Abstract
Peptidylarginine deiminase (PAD) is the enzyme responsible for converting protein-bound arginine residues to citrulline. It has recently been shown that a number of epidermal proteins, including filaggrin, trichohyalin, and keratins, are deiminated by the action of PAD, suggesting a possible role for protein deimination during the final stages of epidermal differentiation. We report here a novel PAD substrate found during the course of identifying deiminated proteins in cultured rat epidermal keratinocytes. We found that a 70-kD protein localized to the periphery of the nucleus was preferentially deiminated after ionomycin treatment in the presence of 2 mM calcium and was associated with apoptotic events in these cells. Furthermore, we discovered that the deimination of nuclear protein could be induced by transfection of a PAD cDNA into rat epidermal keratinocytes. These data suggest that PAD may act on the 70-kD nuclear protein to induce disassembly of the nuclear lamina and promote apoptosis during terminal epidermal differentiation.
T
Although no physiological function has been ascribed to PAD, a number of proteins have been shown to be modified by PAD during the terminal stages of epidermal differentiation (Rogers 1963; Harding and Scott 1983; Senshu et al. 1995). Our recent study showed that the major deiminated proteins in human epidermis are partially degraded disulfide-crosslinked keratin K1 and that the minor components are derived from filaggrin, trichohyalin, and keratin K10 (Senshu et al. 1996). The presence of deiminated K1/K10 and the virtual absence of deiminated K5/K14 suggested that PAD acted preferentially on the differentiation-specific suprabasal keratins during the terminal stages of epidermal differentiation. We therefore speculated that the protein deimination might influence the interaction of basic K1 with its acidic partner K10 and preexistent K5/K14 networks. We further speculated that interactions between filaggrin and keratins might also be influenced by protein deimination. If deiminated filaggrin possesses lower affinity for keratins, as suggested by Harding and Scott (1983), it can be concluded that the deimination of filaggrin impairs its biological function as a keratin-associated protein.
To further explore the role of protein deimination, we attempted to identify the deiminated proteins present in cultured epithelial cells after activation of endogenous PAD by ionomycin treatment in the presence of 2 mM calcium. We have shown that a 70-kD nuclear protein is the major deiminated protein in these cultured cells. We have also shown that transfection of a PAD cDNA causes deimination of the nuclear protein. These findings led us to speculate that PAD might induce nuclear disassembly and promote apoptosis during terminal epidermal differentiation.
Materials and Methods
Ionomycin Treatment of Cultured Cells
Cells from a rat epidermal keratinocyte cell line (kindly provided by Dr. Howard P. Baden of Harvard University, Boston) placed on 12-mm glass coverslips were grown in DMEM (Gibco BRL; Gaithersburg, MD) supplemented with 10% fetal calf serum (Gibco BRL) in a 5% CO2 atmosphere at 37C. These cells were washed three times with prewarmed Locke's solution (0.15 M NaCl, 5 mM KCl, 2 mM CaCl2, 5 mM HEPES, pH 7.1) and were subsequently incubated in 5 μM ionomycin Ca-salt (Sigma; St Louis, MO) in prewarmed Locke's solution for an additional 2 hr.
Transfections
To construct the expression vectors, full-length cDNA of PAD (Watanabe and Senshu 1989) was inserted into the EcoRI cloning site of an expression vector, pECE (Ellis et al. 1986) under the control of the SV40 early promoter. The transfections into rat epidermal keratinocytes were performed by the lipofection method using LipofectAMINE reagent (Gibco BRL) according to the manufacturer's protocol. Briefly, 6 × 104 cultured cells were placed on 12-mm glass coverslips 1 day before transfection. Five μg of vector DNA in 25 μl of OptiMEM (Gibco BRL) was mixed with 2.5 μl of LipofectAMINE reagent in 25 μl of OptiMEM, and was further mixed with 200 μl of OptiMEM 5 min later. The cells were incubated in the above-mentioned transfection mixture for 5 hr. After addition of 250 μl of DMEM with 20% fetal calf serum, the cells were incubated overnight. After the medium was discarded, the cells were further incubated for 24 hr in 500 μl of DMEM with 10% FCS. As a control, the expression vector pCH110 (Pharmacia Biotech; Uppsala, Sweden), containing lacZ gene under the control of the SV40 early promoter, was used.
