Abstract
Heme-binding protein 23 (HBP23), also termed peroxiredoxin (Prx) I, and heme oxygenase-1 (HO-1) are distinct antioxidant stress proteins that are co-ordinately induced by oxidative stress. HBP23/Prx I has thioredoxin-dependent peroxidase activity with high binding affinity for the pro-oxidant heme, while HO-1 is the inducible isoform of the rate-limiting enzyme of heme degradation. We investigated the cellular and subcellular localization of both proteins in rat liver. Whereas by immunohistochemistry (IHC) a uniformly high level of HBP23/Prx I expression was observed in liver parenchymal and different sinusoidal cells, HO-1 expression was restricted to Kupffer cells. By immunoelectron microscopy using the protein A–gold technique, HBP23/Prx I immunoreactivity was detected in cytoplasm, nuclear matrix, mitochondria, and peroxisomes of parenchymal and non-parenchymal liver cell populations. In contrast, the secretory pathway, i.e., the endoplasmic reticulum and Golgi complex, was free of label. As determined by immunocytochemical (ICC) studies in liver cell cultures and by Western and Northern blotting analysis, HBP23/Prx I was highly expressed in cultures of isolated hepatocytes and Kupffer cells. In contrast, HO-1 was constitutively expressed only in Kupffer cell cultures but was also inducible in hepatocytes. These data suggest that HBP23/Prx I and HO-1 may have complementary antioxidant functions in different cell populations in rat liver.
Keywords
H
In normal rat liver, HO-1 is mainly expressed in Kupffer cells (KC) (Goda et al. 1998; Immenschuh et al. 1999b). By contrast, although HBP23/Prx I represents a substantial portion of total hepatic protein (ca. 0.1%), its exact cellular distribution in liver is not known. Moreover, although by cell fractionation HBP23/Prx I has been detected in nucleus and cytoplasm (Kang et al. 1998; Butterfield et al. 1999), its precise subcellular distribution in liver has not been reported. Distribution of antioxidant stress proteins in various cells of an organ and in distinct subcellular compartments is important for the modulation of oxidative damage in diseases such as cancer (Oberley 2002). The major goal of the present study was to investigate the cellular localization and expression of HBP23/Prx I in comparison to that of HO-1 in rat liver. In addition, the subcellular localization of HBP23/Prx I in normal rat liver was analyzed by the protein A–gold technique.
Materials and Methods
Materials
The embedding media for light and electron microscopic immunohistochemical studies, Paraplast plus and LR White, respectively, were obtained from Sherwood Medical (St Louis, MO) and Polysciences (Heidelberg, Germany). Glutaraldehyde and paraformaldehyde were from Sigma (Munich, Germany). Media M199 and RPMI 1640 were obtained from Gibco BRL (Karlsruhe, Germany), and the chemiluminescent detection system for Western blotting, radioisotopes and nitrocellulose filters were from Amersham–Buchler (Braunschweig, Germany). Lab-Tek chamber slides were from Nunc (Naperville, IL), the nucleotide removal kit from Qiagen (Düsseldorf, Germany), and Falcon tissue culture dishes from Becton–Dickinson (Heidelberg, Germany). Protein A was from Pharmacia (Uppsala, Sweden). All other chemicals were purchased from Sigma or Roche Diagnostics.
Antibodies
The monospecificity of the polyclonal rabbit antibody against purified HBP23/Prx I was described previously (Iwahara et al. 1995) and is also shown in Figure 1. The commercial polyclonal rabbit anti-rat HO-1 antibody was obtained from Stress Gene (Victoria, BC, Canada) (Immenschuh et al. 1999b). The monoclonal antibody (MAb) ED2 was purchased from Serotec (Wiesbaden, Germany) and was used as a marker for monocyte–macrophages (Dijkstra et al. 1985). The antibodies for catalase (Beier et al. 1988) and lactate dehydrogenase (Baumgart et al. 1996) have previously been described and were used for positive controls in this study.
Animals
Male Wistar rats (2 months old, body weight 200 g) were used throughout the study. All experiments were approved by the local animal experiment review committee.
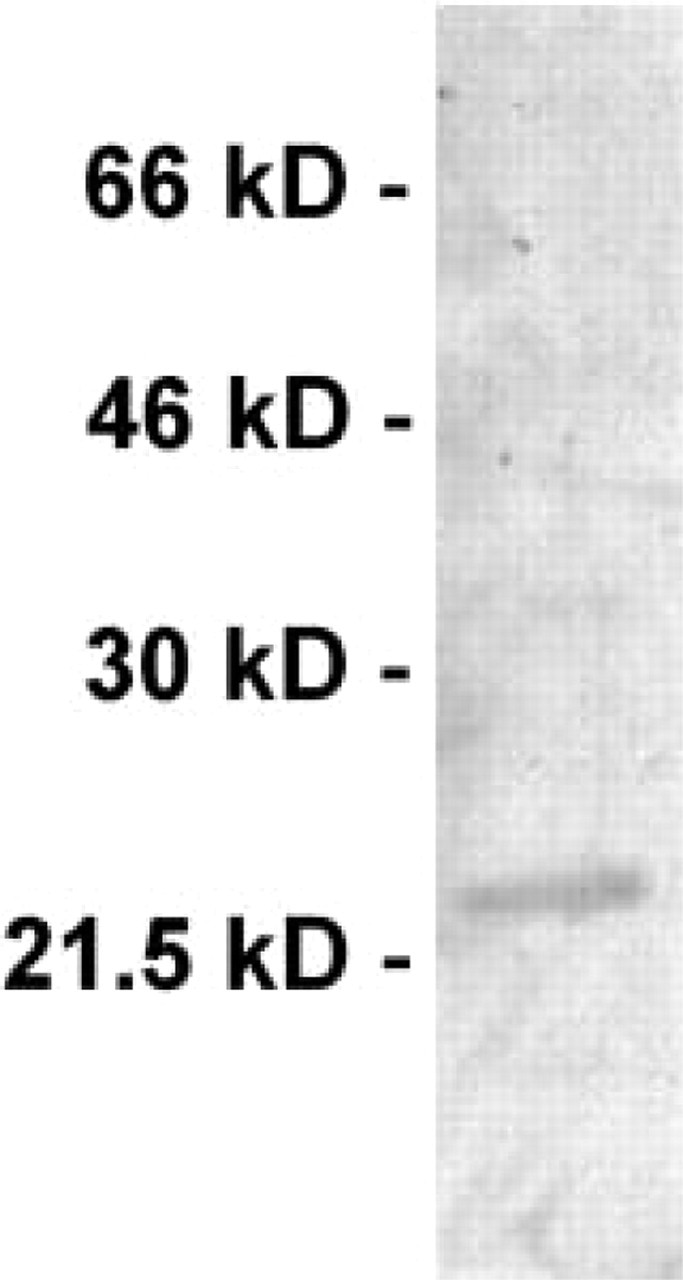
Expression of HBP23/Prx I protein in rat liver. Cytosolic fractions of normal rat liver (50 μg) were subjected to Western blotting analysis and probed with the antibody to HBP23/Prx I used in this study. Molecular mass markers are shown at left. Note the single band at approximately 23 kD.
