Abstract
Examining zebrafish populations for the presence of disease is an integral component of managing fish health in research facilities. Currently, many different strategies are used for zebrafish fish health inspections, which is a scenario that may result in subjective and biased diagnostic evaluations. The goal of this study was to compare the success of pathogen detection between a sample size of randomly selected fish (n = 60) that provides 95% confidence in pathogen detection based on a presumed pathogen prevalence level ≥5%, and other subpopulations and sample numbers commonly submitted for diagnostic testing within a 1000 tank, 30,000 fish, recirculating research system. This included fish collected from a sump tank (n = 53), sentinel fish (n = 11), and fish that were found moribund or freshly dead (n = 18). Additionally, five fish from each subpopulation were collected for histopathologic examination. A second study used retrospective data to examine pathogen distribution between systems (n = 2−5) in multi-system facilities (n = 5) using a sample size of 60 fish per system. For the pathogens detected, results supported the use of representative sample numbers rather than smaller numbers of populations considered more at risk. The exception to this is for the moribund/mortality group, which may be a resource for targeted surveillance of select pathogens. Each system within multi-system facilities should be considered separate units in terms of fish health inspections and biosecurity. Development of these evidence-based standards for fish health inspections in zebrafish systems enhances fish welfare, provides identification of potentially zoonotic pathogens, and ensures scientific integrity and reproducibility of research results.
Introduction
The relatively rapid growth of fish as model organisms in the laboratory exemplifies their value in medical and biological research. Most notable are zebrafish (Danio rerio), although medaka (Oryzias latipes), 1 killifish and mummichogs (Fundulus spp.), 2 goldfish (Carassius auratus), 3 and sticklebacks (Gasterosteus spp.), 4 among others, are commonly utilized. This emergence has led to major scientific discoveries but has also challenged our ability to keep pace and develop proper standards for husbandry of fish in research facilities. Fish health management is important not only from an animal welfare standpoint, but also for ensuring scientific integrity and reproducibility of research results. Inconsistent or improper husbandry of these research models can affect physiologic and immunologic responses that may in turn impart undefined variables into experiments. Thus, adequate time and resources should be put into standardizing and maintaining health within fish populations.
One essential component of managing fish health is monitoring populations for the presence of infections and non-infectious disease. Fish within research systems are at risk for disease, as aquatic conditions favor the growth of disease-causing organisms due to the natural biofilms that harbor pathogens, relatively high fish densities that promote transmission of disease, and elevated fish stress secondary to handling and manipulation. 5 In research fish, there is a growing list of infectious and non-infectious diseases described in laboratory settings, particularly for zebrafish. This includes various Gram-negative bacteria likely representing both primary and opportunistic infections, including Edwardsiella ictaluri, which has been associated with septicemia in previously healthy fish, 6 and Aeromonas spp., which have been linked to skin lesions and subsequent septicemia, likely secondary to water quality issues and increased fish stress. 7 At least six species of mycobacteria have been isolated from zebrafish with varying degrees of chronic morbidity and mortality. 8 Parasitic diseases diagnosed in research facilities include enteritis associated with the nematode Pseudocapillaria tomentosa, 9 skin and gill infections by the dinoflagellate Oodinium pillularis, 10 and internal infections caused by the mxyozoan Myxidium streisingeri 11 and the microsporidians Pseudoloma neurophilia and Pleistophora hyphessobryconis. 12 Non-infectious diseases observed in zebrafish include neoplastic lesions 13 and egg-associated inflammation often in older zebrafish. 14
To date, there are very few guidelines for monitoring pathogens and diseases in zebrafish facilities. Current methods for sampling and testing fish often vary between facilities and lack evidence-based strategies. There is wide variation in the number of fish sampled and the population of fish submitted for testing. Fish number is often arbitrarily determined or centered on disease testing costs. Although practical considerations will always be a part of these decisions, the effect that these biases may have on testing results must also be noted. Based on the Office International des Epizooties (OIE) Aquatic Animal Health Surveillance 15 guidelines and American Fisheries Society 16 guidelines, fish health inspections should use a sample size that provides 95% confidence in pathogen detection using approved diagnostic tests. For practicality, an assumed pathogen prevalence (APPL) of ≤5% is often used. By those standards, to test a population of 2000 fish or more, 60 fish would need to be submitted for testing to provide 95% confidence in pathogen detection at a 5% APPL or 120 fish for a 2% APPL. This does not negate a value in sampling fewer fish. However, negative results need to be interpreted cautiously because of the lower confidence level in pathogen detection. These standards were largely developed for food fish and wild fisheries, and to date, application of these sample sizes have not been examined in zebrafish facilities, which represents a different dynamic in terms of fish husbandry, genetics, and diseases.
Zebrafish sampled for disease testing often come from different subpopulations within the system. This includes randomly sampled fish from the general population, which can provide a wide sample distribution, depending on the number of fish selected. Additionally, facilities may choose to utilize “sentinel” fish housed within isolated tanks or the sump tank that are exposed to effluent water. 17 These programs are designed to evaluate the efficacy of ultraviolet sterilization or as a strategy to monitor fish health within the system. The assumption is that increased exposure to organisms shed from fish in the general population makes sentinel fish more likely to harbor pathogens. More rarely, facilities may utilize moribund or freshly dead fish for risk-based surveillance. There is evidence to suggest that this distribution increases the odds of identifying select pathogens within commercial aquaculture systems. 18 However, this methodology and the effect of subpopulation on success of pathogen prevalence have not been comprehensively explored in zebrafish facilities.
