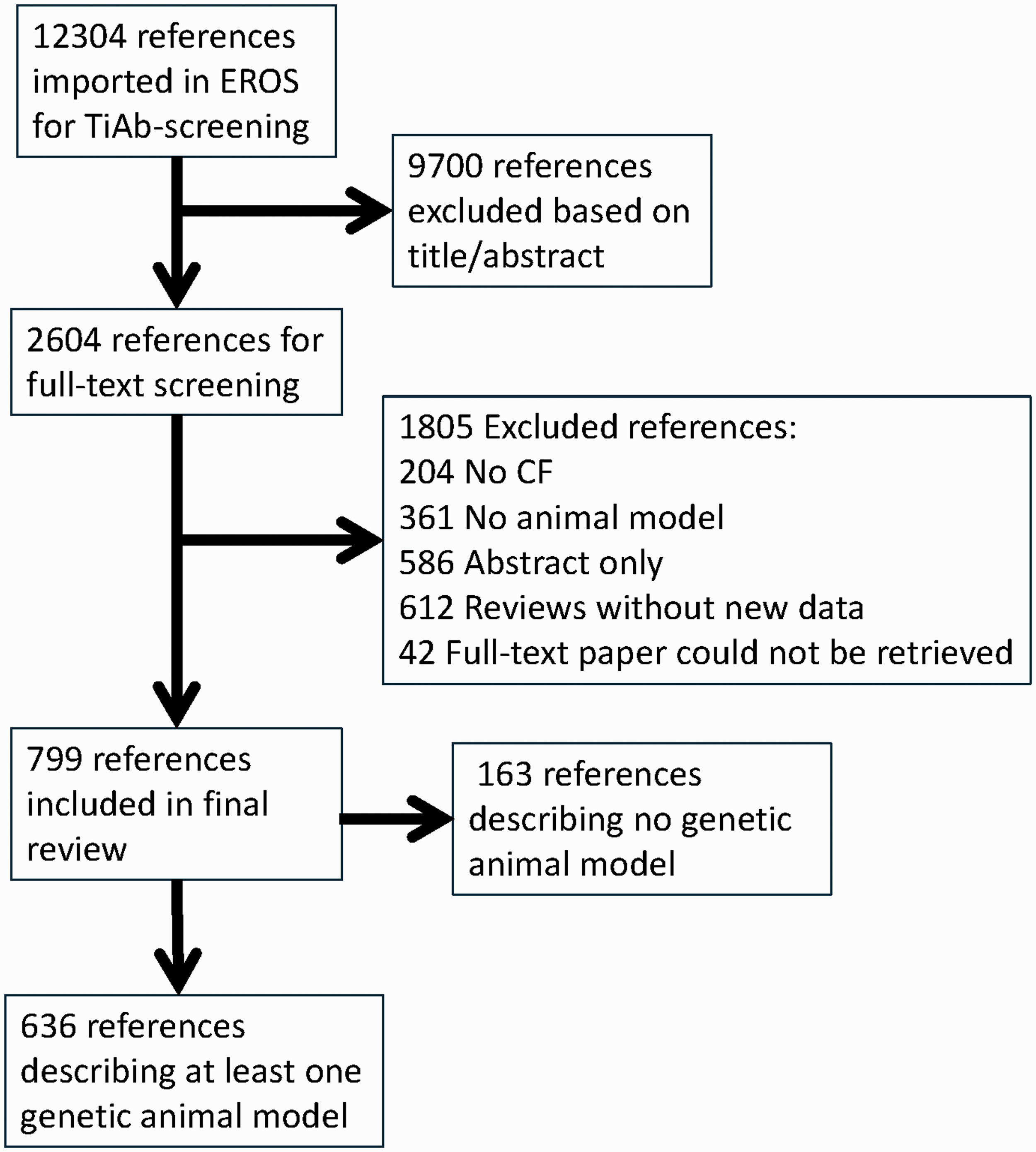
The main results on the genetic models are described in Appendices 1–5 and below. Study identifiers of the 636 references included in this part of the review are provided in Appendix 6 with the corresponding titles. The full references of all included papers and relevant reviews are provided in Appendix 7.