Cell Isolation and Culture
Primary rat hepatocytes were isolated and cultivated as described (Immenschuh et al. 1995). They were cultured under 95% air/5% CO2 in medium M199 with Earle's salts containing 2 g/liter bovine serum albumin (BSA), 20 mM NaHCO3, 10 mM Hepes, 100 U penicillin/ml, 100 μg streptomycin/ml, and 1 nM insulin. In addition, 5% fetal calf serum (FCS) and 10 nM dexamethasone were present during the plating phase up to 4 hr, and cell cultures were incubated in serum-free medium for another 18 hr before treatment. KCs and sinusoidal endothelial cells (SECs) were isolated as previously described (Immenschuh et al. 1999a; Neubauer et al. 2000). Cells were resuspended in culture medium M199 containing 15% FCS, 100 U penicillin/ml, and 100 μg streptomycin/ml. Cell viability was assessed by trypan blue staining. Purity of cell isolations was determined by phase-contrast microscopy, DIL-AC-LDL incorporation of SEC, and ED2 staining of KCs as previously described (Neubauer et al. 1995). KC cultures were more than 99% pure and SECs were contaminated with 2% hepatic stellate cells and with about 10% KCs. Cells were plated on six-well plates (3 × 106 cells/well). After 2 hr, cells were washed for elimination of non-adherent cells. KC cultures were maintained in medium M199 (15% FCS, 100 U penicillin/ml, 100 μg streptomycin/ml) in an atmosphere of 95% air/5% CO2 at 100% humidity. Freshly islolated cells were used for RNA isolation within 1 hr after cell preparation.
Immunohistochemistry and Immunoelectron Microscopy
Rat livers were fixed by perfusion through the portal vein as described (Fahimi 1967). The fixative contained 4% depolymerized paraformaldehyde in PBS, pH 7.4, for light microscopic studies including immunohistochemistry (IHC). For electron microscopy (EM), it consisted of a mixture of 4% depolymerized paraformaldehyde and 0.05% glutaraldehyde in PBS, pH 7.4. After perfusion, livers were cut into small blocks that were kept in the same fixatives for several hours before further processing. Whereas for IHC the tissue blocks were embedded directly in paraffin (Paraplast Plus), for EM studies microslicer sections (c. 100 μm) were embedded in LR White. For IHC studies, 1–3-μm paraffin sections were subjected first to antigen retrieval by digestion with trypsin combined with microwave irradiation as described recently (Grabenbauer et al. 2001), followed by incubation with the specific antibodies to HBP23/Prx I, HO-1, or ED2. The antigen binding sites were detected by a peroxidase-conjugated biotin-straptividin system (Extravidin; Sigma) and visualized by the peroxidase substrate Nova Red (Vector Laboratories). The nuclei were counterstained with hematoxylin. For immunoelectron microscopic studies, ultrathin sections were incubated with the antibody to HBP23/Prx I and the antigen-antibody complexes were visualized with the protein A-gold procedure (Roth 1982), as detailed elsewhere (Baumgart 1994; Fahimi et al. 1996).
Immunocytochemistry on Isolated and Cultured Cells
For ICC studies on isolated cells, cultured cells on Lab-Tek chamber slides were fixed with methanol and acetone. Slides were stored at −20C before processing for immunolocalization. Indirect immunostaining was performed with the immunoperoxidase method with diaminobenzidine (DAB) as substrate (Immenschuh et al. 1999b). In brief, cells were washed with PBS containing 0.1% BSA and were covered with FCS for 30 min. Cells were incubated with the first antibody directed either against HBP23/Prx I, HO-1, or ED2 for 45 min at 37C. After washing, cells were incubated with peroxidase-labeled IgG against murine IgG or rabbit IgG. Cells were washed and incubated with PBS containing DAB (0.5 mg/ml) and H2O2 (0.01%) for 10 min, washed, and counterstained with the nuclear stain hemalaum.
RNA Isolation, Northern Blotting Analysis, and Hybridization
Total RNA for Northern blotting was isolated as previously described (Immenschuh et al. 1999a). Equal quantities of RNA were separated on 1.2% agarose/2.2 M formaldehyde gels. After electrophoresis, RNA was blotted onto nitrocellulose membranes and baked at 80C for 4 hr. After prehybridization for 3–4 hr at 42C, blots were hybridized overnight at 42C with a radiolabeled 883-bp EcoR I-Hind III fragment of pRHO1, which is a plasmid with the full-length rat HO-1 cDNA probe (Shibahara et al. 1985). The cDNA fragment was radioactively labeled with α[32P]-dCTP by random priming using the multiprime DNA labeling kit according to the manufacturer's instructions. To correct for differences in RNA loading of Northern blots, the nitrocellulose filters were stripped and were rehybridized with a 28S rRNA oligonucleotide. The 28S rRNA oligonucleotide was labeled with γ[32P]-dATP at the 3′-end with terminal deoxynucleotide transferase. The hybridization solution contained 6 × SSC, 5 × Denhardt's solution (0.2% Ficoll 400, 0.2% polyvinylpyrrolidone, and 0.2% BSA), 0.5% SDS, 50% formamide, and 100 μg/ml denatured salmon sperm DNA. Blots were washed with 2 × SSC/0.1% SDS (once) and 0.1 × SSC/0.1% SDS (twice) at 65C. Filters were exposed to X-ray films and quantification of the autoradiograms was performed by densitometry.