Immunofluorescence Staining of Deiminated Proteins in Cultured Cells
At 48 hr after the transfection or 2 hr after ionomycin treatment, cells grown on the 12-mm glass coverslips were fixed with 4% paraformaldehyde in PBS at room temperature (RT) for 15 min and washed with PBS. Citrulline residues in the cells were chemically modified for 3 hr at 37C in 0.0125% FeCl3, 2.3 M H2SO4, 1.5 M H3PO4, 0.25% diacetyl monoxime, 0.125% antipyrine, and 0.25 M acetic acid (modification medium) (Senshu et al. 1992). After three washes with PBS, the processed cells were incubated successively with anti-modified citrulline IgG (0.125 μg/ml) (Senshu et al. 1992), biotinylated goat anti-rabbit IgG antibody (Cappel; Durham, NC), and Cy3-conjugated streptavidin (Sigma).
In the control experiments of the transfection study, the cells were first incubated in 0.1% X-gal, 5 mM potassium ferricyanide, 5 mM potassium ferrocyanide, and 2 mM MgCl2 in PBS for 30 min. Later these cells underwent immunofluorescence staining of deiminated proteins as described above.
Protein Extraction and Immunoblot for Detection of Deiminated Proteins
To determine the presence of deiminated proteins, total proteins were extracted 2 hr after the ionomycin treatment described above with a sample buffer of SDS-PAGE. Samples were separated by SDS-PAGE and then transferred to a nitrocellulose membrane to stain total proteins with Fast Green and to detect deiminated proteins. The blot for detection of deiminated proteins was processed as described previously (Senshu et al. 1992). In brief, the blot was soaked in 0.1% ovalbumin in PBS for 15 min at RT and then rinsed lightly with distilled water. The blot was next soaked in 4% paraformaldehyde in PBS for 15 min at RT and rinsed thoroughly with distilled water. Citrulline residues on the blot were chemically modified by overnight incubation at 37C in the modification medium described above. The blot was then incubated successively with anti-modified citrulline IgG (0.125 μg/ml) and peroxidase-labeled anti-rabbit IgG goat antibody (Bio-Rad, Hercules, CA; 5000-fold dilution), followed by silver enhancement (Merchenthaler et al. 1989). The specificity of the immunochemical detection was confirmed by probing equivalent blots incubated in a modification medium free of diacetyl monoxime, antipyrine, and acetic acid.
Detection of Apoptotic Cells
After the ionomycin treatment described above, the apoptotic cells were identified using Apop Tag Plus In Situ Apoptosis Detection Kit-Fluorescence (Oncor; Gaithersburg, MD) according to the manufacturer's protocol. Briefly, 5 × 104 cells grown on the 12-mm glass coverslips were fixed with 4% paraformaldehyde in PBS at RT for 30 min. Fixed cells were incubated in 30 μl of a mixture of terminal deoxynucleotidyl transferase and digoxigenin nucleotides for 1 hr at 37C. These cells were further incubated with 30 μl of the fluorescein-labeled anti-digoxigenin sheep Fab fragment for 30 min at RT.
Results
We showed in our previous report that all-trans retinoic acid as well as calcium increases PAD activity in this cell line, although the mechanism of this activation is largely unknown (Ishigami et al. 1996). To activate endogenous PAD, we increased the intracellular calcium concentration with ionomycin in the presence of 2 mM calcium. Unexpectedly, in all cultured cells we observed a significant degree of immunoreactivity to citrulline around the nucleus after ionomycin treatment (Figures 1a and 1b) but not on the cytoplasm as had been seen previously (Senshu et al. 1995,1996). Moreover, the immunoreactivity was localized to the periphery of the nucleus, as determined by changing the focal plane, and had an irregular cage-like pattern. In contrast, the cells before ionomycin treatment did not reveal any immunoreactivity (Figures 1c and 1d).