Study characteristics
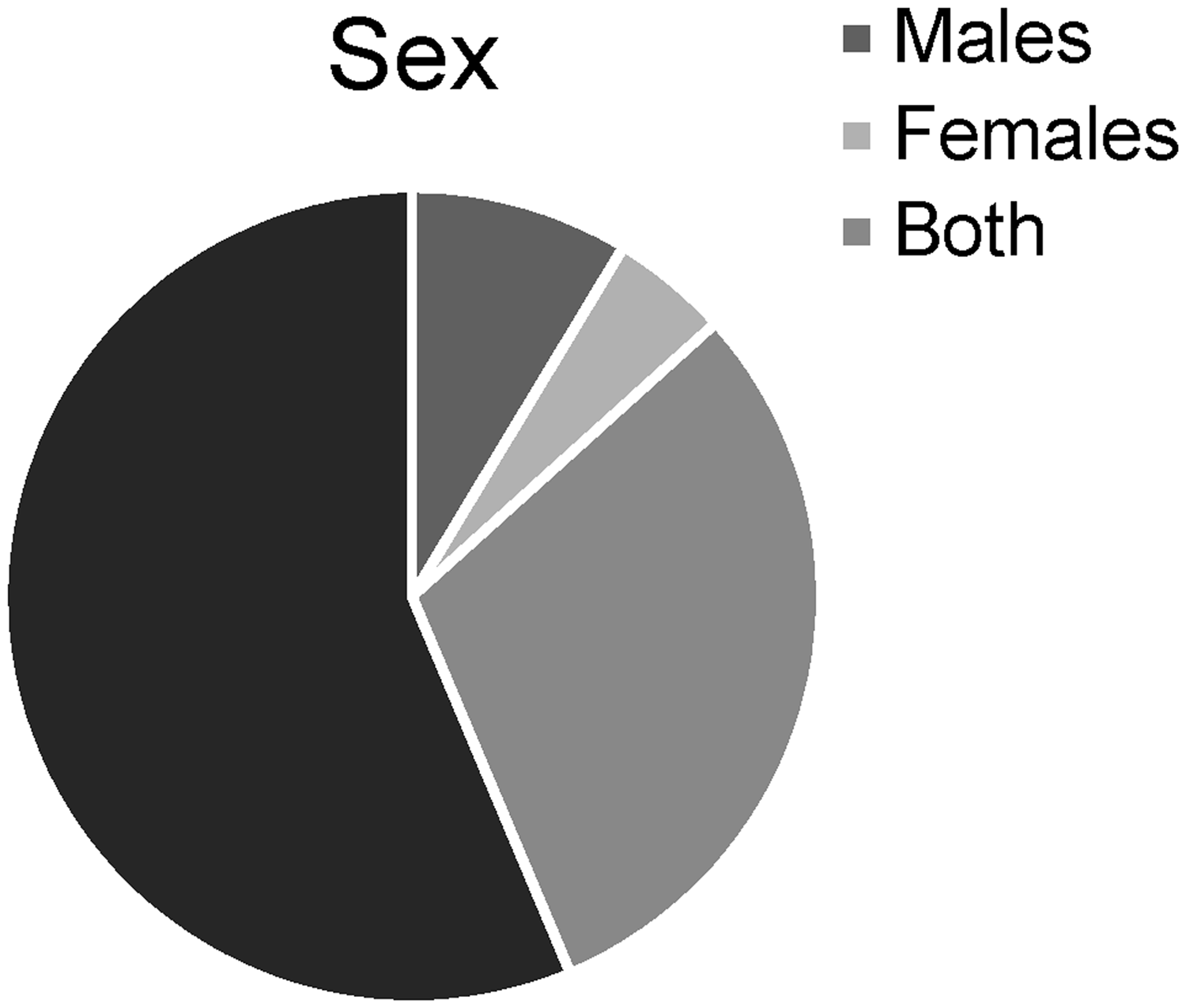
The included 636 publications (Appendix 6) described 751 experimental model groups (excluding groups based on genetic background, which we could not reliably analyse). Of these 751 model groups, 691 comprised mice, 46 pigs, 8 ferrets, 3 rats and 3 zebrafish (Figure 2). The sex of the animals was reported for 328 model groups; 65 were restricted to male animals, 35 to female animals and 228 used both sexes (Figure 3). The percentage of the model animals surviving to a certain time point or experimental stage was provided for 131 of the model groups and ranged from 0% to 100%.
