Abstract
It is well documented that several tissues, including the prostate, are actively involved in the local formation and inactivation of hormonal steroids. To identify the cell types involved in the formation and inactivation of androgens and estrogens in the ventral lobe prostate, we have localized by in situ hybridization (ISH) a large number of steroidogenic as well as steroid-inactivating enzyme mRNAs in the adult mouse prostate. In parallel studies, we also measured enzyme mRNA levels by quantitative real-time PCR (RT-PCR) in ventral lobe prostates. From the results obtained with quantitative RT-PCR, it appears that, with a few exceptions, the enzyme with low mRNA expression could not be detected by ISH. The following enzymes have been localized by ISH: 17β-hydroxysteroid dehydrogenase (17β-HSD) types 1, 2, 3, 4, 7, 8, 9, 10, and 11; 5α-reductase type 2; 5β reductase type 1; P450 7α hydroxylase; estrogen sulfotransferase type 1; 11 β-HSD types 1 and 2; and UDP-glucuronosyltransferase 1A6. All of these mRNAs are expressed in the epithelial cells of prostatic acini. Several enzyme mRNAs were also localized in stromal cells. Types 1, 7, and 10 17β-HSD, estrogen sulfotransferase type 1, and 11 β-HSD types 1 and 2 were found only in epithelial cells. The present results indicate that both epithelial and stromal cells in the mouse prostate play a role in local formation and inactivation of hormonal steroids.
I
In recent years, the existence of a local biosynthesis of sex steroids, or intracrinology (Labrie et al. 1988; Labrie 1991) has been supported by demonstration of the expression of several steroidogenic enzymes in peripheral tissues, including the prostate (Martel et al. 1994; Luu-The 2001; Labrie et al. 2003). We have also reported, using immunocytochemistry and in situ hybridization (ISH), the cellular localization of 3β-hydroxysteroid dehydrogenase (3β-HSD) type 1, 17β-HSD type 5, and 5α-reductase types 1 and 2, the enzymes involved in the conversion of circulating DHEA to DHT in human prostate (Pelletier et al. 1998,1999; El-Alfy et al. 1999). So far, the localization of steroidogenic enzyme mRNAs in the mouse prostate has not been reported.
To define the cell types involved in the local synthesis and degradation of sex steroids in the rodent prostate, we have studied the localization of mRNAs expressing a series of steroidogenic enzymes in the mouse ventral prostate by ISH. For each enzyme, we also measured mRNA levels in mouse ventral prostate using quantitative real-time PCR (RT-PCR).
Materials and Methods
Animals
Fourteen adult male and four female (26–30 g) C57BL6 mice were housed under constant temperature (21 ± 1C) and light (lights on from 06 to 20 hr) regimen. The animals received Purina Chow (Ralston-Purina, St Louis, MO) and tap water ad libitum. For histological procedures, four male and four female mice were perfused between 09 and 10 hr, as described below. For RNA preparation, 10 male animals were killed by cervical dislocation. The ventral prostates were rapidly trimmed and snap frozen in liquid nitrogen and stored in a −80C freezer until processed.