Western Blotting Analysis
After washing of cell cultures twice with 0.9% NaCl, cytosol was prepared essentially as described (Immenschuh et al. 1999a). After addition of 1 ml lysis buffer (0.1% SDS, 10 mM Tris, pH 7.4), cells were boiled for 5 min and homogenized by passing through a 25G needle. The homogenate was centrifuged for 5 min at 4C and the protein content was determined in the supernatant using the Bradford method. A total of 50 μg of total protein was loaded onto a 12% sodium dodecyl sulfate (SDS)-polyacrylamide gel and was blotted onto nitrocellulose membranes by electrophoresis. Membranes were blocked with Tris-buffered saline containing 1% BSA, 10 mM Tris-HCl (pH 7.5), and 0.1% Tween-20 for 1 hr at RT. The primary antibodies for HBP23/Prx I or HO-1 were added at 1:1000 dilution and the blot was incubated for 12 hr at 4C. The secondary anti-rabbit IgG was diluted 1:8000 and a chemiluminescent detection system for Western blotting was used according to the manufacturer's instructions (Amersham-Buchler; Poole, UK). Filters were exposed to X-ray films and quantification of the autoradiograms was performed by densitometry.
Results
IHC Localization of HBP23/Prx I and HO-1 in Rat Liver
High levels of HBP23/Prx I gene expression have previously been demonstrated in liver as compared to kidney, brain or testis (Iwahara et al. 1995). In agreement with those findings, livers probed with a monospecific polyclonal antibody directed against rat HBP23/Prx I (Figure 1) exhibited uniformly strong staining in liver parenchymal cells and in sinusoidal lining cells (SLCs) as well as in bile duct epithelial cells (Figures 2A and 2B). In contrast, staining for HO-1 protein expression was restricted to KCs (Figure 2C). The identity of KCs was confirmed by positive staining with MAb ED2, which is a marker for mononuclear phagocytes (Dijkstra et al. 1985) (data not shown). The control sections processed in the complete incubation medium but without the specific antibodies were consistently negative (Figure 2D).
Immunoelectron Microscopy
The intracellular localization of HBP23/Prx I was also investigated by immunoelectron microscopy using the protein A-gold technique. Gold particles representing antigenic sites for HBP23/Prx I were found in both the nucleus and the cytoplasm of liver parenchymal cells with prominent labeling also of the mitochondrial and peroxisomal matrix (Figures 3A and 3B). In the nucleus, the gold particles were confined to the euchromatin regions and the nucleolus, while the condensed chromatin remained unlabeled (Figure 3A). Moreover, in the cytoplasm the endomembrane system with endoplasmic reticulum and Golgi complex was negative (Figure 3C). In addition to parenchymal cells, gold labeling was also observed in all sinusoidal cells, including endothelial cells (Figure 4A), KCs (Figure 4B), hepatic stellate cells (Figure 4C), and bile duct epithelial cells (Figure 4D). The subcellular distribution of HBP23/Prx I in non-parenchymal cells was essentially similar to that in hepatocytes, with prominent labeling of nucleus and cytoplasm. The identification of these cells was based on their ultrastructural characteristics and their location in the sinusoids (Fahimi 1982) with KCs present mostly in the lumen of sinusoids (Figure 4B), the EC forming the fenestrated lining (Figure 4A), and the HSC with the typical lipid vacuoles in the perisinusoidal space (Figure 4C). Control sections incubated with nonspecific IgG (negative controls) were essentially unlabeled except for very rare gold particles distributed randomly (Figure 3D). The positive controls incubated with antibodies to catalase or lactate dehydrogenase showed specific labeling of peroxisomes (Figure 3E) or the cytoplasm (Figure 3F) as described previously (Beier et al. 1988; Baumgart et al. 1996).
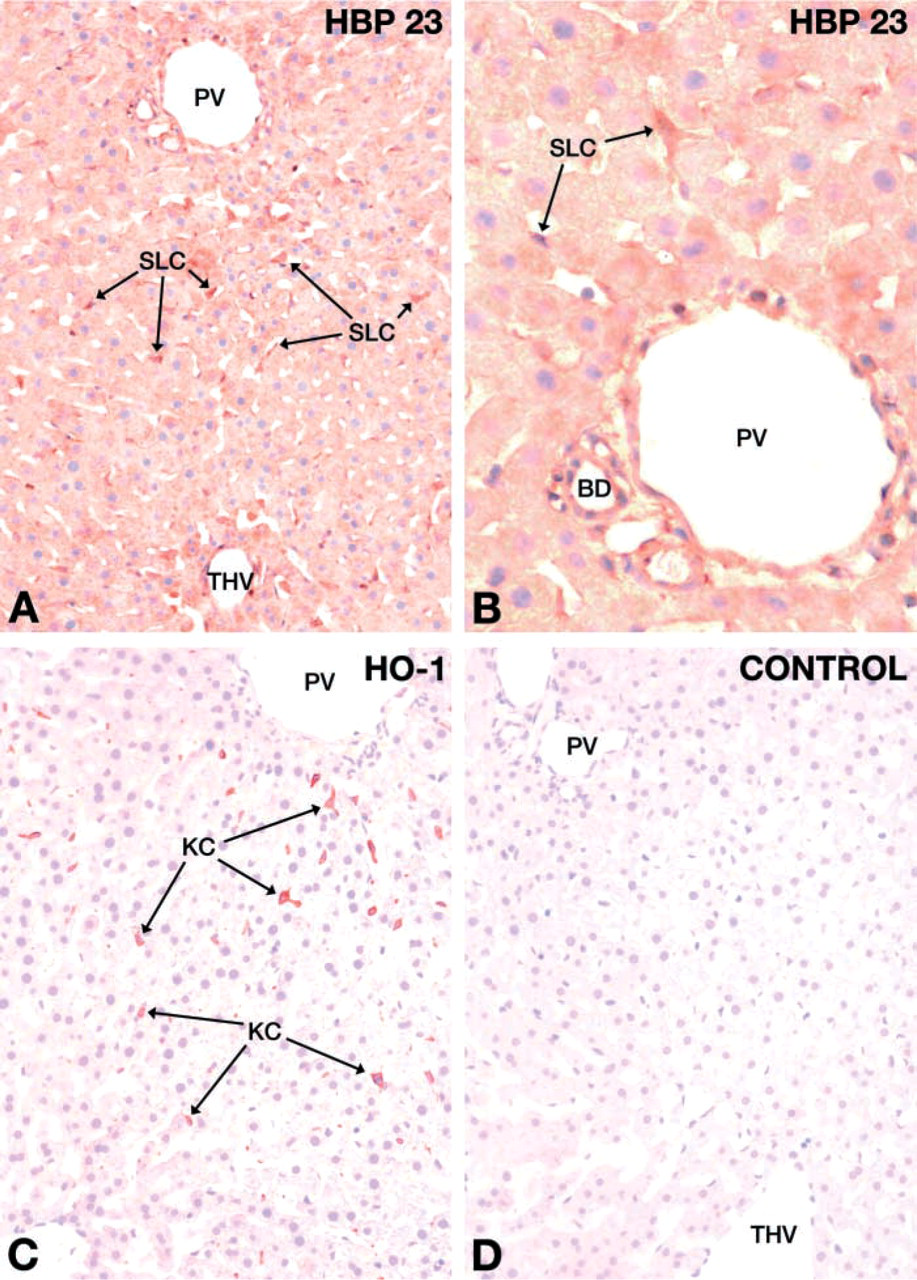
IHC localization of HBP23/Prx I and HO-1 in rat liver. Rat liver sections were incubated with polyclonal rabbit antibodies against rat HBP23/Prx I (
Gene Expression of HBP23/Prx I and HO-1 in Cultures of Isolated Parenchymal and Non-parenchymal Liver Cells
To further substantiate the cellular expression of HBP23/Prx I and HO-1 in different liver cell populations, the expression of HBP23/Prx I and HO-1 was determined by ICC in cell cultures of hepatocytes and KCs. Whereas in 24-hr cultures of parenchymal cells HBP23/Prx I exhibited a uniformly strong diffuse cytoplasmic staining (Figure 5A), HO-1 was expressed only in some hepatocytes, showing weaker and patchy staining pattern (Figure 5B). In KC cultures HBP23/Prx I and HO-1 protein were both detected by ICC, with a stronger signal for the latter (Figures 5C and 5D). HBP23/Prx I and HO-1 protein expression in cultured hepatocytes and KCs was also determined by Western blotting analysis, which confirmed their presence in both cell types (Figure 6A). By quantification of immunoblots, a higher relative level of protein expression was observed for both proteins in KCs than in hepatocytes (Figure 6B).