Fish health inspections outlined in this study utilized a combination of targeted and broad-based diagnostic techniques. Targeted diagnostic techniques provided sensitive and pathogen-specific inspection for previously described zebrafish pathogens, such as Mycobacteria spp. and P. neurophilia, using polymerase chain reaction (PCR) assays. Broad-based methods used strategies to ensure breadth in diagnostic scope with the awareness that the list of known etiologies of zebrafish is still growing. This included bacterial culture, virus isolation, histopathology, and wet-mount cytology.
The objective of this study was to compare diagnostic results between commonly used sampling strategies. A sample size of 60 fish was used as the standard that would allow 95% confidence in pathogen detection based on a presumed pathogen prevalence of 5%. Gross signs of disease and pathogen identification were compared to a relatively small number of sentinel fish, to a larger group of fish collected from the sump tank, and from fish observed as moribund or freshly dead. Additionally, diagnostic results were retrospectively analyzed within facilities that contained multiple independent recirculating systems to examine the boundaries of biosecurity and to guide testing protocols within these potentially more complex establishments. Our goal was to help define the advantages and limitations of targeted and broad-based diagnostic methods currently employed in zebrafish facilities. Examining these facets will help define best practices that provide confidence in testing results without unnecessary economic expenses and fish sacrifice.
Methods
Ethical statement
The use of these animals in these trials was approved by the Institutional Animal Care and Use Committee at Boston Children’s Hospital (IACUC Protocol# 17-08-3511R).
Effect of subpopulation and fish number on success of pathogen detection
Fish health inspection was performed on a recirculating system consisting of 1000 tanks and approximately 30,000 adult zebrafish of mixed genetic lines. A complete description of the husbandry and environmental conditions in housing for the fish used in this study is available as a collection at dx.doi.org/10.17504/protocols.io.mrjc54n. All equipment that comes into contact with research animals or water are cleaned and disinfected prior to and after each use. Disinfection is accomplished by rinsing at a temperature of ≥82.2℃ for a minimum of three minutes, and when objects have been exposed to animals or water continuously for periods in excess of 24 hours, they are disinfected by using a surfactant-free disinfecting solution followed by ≥82.2℃ deionized water or fresh water rinse to remove chemical residues.
Tanks are cleaned/disinfected on average of once every 16 weeks or as needed, and floors are cleaned and disinfected on a weekly basis using Virkon S following manufacturers’ protocols.
Rotifer and Artemia culturing vessels are disinfected with 50% Wescodyne solution with a contact time of five minutes or more between each use.
Fish within the system were divided into four experimental subpopulations. (a) The randomly sampled group consisted of 65 fish sampled by technicians by hand-netting one fish out of 65 different tanks. Based on OIE and AFS Standards, 60 fish sampled from a system of 30,000 fish represents a sample number that provides 95% confidence in pathogen detection, presuming ≥5% pathogen prevalence.15,16 The additional five fish sampled from this group were submitted for histopathology. (b) The sentinel fish group consisted of 16 fish sampled from a single sentinel tank continuously exposed to effluent water for approximately six months. (c) The sump tank group consisted of 58 fish collected from the system sump tank. These fish represented various ages and effluent water exposure time. (d) The morbidity/mortalities group consisted of seven fish found freshly dead with minimal autolysis and 16 fish interpreted as moribund due to either reddened skin, emaciation, anorexia, lethargy, or an inability to maintain constant buoyancy in the water column.
Targeted and broad-based diagnostic methods
Following collection, fish were euthanized with an anesthetic overdose of tricaine methanesulfonate (MS-222; Sigma–Aldrich, St. Louis, MO) at 200 mg/L. 19 Five fish from each group were sampled for histopathology. Fish were fixed whole in 10% neutral-buffered formalin for 72 hours and serially cross-sectioned. Tissues were processed routinely, and 5 µm sections were stained with hematoxylin and eosin and acid-fast stain. Digital slides were produced with a Leica SC2 scanner (FishVet Group, Portland, ME), and tissues were examined using ImageScope software (Leica Biosystems, Richmond, IL) by a U.S. residency and graduate program trained aquatic veterinary pathologist.
Diagnostic assays used to perform pathogen surveillance and screening on sampled fish.