Mouse prostate cell expression of mRNAs encoding the enzymes involved in hormonal steroid formation and inactivation
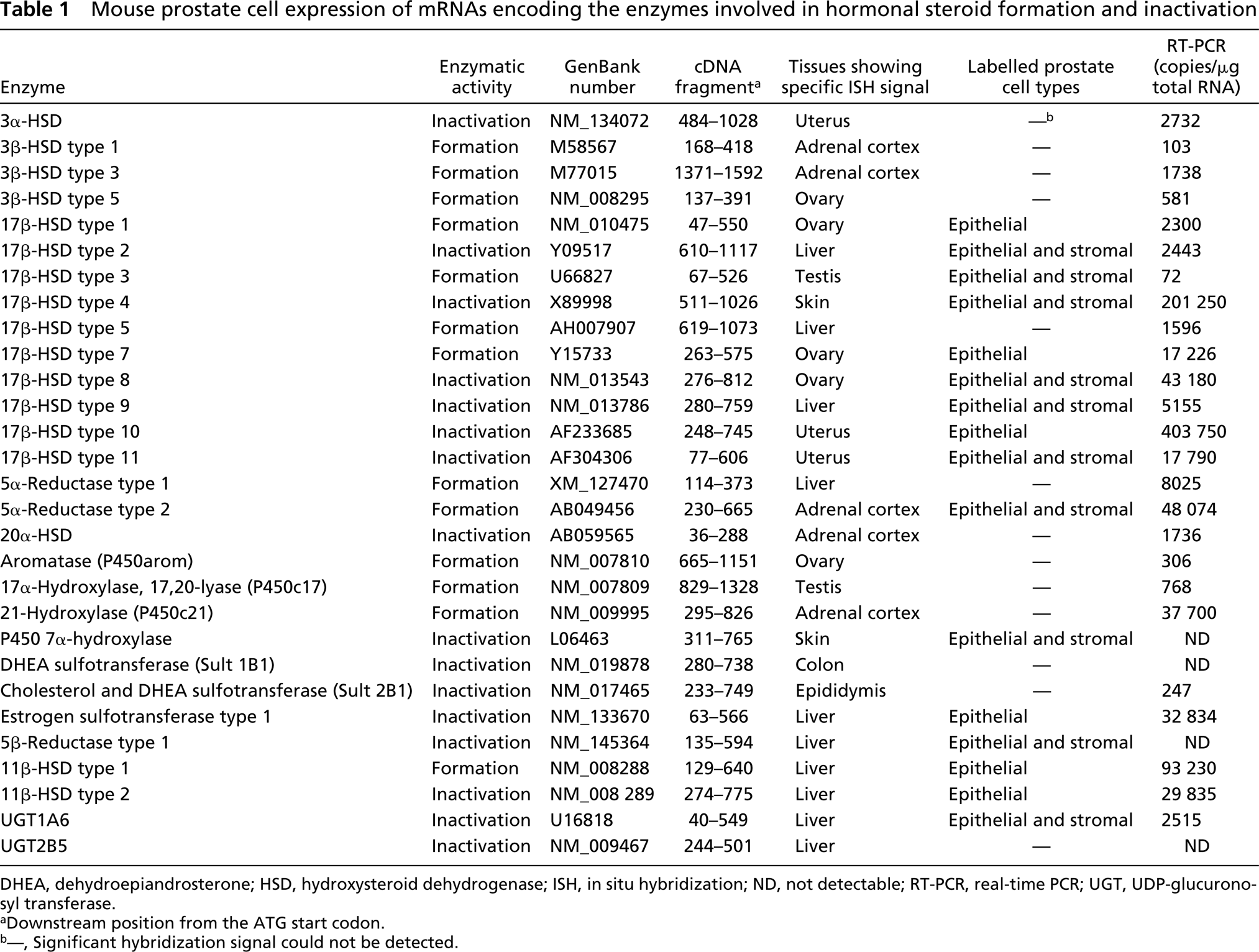
DHEA, dehydroepiandrosterone; HSD, hydroxysteroid dehydrogenase; ISH, in situ hybridization; ND, not detectable; RT-PCR, real-time PCR; UGT, UDP-glucuronosyl transferase.
aDownstream position from the ATG start codon.
b—, Significant hybridization signal could not be detected.
Histological Procedures
All the animals were perfused transcardially with 50 ml 4% (w/v) paraformaldehyde in 0.1 M phosphate buffer (pH 7.4). The different tissues, namely the liver, kidney, lung, adrenal, ovary, testis, mammary gland, uterus, and ventral lobe of the prostate were excised and postfixed in the same fixative for 24 hr at 4C. All the samples were then placed in 15% sucrose in 0.1 M phosphate buffer before being quickly frozen in isopentane chilled in liquid nitrogen.
