Abstract
RORα is an orphan nuclear receptor. A deletion mutation in the RORα gene leads to severe cerebellar defects, known as the staggerer mutant mouse. Although previous in situ hybridization (ISH) studies have shown that RORα is highly expressed in the cerebellum, especially in Purkinje cells, and in the thalamus, sufficient immunohistochemical (IHC) study has not yet been presented. I demonstrate here the IHC analysis of RORα using a specific anti-RORα antibody, in adult and developing mouse nervous system. RORα immunoreactivity was observed in the Purkinje cell and molecular layers of the cerebellum. The co-localization of RORα with calbindin D28K (CaBP) and parvalbumin indicates that RORα-positive cells were Purkinje cells, stellate cells, and basket cells. In addition to the cerebellum, strong to medium RORα immunoreactivity was found in the thalamus, cerebral cortex (mainly in the layer IV), dorsal cochlear nucleus (DCN), suprachiasmatic nucleus (SCN), superior colliculus, spinal trigeminal nucleus, and retina. The immunostaining was restricted in nuclei of neurons. Developmentally, RORα immunoreactivity was observed in the cerebellum and thalamus from embryonal day 16 (E16). The distribution of RORα immunoreactivity and RORα mRNA hybridization signal was almost coincident. However, the intensity of hybridization signal was not always parallel to that of immunoreactivity.
Keywords
N
Homozygous staggerer mutant mice show severe cerebellar defects, including degeneration of granule cells and ectopic Purkinje cells reduced in number and size with poor dendritic arbors (Sidman et al. 1962). Evidence suggests that the effect of the staggerer gene mutation appears intrinsically in Purkinje cells and that the degeneration of granule cells is a secondary phenomenon (Herrup and Mullen 1979,1981; Herrup 1983). Staggerer mice carry a deletion mutation in the RORα gene that prevents translation of the ligand-binding homology domain (Hamilton et al. 1996; Matysiak-Scholze and Nehls 1997), and RORα-knockout mice show similar symptoms to staggerer mice (Dussault et al. 1998; Steinmayr et al. 1998).
Distribution of RORα mRNA expression is revealed by Northern blotting and ISH analyses in the mouse brain (Matsui et al. 1995). RORα is expressed in specific areas of the brain, including the cerebellum, thalamus, and olfactory bulb. By E14-E15, RORα is already highly expressed in Purkinje cells (Hamilton et al. 1996; Nakagawa et al. 1997). In contrast, RORβ is highly expressed in the retina, SCN, and pineal body (Schaeren-Wiemers et al. 1997). RORγ is highly expressed in the skeletal muscle and thymus but not in the nervous system (Hirose et al. 1994). RORα has multiple isoforms produced by alternative RNA processing at the amino-terminal region (Becker-André et al. 1993; Giguère et al. 1994; Matysiak-Scholze and Nehls 1997).
Although these data analyzing mRNA expression levels are impressive, they provide no information on the protein itself. It has remained unknown whether mRNA expression levels correlate with protein levels temporally and spatially. Subcellular localization of the protein is also unknown. I demonstrate here the IHC analysis of RORα in the adult and developing mouse brain, examining the validity of IHC data by comparison with ISH data.
Materials and Methods
Preparation of Fusion Proteins
Mouse RORα (nt 1142–1587), rat RORβ (nt 1381–1765), and mouse RORγ (nt 1213–1618) cDNA fragments containing the carboxy-terminal regions were prepared by reverse transcriptase-polymerase chain reaction (RT-PCR) according to the sequence data (GenBank accession numbers S82720, L14610, and U43508, respectively) and subcloned into pGEX-6P (Amersham Biosciences; Piscataway, NJ). RORα (aa 347–467), RORβ (aa 337–459), and RORγ (aa 393–516) glutathione S-transferase (GST) fusion proteins were prepared and affinity-purified with glutathione-Sepharose (Amersham) according to the manufacturer's protocol.
Preparation of Brain Extracts
Brain tissues collected from adult ddY albino mice were homogenized with a Teflon-glass homogenizer in one volume of ice-cold 2%SDS, 2% Triton X-100, 1 mM EDTA in 50 mM Tris-HCl, pH 7.6, or 1 mM EDTA in 5 M guanidine HCl.
Western Blotting Analysis
The brain extracts (50 μg) and aliquots of the GST fusion proteins were denatured, applied to 10% SDS-PAGE, and blotted onto polyvinylidene difluoride filters (Immobilon P; Millipore, Bedford, MA). The filters were incubated with the goat polyclonal anti-RORα1 antibody (C-16; Santa Cruz Biotechnology, Santa Cruz, CA; 1:1000) or mouse monoclonal anti-GST antibody (B14; Santa Cruz; 1:1000), followed by incubation with horseradish peroxidase-conjugated anti-goat IgG(H + L) or anti-mouse IgG(H + L) antibody (Vector Laboratories, Burlingame, CA; 1:2000). Immunoreactivity was visualized with the ECL Plus Western blotting detection reagents (Amersham).
Preparation of Tissue Sections
Adult, neonatal, and fetal ddY albino mice were used. Tissues were prepared as described previously (Ino 2003). Briefly, adult and neonatal mice were perfused with 4% paraformaldehyde in 0.1 M phosphate buffer, pH 7.5, via the heart under pentobarbital anesthesia. Brains sliced at approximately 5-mm thickness were further fixed in the same solution at 4C for 2 days. Fetal mice (E16) were rinsed with saline and cut into two or three pieces and fixed in the same solution at 4C for 2 days. The tissues were then transferred to distilled water and incubated at 4C overnight. The tissues were boiled in distilled water for 1.5–3 min for antigen retrieval and then immersed in 30% sucrose in PBS at 4C overnight. The antigen retrieval procedure was critical for successful immunostaining for RORα (Ino 2003). The tissues were frozen in crushed dry ice. Cryostat sections (10-μm) were prepared, placed on 0.02% poly-
Preparation of Probes
Mouse RORα (nt 617–1587), rat RORβ (nt 737–1765), and mouse RORγ (nt 647–1618) cDNAs were prepared by RT-PCR and subcloned into pGEM-T (Promega; Madison, WI) or pBluescript II (Stratagene; La Jolla, CA). Digoxigenin (DIG)-labeled antisense and sense riboprobes were prepared with DIG RNA labeling mix (Roche Diagnostics; Mannheim, Germany) and SP6, T3 or T7 RNA polymerase using linearized plasmids as templates. Riboprobes were hydrolyzed with alkaline (in 40 mM sodium bicarbonate, 60 mM sodium carbonate, and 5 mM dithiothreitol at 60C for 20 min) to an average size of 300 nucleotides.