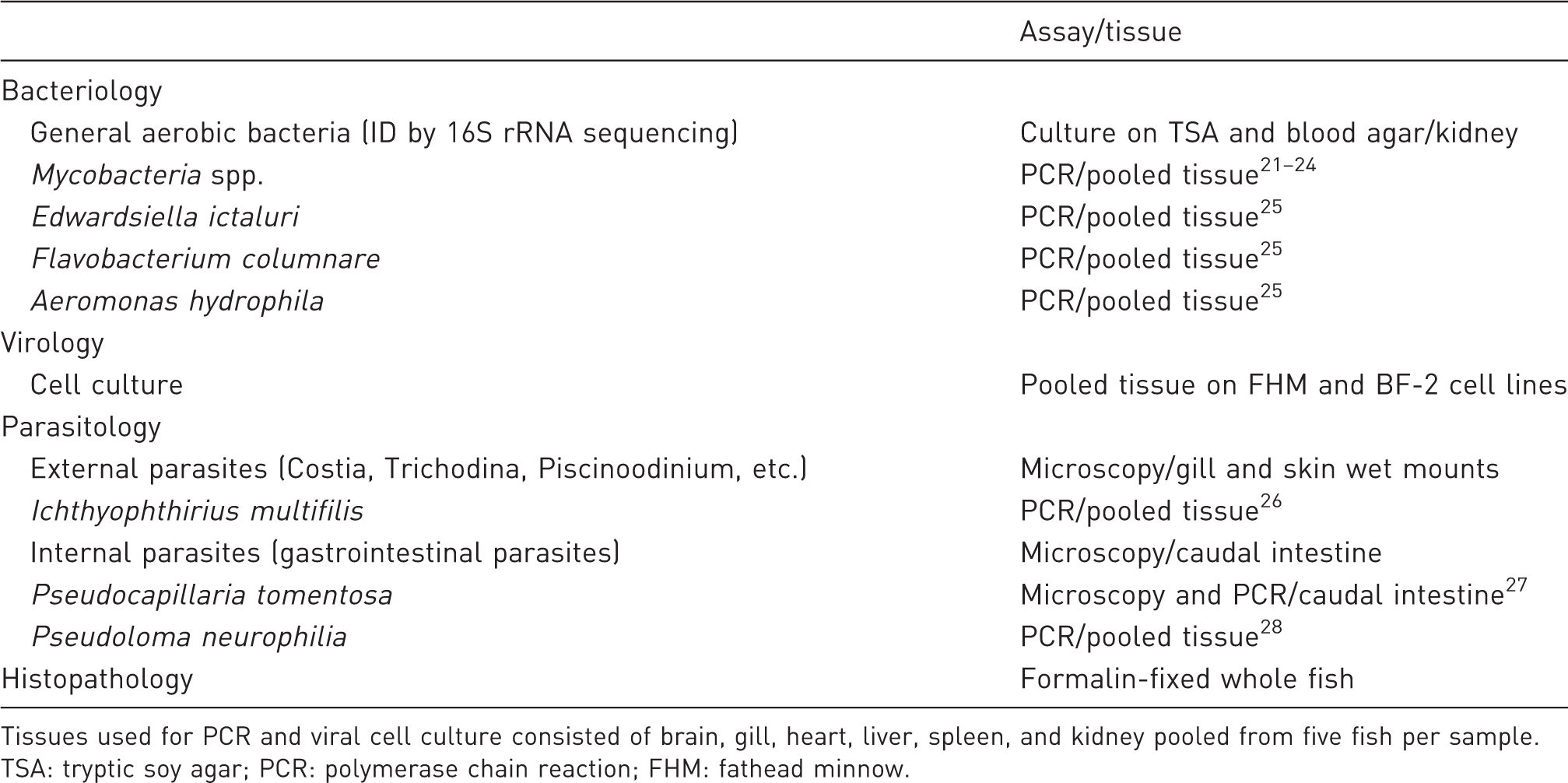

Tissues used for PCR and viral cell culture consisted of brain, gill, heart, liver, spleen, and kidney pooled from five fish per sample.
TSA: tryptic soy agar; PCR: polymerase chain reaction; FHM: fathead minnow.
Virology and pathogen-specific PCR was performed on homogenates of the brain, gill, heart, liver, spleen, and kidney pooled from five fish per sample. Homogenate (100 mg) was used for virus isolation on epithelioma papulosum cyprini and fathead minnow cell lines incubated at 30℃ for 28 days with a blind pass at 14 days. Tissue homogenate (25 mg) was used for PCR for targeted disease surveillance of select pathogens of zebrafish outlined in Table 1 following published protocols. Each assay was verified for adequate sensitivity and run with appropriate positive and negative controls. All positive PCR results for P. tomentosa were confirmed by microscopy.
Effect of system on diagnostic results in multi-system facilities
A retrospective analysis was performed on data collected from fish health inspections in five facilities containing two or more systems with 1000–25,000 fish per system (Table 5). Sixty-five fish were randomly sampled from each system, and diagnostic testing was performed as described above.
Statistical analysis
A univariate odds ratio was used to compare the likelihood of detecting each observed disease sign and pathogen within the experimental subpopulations compared to the randomly sampled group. The effect of sample size on success of pathogen detection was calculated using epitools (http://epitools.ausvet.com.au/). The pathogen prevalence needed to provide a 95% confidence interval based on sample size of each group was determined, as previously described by Murray et al., 29 with an assumed assay sensitivity of 90–95%.
For the retrospective analysis exploring pathogen detection between systems in multi-system facilities, a proportions test (prop.test function in R; The R Foundation, Vienna, Austria) was used to explore pathogen distribution and determine whether the proportions were equal. Additionally, a simple linear regression analysis (lm() function in R) was used to determine the relationship between number of fish per system and total number of pathogens detected in a system to asses if there is an increased risk of pathogen presence in systems with more fish. All statistics were run with a significance level of p < 0.05.