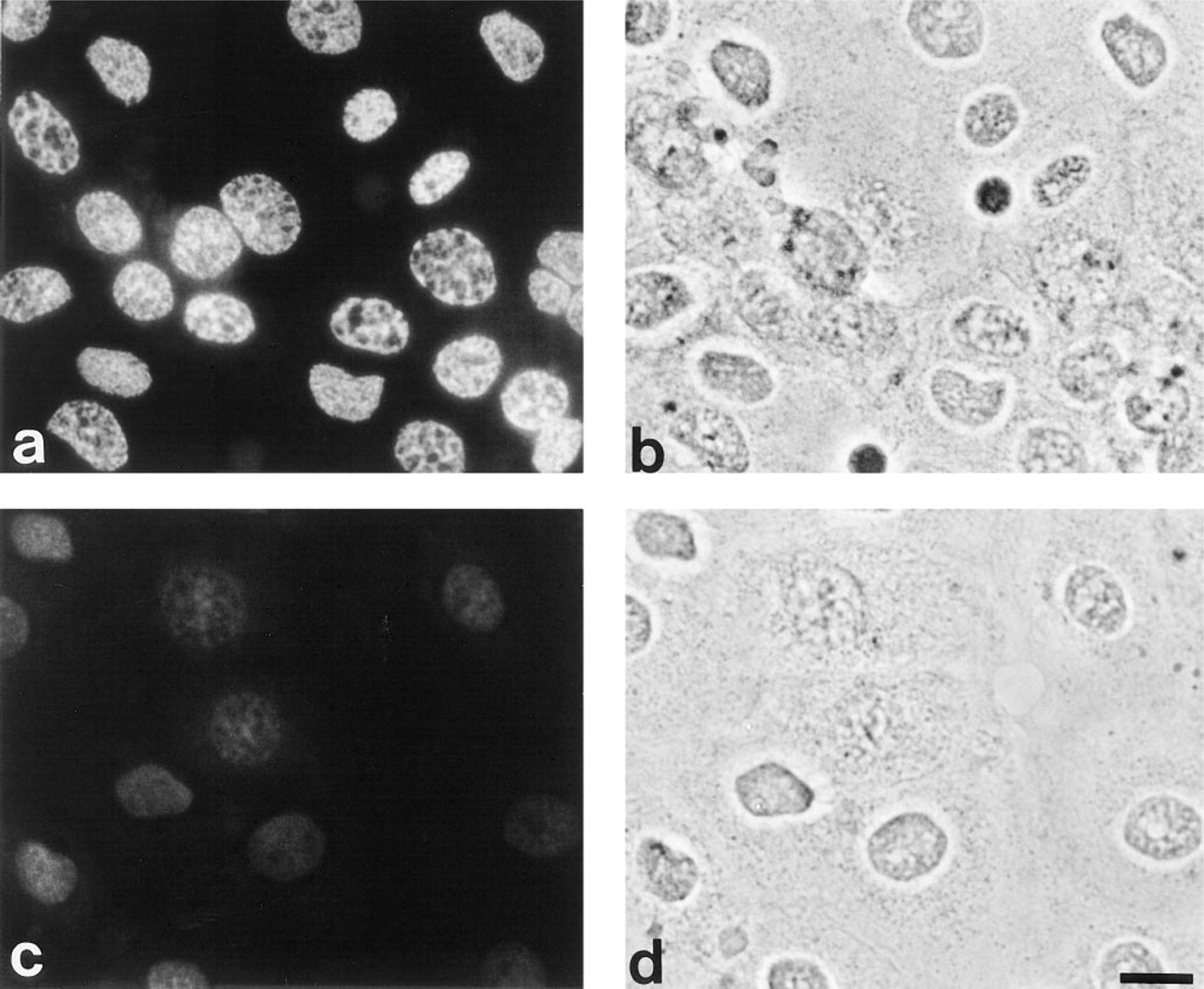
Localization of deiminated proteins in cultured cells with
The immunofluorescence analysis data raised the possibility that this antibody was recognizing a previously unknown PAD substrate. To explore this possibility in greater detail, samples obtained from cells with and without ionomycin treatment were resolved by SDS-PAGE and Western blotted to a nitrocellulose membrane for staining total proteins with Fast Green (Figure 2, Lane 1) as well as detecting deiminated proteins with anti-modified citrulline antibody (Figure 2, Lanes 2-4). The amounts of proteins applied to Lanes 3 and 4 were adjusted so that they were approximately equal to the overall staining intensity in the keratin region of samples in Lanes 1 and 2. Gels were calibrated with a molecular weight marker mixture (Bio-Rad #161-0304). Notably, a major band appeared at the molecular weight of 70 kD after incubation with ionomycin in the presence of 2 mM calcium for 2 hr (Figure 2, Lane 2), and a faint, minor band appeared with a higher and lower molecular weight (Figure 2, Lane 2). Although we do not have enough evidence to explain the nature of these molecules, one possibility is that the higher molecule may be the precursor form and the lower molecule might be the degradation product or less phosphorylated form. Virtually no appreciable amount of deiminated proteins was detected in the sample obtained from the cells without the ionomycin treatment (Figure 2, Lane 3) or from those without incubation in the modification medium (Figure 2, Lane 4).