Species used in the 751 included experimental groups. Sex used in the 751 included experimental groups.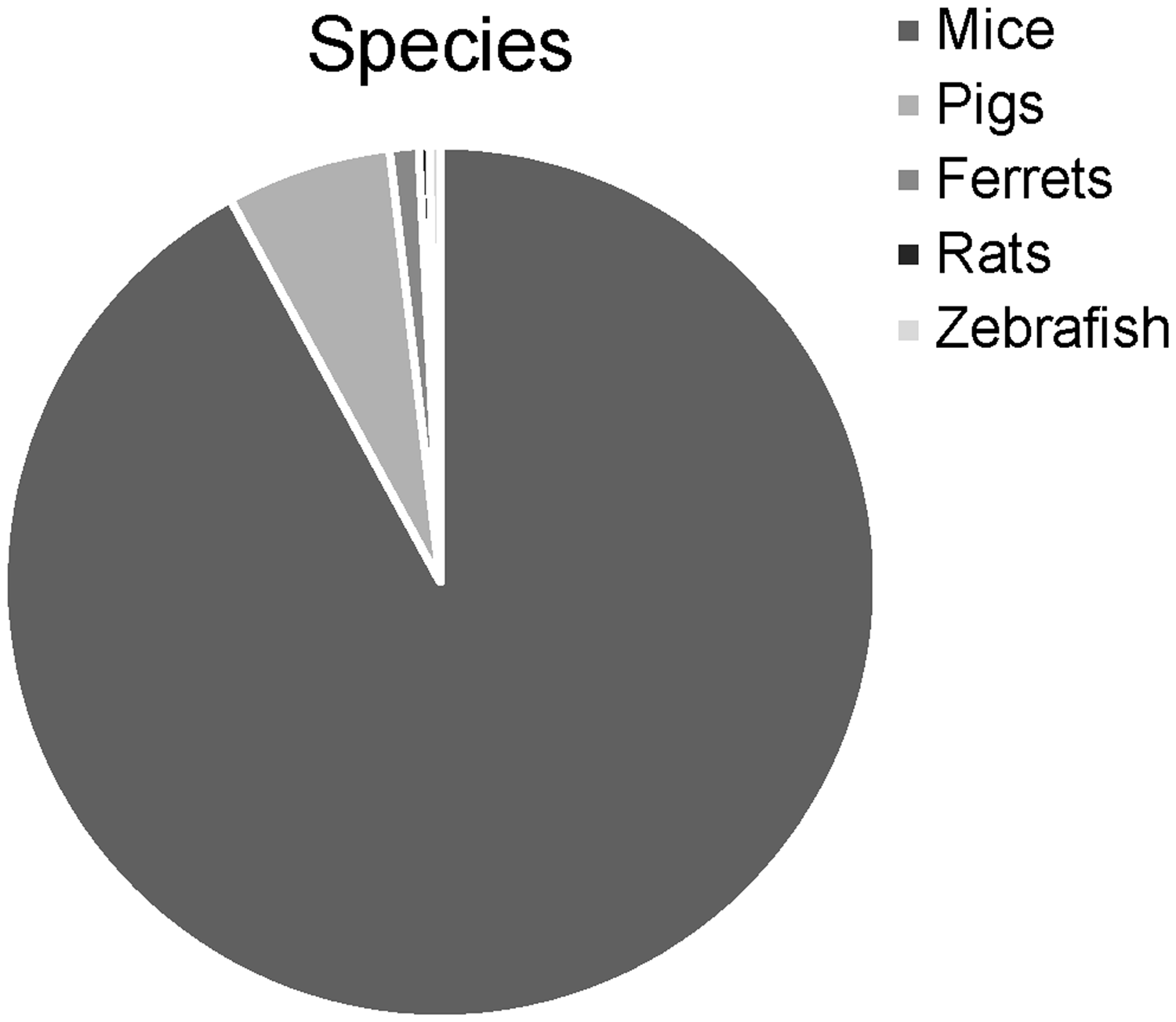
Reporting quality in subsample
Reporting of experimental detail in the subset of 166 papers with additional data extraction was highest for the number of animals used and their age: 85% of papers provided information on group size, and 70% described the age of the animals. The supplier (or in-house breeding) was provided in 55% of the papers. Only 21% described the weight of the animals. Intestinal obstruction is common in mouse models for CF, and can be limited by adapted diets, administering laxatives or using special bedding. We thus analysed reporting of diet (53% reported), the use of laxatives (19%) and special bedding (12%).
