Abstract
Bacteria are relevant in rodent quality assurance programmes if (a) the animals are at risk and (b) presence in the animals makes a difference for animal research or welfare, for example because the agent regulates clinical disease progression or impacts its host in other ways. Furthermore, zoonoses are relevant. Some bacterial species internationally recommended for the health monitoring of rats and mice, that is, Citrobacter rodentium, Corynebacterium kutscheri, Salmonella spp. and Streptococcus pneumonia, are no longer found in either laboratory or pet shop rats or mice, while there is still a real risk of impact on animal research and welfare from Filobacterium rodentium, Clostridium piliforme, Mycoplasma spp., Helicobacter spp. and Rodentibacter spp., while Streptobacillus moniliformis may be considered a serious zoonotic agent in spite of a very low risk. Modern molecular techniques have revealed that there may, depending on the research type, be equally good reasons for knowing the colony status of some commensal bacteria that are essential for the induction of specific rodent models, such as Alistipes spp., Akkermansia muciniphila, Bifidobacterium spp., Bacteroides fragilis, Bacteroides vulgatus, Faecalibacterium prausnitzii, Prevotella copri and segmented filamentous bacteria. In future, research groups should therefore consider the presence or absence of a short list of defined bacterial species relevant for their models. This list can be tested by cost-effective sequencing or even a simple multiple polymerase chain reaction approach, which is likely to be cost-neutral compared to more traditional screening methods.
Introduction
Mice and rats are quality assured by screening for the absence of specific pathogens, for example as recommended in guidelines issued by the Federation of European Laboratory Animal Associations (FELASA). 1 However, many of those bacteria screened for are seldom found in barrier-protected rodents, 2 while the access to modern molecular techniques has revealed other and often non-cultivable bacteria as serious modulators of animal disease models. It has therefore been proposed that symbiotic bacteria with an impact on rodent models are also made part of routine quality assurance, 3 and furthermore, it has been discussed whether some animal models, for example within immunology, would actually be improved if the immune system had been challenged with a more diverse microbiota, even if this included some pathogens. 4
The FELASA pathogens
Bacterial agents recommended by FELASA for routine screening in breeding colonies of rats and mice 1 and the prevalence with which they have been diagnosed globally in recent surveys from laboratory animal colonies, pet shops and single cases.
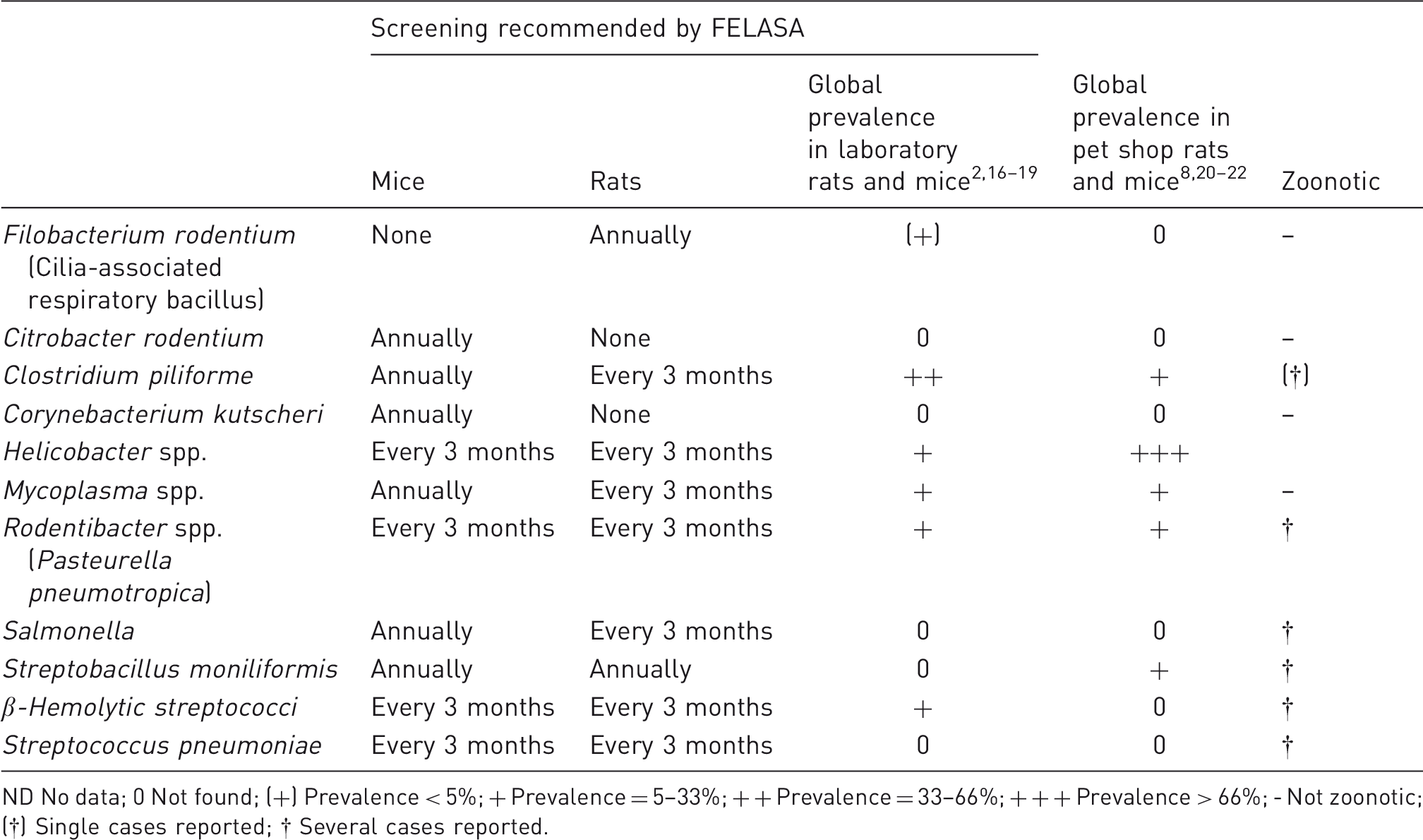
ND No data; 0 Not found; (+) Prevalence < 5%; + Prevalence = 5–33%; + + Prevalence = 33–66%; + + + Prevalence > 66%; - Not zoonotic; (†) Single cases reported; † Several cases reported.
