Abstract
Human papillomavirus (HPV) is believed to promote the oncogenic process, and the correlation between viral oncoproteins and dysfunction of p16INK4A tumor suppressor protein in oral lesions is controversial. To test the hypothesis that anogenital HPV types participate in disruption of the regulation of p16INK4A suppressor protein in oral lesions, we analyzed 46 oral biopsy specimens for the presence of HPV 6/11 and 16/18 by in situ hybridization (ISH) and for p16INK4A expression by immunohistochemistry (IHC). Eighteen (39%) of the 46 oral lesions were HPV-positive and 28 (61%) were HPV-negative. HPV 6/11 DNA was found in 5 (11%) and HPV 16/18 in 13 (28%) of 46 biopsies. Nine of the 18 HPV-positive oral lesions (50%), assessed by catalyzed signal amplification coupled to ISH (CSA–ISH), gave high-intensity p16INK4A immunostaining. Focal and diffuse patterns were observed in 11/13 (77%) lesions with HPV 16/18, focal immunopositivity in 3/5 (80%) with HPV 6/11, and negative or sporadic p16-labeling in 18/28 (64%) without the presence of HPV DNA. These results showed a strong association between overexpression of p16 protein and malignant oral lesions, mainly those infected by HPV 16/18. We can conclude that high-risk HPV types are associated with p16 overexpression, and p16 may serve as a biomarker in oral cancer related to high-risk HPV infection.
O
Tumor cells are therefore believed to result from deregulation of two major pathways of cell cycle control: the p53 and pRB pathways mediated by p16INK4A (Pande et al. 1998; Robles and Adami 1998; Schoelch et al. 1999). Recently, mutation of p16INK4a was observed in some HPV-positive head and neck cancers, and such a mutation was also reported in tumors having inactivated p53 and RB genes (Saito et al. 1999). However, deletions or mutations of the p16 gene may not participate significantly in tumorigenesis of cervical cancer and other tumors (Kim et al. 1998). To gain a better understanding of the association between p16INK4A protein and deregulation of cell cycle control in oral cancer, we analyzed IHC expression of p16 protein and the presence of some anogenital HPV types in benign, premalignant, and malignant lesions.
Materials and Methods
Tissue Specimens
Forty-six formalin-fixed and paraffin-embedded oral biopsy specimens that had been taken for routine pathological diagnosis were obtained from the School of Dentistry archives, University of São Paulo State (UNESP), Brazil and approved by the Brazilian institutional ethical committee on human experimentation. All the samples were archival specimens graded according to the criteria recently proposed for oral white lesions (Van der Waal et al. 2000). Tissues were sectioned and evaluated histologically after staining in hematoxylin and eosin and their histopathological diagnoses were reconfirmed by two experienced professionals in a double-blind protocol. Of the 46 lesions, five were diagnosed as oral hyperplasia (OH), 14 were oral squamous papilloma (OSP), 10 were oral premalignant lesions (OPL) with mild to severe dysplasia, and 17 were oral squamous cell carcinoma (OSCC). Thin sections of 5 μm were cut, placed on organosilane-pretreated slides, and submitted to ISH and IHC examination.
DNA-DNA In Situ Hybridization
Sections were dewaxed in xylene, passed through an alcohol series, and air-dried. The slides were washed in standard saline citrate (2 × SSC = 0.3 M NaCl and 0.03 M C6H5Na3O7), pH 7.0, 0.2 N HCl, and a second 2 × SSC bath. The sections were incubated with 20 μg/ml proteinase K (GIBCO; Gaithersburg, MD) at 37C for 4 min, washed in fresh 2 × SSC buffer, 0.1 M triethanolamine solution containing 1% acetic anhydride (for 10 min at room temperature), and finally 2 × SSC buffer. ISH was performed with wide-spectrum 6/11 and 16/18 nonradioactively labeled HPV DNA probes (DAKO; Glostrup, Denmark) and a commercially available hybridization kit with a catalyzed system of amplification (CSA), Genpoint (DAKO). The probe solution was dropped on the sections, which were covered with coverslips, and cellular DNA was denatured on a hot plate at 95C for 10 min. The sections were briefly chilled in an iced chamber and incubated at 37C overnight in a humidified chamber. After hybridization, the sections were washed in 0.05 M Tris-HCl buffer (at RT), 2 × SSC, and 0.5 × SSC buffers. Hybridized DNA was detected with the Genpoint system (DAKO) according to the manufacturer' instructions and the sections were incubated in 40 mg of diaminobenzidine and 20 μl of peroxidase in 100 ml of saline PBS. Finally, the sections were rinsed in distilled water, counterstained with Meyer' hematoxylin, and washed in dilute aqueous ammonia and distilled water. After that, the sections were dehydrated in graded alcohol, allowed to dry, and mounted in Entellan mounting medium (Merck; Darmstadt, Germany). For each reaction three controls were used: one positive control (commercial slide in the Genpoint kit), one negative control (commercial slide in the Genpoint kit without DNA probe), and one normal oral tissue control. DNA extraction and β-globin gene amplification were performed to verify the DNA quality of the tissue samples.