In Situ Hybridization
ISH was performed as described previously with some alterations (Ino et al. 1994). Sections were immersed in 0.3% Triton X-100 in PBS at room temperature (RT) for 2 hr. The adult tissue sections were incubated with 1 μg/ml proteinase K in PBS at 37C for 10 min. After washing with PBS, the sections were immersed in 4% paraformaldehyde in 0.1 M phosphate buffer, pH 7.5, at RT for 10 min. After washing with PBS, the sections were immersed in 0.2 N HCl at RT for 10 min, immersed in 0.1 M triethanolamine HCl, pH 8.0, at RT for 5 min, and immersed in freshly prepared 0.25% acetic anhydrate in 0.1 M triethanolamine-HCl, pH 8.0, at RT for 10 min. After washing with PBS, the sections were incubated in 50% formamide in 2 × SSC at RT for 2 hr. The sections were hybridized with approximately 0.5 μg/ml DIG-labeled riboprobes in the hybridization solution (50% formamide, 0.2 mg/ml
Immunohistochemistry
IHC was performed as described previously (Ino 2003). Briefly, sections were immersed in 0.3% Triton X-100 in PBS at RT for 2–3 hr and blocked in 5% skim milk in PBS at RT for several hours. Immunoreaction was performed with primary antibodies in the blocking solution at RT overnight. Primary antibodies for IHC were goat polyclonal anti-RORα1 (C-16; 1:1000) and mouse monoclonals anti-CaBP (CB-955; Sigma, St. Louis, MO; 1:1000), anti-parvalbumin (PARV-19; Sigma; 1:1000), and anti-neuronal nuclei (NeuN; Chemicon, Temecula, CA; 1:1000). After washing with PBS, the sections were incubated with biotin-conjugated anti-goat IgG(H+L) antibody (Vector; 1:200), followed by reaction with the Vectastain ABC kit (Vector). The reaction was developed with 3,3'-diaminobenzidine, nickel sulfate, and hydrogen peroxide. For double staining after incubation with primary antibodies, the sections were incubated with biotin-conjugated anti-goat IgG(H+L) antibody (Vector; 1:200), followed by incubation with Alexa Fluor488-conjugated streptavidin (Molecular Probes, Eugene, OR; 1:400) and Texas Red-conjugated anti-mouse IgG(H+L) antibody (Vector; 1:200). The sections were observed by fluorescence microscopy.
Results
Western Blotting Analysis with the Anti-RORα Antibody
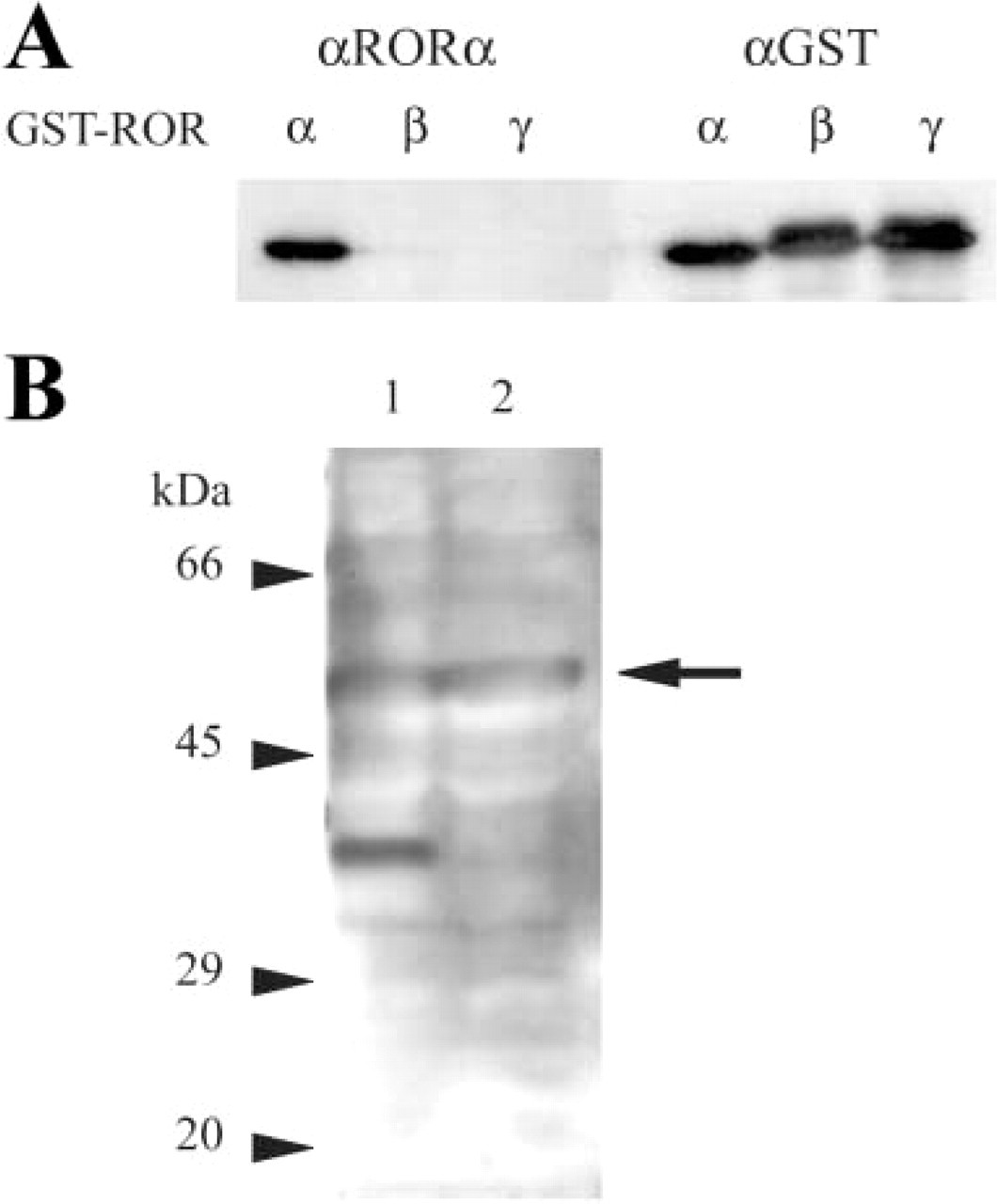
First I examined the specificity of the anti-RORα antibody (C-16) used in this study because the carboxy-terminal region of RORα, which is recognized by this antibody, is partly identical to those of RORβ and RORγ. Figure 1A shows that this antibody was specific for RORα and crossreacted neither for RORβ nor RORγ.
In brain proteins extracted with 2% SDS, 2% Triton X-100, 1 mM EDTA in 50 mM Tris HCl, pH 7.6, two bands (53–59 kD and 38–40 kD) were recognized by the anti-RORα antibody (Figure 1B, Lane 1). In contrast, in brain proteins extracted with 1 mM EDTA in 5 M guanidine-HCl, only the upper band was recognized (Figure 1B, Lane 2). The lower band in Lane 1 may correspond to degradation products. Because the upper band is broad, it may be composed of multiple protein species. The upper band shows a good correlation with the theoretical molecular weights of mouse RORα1 (58.8 kD) and RORα4 (53.4 kD).