Results
Association of subpopulation and disease signs
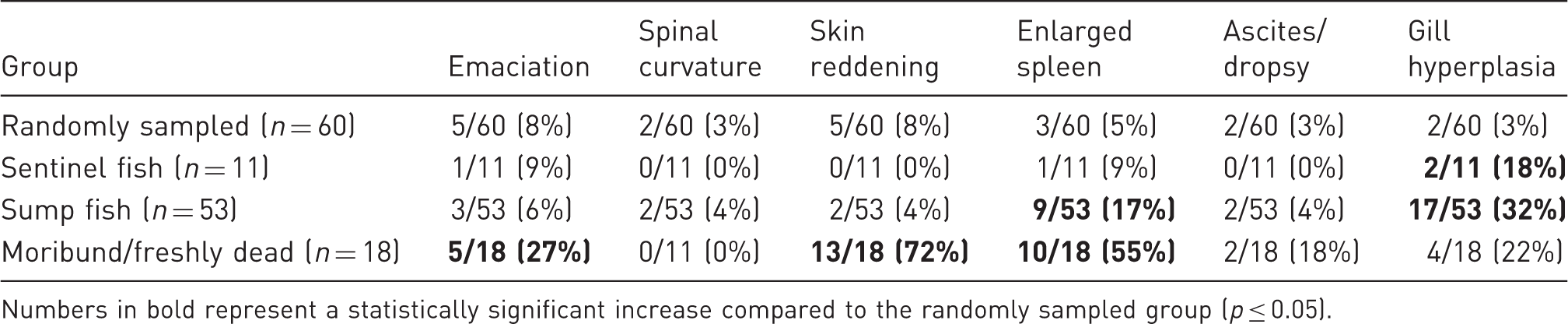
Gross signs of disease observed during necropsy included emaciation, spinal curvature, skin reddening, splenomegaly, ascites/dropsy, and gill epithelial hyperplasia (Table 2 and Figure 1(a) and (b)). The incidence of disease signs was significantly different among the various subpopulations. Emaciation and skin reddening were 4.2 times and 28.6 times, respectively, more likely to be seen in the morbidity/mortality group compared to the randomly sampled group. Splenomegaly was 23.8 times more likely to occur in the morbidity/mortality group and 3.9 times more likely to occur in the sump fish group compared to the randomly sampled group. Gill hyperplasia was 14.9 times more likely to occur in the sump group and 10.9 times more likely to occur in the sentinel group than the randomly sampled group. Spinal curvature, ascites, and dropsy were not significantly associated with any one experimental group.
Gross and microscopic findings in zebrafish. (a) Zebrafish demonstrating generalized dropsy and reddening of the skin. (b) Splenomegally (arrow). (c) Multifocal, variably sized granulomas (arrowheads) throughout the kidney. 10× magnification, bar = 200 µm. (d) Intraspinal pseudocyst filled with microsporidia consistent with Pseudoloma neurophilia. 40× magnification, bar = 50 µm. Incidence of disease signs observed on gross necropsy for each zebrafish subpopulation. Numbers in bold represent a statistically significant increase compared to the randomly sampled group (p ≤ 0.05).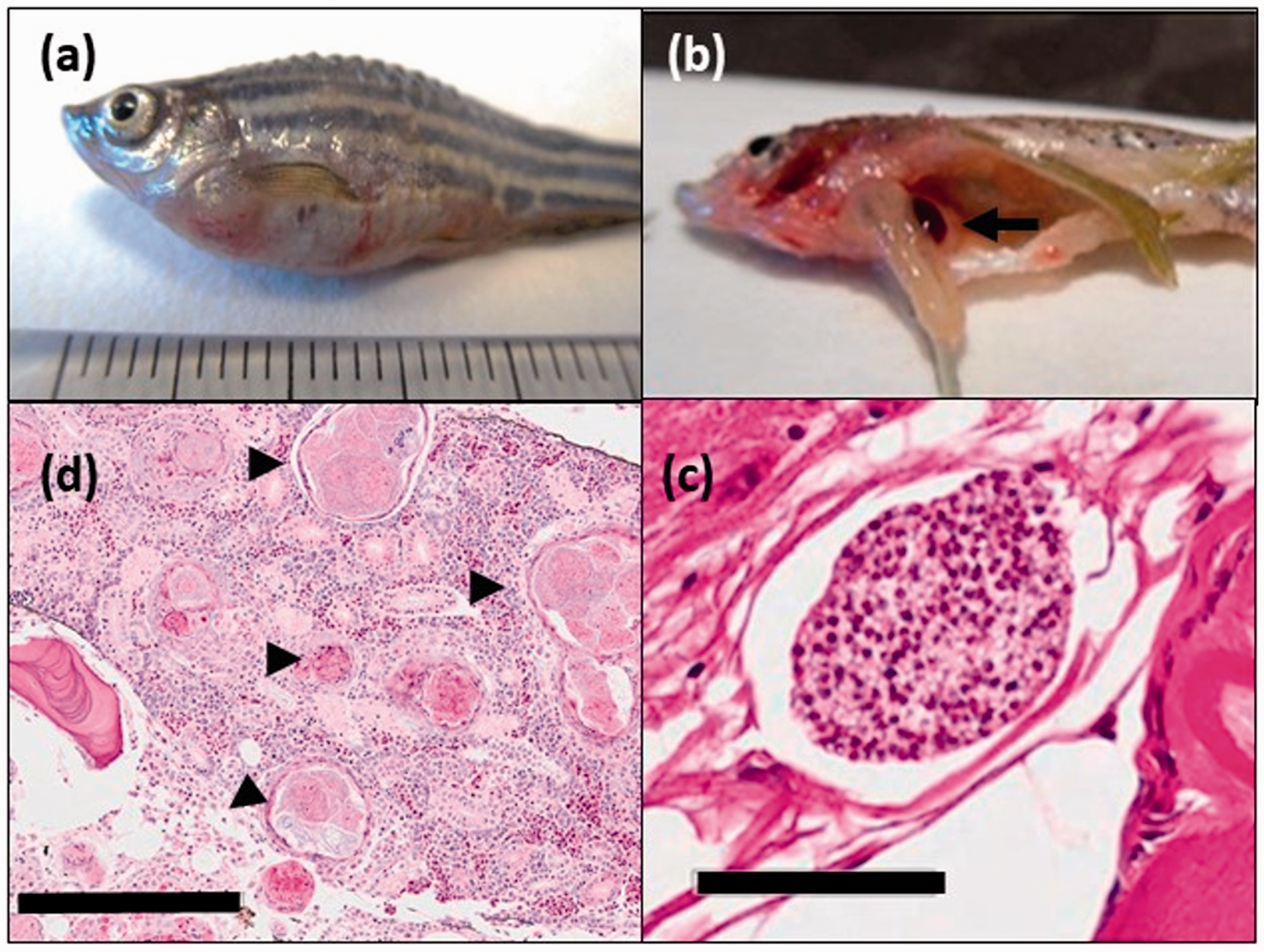
Association between success of pathogen detection, subpopulation, and sample size
Organisms detected in each subpopulation of zebrafish within the same system.