Immunohistochemistry for p16INK4A Protein
Immunostaining Procedures. The sections were dewaxed in xylene, rehydrated in graded alcohol, and rinsed in water. To inhibit endogenous peroxidase activity, the sections were treated by immersion in five changes of 0.3% H2O2 in absolute methanol (5 min each change) and rinsing in water. For antigen retrieval, the sections were immersed in 10 mM sodium citrate buffer (pH 6.0) and boiled twice for 12 min in a high-intensity microwave oven. To detect p16INK4A protein, an IHC assay was performed using the streptavidin-biotin system (LSAB; DAKO). The slides were incubated with primary monoclonal antibody for p16 protein (p16/Abs4, diluted 1:40; Labvision, Fremont, CA) in a humidified chamber at 4C overnight. The sections were then incubated with biotinylated anti-rabbit antibody and streptavidin-peroxidase complex (LSAB system; DAKO) at 37C for 30 min. Careful rinses were performed, with several changes of PBS between each stage of the procedure. The sections were incubated with diaminobenzidine (Gibco), then counterstained lightly with Carrazzi' hematoxylin, dehydrated in graded alcohol, allowed to dry in air, and mounted with Permount mounting medium (Merck).
Controls. Positive controls were included in each assay, comprising cell block sections of HeLa cell line, negative controls in which the primary antibody was replaced with PBS, and a control of normal oral tissue. The immunoreaction cut-off was established by quantifying nuclear and cytoplasmic positivity in normal tissue, which ranged from zero to 5%.
Quantitative Evaluation of p16INK4A Immunostaining
The IHC expression of p16 was quantified in a double-blind protocol and classified according to nuclear and cytoplasmic positivity. The biopsies were scored as positive when more than 5% cells (cut-off) stained positively, and were separately graded as described by Klaes et al. (2001): negative (0–5% of nuclei and cytoplasm positive), sporadic (5–10% of nuclei and cytoplasm with weak and scattered positivity), focal (>10–30% of labeled nuclei and cytoplasm strongly positive, spreading in one tissue area), and diffuse (>30–85% of labeled cells with strong positivity, spreading in several tissue areas). Biopsies that exhibited a diffuse pattern were considered to have high IHC expression of p16INK4A. Focal distribution was interpreted as moderate expression and sporadic positivity as low expression.
Statistical Analysis
The p16INK4A IHC expression results were analyzed by the χ2 probability test, with p<0.05 being considered statistically significant.