Localization of RORα in the Adult Mouse Nervous System
I demonstrate the localization of RORα mRNA and protein by ISH and IHC with the specific antisense riboprobe and antibody. RORα mRNA has been al ready reported to be abundantly expressed in the cerebellum, especially in Purkinje cells, and thalamus (Matsui et al. 1995).
Specificity of the anti-RORα antibody (
In the cerebellum, strong hybridization signal was observed in Purkinje cells. Hybridization signal was also observed in the molecular layer, although the intensity was weaker than in Purkinje cells, but not in the granule cell layer (Figure 2B). IHC showed strong RORα immunoreactivity in Purkinje cells and in cells of the molecular layer (Figure 2A). In either case, immunostaining was located in nuclei. Although the mRNA level in Purkinje cells was much greater than that in cells of the molecular layer, a difference in RORα immunoreactivity between them was less apparent. Strong hybridization signal was also observed in the thalamus, and simultaneously strong nuclear RORα immunoreactivity was located in this region, except for the reticular thalamic nucleus (Figures 2C and 2D). In the cerebral cortex, RORα mRNA and protein were found mainly in layer IV (Figures 2E and F). They were also observed in the rostral part of the piriform cortex but not in the caudal part, and in the entrhinal cortex (Table 1). RORα-positive cells in the piriform cortex mainly existed in layer II (data not shown).
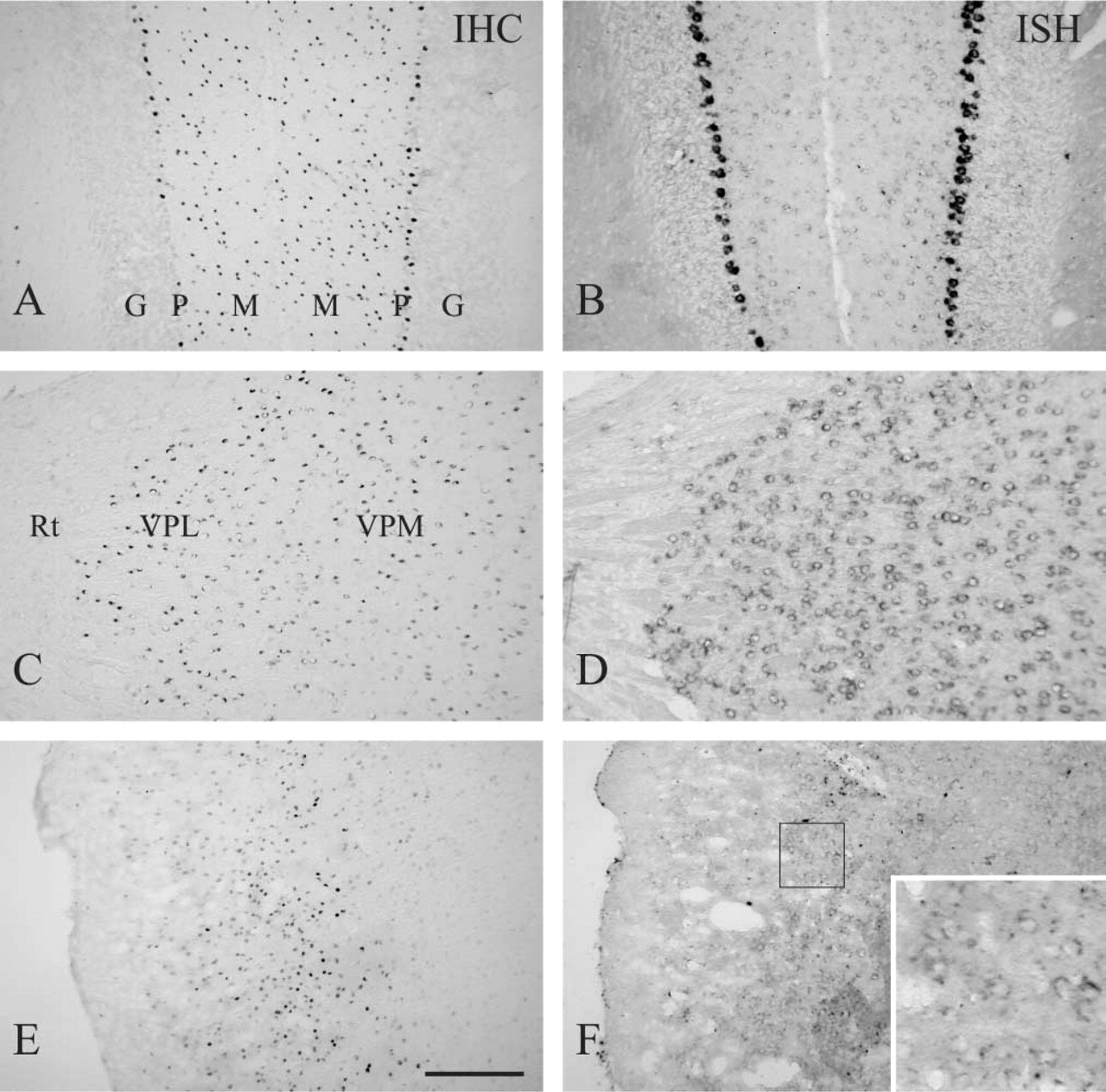
RORα in adult mouse cerebellum, thalamus, and cerebral cortex. Parasagittal sections of the cerebellum (
In addition to the cerebellum, thalamus and cerebral cortex, conspicuous RORα immunoreactivity was observed in the DCN (Figure 3A). Although RORα mRNA expression was also observed in this region, the level was lower than in Purkinje cells and in the thalamus and was approximately equivalent to that in the molecular layer of the cerebellum (Figure 3B). Cells showing strong RORα immunoreactivity were located in the superficial region of the DCN.