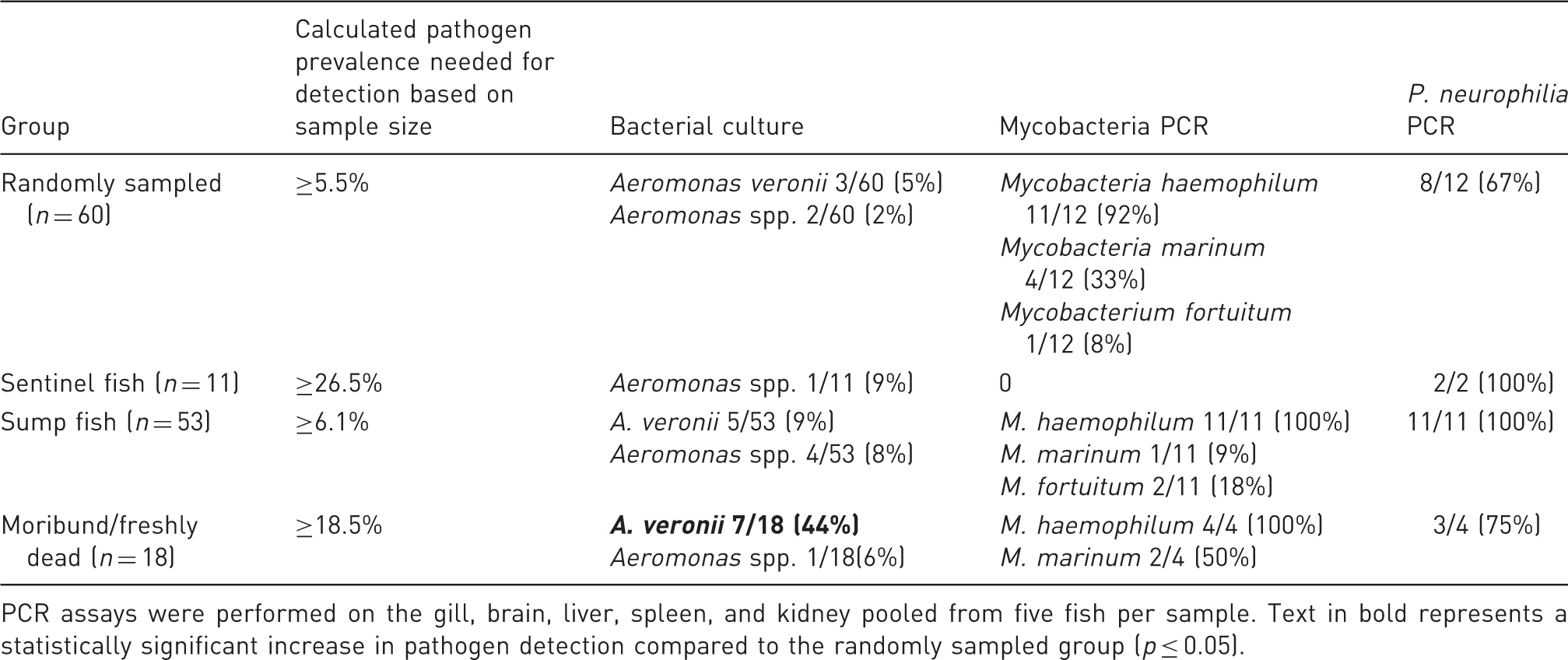

PCR assays were performed on the gill, brain, liver, spleen, and kidney pooled from five fish per sample. Text in bold represents a statistically significant increase in pathogen detection compared to the randomly sampled group (p ≤ 0.05).
Organisms detected within the randomly sampled subpopulation (n = 60) broken into sequential increments of 10 fish to examine the effect of sample number on detection success.