However, it is questionable whether the bacteria currently listed represent the major risk factor for the unsuccessful use of rodents as models for human disease, since some of these, i.e. Citrobacter rodentium, Corynebacterium kutscheri and Streptococus pneumonia, have not been found in any microbiological surveys published within the last 15 years. This is also the case for Streptobacillus moniliformis and Salmonella spp., but S. moniliformis was isolated from barrier-bred laboratory mice back in 199014 and from pet rats 13 and human patients15,16 as late as 2017, and Salmonella spp. is present with a low prevalence among wild rodents17,18 and is one of the most common food-borne infections. 19 The majority of those agents found cause mostly latent infections, but their routine screening can be argued on their ability to interfere with various models even in the absence of clinical manifestations. Rodentibacter spp. (previously known as Pasteurella pneumotropica but now reclassified as eight different species 20 ), while probably unable to cause pneumonia in immune-competent mice,21,22 may be isolated from abscesses,23,24 may alter cytokine expressions, 25 and may have some additional impact on immune-deficient mice.22,26 A few of the numerously described Helicobacter spp. may cause clinical disease.27–32 Clostridium piliforme may cause Tyzzer's disease,33–36 and although infections with it are often latent due to genetic traits in rats 37 and mice, 38 research interference may still be expected due to dysfunction of the liver. 39 Mycoplasma pulmonis and M. arthritidis may cause respiratory, eye and joint-related conditions and may affect reproduction, at least if enhanced by other factors.21,40–45 However, even in the absence of clinical symptoms, it may have impact on certain models.46–50 Filobacterium rodentium (formerly known as cilia-associated respiratory (CAR) bacillus 51 ), while usually asymptomatic, 52 is the cause of chronic respiratory disease in rats,53,54 and BALB/c mice.54,55 There is only one reported case that infection with group B β-haemolytic streptococci, S. agalactiae, has caused general abscesses and mortality, 56 and, therefore, this is an example of an agent that does not necessarily require routine monitoring. Rodentibacter spp., 57 Helicobacter spp., 58 C. piliforme, 59 M. pulmonis, 60 and F. rodentium 61 can be diagnosed by polymerase chain reaction (PCR), and although serology may be considered less sensitive and specific it is also commonly applied for C. piliforme, 35 M. pulmonis 62 and F. rodentium. 63 β-haemolytic streptococci need cultivation on a blood agar prior to serotyping or PCR of β-haemolytic colonies. 64
Other facultative pathogens
Gut commensals with an impact on mouse and rat models
Alistipes
Alistipes spp. appear to impact neuropsychiatric mouse models. Increased abundances have been correlated with a low level of serotonin in the ileum in autistic mice 94 and the expression of depressive behaviour in grid floor housed mice. 95
Akkermansia
Akkermansia muciniphila is the only well-described member of the genus, and the only one observed in rodents to date. 96 It feeds on mucin in the gut. 97 In both humans and mice, if present in the colony, it constitutes 0.1–3% of the total gut microbiota, but it may only be present in young mice. 88 Its anti-inflammatory potential is linked to the reduced incidence or severity of disease, as in murine models for type 1 diabetes98,99 and diet-induced obesity, 100 while it seems to have a co-inducing impact on the development of azoxymethane-induced colon tumours in mice. 101 It is a potential target of dietary intervention studies, 102 for example feeding with the popular prebiotic xylooligosaccharide has been found to propagate A. muciniphila in mice. 103 Therefore, it can be problematic to use mice from colonies free of A. muciniphila in studies of prebiotics. 98
Bifidobacterium