Distribution of RORα and RORβ in the mouse nervous system and pituitary gland a
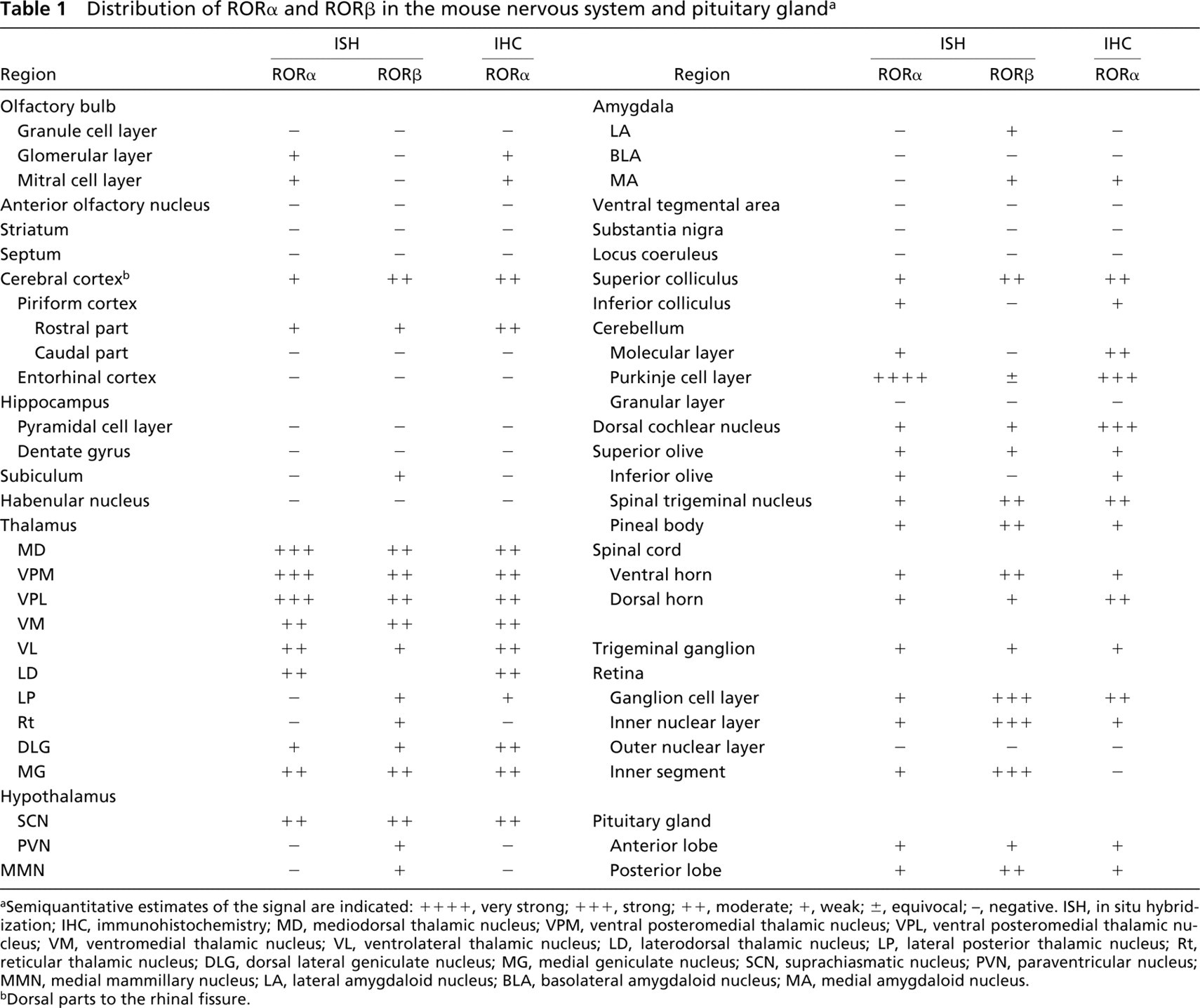
aSemiquantitative estimates of the signal are indicated: + + + +, very strong; + + +, strong; ++, moderate; +, weak; ±, equivocal; −, negative. ISH, in situ hybridization; IHC, immunohistochemistry; MD, mediodorsal thalamic nucleus; VPM, ventral posteromedial thalamic nucleus; VPL, ventral posteromedial thalamic nucleus; VM, ventromedial thalamic nucleus; VL, ventrolateral thalamic nucleus; LD, laterodorsal thalamic nucleus; LP, lateral posterior thalamic nucleus; Rt, reticular thalamic nucleus; DLG, dorsal lateral geniculate nucleus; MG, medial geniculate nucleus; SCN, suprachiasmatic nucleus; PVN, paraventricular nucleus; MMN, medial mammillary nucleus; LA, lateral amygdaloid nucleus; BLA, basolateral amygdaloid nucleus; MA, medial amygdaloid nucleus.
bDorsal parts to the rhinal fissure.
In addition to the above regions, moderate RORα immunoreactivity was observed in the suprachiasmatic nucleus (SCN), superior colliculus, spinal trigeminal nucleus, and retina (Figures 3C, 3E, and 3G; Table 1). In the retina, ganglion cells and cells of the inner nuclear layer showed positive immunoreactivity. In either case, moderate to low levels of RORα mRNA expression were observed (Figures 3D, 3F, 3H; Table 1). In any portion, RORα immunoreactivity was observed in nuclei.
The data are summarized in Table 1. By comparison, the distribution of RORβ mRNA expression is also shown. RORγ mRNA expression was not observed in the mouse nervous system and pituitary gland (data not shown). No hybridization signal was observed with the sense riboprobes (data not shown).
Classification of RORα-positive Cells
I performed the classification of cells showing RORα immunoreactivity in the cerebellum, DCN, and thalamus by double fluorescence immunostaining. Among cerebellar neurons, Purkinje cells are CaBP- and parvalbumin-double positive, and stellate cells and basket cells are parvalbumin-positive but CaBP-negative (Celio and Heinzmann 1981; Celio 1990). Therefore, CaBP and parvalbumin were used as cell markers for the classification of cell types in the cerebellum. Purkinje cells were clearly RORα-positive (Figures 4A–4C, arrows). In the molecular layer, all RORα-positive cells were parvalbumin-positive (Figures 4D–4F, arrowheads). Therefore, these RORα-positive cells were stellate cells or basket cells. In the granule cell layer, no RORα-immunoreactive cells were found, which indicates that granule cells as well as Golgi cells were RORα-negative.