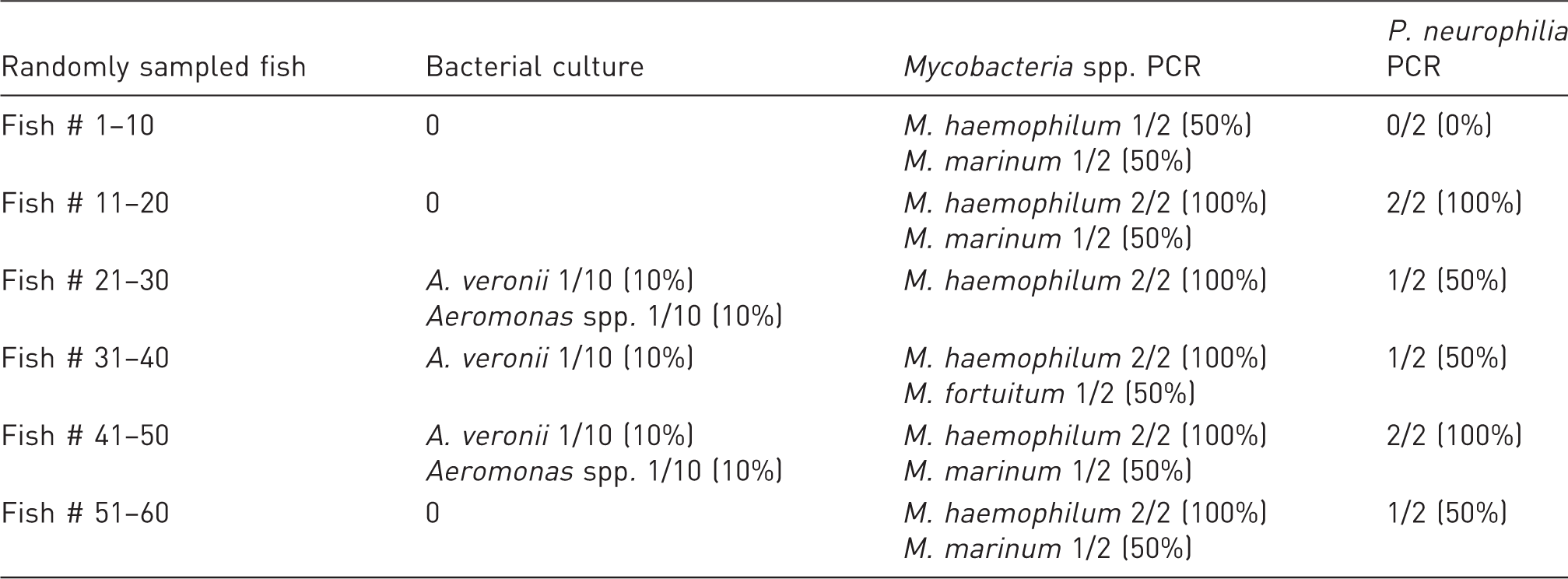

Within the subpopulations with a relatively small sample size, sampling 11 zebrafish within the sentinel group should provide detection of pathogens that had ≥26.5% prevalence. In this group, only Aeromomas spp. and P. neurophilia were detected. There was lack of detection of the three Mycobacterium spp. or for Aeromomas veronii. Within the moribund/mortality group, a sample size of 18 fish should provide detection for pathogens with ≥18.5% pathogen prevalence. Both Aeromonad isolates, M. haemophilum, M. marinum, and P. neurophilia were identified, while M. fortuitum was not found. Relative prevalence of A. veronii was higher in the morbidity/mortality group (5/13 moribund fish and 2/5 dead fish; total of 7/18 fish, 44%), and there was a 12.1 times greater risk of infection for A. veronii in the morbidity/mortality group compared to the randomly sampled group.
Histopathology
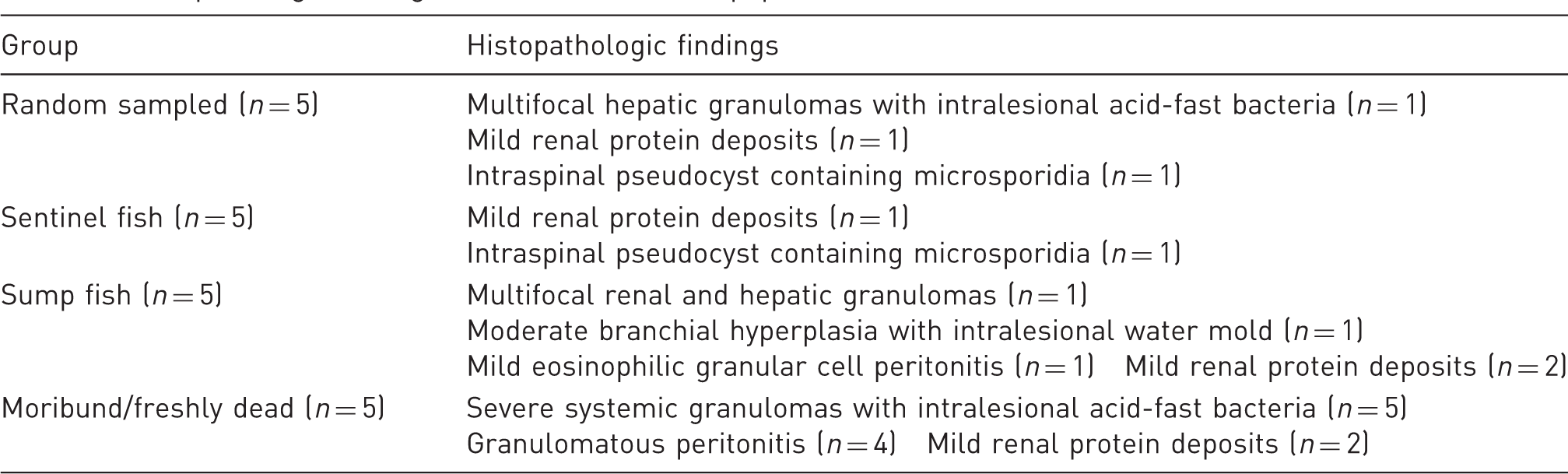
Histopathologic findings outlined in Table 5 included multi-organ granulomas found in 7/20 (28%) fish (Figure 1(c)). These were found in at least one fish from each group except for the sentinel group. Granulomas were associated with intracellular acid-fast organisms consistent with mycobacteria in 3/7 (43%) fish. Also present on histopathology was a granulomatous (n = 1) and eosinophilic granular cell peritonitis (n = 1) not overtly associated with any observed pathogens and branchial hyperplasia (n = 1) associated with water mold. Intraspinal pseudocysts containing microsporidia consistent with P. neurophilia were present in one fish in each of the randomly sampled group and the sentinel group (Figure 1(d)). There were no observations of microspores in the sump fish group or the moribund/mortality group. Mild, multifocal, renal protein deposits were present in 6/20 (30%) fish and were represented in all groups.