While general husbandry conditions may not have major effects on the outcomes of genetic models, they should be reported according to the ARRIVE guidelines. In our subset, they were poorly reported: laboratory temperature in 13%, relative humidity in 4% and the timing of the light/dark cycle in 3% of the papers. Experimental design was also poorly reported: only one paper reported performing a power analysis, 11% described randomisation of part of the protocol and 15% described implementing blinding for at least part of the experiment. The number of animals taken out of the experiment before the final measurements was described in 6% of the papers.
Genetic models of CF
In clinical settings, several mutations in the Cftr gene are known to cause CF. Most genetic animal models for CF have mutations in the Cftr gene, comparable to human CF patients. The clinical variability is matched by a variety in the available animal models. For this review, we harmonised the different mutations on different backgrounds as described in Table 1. The models based on a single mutation in the Cftr gene in ferrets, mice, pigs, rats and zebrafish are provided in Appendix 1, together with the outcome measures assessed.
The outcome measures assessed cover a wide range of phenomena. Many biochemical parameters have been assessed (e.g. cholesterol, triglycerides, glucose, insulin, pancreatic enzymes, cytokines and antioxidants), and many organs have been histologically or morphologically investigated (e.g. the lungs, biliary tract and pancreas). Furthermore, a vast proportion of the outcome measures covers electrophysiological parameters, reflecting the mechanistic change in function of the CFTR channel in CF. The main tissues for the electrophysiological measurements are nasal epithelium, trachea and lung, intestines and pancreas. Inflammatory cells and cytokines are often quantified in bronchioalveolar lavage fluid. Moreover, mucus from various locations has been investigated in detail.
Our search of the COMET database showed that no core outcome measures for CF have been defined yet, but different initiatives, including consensus meetings and SRs, are in progress to develop them. According to the literature, currently available and accepted clinical end points comprise pulmonary function as assessed by lung spirometry (e.g. forced expiratory volume), quality of life, pulmonary exacerbations, microbiology, inflammatory markers, nasopotential difference and body mass–based measures.15,16 Both these outcome classes have their parallels in the listed animal models.
The life expectancy of CF patients has increased from 10 to 40 years over the last three decades. 16 Hence, new outcome measures are sought after. Those being considered are, for example, the washout of inert gasses during breathing, imaging (x-ray, computed tomography, magnetic resonance imaging and positron emission tomography), mucociliary clearance, other potential differences and/or currents, sweat chloride content, intestinal transit time and intestinal pH (via wireless capsules), malabsorption, and biochemical markers mainly in the sputum and blood.15,17 Except for the quality of life, which is measured with questionnaires, all these considered outcome classes have their parallels in the listed animal models.
Many papers described two or more models. The most observed combination is of a complete loss-of-function mutation (Cftr–/–) and the ΔF508 mutation. In addition, many authors describe the use of gut-corrected mouse models for CF for both ΔF508 and Cftr–/– mutations. In these gut-corrected models, the functional Cftr protein is present in the gastrointestinal tract, alleviating intestinal blockage common to the non-corrected models. This gut correction is induced by the FABP-Cftr transgene, with the rat fatty acid binding protein 2, intestinal promoter (Fabp2) driving the expression of Cftr. The gut-corrected models are listed in Appendix 2.
Mutations in several other genes have resulted in phenotypes that resemble part of CF pathology. Genetic mouse models for CF other than those in the Cftr are most commonly those with alterations in the β-ENaC gene and the IL-10 gene. Mutations in other genes have also been used to model CF. Papers describing the CF-like phenotype-inducing mutations in genes other than Cftr are listed in the table in Appendix 3.
The oldest genetic animal model for CF is not the first targeted Cftr mutant. In 1974, Kaiser et al. described the Cri/cri mouse. We are not aware of publications on this model since a publication from 1991, 18 which was excluded from our review because the full-text paper was not available.
Seven papers describe eight models with double mutations, which are provided in Appendix 4. Two of these combine mutations in Cftr with ENaC overexpression, four combine mutations in Cftr with another gene, two combine ENaC overexpression with mutations in a gene other than Cftr. One paper described a conditional mutation, and two papers described the use of antisense RNA, all targeting Cftr. The outcomes are provided in Appendix 5.
How to use the tables
There are several options for using the tables. For finding models in which a specific outcome measure has been reported, a text search of Appendices 1–5 would be the optimal strategy. For retrieving references on specific types of models, the relevant table(s) can be screened line by line. Appendices 1–5 are sorted by species and mutation. For all types of questions, a text search for words in the titles can be performed in Appendix 6 or 7.