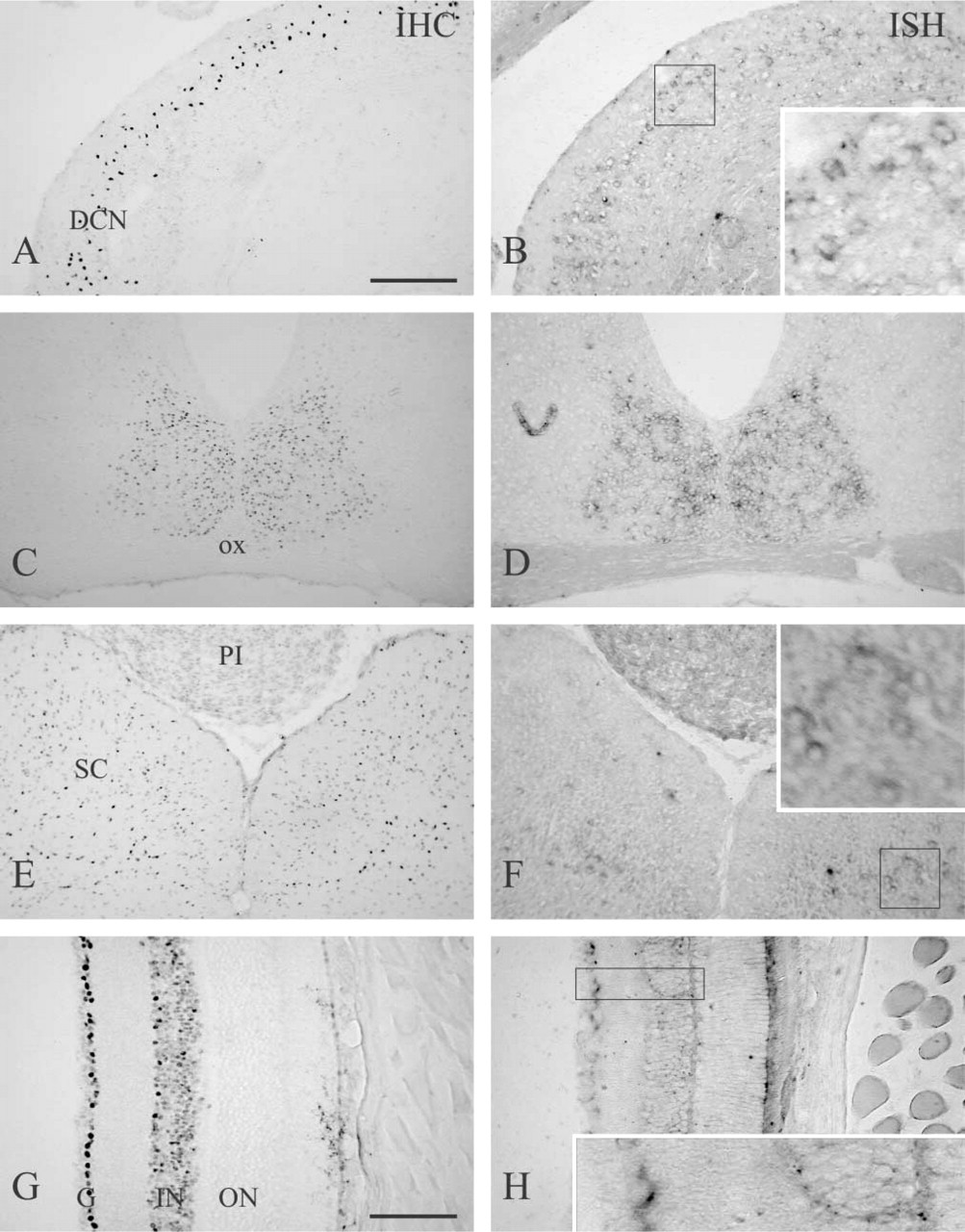
RORα in the adult mouse DCN, SCN, superior colliculus, and retina. Coronal sections of the DCN (
In the DCN, cells sporadically found in the superficial region showed intense CaBP immunoreactivity; these are ectopic Purkinje cells (Purkinje-like cells, Figure 4H) (Celio 1990; Hurd and Feldman 1994). These cells were RORα-positive (Figures 4G and 4I, arrows). Except for ectopic Purkinje cells, no CaBP immunoreactivity was found. In contrast, many cells showed moderate parvalbumin immunoreactivity, and these parvalbumin-positive cells were RORα-positive (Figures 4J–4L).
The classification of RORα-positive cells in the thalamus using NeuN as a specific neuronal marker was performed. Figures 5A–5C clearly show that RORα immunoreactivity was found only in NeuN-positive neurons, but not all NeuN-positive neurons were RORα-positive. NeuN-positive and parvalbumin-positive neurons in the reticular thalamic nucleus were RORα-negative (Figures 5D–5F), which indicates that parvalbumin is independent of RORα expression.
Localization of RORα in the Developing Mouse Nervous System
I examined the localization of RORα mRNA and immunoreactivity in the developing mouse nervous system using mice from E16 to postnatal day 21 (P21). As early as E16, both strong hybridization signal and immunoreactivity were found in the cerebellum and thalamus (Figures 6A–6D). At E16 and P0, RORα-positive immature Purkinje cells with small- to medium-sized nuclei were not yet arranged in a single layer (Figures 6A and 6E). At P7, RORα immunoreactivity was observed in Purkinje cells having large nuclei and forming the monolayer architecture (Figure 6F). In addition to Purkinje cells, RORα-positive cells with small nuclei were observed in the molecular layer in the vicinity of the Purkinje cell layer (Figure 6F). In rats, the generation of basket cells peaks around P7 and the generation of stellate cells is delayed several days (Altman 1972). Given that the timetable of the cerebellar neurogenesis in rats is similar to that in mice, the majority of these RORα-positive cells in the molecular layer are basket cells. At E16, P0, or P7, no RORα immunoreactivity was observed in the external germinal layer. At P14 and P21, RORα-positive cells in the molecular layer increased in number and spread over the entire area of the molecular layer (Figures 6G and 6H). The distribution of RORα-positive cells was almost identical to that of adult mice.
Using parvalbumin as a cell marker, a detailed examination of the development of RORα-positive cells in the cerebellum was performed (Figure 7). At E16 and P0, no parvalbumin was detected in the cerebellum (data not shown). At P7, parvalbumin was detected only in Purkinje cells, and therefore RORα-positive cells in the molecular layer were parvalbumin-negative (Figures 7A–7C, arrowheads). At P14, several but not all RORα-positive cells in the molecular layer exhibited parvalbumin immunoreactivity (Figures 7D–7F, arrows), and at P21 the number of RORα- and parvalbumin-double positive cells in the molecular layer further increased (Figures 7G-7I, arrows). As shown in Figures 4D-4F, almost all RORα and parvalbumin immunoreactivity in the molecular layer was overlapped in the adult. The extent of the distribution of RORα-positive cells in the molecular layer appeared to be restricted within the arborization of Purkinje dendrites, and this extent spread in accordance with the maturation of Purkinje cells during development.
Discussion
Distribution of RORα Immunoreactivity
I demonstrated here the successful IHC staining of RORα, whose distribution was closely coincident to that of RORα mRNA expression. For the IHC detection of RORα, antigen retrieval was critical. I used the antigen retrieval method in which tissues were boiled en bloc in distilled water before preparation of frozen sections (Ino 2003). RORα immunoreactivity was located in cell nuclei and was mainly distributed in the cerebellum (especially in Purkinje cells, stellate cells, and basket cells), thalamus, and DCN. Although the distributions of RORα mRNA and protein closely coincided with each other, their levels were not parallel. Both strong hybridization signal for RORα mRNA and strong immunoreactivity of RORα protein were seen in Purkinje cells, whereas strong immunoreactivity of the protein but weak hybridization signal of the mRNA were observed in stellate cells and basket cells of the cerebellum and in cells of the DCN. Although the hybridization signal of the mRNA in the DCN was weaker than that in the thalamus, the immunoreactivity of the protein was conversely stronger. It is unknown why there was an imbalance between mRNA and protein levels. Translation levels may be different from cell to cell.