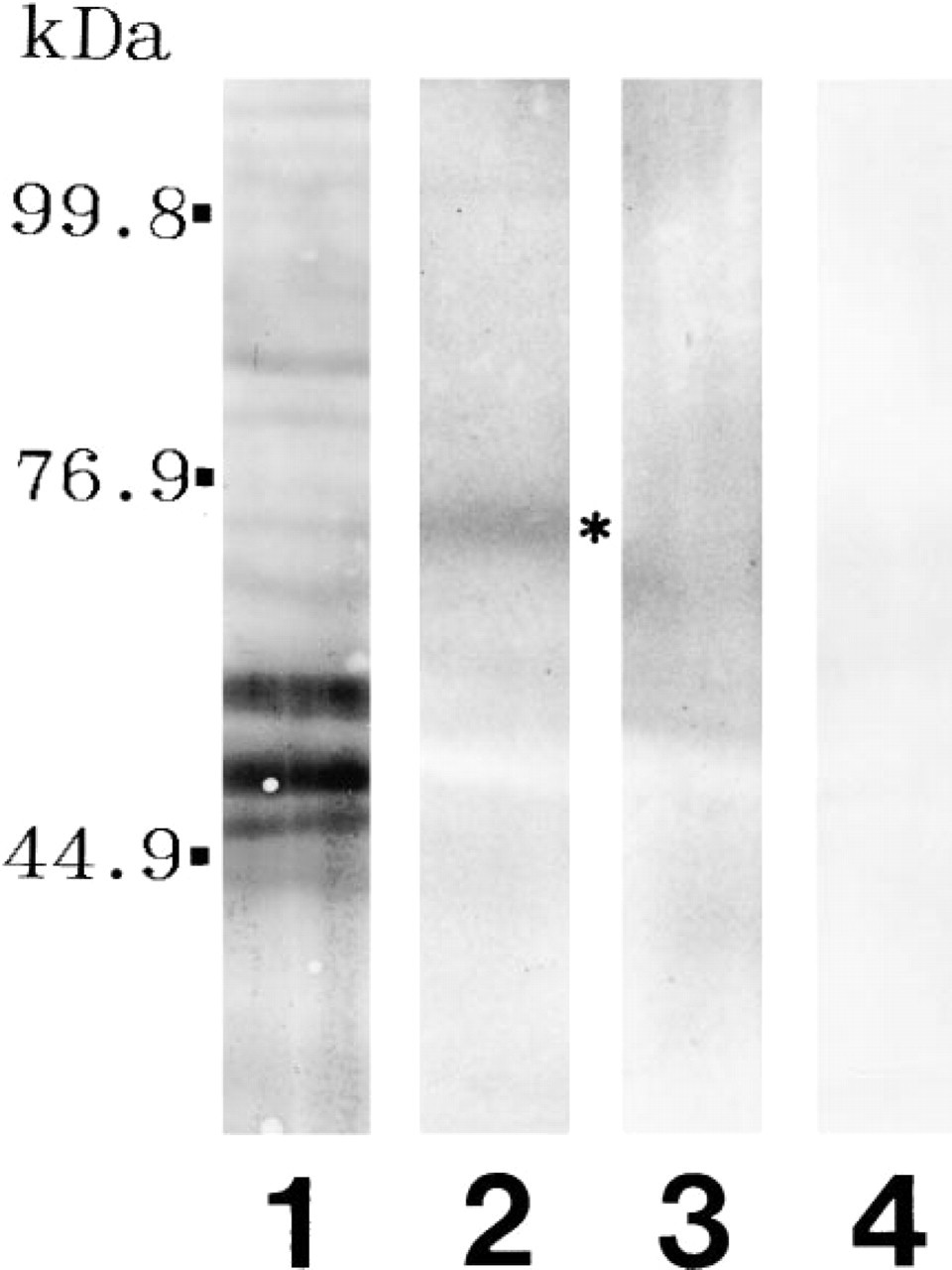
Identification of deiminated proteins in the extracts from cultured cells with (Lane 2; asterisk shows 70-kD protein) or without (Lane 3) ionomycin treatment. Lane 1 shows total proteins revealed by Fast Green staining, and Lane 4 shows deiminated proteins in the extracts from cultured cells with ionomycin treatment without incubation in the modification medium.
The process of terminal epidermal differentiation is regulated in a highly complicated fashion. During the transition from granular cells to cornified cells, the cell membrane becomes more permeable, allowing a calcium influx that induces several terminal differentiation-related events, including apoptosis. Because the DNA of cells undergoing apoptosis may be fragmented, even though the nuclear envelope remains intact under the phase-contrast view, we assayed for the presence of apoptotic cells to verify that ionomycin treatment induces DNA fragmentation in these cells. As shown in Figures 3a and 3b, all cultured cells treated with ionomycin have shown the incorporation of digoxigenin-labeled nucleotides in their nuclei, suggesting DNA fragmentation in these cells. In contrast, no cells undergo the apoptotic processes before ionomycin treatment (Figures 3c and 3d).
The biological effects on cultured keratinocytes after ionomycin treatment were unexpected and diverse. We were especially surprised to observe that calcium influx induced protein deimination in the nuclear protein concomitant with DNA fragmentation. These findings suggest that the deimination of this nuclear protein has an important role in apoptotic events. To examine this directly, we transiently transfected rat epidermal keratinocytes with the expression vector containing the PAD cDNA under the control of the SV-40 promoter and analyzed the cells by indirect immunofluorescence staining using anti-modified citrulline antibody. The transfection efficiency was approximately 3-5% under our experimental conditions. In the transfected cells, the citrulline immunoreactivity was predominantly localized in the nucleus, whereas it was not detected in the surrounding cells (Figures 4a and 4b). Furthermore, the transfected cells displayed nuclear disruption with a compact cytoplasm, and had rounded up and lifted from the substrate. In contrast, neither citrulline immunoreactivity nor any morphological change was observed in cells transfected with the expression vector pCH110 containing the lacZ gene under the control of the SV40 early promoter (Figures 4c and 4d).