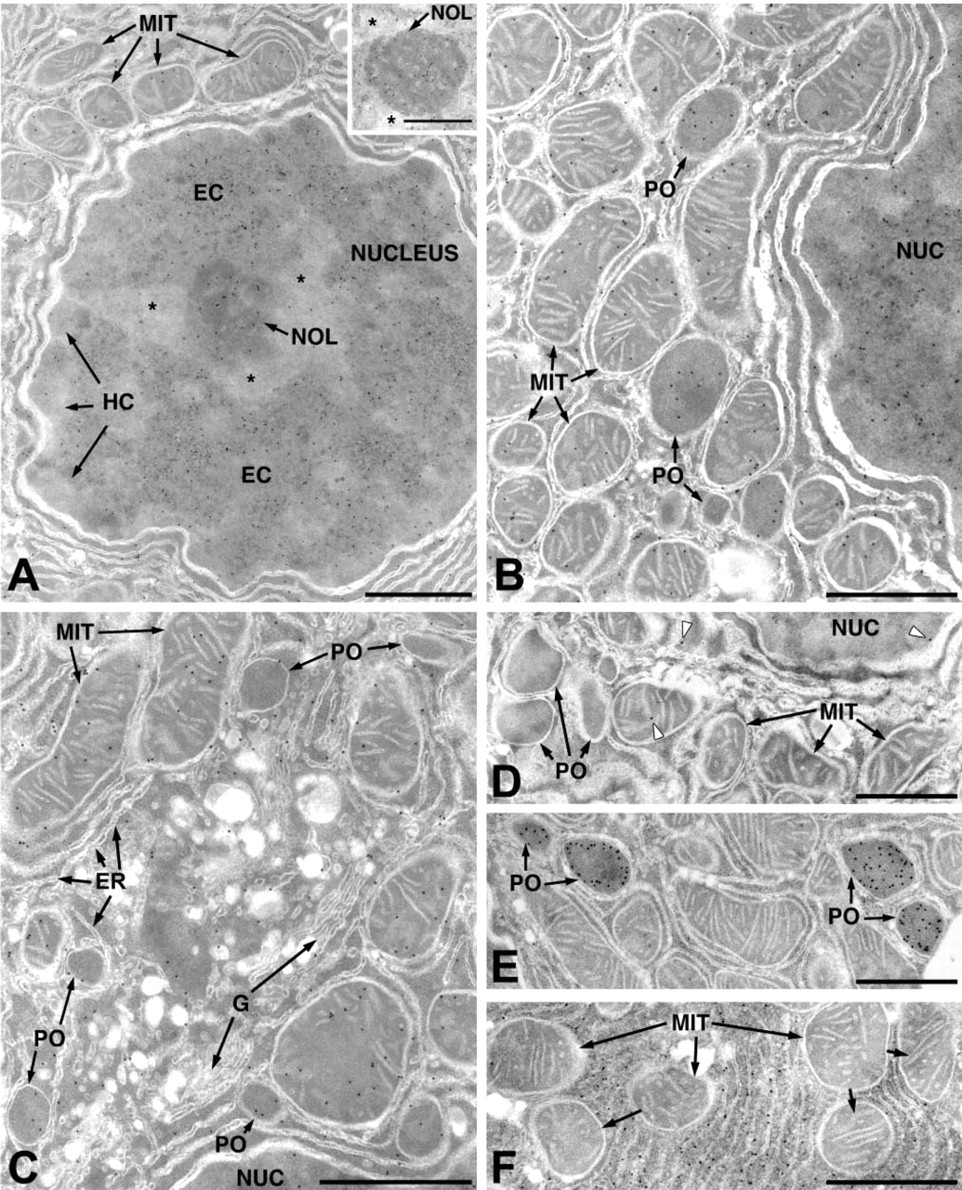
Electron microscopic immunocytochemical localization of HBP23/Prx I in parenchymal cells of rat liver. (
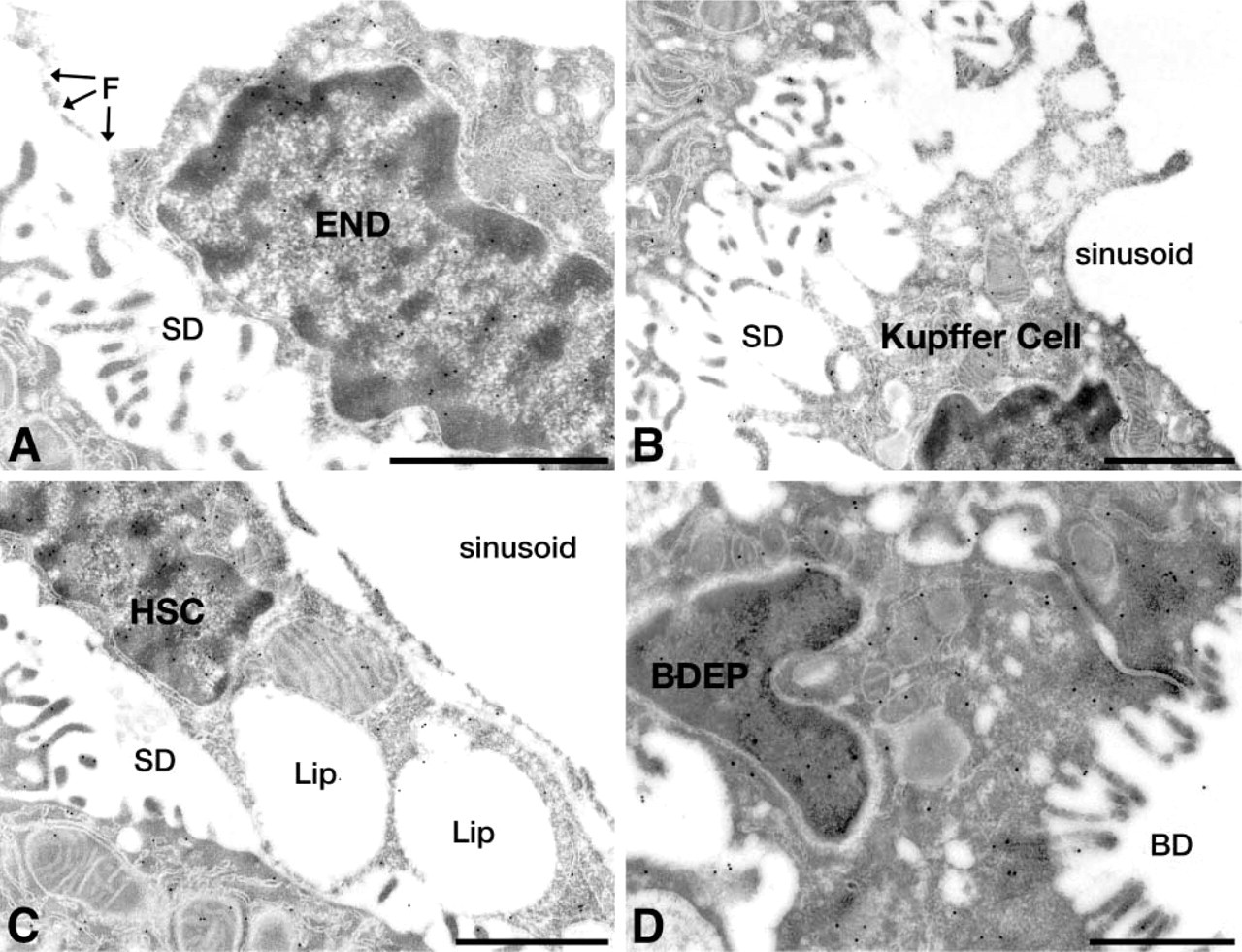
Electron microscopic immunocytochemical localization of HBP23/Prx I in non-parenchymal cells of rat liver. Ultrathin sections were incubated with a polyclonal rabbit antibody to HBP23/Prx I, followed by the protein A-gold procedure. Gold particles representing antigenic sites for HBP23/Prx I are observed (
As determined by Northern blotting analysis, HBP23/Prx I mRNA levels were similar in freshly isolated hepatocytes and KCs, whereas a higher level of mRNA expression was observed in freshly isolated SECs (Figure 7). By contrast, HO-1 mRNA expression was high only in freshly isolated KCs, being hardly detectable in hepatocytes and SEC (Figure 7). Primary hepatocytes and KCs were cultured for up to 5 days and mRNA steady-state levels were analyzed after various times of cell culture. HBP23/Prx I mRNA was strongly expressed for up to 5 days in both hepatocytes and KCs (Figure 8). A different mRNA expression pattern was observed for HO-1, which gave a strong signal in parenchymal cells cultured for 24 hr, with decline of mRNA levels up to day 5. By contrast, HO-1 mRNA expression levels in KC remained consistently high throughout 5 days of cell culture, which is in agreement with earlier observations from our laboratory (Immenschuh et al. 1999b).
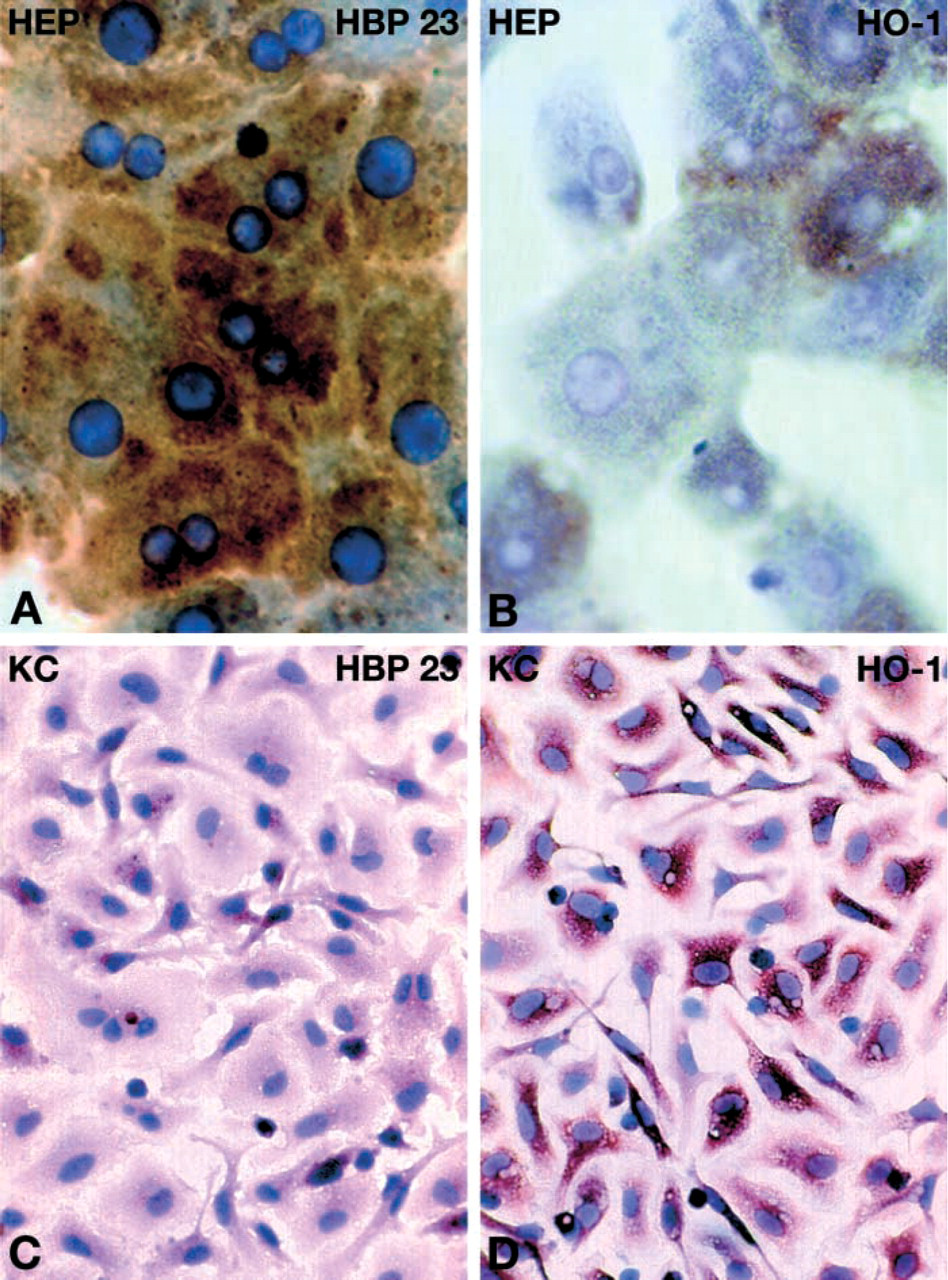
ICC localization of HBP23/Prx I and HO-1 in cultures of primary rat hepatocytes and KCs. Hepatocytes were prepared from livers of in vivo-perfused rats and were cultured as described in Materials and Methods. After 24 hr of cell culture, hepatocytes (
Discussion