Bifidobacterium spp. are anaerobic and frequently branched bacteria, and are major members of the colon microbiota of mammals. 104 High abundances of bifidobacteria are strongly correlated to low levels of inflammation in mice 89 and leptin in rats. 105 They are among the most popular probiotics for humans, and mice are used for the development of them.106–108 Prebiotics tested in mice often act directly on the abundance of Bifidobacterium spp,89,109,110 and it is therefore crucial that mice in such studies carry this organism. 104
Bacteroides
B. fragilis is normally considered a gut commensal, but toxin-producing strains may be regarded as pathogenic, for example they may induce colonic tumours in multiple intestinal neoplasia mice, 111 and feeding gnotobiotic mice B. fragilis toxins causes symptoms of diarrhoea. 112 In contrast to this, the polysaccharide A (PSA) from B. fragilis protects against Helicobacter hepaticus-induced colitis in mice; most likely because it prevents IL-17 secretion. 113 Through its stimulation of the dendritic cells, PSA is important for gut immune maturation and the balance between T helper cells type 1 and 2, as well as for the gut response to pathogens. 114 B. fragilis is also known to protect against intestinal natural killer T cell-mediated oxazolone-induced colitis in mice. 115 Feeding B. fragilis to the maternal immune activation mouse model of autism reduces symptoms, which may be due to the normalization of one specific gut metabolite, 4-ethylphenylsulfate. 116 B. vulgatus seems to enhance IBD in HLA-B27 transgenic rats 117 and IL-10 knockout mice, 118 and it increases in abundance in old mice. 119
Faecalibacterium
Faecalibacterium prausnitzii is a clostridium-related bacterium that is among the most abundant members of the human gut microbiota, where it is an important producer of butyrate and appears to play an important role in the breakdown of complex carbohydrates.120,121 In mice, it reduces the severity of various models of IBD, and it may be a key target in IBD intervention studies.122–124 It is found spontaneously in some mouse colonies, 125 while it is absent in others. 126 In addition, other but less well-defined Clostridium spp. are known to induce colonic regulatory T cells. 115
Prevotella
Gram-negative mostly anaerobic Prevotella spp. are observed in the gastrointestinal tract, including the oral cavity. P. copri may increase the severity of dextran sodium sulfate (DSS)-induced colitis in mice. 127 The lower response to DSS induction observed in Caspase-3 knockout mice may be counteracted by cohousing with wild type mice, which significantly increases the abundance of Prevotella spp. in the knockout mice. 128 Tumour-bearing mice with azoxymethane/DSS-induced colorectal cancer may have decreased abundances of Prevotellaceae, 101 while a high abundance of Prevotellaceae in the gut of leptin-deficient obese mice correlates to an impaired glucose tolerance. 88
Proteus
Proteus mirabilis is a common inhabitant of the murine gut. So far, it has been considered a commensal; however, it seems to increase in abundance in mouse models of Parkinson's disease, and the transfer of P. mirabilis isolated from such models to other mice damages dopaminergic neurons and motor functions. 129
SFBs (Candidatus Savagella)
SFB is a more popular term for the Clostridium cluster I-related Candidatus Savagella. They are sometimes wrongly referred to as Candidatus Arthromitus, but these are Lachnospiraceae-related bacteria found in arthropods and not in mammals.
130
Species found in mice and rats are closely related but different from one another.