Results
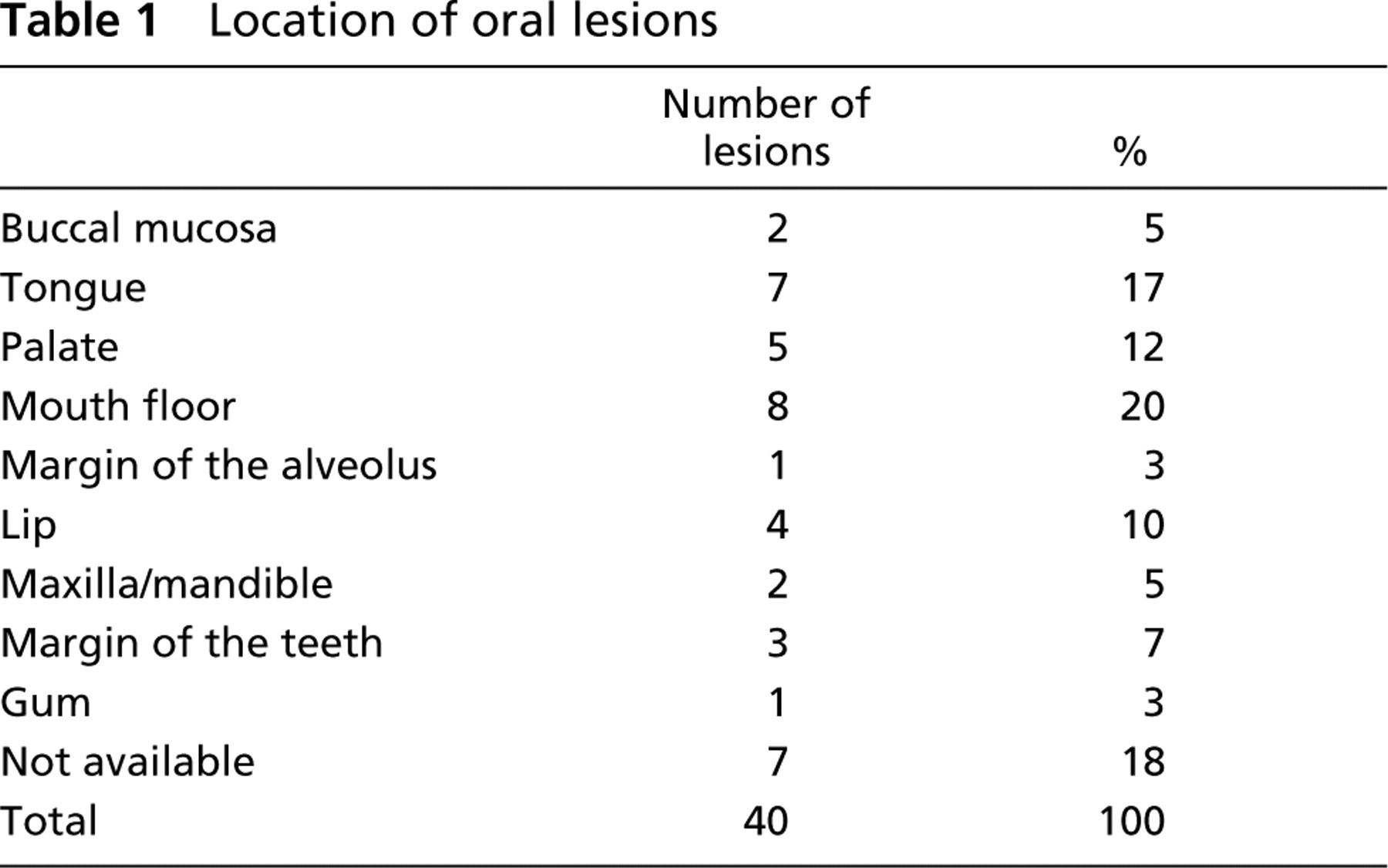
The oral lesions used in the present study (Table 1) were most commonly located on the mouth floor (20%), tongue (17%), and palate (12%).
Table 2 reports the frequency of HPV ISH expression in oral lesions. Eighteen (39%) of the 46 oral lesions were HPV-positive and 28 (61%) were HPV-negative. HPV 6/11 DNA was found in two (14%) of 14 OSPs, two (20%) of 10 OPLs, and one (6%) of 17 OSCCs. HPV 16/18 was found in one (20%) of 5 OHs, three (22%) of 14 OSPs, four (40%) of 10 OPLs, and five (29%) of 17 OSCCs. No oral lesion was simultaneously positive for HPV 6/11 and 16/18. These results show a strong correlation of the premalignant and malignant lesions with HPV 16/18 positivity.
Table 3 shows the data of p16INK4A IHC expression to histopathological diagnosis in oral biopsies. p16INK4a focal and diffuse patterns of positivity were found mainly in premalignant and malignant oral lesions (OPL and OSCC), whereas sporadic pattern was associated with benign lesions (OH and OSP). These results showed a strong association between overexpression of p16 protein and oral lesions that were malignant or with high malignancy potential (p = 5.0 E–03; p<0.05). Interestingly, the OH lesion (n=1) infected with HPV DNA 16/18 gave a diffuse pattern. On the other hand, some premalignant (n=3) and malignant (n=2) lesions, HPV-negative, had negative or sporadic p16 scores. These results may indicate a close association between high-risk HPV infection and specific p16 IHC patterns.