In Situ Hybridization
Frozen sections (10 μm thick) were serially cut at −20C and mounted onto gelatin- and poly-
The cDNA fragments were obtained by amplification using polymerase chain reaction. ISH with the antisense and sense 35S-labeled cRNA probes was performed as previously described (Givalois et al. 1997). Briefly, the sections were prehybridized at room temperature in a humid chamber for 2 hr in 450 μl/slide of a prehybridization buffer containing 50% formamide, 5× SSPE (1× SSPE being 0.1 M NaCl, 10 mM NaH2PO4, pH 7.4, 0.8 mM EDTA), 5× Denhart's buffer, 200 mg/ml, denatured salmon testis DNA (Sigma, St Louis, MO), 200 μg/ml yeast tRNA, 2 μg/ml Poly A (Boehringer-Mannheim, Montreal, Canada), and 4% dextran sulfate. After prehybridization treatment, 100 μl hybridization mixture (prehybridization buffer containing 10 mM dithiothreitol and the 35S-labeled cRNA probe at a concentration of 10 × 106 cpm/ml) was spotted on each slide, sealed under a coverslip, and incubated at 37C overnight (15–20 hr) in a humid chamber.
(
After hybridization, coverslips were removed and slides were rinsed in 2× SSC at room temperature for 30 min. Sections were then digested by RNase A (20 μg/ml in 2× SSC) at 37C for 30 min, rinsed in decreasing concentrations of SSC (2× SSC and 1 × SSC) for 30 min at room temperature, washed in 0.5× SSC for 30 min at 37C, followed by 90 min at room temperature in 0.5 × SSC, and finally for 30 min at room temperature in 0.1 × SSC.
The sections were then dehydrated and coated with liquid photographic emulsion (Kodak-NTB2; diluted 1:1 with water). Slides were exposed for 7–45 days, developed in a Dektol developer (Kodak, Rochester, NY) for 2 min, and fixed in rapid fixer (Kodak) for 4 min. Thereafter, the sections were rinsed and stained with haematoxylin.
RNA Preparation and Quantification of mRNA Expression Levels
Total RNA was isolated from a pool of 10 prostates with Trizol (Invitrogen, Burlington, Ontario). Twenty μg of total RNA was converted to cDNA by incubation with 400 U SuperScript II reverse transcriptase (Invitrogen), T7-oligo-d (T)24 as primer (5′-GGC-CAG-TGA-GTA-ATA-CGA-CTC-ACT-ATA-GGG-AGG-CGG-(dT)24–3′), 1× first-strand buffer (50 mM Tris-HCl, pH 8.3, 75 mM KCl, 3 mM MgCl2, 10 mM dithiothreitol) and 0.5 mM deoxynucleotide triphosphate at 42C for 1 hr.
Quantification of mRNA levels was performed using a quantitative RT-PCR method that permits monitoring the efficiency of the PCR amplification process by calculating the efficiency coefficient of corresponding standard curves. The LightCycler Realtime PCR apparatus as well as reagents were from Hoffman-La Roche, Inc. (Nutley, NJ). The reaction was performed using the amount of cDNA corresponding to 30 pg of initial total RNA. Oligoprimer pairs that allow the amplification of ~200 bp of indicated mRNA were designed by GeneTools software, and their specificity was checked by blasting in the GenBank database. To avoid errors due to RNA and cDNA preparation and loading, we normalized with a housekeeping gene Atp5o (subunit O of ATPase) at each assay. Atp5o has been shown to be a gene that has stable expression levels from embryonic life through adulthood in various tissues (Warrington et al. 2000). The mRNA levels are expressed as number of copies/μg total RNA that are calculated using a standard results curve of Atp5o.