HBP23/Prx I and HO-1 are antioxidant proteins that are coordinately induced by oxidative stress in different cell culture systems (Immenschuh et al. 1995, 1999a; Siow et al. 1995; Nakaso et al. 2000). Here, the cellular expression patterns of both proteins were investigated in rat liver. Our results reveal a differential gene expression pattern in parenchymal and non-parenchymal cells of rat liver.
Localization of HBP23/Prx I and HO-1 Gene Expression in Various Liver Cell Populations
Gene expression of HBP23/Prx I was uniformly strong in both parenchymal cells and non-parenchymal cells of rat liver (Figure 2). The hepatic gene expression pattern of HBP23/Prx I in whole liver was also confirmed quantitatively by mRNA expression analysis and Western blotting in freshly isolated and cultured liver cell populations (Figures 6–8). HBP23/Prx I was consistently expressed over 5 days of cell culture in hepatocytes and exhibited an even higher expression level in cultured KCs (Figure 8). A cell type-specific expression of HBP23/Prx I mRNA and protein in different rat organs has recently been shown. Therefore, HBP23/Prx I protein was widely detectable in the peripheral and central nervous systems of rat, with the strongest expression in oligodendrocytes and Schwann cells (Mizusawa et al. 2000). In mouse intestine, HBP23/Prx I is localized in columnar epithelial cells of the middle to lower part of the intestinal villi, whereas less intense immunoreactivity for HBP23/Prx I is observed in Paneth cells, goblet cells, and myofibroblasts (Ishii et al. 2000). In rat kidney, Oberley et al. (2001) found heavy immunostaining for HBP23/Prx I in proximal tubules, with weaker reaction in glomeruli, distal tubules, and transitional epithelium of renal pelvis.
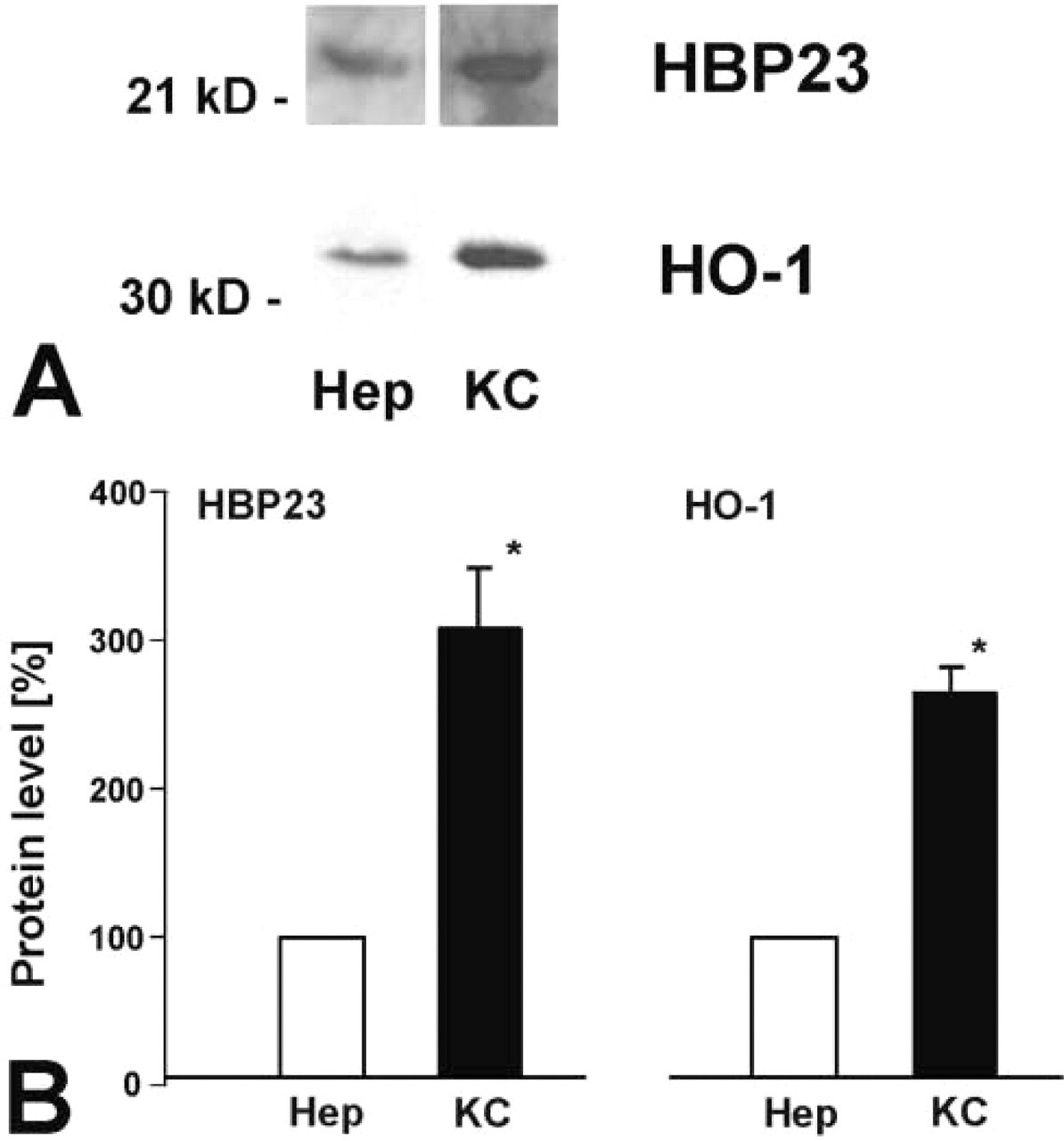
HBP23/Prx I and HO-1 protein levels in cultured hepatocytes and KCs of rat liver. (