When searching for text words, we highly recommend searching for all known synonyms of a term of interest and searching, for example, for the first letters of the term to retrieve both the singular and the plural use and different spellings. For all searches, we advise a search in at least one literature database such as PubMed, EmBase or Web of Science besides searching our tables in order to check for more recent publications on the topic. Only references after 23 December 2015 need to be screened. The searches provided in Table 1 for CF and the previously published filters19,20 for animal studies can be used.
Discussion
In this systematic mapping review, we provide an overview of the published genetic animal models for CF up to December 2015. Most genetic animal models for CF have altered Cftr, either with an artificial premature stop codon or with a mutation resembling one observed in a large proportion of CF patients. In the Appendices, we present the outcome measures measured in these models, with a direction of the effects when compared to a wild-type or heterozygous control group. To the best of our knowledge, this mapping review is the largest and most systematic inventory to date on animal models for CF (and one of the largest within the field of laboratory animal sciences). It is a convenient tool for any CF researcher who wants to know what outcomes have been measured in specific genetic CF animal models.
Our comprehensive search for this review retrieved more than 12,000 references. Several narrative reviews with a smaller scope have, for example, described the disease phenotype in genetic mouse models.4,21,22 In addition, mouse phenotypes have been compared to those in larger animal models.23–25 Some reviews focus on specific aspects of this multifaceted disease; for example, Olivier et al. describe gastrointestinal, pancreatic and hepatobiliary phenotypes in CF animal models. 3
For the identified models, we provide an overview of the basic characteristics and outcome measures with the direction of a potential difference with control animals. These data show what has been measured in animal models of CF up to December 2015. More recent and future primary studies may measure other outcomes. No clear knowledge gaps were observed; most aspects of human CF pathology have been addressed. The tables provided with this review can be used as a basis to find the relevant literature to design future experiments.
Our tables list the outcome measures used in the primary studies with the direction of the effect in model compared to healthy animals. Outcome measures selected for clinical trials should ideally be accurate, precise, reliable, non-invasive, easy to measure and inexpensive. 16 Furthermore, they should be informative for distinguishing patients from controls and effective from non-effective treatments. For preclinical studies, no equivalent to the COMET initiative exists but for the currently accepted and considered outcome measures in humans, Equivalent measures in animal have been tested. To improve animal-to-human translation, the preclinically selected end points should relate to clinical outcome measures. A full comparison of the outcome measures in their ability to distinguish model from control animals and effective from non-effective treatments is beyond the scope of this mapping review.
Our search was complicated by the fact that ‘cystic fibrosis’ is often abbreviated to CF, and that excluding the abbreviation from our searches would have resulted in missing relevant papers (which we observed in scoping searches). Unfortunately, the abbreviation ‘CF’ is used for many unrelated phenomena (e.g. conferre, chow-fed, chocolate-fed, cell-free, the CF-1 rat strain, coronary flow, colonisation factor, cerebrospinal fluid, complement fixation, cytosolic fraction, chloroform, corn fibre, centrifugal fluid, capacity of filtration, chicken fat, catecholamine fluorescence, etc.), resulting in false hits. We attempted to limit the number of irrelevant hits by combining the abbreviation ‘CF’ with search terms for CF-relevant organ systems such as the lungs, pancreas, sinuses and so on. During protocol development, all search terms were individually tested. Our review protocol and search were developed in collaboration with an information specialist, clinician and anonymous patient. All of them have contributed unique terms to our search.
A limitation of our review results from the search date of 28 December 2015; the number of references retrieved was so substantial that we needed more than two years for analysis. We thus do not list the more recently developed CF animal models. Since then, there have been important developments in genetic animal models, for example the development of a CF model in sheep using CRISPR/Cas9. 26 We therefore urge those interested to combine the use of this review with a search for the literature published since December 2015, which will be aided by our search strings.
Performing an update of our search was not feasible. SRs are notoriously strenuous. 27 Mainly, the screening, data extraction and data analyses become more time-consuming with larger reviews. The field of CF animal models is not the easiest topic for a full SR, as there are many models, mutations and organs affected. We included 799 of the 12,304 originally screened papers in our overall systematic map. This size of review is only manageable by teamwork; paper selection was performed by seven different scientists, and data extraction by nine. The EROS platform enabled easy discrepancy resolution in the paper-selection phases.