Discussion
The biological significance of protein deimination has not been well studied, partly because the methods to detect deiminated proteins are not sufficiently sensitive. We have recently developed a sensitive method for the detection of deiminated proteins based on the chemical modification of citrulline residues and subsequent probing of the modified residues with a monospecific antibody (Senshu et al. 1992). This method enabled us convincingly to detect the major deiminated proteins by immunofluorescence and immunoblot analysis. To identify the PAD substrates that are subjected to protein deimination in cultured cells, we conducted indirect immunofluorescence staining using the newly developed method on rat epidermal keratinocytes under distinctive experimental conditions. Unexpectedly, we found that a protein at the periphery of the nucleus is deiminated in calcium-induced apoptotic events and that the overexpression of PAD caused the deimination and disintegration of the nucleus in transfected cells. These findings suggest that PAD should be added to the growing list of apoptosis-related proteins. However, it remains unclear why protein deimination was not observed on the cytoplasmic keratins as we have reported in our previous study (Senshu et al. 1995,1996). One possibility is that PAD might preferentially act on the skin-type differentiation-specific keratins K1/K10, but not on basal keratins.
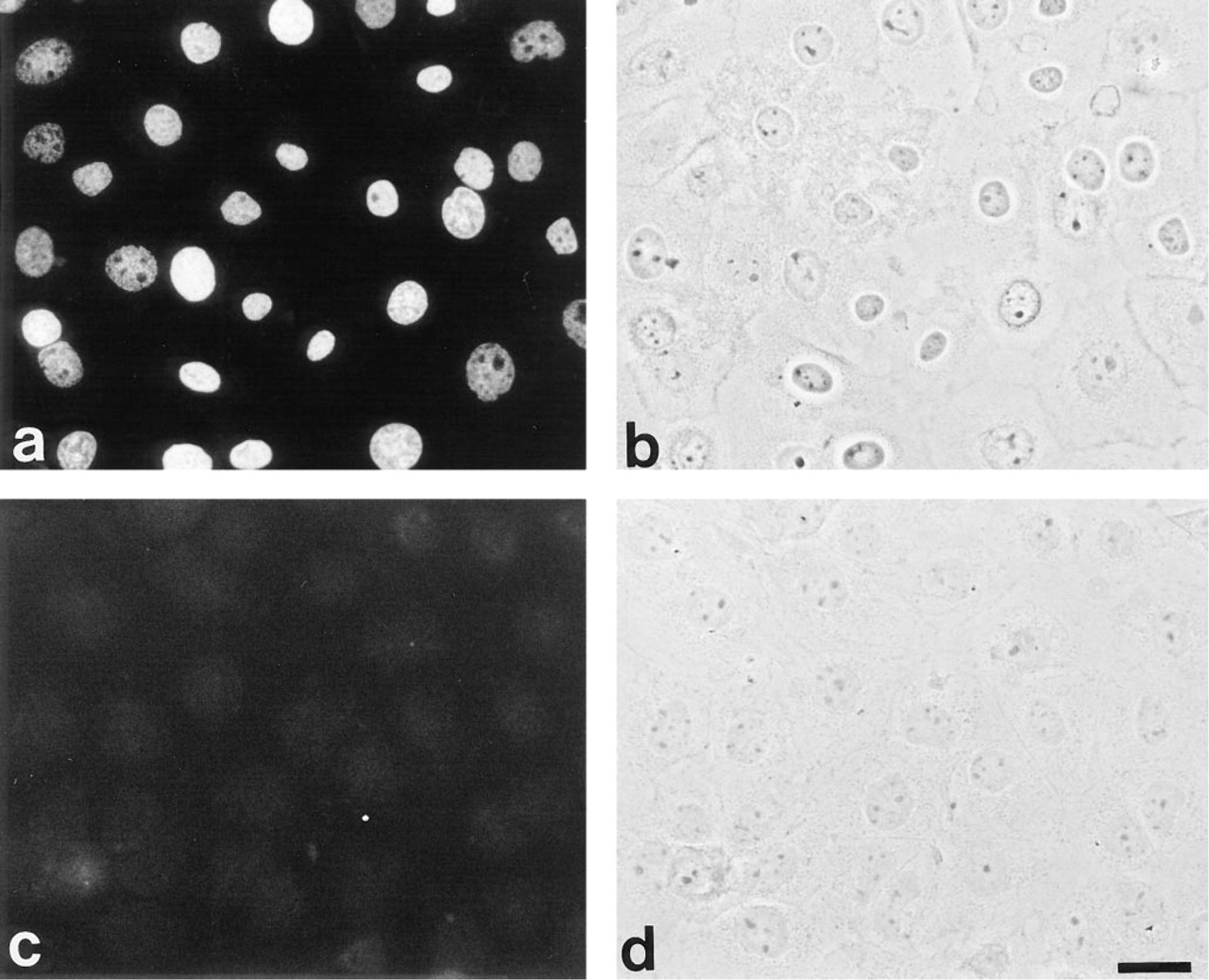
Detection of apoptotic cells with