Effect of system and fish number on pathogen prevalence in multi-system facilities
Histopathologic findings in each zebrafish subpopulation.

Organisms isolated from zebrafish sampled from multiple independent recirculating systems within the same facility.
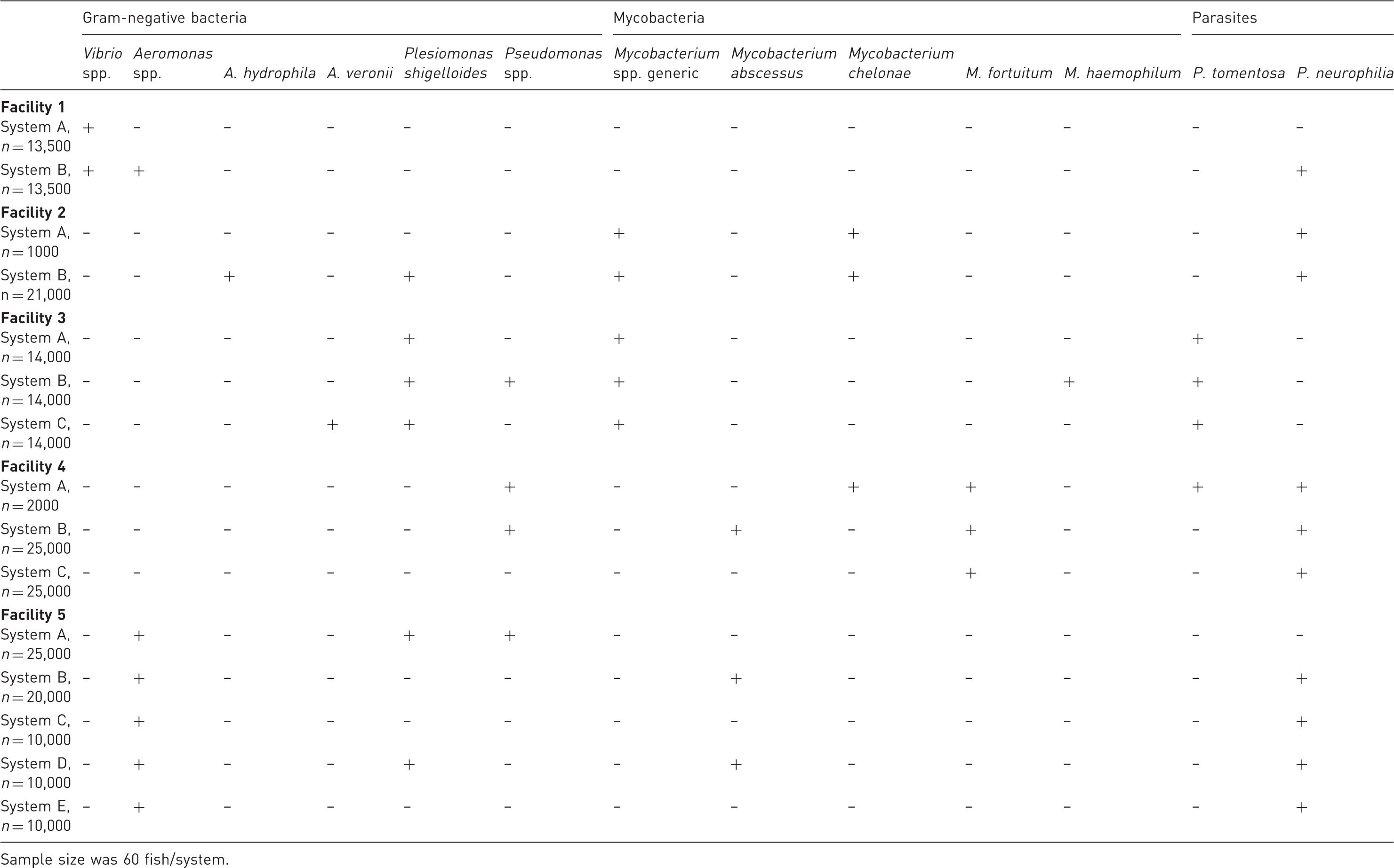

Sample size was 60 fish/system.
Discussion
The goal of this study was to evaluate how the success of pathogen detection varies between diagnostic sampling strategies within zebrafish research systems. We used a combination of targeted and broad-based diagnostic methodologies on different subpopulations of fish using sample numbers that are commonly submitted for diagnostic testing. Results demonstrated that within this system and for the specific pathogens isolated, there are significant differences in disease signs and success of pathogen detection, depending on the sampling strategy. Results suggest that for most pathogens, the number of fish sampled may be more important to diagnostic success than the subpopulation. The exception to this was for an Aeromonas spp., where increased risk of infection in the moribund/mortality group suggests more targeted surveillance for the Aeromonas spp. can be achieved by sampling fish that are moribund or freshly dead. This may be a result of the role of Aeromonads in morbidity and mortality as either an opportunistic or primary pathogen. 30
The relatively higher detection rate of M. haemophilum and, to a lesser extent, M. marinum was consistent with previous studies that describe high prevalence levels for these species within zebrafish systems. 31 M. haemophilum and M. marinum have relatively high pathogenicity in zebrafish compared to other mycobacterium species,8,32 which may also result in increased transmission and persistence within the zebrafish population. In theory, this would result in a higher detection rate, as demonstrated in this study. It is unknown why positive identification of mycobacteria was lacking in fish in the sentinel group. Based on the high prevalence rates of M. haemophilum and M. marinum in other groups, it would have been assumed that prevalence was higher than the 26.5% threshold that statistically limited the sample size of 11 fish.