Our overview of the models up to December 2015 may not be fully complete, as we did not screen reference lists of included studies and relevant reviews as planned (because of the huge number of publications already retrieved). One of our previous SRs shows that reviewing reference lists only adds few references to a SR. 28 The number of studies retrieved by snowballing depends on the research field, the proper use of title and abstract terminology and keywords by the authors of primary studies, and the number of studies published in non-indexed journals. We expect that most available models were retrieved by our extensive searches.
Data extraction was performed by nine different scientists, which regularly resulted in different wording. Data entries from these nine scientists were collated and harmonised to the best of our abilities. For future reviews of this size, with several scientists collaborating on data extraction, we highly recommend ensuring a consistent format of the data upfront, for example by using drop-down lists and restricted entry formats. Besides, we advise against extraction of complex data with multiple synonyms such as background strain and exact mutation for large mapping reviews.
Many authors of studies on genetic animal models for CF did not report the specific mutation and background used. We therefore decided to pool different mutations, as described in Table 1. Due to this pooling, the Cftr–/– category can comprise all models referred to as knock-outs, which are based on premature stop codons and other null mutations, mainly in exons 10, 1 and 2. We kept the mutations that are specifically clinically relevant, for example the ΔF508, G551D and G480C mutations, separate in our tables wherever possible. Our pooling of mutations does, however, result in a very heterogeneous group of Cftr–/– models. Phenotypic differences between the genotypes are substantial,4,22 and variation can be increased even further by the genetic background. 6 Of note, most models with mutations corresponding to the human disease use the ΔF508 mutation. This is no surprise, as the prevalence of this mutation is highest among CF patients. However, there currently is an unmet need of treatment for patients with other mutations, who have expressed feeling neglected in medical research.
In general, systematic maps do not comprise a risk-of-bias analysis. Tools are available to analyse the risk of bias specifically in animal studies.13,29 These tools are developed for analysing studies comparing a treatment intervention to a control group and can easily be adapted for studies comparing model animals to corresponding controls. Alternatively, we could have evaluated the quality of the included publications by comparing them with the ARRIVE guidelines for reporting animal studies. 30 We decided against a full risk of bias and reporting analysis because of time restraints. We did analyse the reporting of several study quality indicators in a non-random sample of the included papers.
In this sample, reporting of experimental details was poor. While study quality may exceed reporting quality, we are concerned about the quality of preclinical research. While randomisation of genotype is not possible, blinding of outcome assessment often is. Only 15% of our sample of studies reports any blinding in the experimental design. Our main concern is that we observed many publications in which mutant mice were compared with non-littermate control groups, which results in confounding of the effect of the mutation by other differences in baseline characteristics. Furthermore, the strains were often poorly described, and it was often unclear if the mutant mice had been backcrossed at least 10 times to ensure a stable genetic background.
We envision researchers searching our tables for their models and outcomes of interest to find the relevant research, which should save them substantial time compared to searching PubMed and Embase thoroughly, while providing a complete overview of the available literature up to the end of 2015. We highly encourage other preclinical researchers to perform a more detailed SR of a subset of these models by extracting and analysing additional data, in order to analyse the quality and translational value of these CF models in more depth. Using this mapping review as a starting point could save a substantial amount of time.
Supplemental Material
Supplemental material for Animal models for cystic fibrosis: A systematic search and mapping review of the literature – Part 1: genetic models
Supplemental Material for Animal models for cystic fibrosis: A systematic search and mapping review of the literature – Part 1: genetic models by Cathalijn HC Leenaars, Rob BM De Vries, Anna Heming, Damian Visser, David Holthaus, Joey Reijmer, Janneke Elzinga, Rosalie WM Kempkes, Carine Punt, Wouter Beumer, Franck LB Meijboom and Merel Ritskes-Hoitinga in Laboratory Animals
Footnotes
Acknowledgements
The authors would like to acknowledge Alice Tillema (information specialist), Peter Mercus (clinician) and an anonymous patient for reviewing our protocol, and the anonymous patient for reviewing a draft of this publication.
Declaration of Conflicting Interests
The author(s) declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: Two of the authors (CP and WB) work for ProQR Therapeutics, a company developing an investigational product (eluforsen) for treating CF. This does in our opinion not result in competing interests for a systematic review of animal models not describing therapeutic interventions.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by NWO (grant number 313-99-310), R2N, Federal State of Lower Saxony and the DFG (grant number FOR2591, BL 953/11-1).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