Identifying the substrates of PAD will be the key to understanding the role of protein deimination in vivo. Given its molecular weight and its localization, it is attractive to think that the protein identified as a PAD substrate in this study may be one of the nuclear lamins. Unfortunately, detailed identification of the deiminated proteins was not accomplished in the present study because of the limited availability of antibodies specific enough to rat nuclear lamins. Further studies are necessary for characterization of the 70-kD deiminated nuclear protein and clarification of its role in normal and apoptotic cells. It will also be important to examine whether the deimination of the 70-kD nuclear protein is followed by the disintegration of the nucleus in association with membrane blebbing and DNA fragmentation in a stable transformant. If such is the case, we will be able to obtain biochemical evidence of apoptotic events that could not be shown in transiently transfected cells because of the limited quantity of available materials.
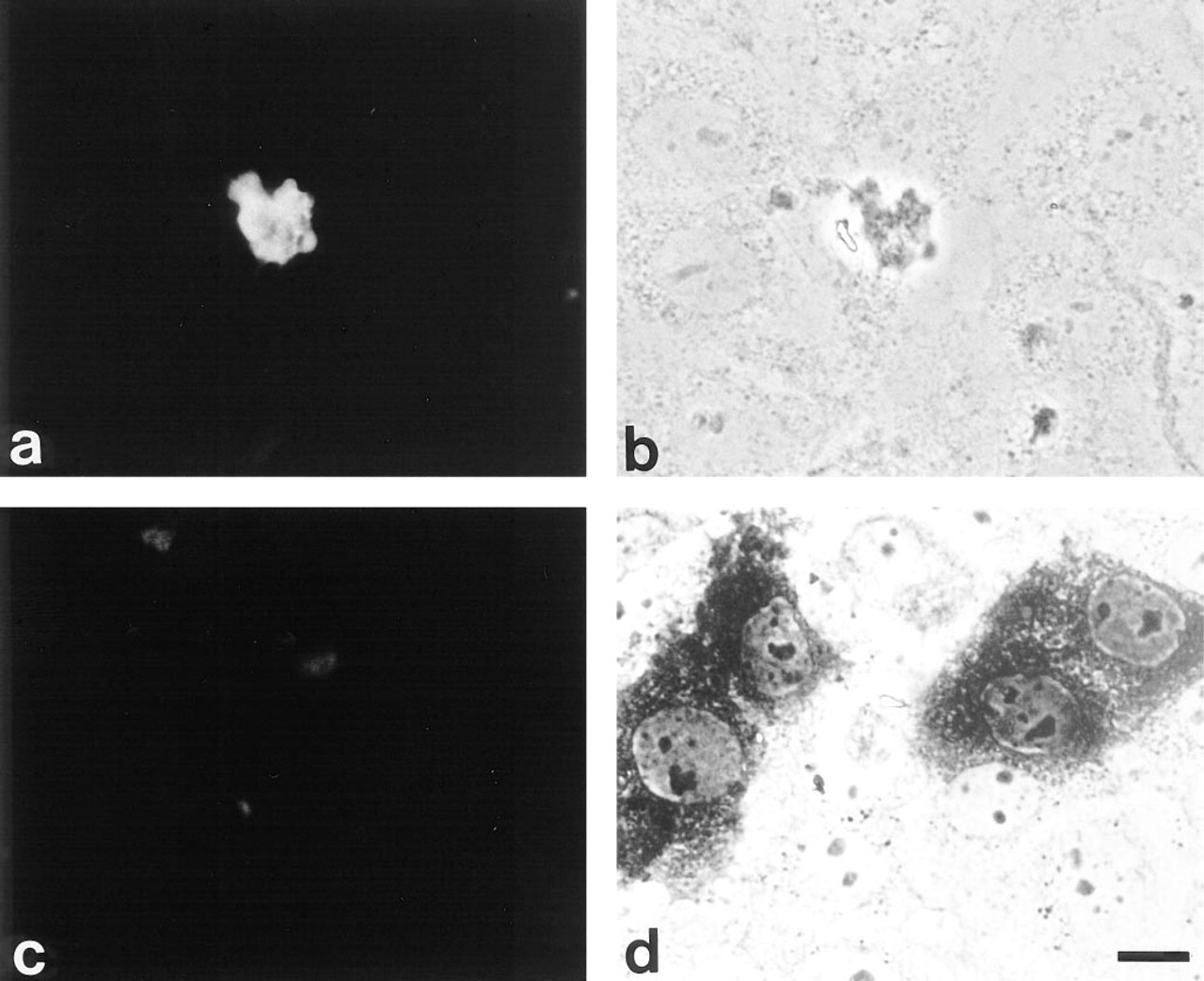
Localization of deiminated proteins in cultured cells transfected with PAD cDNA
Although the detailed mechanisms of nuclear degradation by protein deimination remain obscure, this study suggests a possible role for PAD in apoptotic events. Apoptosis is a genetically regulated mechanism of controlled cell death commonly observed during development, disease, and viral infections (reviewed in Martin and Green 1995; White 1996). Several genes and gene products associated with apoptosis have been identified, including the multiple downstream substrates of the ICE-related cysteine proteases. Major substrates identified include poly (ADP-ribose) polymerase, lamins