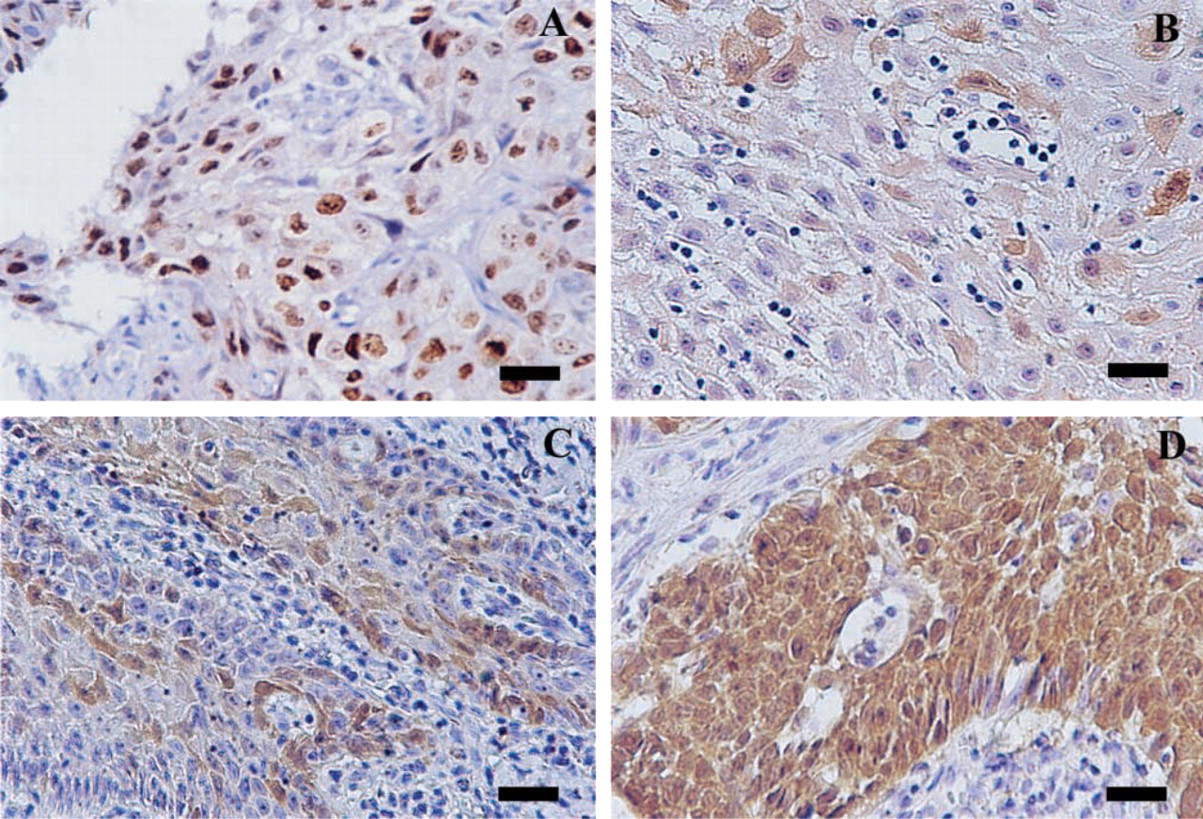
Table 4 matches p16INK4A expression in oral lesions against infection by HPV. Nine of the 18 HPV-positive oral lesions (50%), assessed by CSA-ISH, gave intense p16INK4A immunostaining in a diffuse pattern and five (28%) a focal pattern of staining. Focal and diffuse patterns were observed in 11 of 13 (77%) HPV 16/18-infected lesions, focal immunopositivity in three of five (80%) with HPV 6/11 DNA, and negative or sporadic p16 labeling in 18 of 28 (64%) with no HPV DNA (Figure 1). These results also demonstrate that different patterns of p16 immunopositivity could be associated with low- or high-risk HPV infection in oral tissues (p = 1.0 E–04; p<0.05).