Compared to M. haemophilum and M. marinum, M. fortuitum is believed to be relatively less virulent and may act as more of an opportunistic pathogen. 31 The lower pathogenicity of M. fortuitum may result in lower prevalence within the system, thus making detection more challenging compared to more pathogenic mycobacterial species. This was demonstrated by a lack of positive identification of M. fortuitum within the sentinel and moribund/mortality groups or within the first 30 fish tested in the randomly sampled group. This highlights the importance of sample size in adequately characterizing less prevalent pathogens in zebrafish systems.
Granulomatous inflammation and acid-fast bacteria consistent with mycobacterial infection were seen on histopathology in each subpopulation of fish except fish in the sentinel group, which is consistent with PCR results. Histopathology provided evidence of manifested mycobacteriosis and lesion development, but alone, it did not provide a diagnosis of which mycobacterial species were present in the fish. Molecular testing for mycobacteria provided speciation of pathogens present but only inference of disease risks and impact on the health of the fish. However, together, molecular testing and histopathology were valuable in fully characterizing the scope and impact of infection within the system. We suggest that histopathology be used as a tool to confirm and characterize disease due to infectious agents and as a means for screening for non-infectious disease, but not as a method for targeted pathogen surveillance. The relative low sensitivity of histology and relatively higher costs compared to being able to pool five fish per sample for PCR renders it a less optimal strategy.
The chronic pathogenicity 33 and widespread prevalence 29 of P. neurophilia provided a largely uniform diagnosis across all subpopulations using PCR and five fish/sample pools. Positive identification of the microsporidium was lower using histopathology, with negative results for the sump group and morbidity/mortality group. This was not unexpected, as cross-sectioning the fish limits the coverage of the spinal cord, although in the authors’ opinion, it is the preferred method for ensuring adequate visualization of other major organs.
There were six distinct Gram-negative bacteria isolated from kidney swabs within the subpopulation study and the retrospective analysis of data from multi-system facilities. Bacterial culture cannot differentiate between colonization and true tissue invasion, and postmortem interval may have affected bacteriology results for dead fish within the moribund/mortality group. With the exception of highly virulent E. ictaluri, 6 the prevalence and significance of Gram-negative bacteria in zebrafish systems are largely unknown. Based on limited reports in zebrafish and what has been described in fish in more traditional aquaculture settings, A. hydrophila, A. veronii, P. shigelloides, and Pseudmonas spp. are believed to reflect the microflora of the aquatic habitat but with roles as primary and opportunistic pathogens.7,34–38 A statistically increased detection rate of Aeromonas spp. in the moribund/mortality group, including isolation from 38% of tested moribund fish, suggests its role as a pathogen. Adequate interpretation of these findings requires further research to provide genomic comparisons to previously described fish pathogens and to fulfill Koch’s postulates through challenge studies to describe pathogenicity. Isolates deemed to pose substantial risks to zebrafish health could then be prioritized for development of more sensitive methodologies for targeted surveillance.
Fish health inspections may also be useful for identifying potentially zoonotic pathogens present in the system. Mycobacteria and Gram-negative bacteria isolated in this study, specifically, M. marinum, M. fortuitum and M. chelonae, as well as Vibrio, Aeromonad, Pseudomonad, and Plesiomonas species, have been linked to infections in humans through exposure to fish or freshwater.39–45 Identification of potentially zoonotic pathogens provides risk assessment data for technical and research personnel who work within the systems, and it also promotes the use of personal protection equipment.
Within multi-system facilities, the pathogen landscape was different between systems more often than it was the same. When pathogens were present within the same facility, there were relatively few differences in disease detection rates, with the exception of P. tomentosa in one system. This trend provides evidence that each system within a facility should be considered unique with regards to disease testing and biosecurity. There is limited information identifying biosecurity risks within zebrafish facilities, but based on fundamental principles of aquaculture, fish movement, equipment disinfection, handling of sick and dead animals, and reducing fish stress 46 all likely play major roles in the spread and impact of infectious disease on zebrafish recirculating systems. Results from this study did not find that there was an association between the number of fish in a system and the subsequent number of pathogens, although larger systems may pose different biosecurity challenges compared to smaller systems. However, a more comprehensive and multi-faceted study is needed to characterize risk factors such as tank number, fish density, movement of fish, and so on that may affect infectious disease dynamics in zebrafish systems.
Through studies that determine best practices, we can begin to utilize resources to support fish health more efficiently. In the absence of established guidelines, the evaluation of each health program is subjective, which makes it difficult to share data and support movement of fish between facilities. The first step to developing these guidelines is to understand the disease landscape to help prioritize pathogens for elimination and exclusion. Based on results from this study, we suggest that this be performed for each system within multi-system facilities and that OIE standards for fish health inspections be used that allow >95% confidence in pathogen detection with a presumed pathogen prevalence of 5%. The use of small numbers of fish, even if sentinel, moribund, or freshly dead, were not sufficient to characterize system-wide fish health, especially when pathogen prevalence was relatively low. We also suggest that histopathology be utilized to confirm infectious disease and characterize its effect on fish health, in addition to screening fish for non-infectious causes of disease.
Expansion of this research to include additional pathogens and replicate systems will further elucidate how best to utilize resources for diagnosing infectious disease in zebrafish systems and set standards for exploring these dynamics in other laboratory models of fish. Our hope is that this will lead to more standardized and efficient guidelines for disease testing.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