A, B, and C, α-fodrin, topoisomerase I, and β-actin. Although the role of any of these substrates in apoptosis during terminal epidermal differentiation is still not clear, in considering our present findings and other previous findings (Gavrieli et al. 1992; Senshu et al. 1995, 1996), we can offer the following speculation as to the possible role for PAD in apoptotic events. PAD initially deiminates the 70-kD nuclear protein to facilitate nuclear breakdown by opening up the nucleus and permitting degradation of other nuclear substrates, as well as rendering the nucleases accessible to activation by cytoplasmic factors. Subsequently, the keratins appear to be deiminated, resulting in the collapse of the filamentous keratin network forming a densely packed filamentous structure which is known morphologically as a “keratin pattern.” Such speculation might be elucidated by studying the correlation between the appearance of deiminated proteins and the morphological changes in the transitional layers between the granular layers and the cornified layers in the slowly keratinizing epithelium. By identifying nuclear proteins involved in these processes, the mysteries underlying the dynamic roles of PAD during terminal epidermal differentiation should become increasingly apparent.
Footnotes
Acknowledgments
Supported by a grant (#09670896) from the Ministry of Education, Science, Sports and Culture of Japan.
We thank Dr Joseph A. Rothnagel (University of Queensland, Australia) for his useful discussion and Dr Howard P. Baden (Harvard University, Boston) for the rat epidermal keratinocyte cell line.