Location of oral lesions

Presence of human papillomavirus (HPV) in 46 oral lesions detected by in situ hybridization a
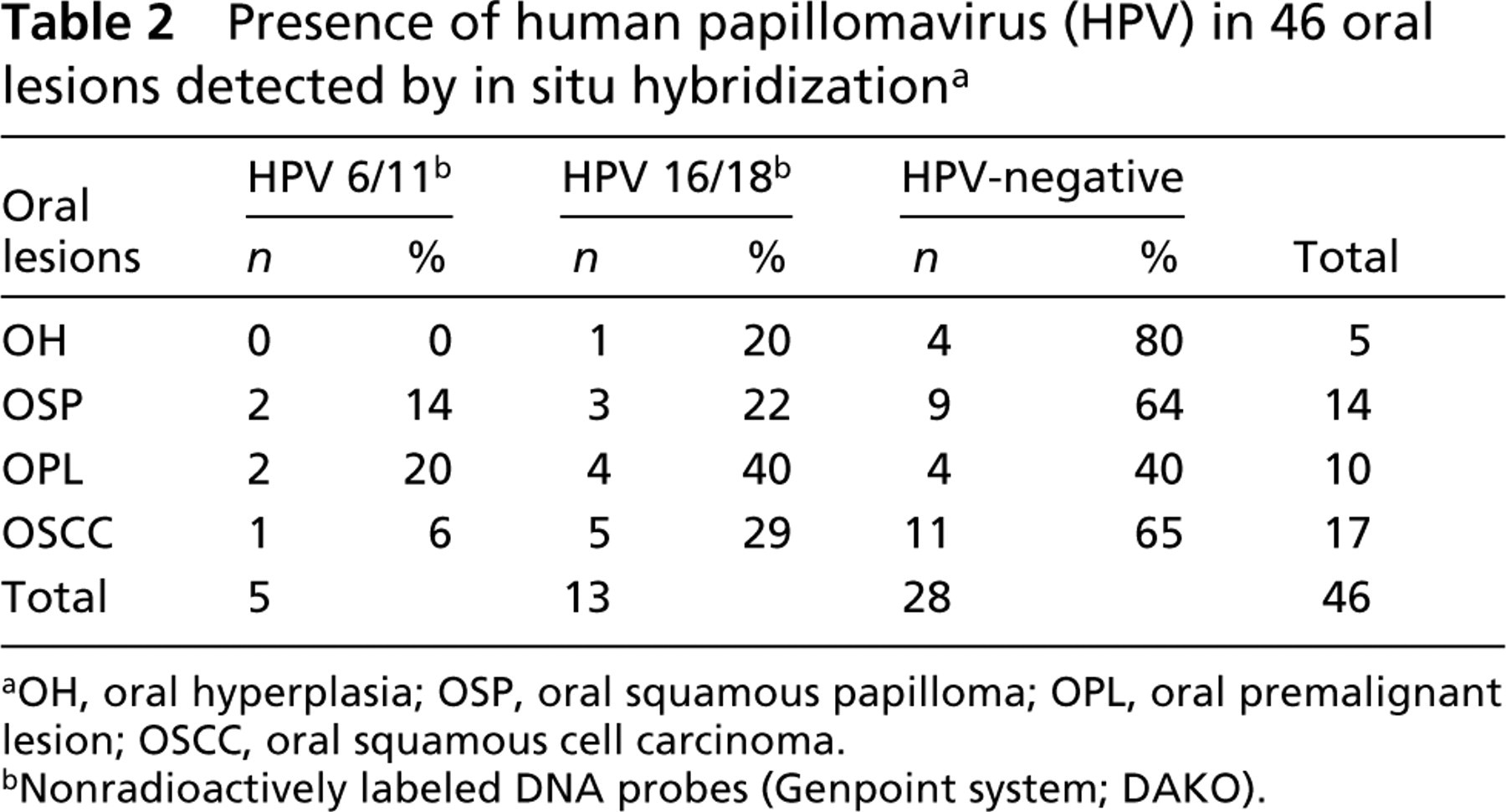
OH, oral hyperplasia; OSP, oral squamous papilloma; OPL, oral premalignant lesion; OSCC, oral squamous cell carcinoma.