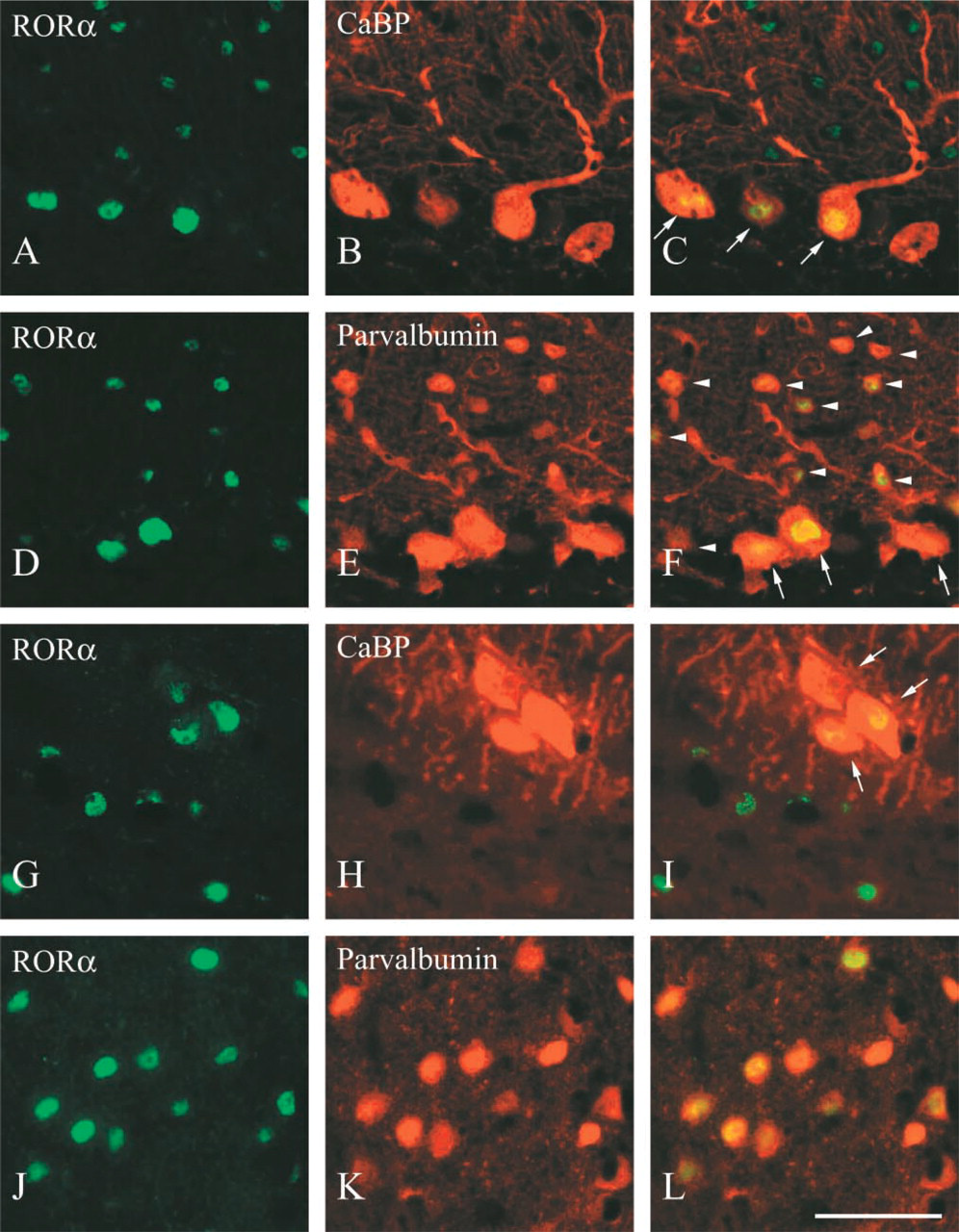
Co-localization of RORα with CaBP and parvalbumin in adult mouse cerebellum and DCN. Parasagittal sections of the cerebellum (
I further demonstrated the existence of RORα protein in the retina (especially in ganglion cells and in the inner molecular layer) and SCN. An experiment using RORα-knockout mice, in which truncated RORα fused with β-galactosidase lacking normal RORα function is generated instead of RORα, showed that β-galactosidase activity is detected not only in the Purkinje cells and thalamus but also in the retinal ganglion cell and SCN (Steinmayr et al. 1998). The present data are consistent with their study in these points. In that study, however, β-galactosidase activity in the molecular layer of the cerebellum was not shown. RORα mRNA expression is observed in the molecular layer of normal developing mice from P14, when stellate cells and basket cells have already been generated (Nakagawa et al. 1997). Because RORα mRNA expression in the molecular layer is low compared to Purkinje cells, β-galactosidase activity may have not been detected in the RORα-knockout mice. I believe that RORα protein exists in stellate cells and basket cells of normal mice. In addition, Steinmayr et al. (1998) demonstrated β-galactosidase activity in the testis, epididymis, and skin (epidermis, hair follicle, and sebaceous gland). I observed RORα immunoreactivity in the epidermis, hair follicle, and sebaceous gland, but not in the testis and epididymis (data not shown). Although the reason for the discrepancy is unknown, further pursuit of this problem was not performed in the present study.
There are at least four RORα isoforms in humans (Becker-André et al. 1993; Giguère et al. 1994) and at least two in mice (Matysiak-Scholze and Nehls 1997). These isoforms have distinct amino-termini, produced by differential promoter usage and alternative slicing. The present ISH data did not distinguish these isoforms because I used the common region for the probe, and the IHC data also did not distinguish the isoforms because the antibody recognizes the common carboxy-terminus.
Cells Showing RORα Immunoreactivity
In the cerebellum, the RORα immunoreactivity was observed in Purkinje cells, stellate cells, and basket cells, which was clearly demonstrated by the coexistence with CaBP and parvalbumin. Granule cells and Golgi cells were RORα-negative. In the DCN, RORα exists in ectopic Purkinje cells. From the size of RORα-positive nuclei and their distribution, most RORα-positive cells in the DCN may be cartwheel cells (Wouterlood and Mugnaini 1984), and stellate cells may be also RORα-positive (Wouterlood et al. 1984). However, further reliable classification of the cell type in the DCN was difficult because no useful cell marker to distinguish them was available and the cytoarchitecture is not so clear as in the cerebellum. Cartwheel cells share some features common to Purkinje cells, such as cerebellin immunoreactivity (Mugnaini and Morgan 1987).
Studies on Staggerer Mutant Mice and Possible Physiological Roles of RORα
Analyses of staggerer mutant mice have provided suggestions about the physiological role of RORα. Studies using chimera mice suggest that the defects intrinsically exist in Purkinje cells but not in granule cells (Herrup and Mullen 1979,1981; Herrup 1983; Soha and Herrup 1995), which is supported by the distribution of RORα mRNA and protein.