131
Mouse colonies differ in terms of SFB status,93,132 which is crucial to know, as SFBs strongly influence immune functions such as IgA production, T-cell response and intestinal T helper cell type 17 (T
Discussion
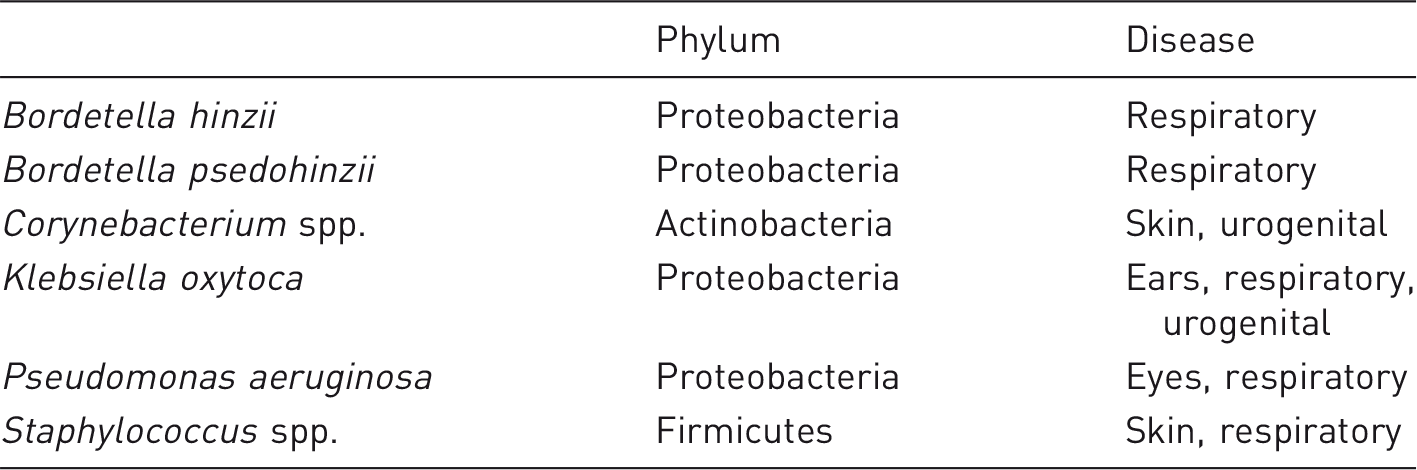
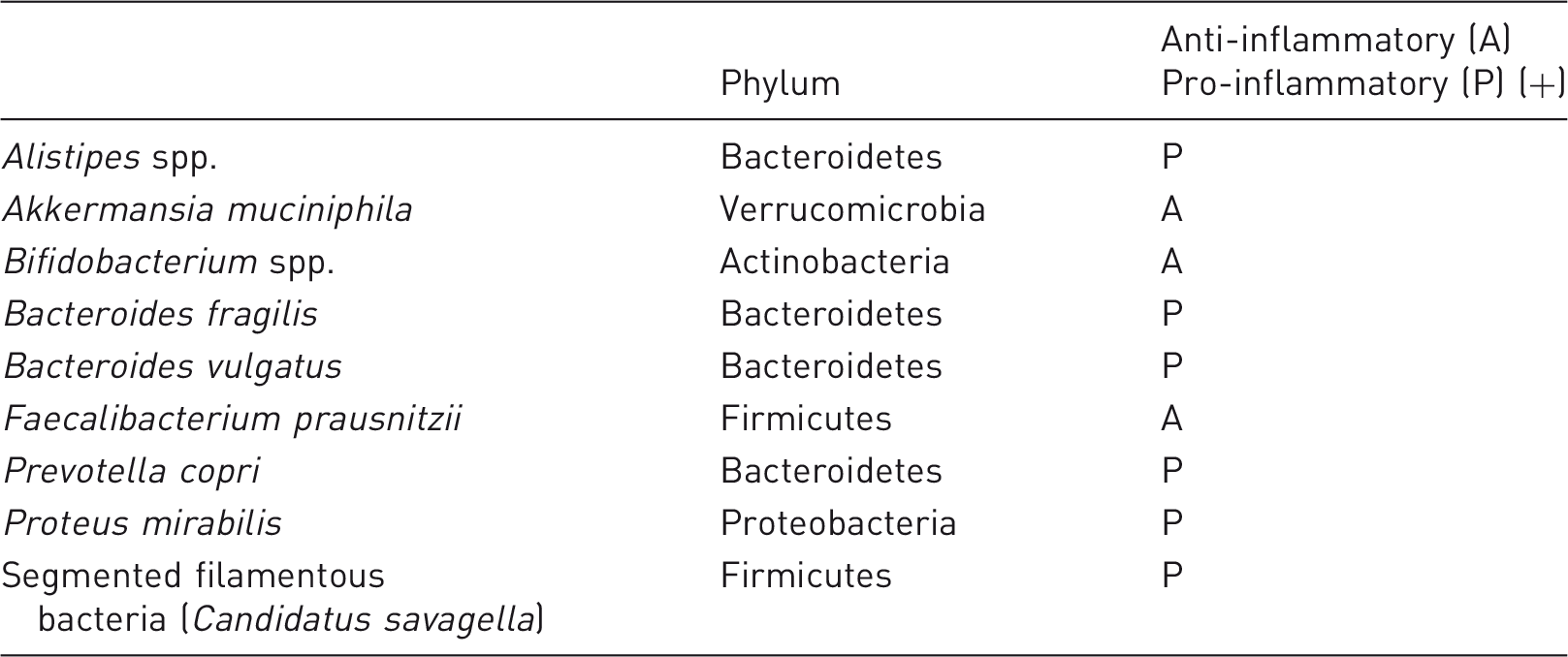
To select specific bacteria for consideration before using animals for a specific study, it seems relevant to consider agents if (a) the animals are in reality at risk and (b) if infections in the animals will make a difference for animal research or welfare, for example because the agent regulates clinical disease progression or in other ways has an impact on animal models. To this should be added that it is relevant to consider a zoonotic potential. This also implies that if neither (a) or (b) are fulfilled, then there is no need for specific screening. C. rodentium, C. kutscheri, Salmonella spp. and S. pneumonia are, in reality, absent in laboratory rats and mice. Furthermore, a single reported case on β-haemolytic streptococcal disease 56 is questionable as a justification for frequent global screening in favour of some of the other agents presented here. All of the bacteria listed in Tables 2 and 3 fulfil requirements (a) and (b), but they are obviously not equally relevant for all types of research and, therefore, it may be up to the individual facilities and the needs of their research groups to decide which ones should be routinely screened for. However, it is no longer sufficient only to secure the sole absence of pathogens. Some bacteria are crucial for some models but ruin other models, for example SFBs facilitate IBD 132 but counteract type 1 diabetes, 139 F. prausnitzii and A. muciniphila reduce severity in IBD 124 and type 1 diabetes models, 98 respectively, but this is precisely why they are targets for intervention.99,123 F. rodentium, C. piliforme, Mycoplasma spp., Helicobacter spp. and Rodentibacter spp. also fulfil requirements (a) and (b). In addition, it may be argued that F. rodentium, C. piliforme, and Mycoplasma spp. represent organisms for which there is a real risk of outbreak of clinical disease with a high prevalence in a colony. It may also be argued that S. aureus is as important as Rodentibacter spp., since reports about abscesses are more or less similar. 