Nonradioactively labeled DNA probes (Genpoint system; DAKO).
p16INK4A immunohistochemical expression in 46 biopsies of benign, premalignant, and malignant oral lesions a
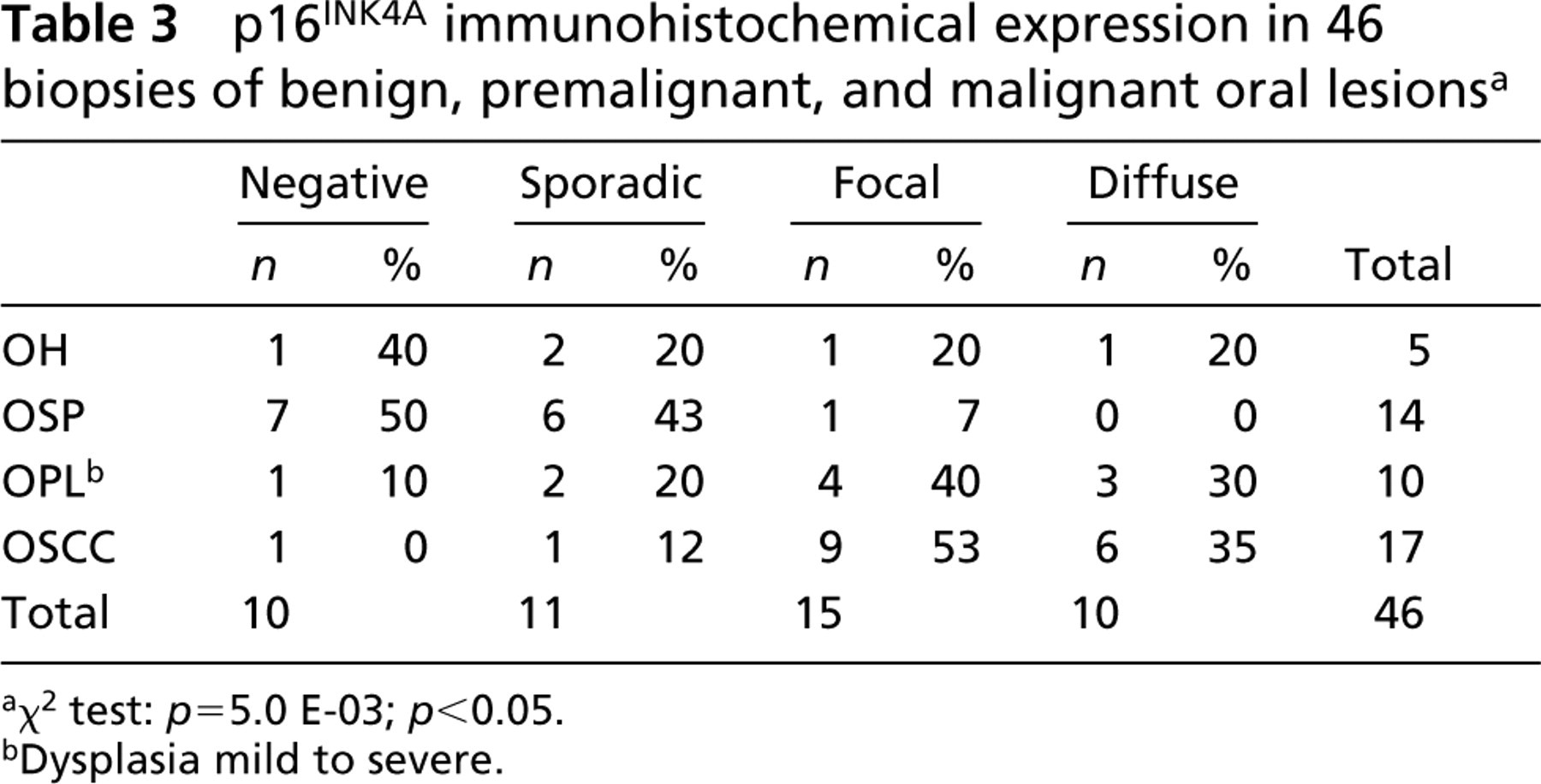
χ2 test: p = 5.0 E-03; p<0.05.
Dysplasia mild to severe.