Purkinje cells of staggerer mice are reduced in size and number, ectopic in location, and rudimentary in dendritic arborization. However, the neurogenesis of Purkinje cells is not affected in staggerer mice and the reduced number is due to cell death after differentiation (Vogel et al. 2000). Purkinje cells of staggerer mice show characteristic features of embryonal Purkinje cells, such as embryonal cell-surface carbohydrate patterns (Hatten and Messer 1978; Trenkner 1979; Edelman and Chuong 1982), multiple innervation by climbing fibers (Crepel et al. 1980; Mariani and Changeux 1980), and NMDA responses (Dupont et al. 1984). It is likely that RORα is essential for the maturation of Purkinje cells during development. Although the role of RORα after maturation remains unknown, RORα may be necessary for survival of mature Purkinje cells, because heterozygous staggerer mice, despite a lack of overt clinical phenotype, show progressive Purkinje cell degeneration with age (Zanjani et al. 1991; Doulazmi et al., 1999; Hadj-Sahraoui et al. 2001).
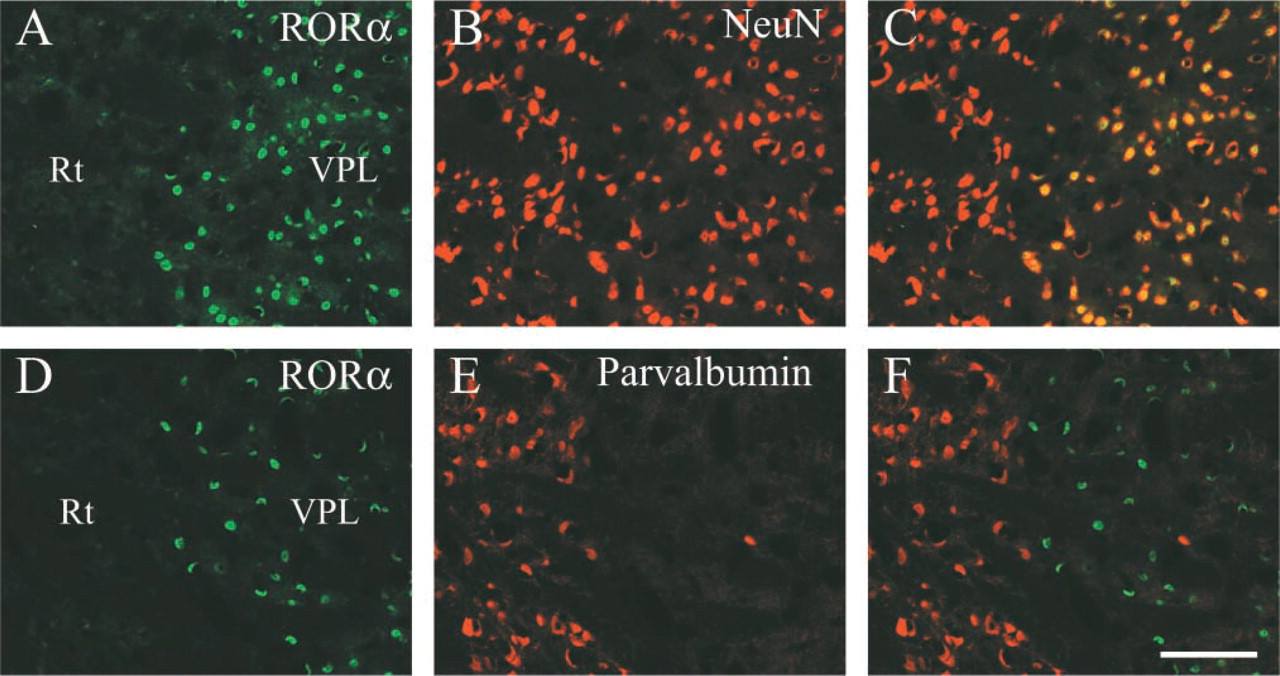
Co-localization of RORα with NeuN in the adult mouse thalamus. Coronal sections of the thalamus around the reticular thalamic nucleus (Rt)-ventral posterolateral thalamic nucleus (VPL) boundary. Localization of RORα (
In addition to Purkinje cells, defects in staggerer mice have been reported in the DCN (Berrebi et al. 1990), inferior olive (Shojaeian et al. 1985), and olfactory bulb (Monnier et al. 1999). Cartwheel cells are eliminated in the staggerer DCN. Reduction in size of glomerular and external and internal plexiform layers and reduction in number of mitral cells are observed in the staggerer olfactory bulb. Decreased cell numbers are found in the staggerer inferior olive. These abnormalities in the central nervous system are likely to be directly influenced by the absence of RORα in these portions, because both RORα mRNA and protein are detected there. Alternatively, the cell loss may be due to an indirect effect, as in the case of granule cells of the cerebellum.
In addition to the central nervous system, abnormalities have been also observed in the staggerer immune system, including the thymus and spleen (Trenkner and Hoffmann 1986). In contrast to the central nervous system, I found neither RORα mRNA nor protein in the thymus and spleen of adult and young mice (data not shown). There is a possibility that a distinct genetic locus, a small thymus located between the staggerer and shorter-ear loci on chromosome 9 (Heinlein and Wolle 1992), accounts for the immuno-dysfunction of staggerer mice. However, this does not exclude another possibility, that RORα is involved in normal development of the immune system. It would be interesting to examine whether there are any abnormalities in the RORα-knockout mouse immune system.
Although staggerer mutant mice display severe defects in Purkinje cells, no apparent change was observed in the thalamus, SCN, and retina, where high to moderate levels of RORα mRNA and protein are observed. This fact has been attracting attention because a high level of RORα mRNA expression in the thalamus has been previously known. One possible explanation is that RORβ, highly expressed in the thalamus, SCN, and retina, compensates the role of RORα and conceals the staggerer phenotype in these portions. This is plausible; however, evidence has not been presented. In addition, I showed RORα immunoreactivity in the stellate cells and basket cells of the cerebellum with no detectable RORβ mRNA expression. Because stellate cells and basket cells of staggerer mice appear to be unaffected (Sotelo and Changeux 1974; Landis and Sidman 1978), the above explanation is not sufficient. Possibly the absence of RORα appears as phenotype only in restricted cell types. At least, RORα and RORβ are not always necessary for survival or function of neurons, because many neurons, such as hippocampal neurons, lack both RORα and RORβ (RORγ is not expressed in the nervous system).