In contrast to HBP23/Prx I, a different pattern of hepatic gene expression was observed for HO-1. Strong HO-1 immunoreactivity was detected in KCs, but not in parenchymal cells of normal rat liver (Figure 2C), thus confirming earlier observations (Bauer et al. 1998; Goda et al. 1998; Immenschuh et al. 1999b). Different levels of HBP23/Prx I and HO-1 mRNA expression were observed in freshly isolated and cultured liver cell populations (Figure 8). HBP23/Prx I gene expression was consistently high in cell cultures of hepatocytes and KCs for up to 5 days, whereas gene expression of HO-1 decreased in hepatocytes within 5 days in culture. These findings essentially agree with reports that have consistently shown higher levels of HO-1 expression in KCs, in contrast to lower HO-1 gene expression in other liver cell populations (Bissell et al. 1972; Bauer et al. 1998; Goda et al. 1998; Immenschuh et al. 1999b). The kinetics of HO-1 mRNA expression in cultured hepatocytes, with a marked increase from day 0 to day 1 and a continous decrease thereafter, might be explained by the cell isolation procedure that can be considered a stress stimulus per se that may induce HO-1 gene expression. In addition, Schuetz et al. (1988) have previously shown that expression of various genes can be markedly affected by the hepatocyte culture conditions, such as the use of various basement membrane matrices. Nevertheless, evidence of HO-1 immunostaining was observed in liver parenchymal cells of oxidant-stressed livers (unpublished observations), confirming the inducibility of HO-1 expression also in hepatocytes (Immenschuh et al. 1995; Bauer et al. 1998).
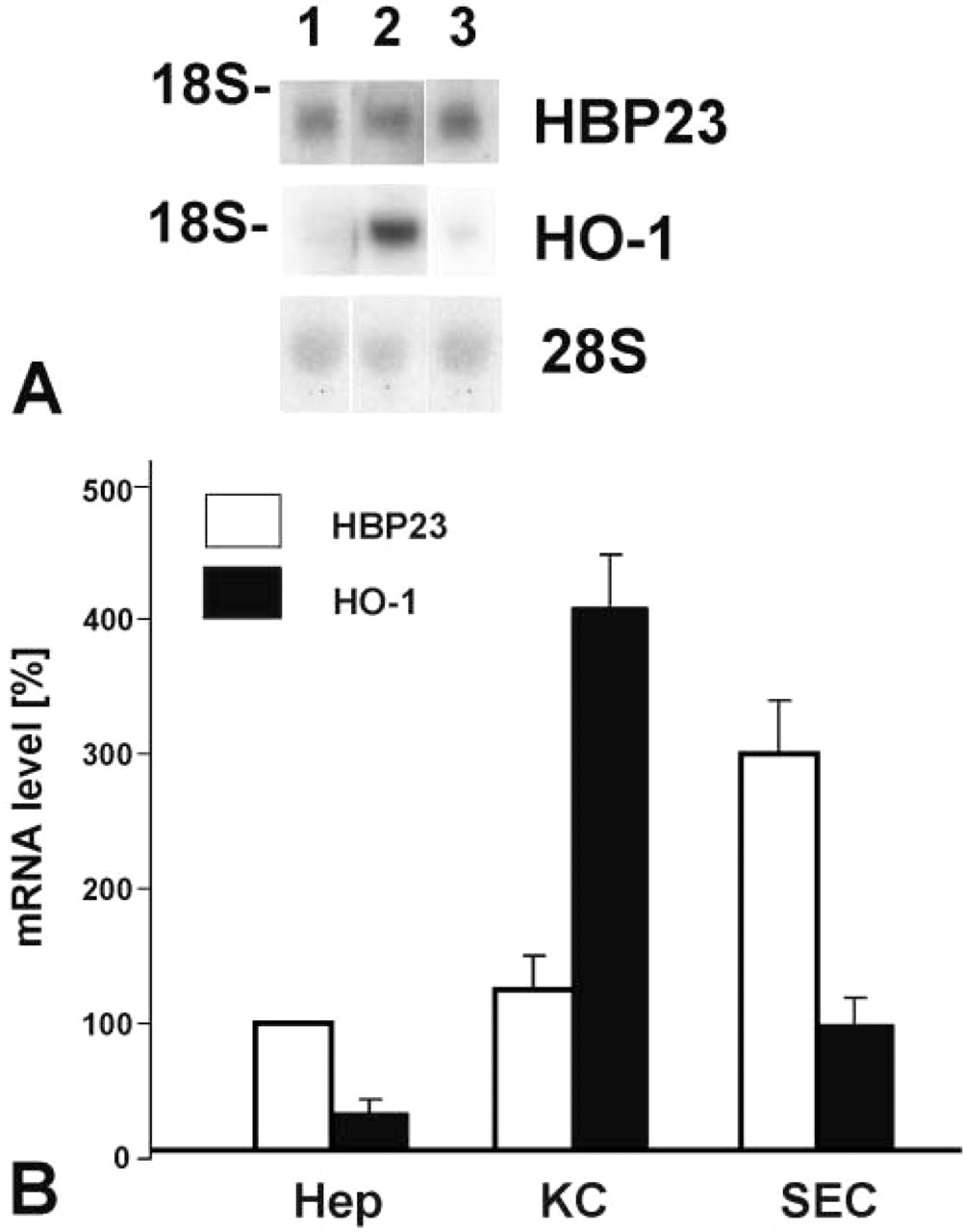
HBP23/Prx I and HO-1 mRNA expression levels in freshly isolated cells of rat liver. (
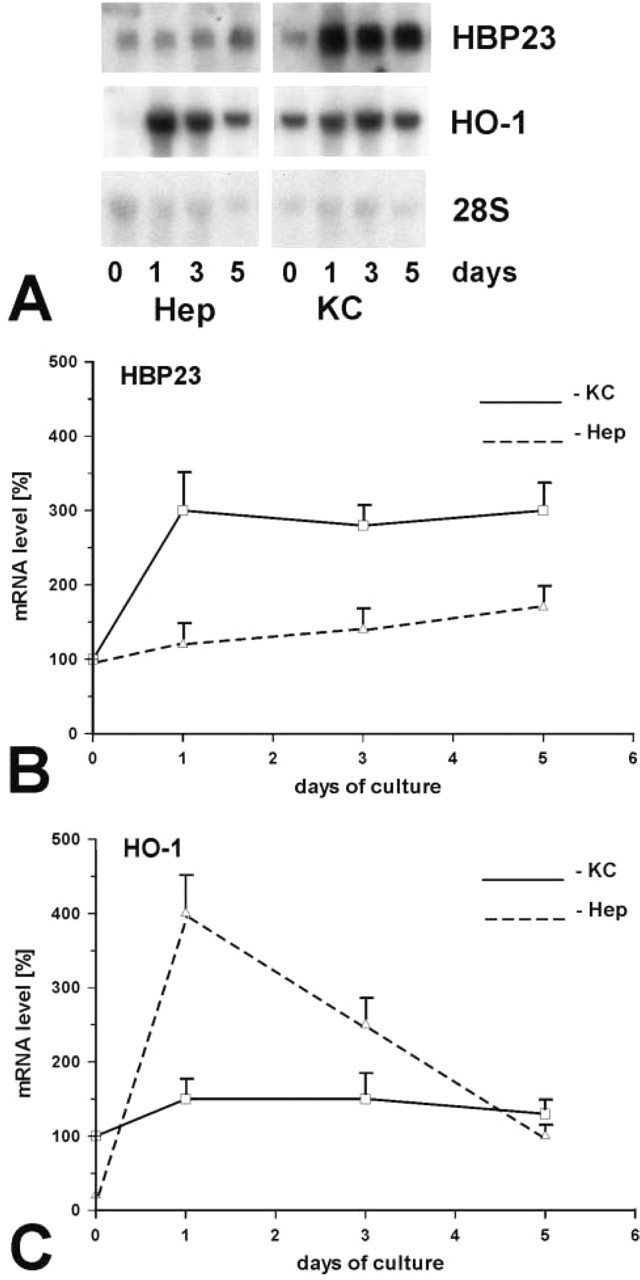
HBP23/Prx I and HO-1 mRNA expression levels in cultured hepatocytes and KCs of rat liver. (