However, we found focal p16 expression in nine (32%) of the oral lesions that were negative for HPV DNA, of which eight (29%) were diagnosed as OSCC and one (3.5%) as OPL. This result suggests that moderate expression in the absence of viral DNA could be associated with histological type and that p16 gene dysfunction could be caused by other carcinogenic co-factors. On the other hand, we found biopsies with HPV 6/11 and 16/18, respectively, with p16 pattern negative (n=2) or sporadic (n=2). Lesions with HPV 6/11 and 16/18, but p16-negative, were diagnosed as OSP, whereas those combining HPV 6/11 and 16/18 DNA with sporadic p16 expression were classified as, respectively, OSP and OSCC. These results also suggest that, in some lesions, HPV appears not to be involved in p16INK4A deregulation. It is possible that high-risk viral presence alone does not give E6/E7 viral protein expression and therefore results in normal p16 function. Further carcinogenic mechanisms and/or expression of other oncoproteins may therefore have to be considered for a more complete understanding of the rapid cell proliferation or malignancy transformation.
p16INK4A overexpression in oral lesions infected by anogenital HPV a
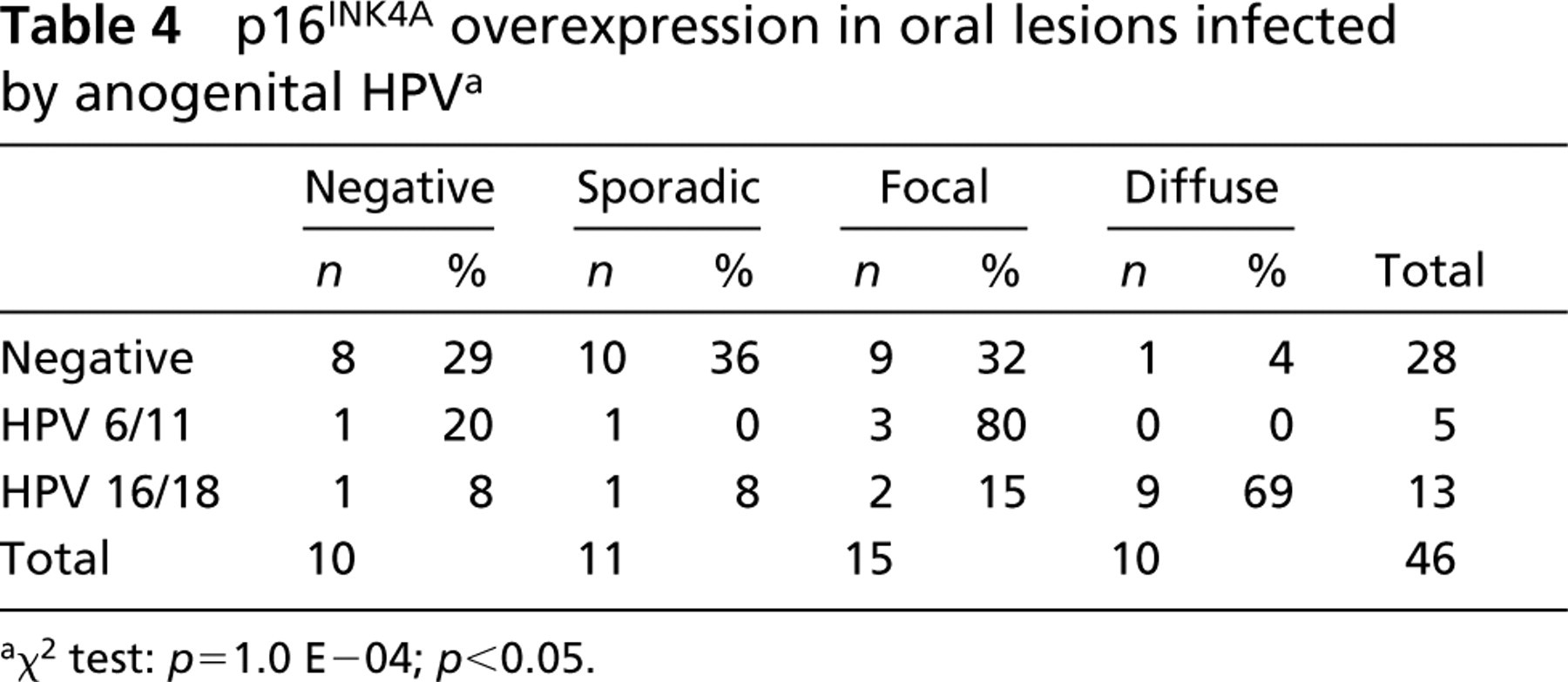
χ2 test: p = 1.0 E-04; p<0.05.