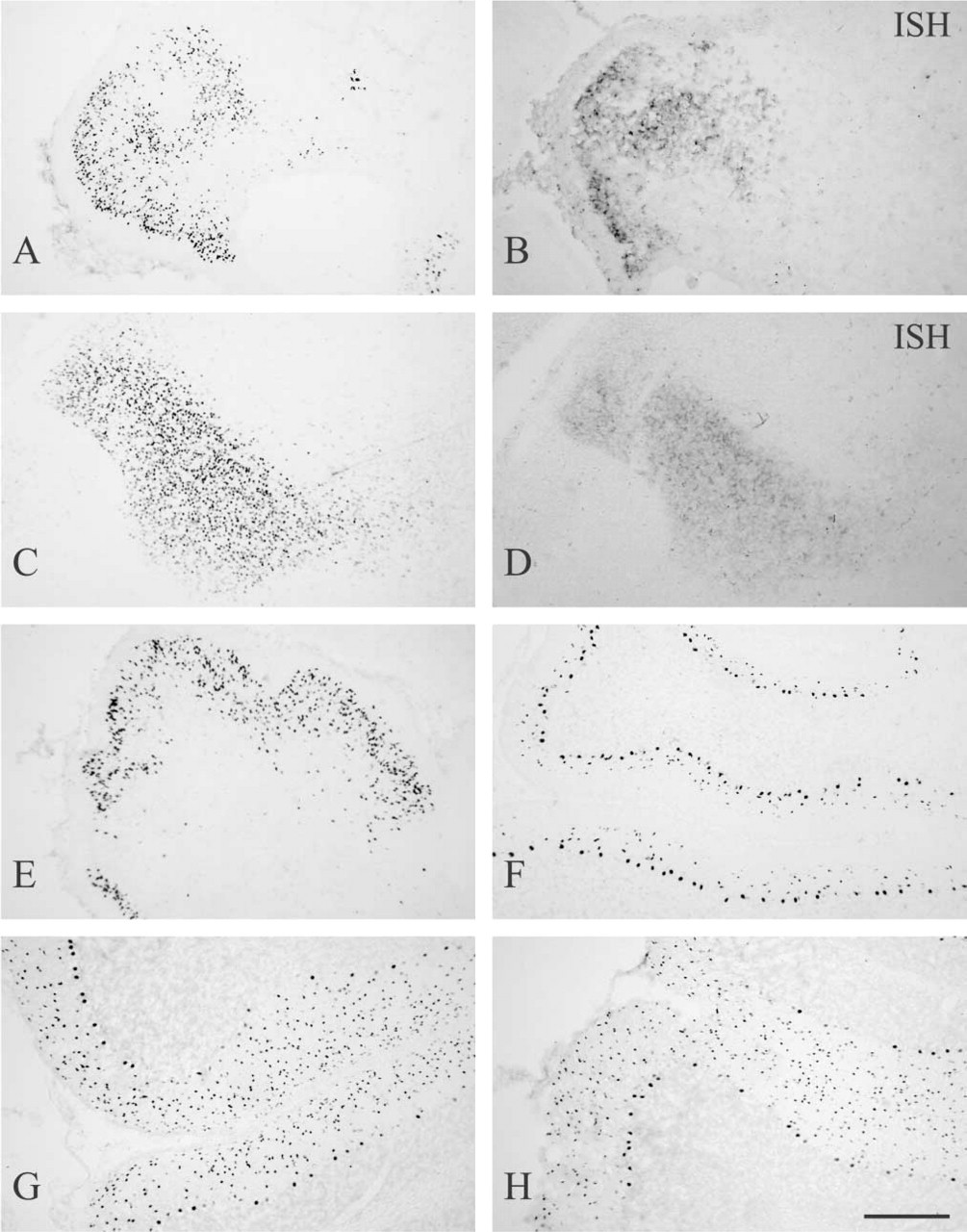
RORα in developing mouse brain. Parasagittal sections of cerebellum (
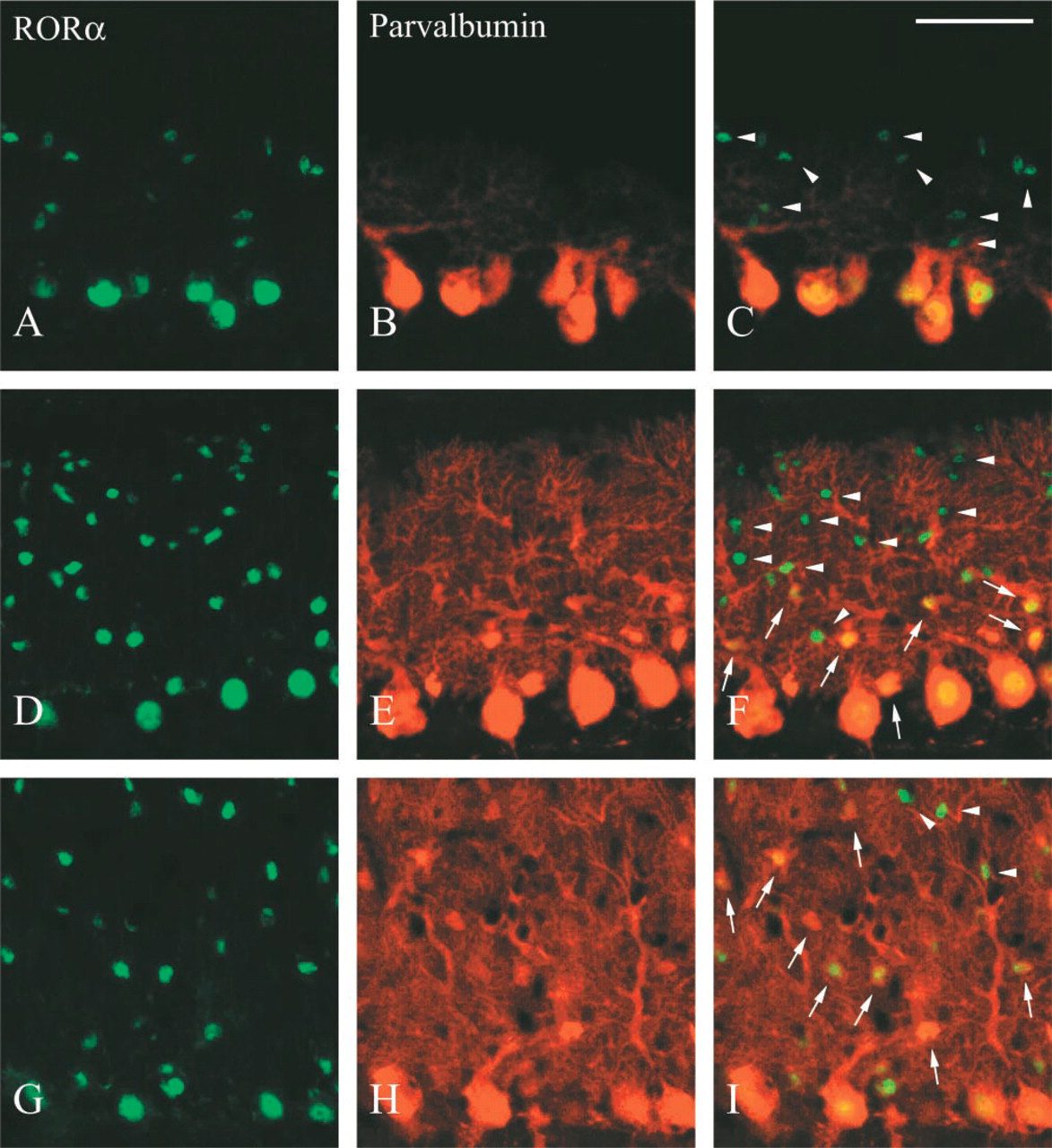
Co-localization of RORα with parvalbumin in developing mouse cerebellum. Parasaggital sections of the cerebellum from P7 (