Physiological Functions of HBP23/Prx I and HO-1 in Liver
Prxs are multifunctional antioxidant proteins that exhibit peroxidase enzyme activity, as demonstrated by overexpression in HeLa cells (Kang et al. 1998). Therefore, Prxs similar to other antioxidant enzymes may complement the cellular defense mechanisms against oxidative stress (Scandalios 1997; Oberley 2002; Shen and Nathan 2002). The antioxidant properties of HBP23/Prx I may also be related to its high binding affinity for the pro-oxidant heme. Because non-protein bound heme produces free radicals, HBP23/Prx I might contain the pro-oxidant function of heme by non-covalent binding (Vincent 1989). Alternatively, HBP23/Prx I may serve as an intracellular transporter of heme, as was previously shown for glutathione-S-transferases (Muller–Eberhard and Nikkila 1989; Boyer and Olsen 1991). The localization of HBP23/Prx I in mitochondria and peroxisomes (Figures 3 and 4), which contain substantial amounts of cytochromes and catalase, two hemoproteins, would support either one of the above-mentioned notions. Moreover, Prxs have been shown to regulate signal transduction pathways and to modulate cell proliferation and differentiation (Jin et al. 1997; Wen and Van Etten 1997; Kang et al. 1998; Seo et al. 2000; Chang et al. 2002).
The major physiological role of hepatic HO-1 appears to be the regulation of cellular heme homeostasis (Maines 1997), and it has been shown in vivo that HO can counteract heme-induced inflammation in liver (Wagener et al. 2001). In addition, HO-1-derived carbon monoxide is considered an important signaling gas in liver (Suematsu and Ishimura 2000), with the HO-product bilirubin functioning as a radical scavenger in vitro (Stocker et al. 1987).
Subcellular Localization of HBP23/Prx I in Liver Parenchymal Cells