Results
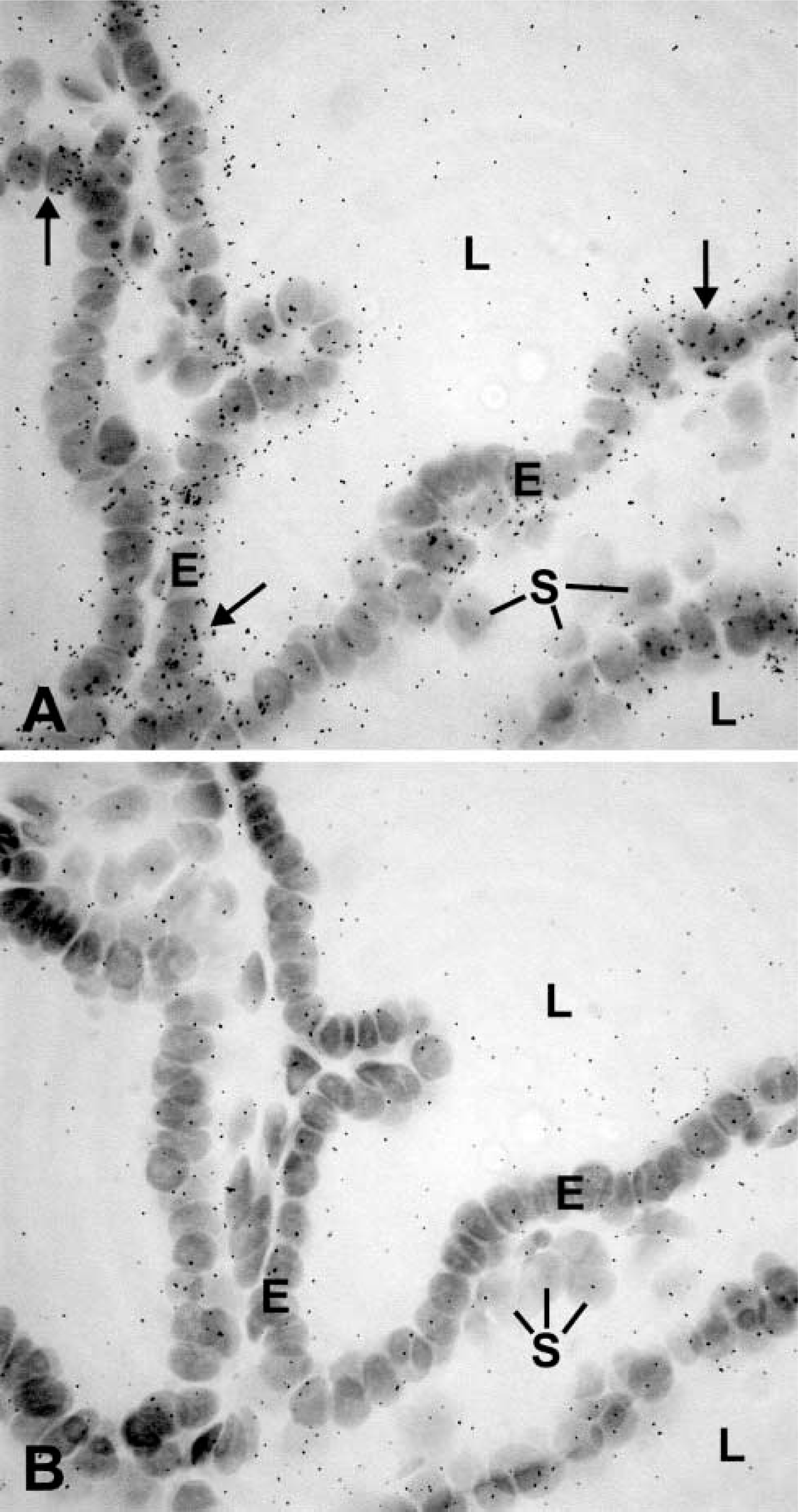
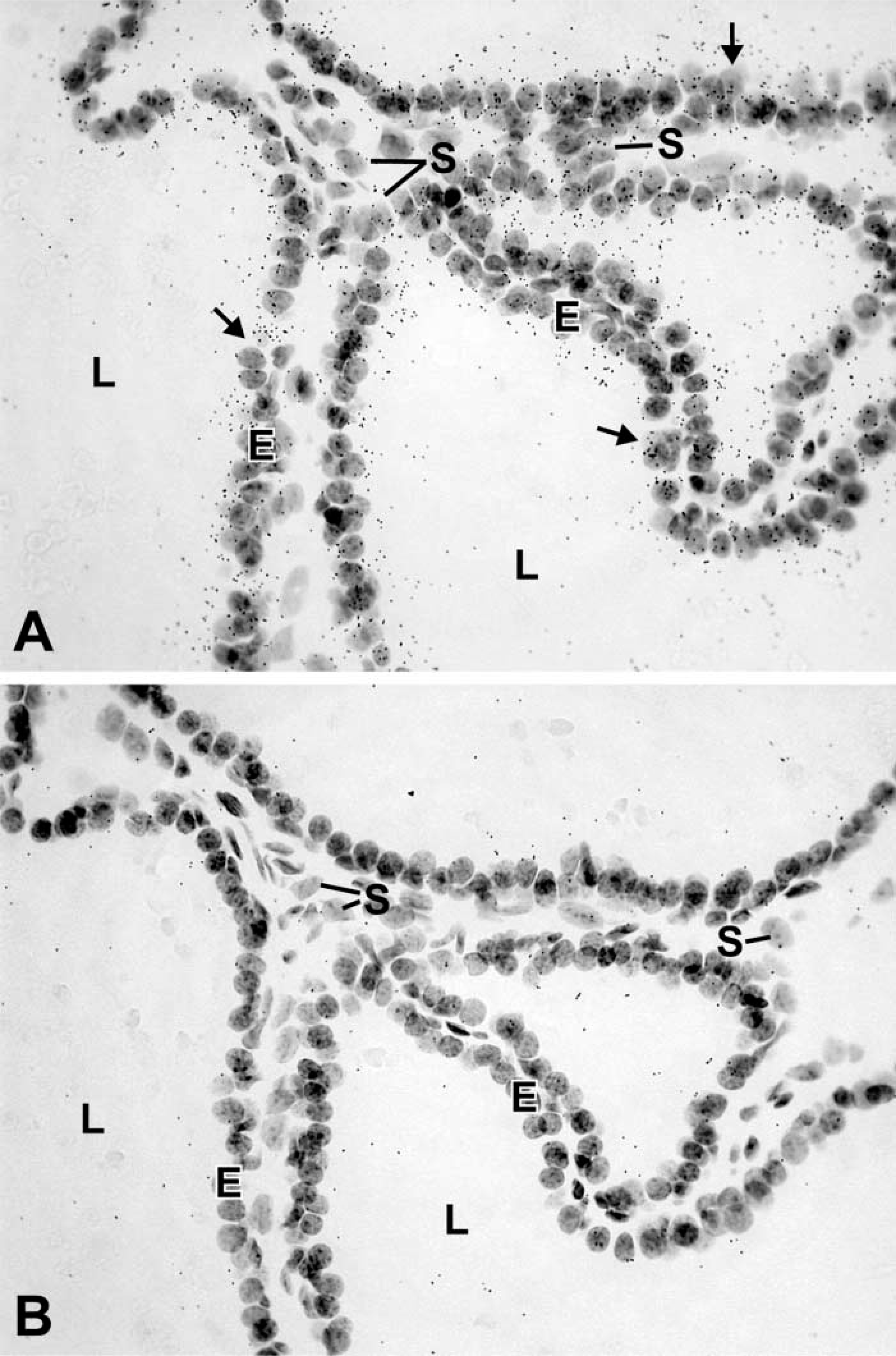
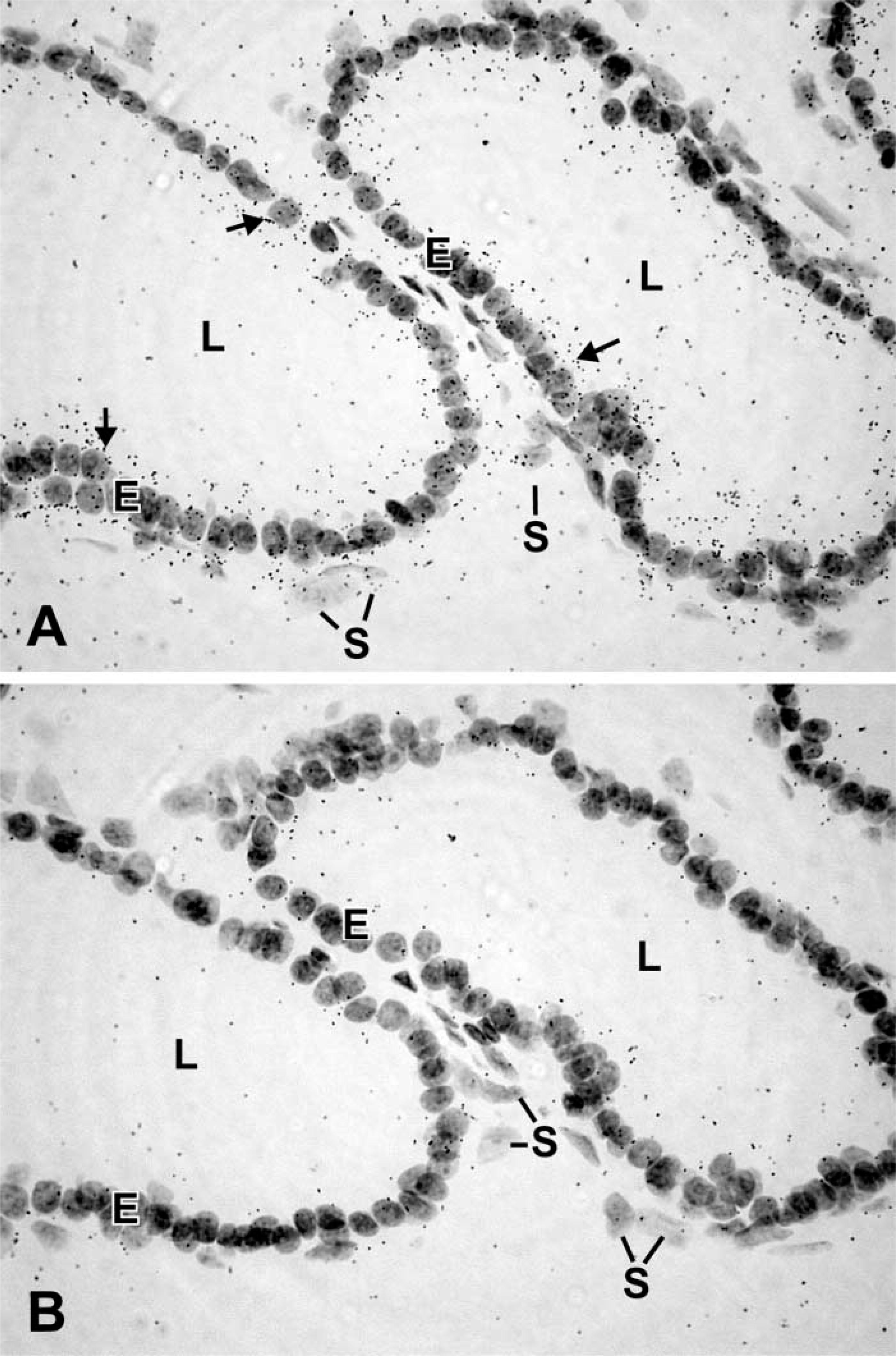
As positive controls, enzyme mRNAs were localized by ISH in tissues known to express the enzyme. For all the enzymes studied, positive data were obtained in at least one tissue excluding the prostate (Table 1). Specific labeling was detected in the prostate for the following enzymes involved in either formation or degradation of hormonal steroids: 17β-HSD types 1, 2, 3, 4, 7, 8, 9, 10, and 11; P450 7α-hydroxylase; 5α-reductase type 2; 5β-reductase type 1; estrogen sulfotransferase type 1; 11β-HSD types 1 and 2, and UDP-glucuronosyltransferase 1A6 (UGT1A6) (Table 1). The enzyme mRNAs have been found to be expressed in epithelial cells of the prostatic acini (Table 1; Figures 1–3). In several cases, stromal cells were also specifically labeled (Figures 1–3). Types 1, 7, and 10 17β-HSD, 11β-HSD types 1 and 2, as well as estrogen sulfotransferase type 1 were only detected in epithelial cells. Several enzymes could not be detected by ISH, even following long exposures (up to 45 days). As shown in Table 1, quantitative RT-PCR data show a wide variation in the expression of the different enzyme mRNAs. In general, it could be observed that when the amount of mRNA copies/μg total RNA was less than 2000, no hybridization could be detected. When the radiolabeled sense probes were used for hybridization in consecutive sections, only weak and diffuse labeling corresponding to background could be detected in all cases (Figures 1–3).