140 However, laboratory rodents probably catch this agent from their human caretakers, as it is extremely rare in wild rodents, 141 and, therefore, it is far more difficult to keep out of barrier facilities. Zoonotic agents, such as S. moniliformis, despite presenting a low risk, may be included; however, it could be argued that Leptospira spp., also latently present in the wild rodent population,142,143 are equally important as a zoonosis, and it is unlikely to isolate any of these two agents in modern animal facilities.
Sequencing costs are rapidly declining, and the fast, cost-effective screening of samples for unknown species using portable, real-time sequencers will soon change molecular diagnostics.103,144 Even if sequencing is not an option, most bacteria can still be monitored simply by using PCR. Since it is likely that the list of bacteria with a research impact will continue to grow, it may well be best to reduce the number of organisms routinely screened for at breeders and to allow local needs to direct what should be included in further testing. On the other hand, with the use of multiplex PCRs it may be possible to screen for a longer list of bacteria, routinely or on request, which may be seen as a competitive move for a commercial breeder. As positive results for wanted organisms need less frequent re-confirmation and as laborious cultivations are no longer required, which also eliminates the need for transporting live animals to laboratories, the costs of bacteriology will be reduced dramatically.
In conclusion, research groups may benefit from having a short list of the presence or absence of defined and relevant bacterial species for the animals they use. The list will probably need current revision, as new bacteria with an impact will be involved, and as increase in knowledge and technology will lead to a higher resolution in identification. Routine screening can be performed using cost-effective sequencing or simple multiple PCR approaches. With fewer pathogens on the list and with the transition from cultivation to molecular methods, this is likely to be cost-neutral.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.