Cell fractionation has been used to detect subcellular distribution of Prxs and, in most cases (five of six Prxs), the proteins were recovered in cytosol except for Prx III, which was found exclusively in mitochondria (for review see Rhee et al. 1999; Fujii and Ikeda 2002). In rat kidney, Oberley et al. (2001) have shown by immunoelectron microscopy that Prxs and thioredoxins exhibit a distinct localization pattern with Prx I and II in nucleus and cytoplasm, Prx III and V in mitochondria and Prx IV in lysosomes. Independently, a similar subcellular distribution of Prxs has recently been demonstrated in human mesotheliomas (Kinnula et al. 2002). Our observations, using the immunogold technique, have clearly revealed the presence of HBP23/Prx I not only in the cytoplasm and nucleus but also in the matrix of mitochondria and peroxisomes of liver parenchymal cells (Figures 3 and 4). This further emphasizes the important anti-oxidative role of HBP23/Prx I because substantial amounts of reactive oxygen species are generated in mitochondria (Chance et al. 1979) and peroxisomes (Singh 1996). By contrast, HO-1 has previously been demonstrated to be localized in the endoplasmic reticulum (Shibahara et al. 1980) which, according to our observations, appears to lack HBP23/Prx I in hepatocytes.
In conclusion, the data show a differential localization of HBP23/Prx I and HO-1 in various liver cell populations. The expression patterns of these antioxidant stress proteins suggest a complementary physiological role in normal liver.
Footnotes
Acknowledgements
We wish to acknowledge grant supports: from the Deutsche Forschungsgemeinschaft, Bonn, Germany, SFB 402 A8 (SI), C6 (GR), and SFB 601 B1 (EB, HDF).
We wish to thank Dr A. Schad for electronic digital image processing. The technical assistance of Heike Steininger, Gabi Krämer, E. Richter, W. Stöchmann, and Inge Frommer is gratefully acknowledged.