(
(
Discussion
The present results indicate that several enzymes involved in the biosynthesis and inactivation of hormonal steroids can be localized in the mouse prostate by ISH. All the enzymes that we have been able to detect were expressed in epithelial cells. Several enzymes were also found to be expressed in both epithelial and stromal cells. In mouse prostatic acini, the majority of epithelial cells are luminal cells and very few basal cells are observed. In fact, the ratio of luminal to basal cells is ~10:1 (El-Alfy et al. 2000). Although it was not possible to clearly identify basal cells in frozen sections, it can be assumed that most of the specific labeling detected by ISH over acini originates from the luminal cells. In human prostate acini, the basal cells are abundant and form a continuous layer resting on the basement membrane; the ratio of luminal to basal cells is 1:1 (El-Alfy et al. 2000). Human basal cells have been shown to express steroidogenic enzymes involved in the conversion of DHEA to DHT, namely 3β-HSD type 1, 17β-HSD type 5, and 5α-reductase type 2 (Eicheler et al. 1994; Pelletier et al. 1998; El-Alfy et al. 1999; Pelletier and El-Alfy 2000). It is noteworthy that none of the steroidogenic enzymes involved in the conversion of DHEA to testosterone that are expressed in the human prostate basal cells could be detected by ISH in the mouse prostate. This might be related to the absence of DHEA secretion by the adrenal cortex in rodents (Miller 1988). On the other hand, the presence of 5α-reductase type 2 mRNA in the mouse prostate indicates that there is a local conversion of the weak androgen testosterone into the most potent natural androgen, DHT.
One cannot conclude, however, that when a hybridization signal cannot be detected in the mouse prostatic tissue, the enzyme mRNA is not expressed. The results obtained by quantitative RT-PCR indicate that with a few exceptions, there is a good correlation between the number of mRNA copies/μg total RNA and the detection of ISH signal, which was absent when the number of copies was below 2000. Each probe that provided negative results in the mouse prostatic tissue was shown to give positive results in at least one other tissue, indicating the effectiveness of each probe in detecting specific mRNAs under our experimental conditions. Altogether, the present results indicate that the failure to obtain an ISH signal for some enzymes is likely due to low levels of expression of those enzyme mRNAs.
With the approach involving the use of frozen sections, we could not localize any enzyme in the walls of blood vessels. In previous studies performed in the paraffin-embedded human prostate, we could detect immunoreactive 17β-HSD type 5 in endothelial cells of capillaries, veins, and arteries (El-Alfy et al. 1999). Other studies involving the use of specific antibodies raised against mouse steroidogenic enzymes for immunocytochemical localization should provide additional information on the cell types expressing those enzymes in the mouse prostate.
From the present results, it clearly appears that the enzymes involved in the biosynthesis (17β-HSD types 1, 3, and 7; 5α-reductase type 2; and 11β-HSD type 1), as well as the inactivation (17β-HSD types 2, 4, 8, 9, 10, and 11; 7α-hydroxylase; estrogen sulfotransferase type 1; 11β-HSD type 2; and UGT1A6), of hormonal steroids are all expressed by the epithelial cells, and several of them are also detected in stromal cells.
Footnotes
Acknowledgements
Supported by Genome Canada and Genome Québec.