(
Discussion
The choice of a suitable method for detection of HPV DNA has become increasingly complex (De Villiers 1997). Whereas PCR has been considered the most appropriate method for viral detection, ISH is widely used in routine diagnosis or scientific investigations. Although considered a less sensitive method, ISH has proved to be a very important molecular tool in diagnosis and research and has significantly advanced the study of gene structure and expression at the level of individual cells. The usefulness of conventional ISH was occasionally limited by low detection sensitivity (Faulkner–Jones et al. 1990; Schneider et al. 1991), an obstacle that may now be overcome by new techniques. The strategies to improve threshold levels for detection with ISH include protocols that either amplify the detection signals, using several antibody steps, or increase the absolute amount of hybridized probes. Moreover, new commercially available amplification systems really do improve viral DNA detection and allow the localization of the viral genome or its transcripts within the morphology of the lesions (De Villiers 1997; Lizard et al. 2001). In this study we used ISH with a catalyzed signal amplification system (CSA–ISH), a novel signal amplification method based on the catalyzed amplification of positive hybridization signals using biotinyl–tyramide complexes. CSA–ISH gives much better results, providing higher signal intensity and sensitivity than conventional immunoenzymatic revelation procedures (Birner et al. 2001). A recent study showed that the Genpoint system is sensitive enough to detect one or two copies of HPV DNA in isolated CaSki line cells (Lizard et al. 2001). Using CSA amplification, we found 39% of oral lesions to be HPV 6/11- and 16/18-positive, with a higher prevalence of HPV 16/18 in both oral dysplasias and carcinomas. Various ranges of HPV DNA positivity (Mao 1995; Fornatora et al. 1996; D'Costa et al. 1998; Miguel et al. 1998) have been reported in oral lesions, with a mean frequency (Miller and Johnstone 2001; Soares et al. 2003) of 20–30%. Moreover, recent reports have shown a higher percentage (50%) of positivity of high-risk HPV in head and neck cancers (Miller and White 1996; Sand et al. 2000). Therefore, our findings may indicate a stronger etiological link between high-risk HPV and OSCC.
We also found a close association between low-risk HPV and oral squamous papilloma, whose prognostic behavior is usually benign. However, we found HPV 16/18 in one (20%) of five OHs and in three (22%) out of 14 OSPs, suggesting that HPV can lead benign cells to malignant progression. A high prevalence of HPV 16/18 has been found in OSP, suggesting viral involvement in oral carcinogenesis, and recent studies have shown a close association between oral hyperplasia and/or OSP HPV 6/11 and HPV16/18 (Paparotto Lopes and Meeks 2001). Therefore, early detection of HPV in these lesions is highly recommended and may offer new directions for oral cancer therapy and prevention.
Abnormalities in various components of the cell cycle regulatory machine and disruption of cell cycle control are common features of many cancers, including oral cancers (Geradts et al. 1995; Tanaka et al. 2001; Tsai et al. 2001; Yoo et al. 2002; Soares et al. 2003). Recent advances in cell biology have elucidated mechanisms of cell cycle control that not only involve cyclin and cyclin-dependent kinase (CDK) complexes but also depend on CDK inhibitors, such as p16INK4A (Zur Hausen 2002).
In the present study, p16 positivity patterns were found to correlate with the stage of tumor progression, focal and diffuse features being associated, respectively, with premalignant and malignant oral lesions, and negative or sporadic labeling with benign lesions. Some studies on cervical cancer have demonstrated that p16INK4A may serve as a marker to differentiate neoplastic lesions from hyperplastic or reactive lesions (Sano et al. 1998; Bibbo et al. 2002). However, few studies have used IHC tests to compare p16 expression in oral cancer with tumor grading (Papadimi-trakopoulou et al. 1997; Pande et al. 1998; Chen et al. 1999). Our findings are in accordance with a previous study indicating that p16 IHC expression could be used as a helpful marker in oral tumor progression (Chen et al. 1999). Surprisingly, some premalignant and malignant lesions in the present study exhibited negative or sporadic p16 positivity, whereas one hyperplastic lesion had diffuse immunostaining. An OH lesion was infected with HPV DNA 16/18 and some premalignant and malignant lesions were HPV-negative. OHs infected with HPV 16/18 and showing p16INK4A overexpression suggest that some lesions classified as benign could hide a malignant potential when infected with high-risk HPV. The present study also demonstrated distinct p16-labeling patterns in low- and high-risk HPV lesions, which were frequently focal in HPV 6/11 and diffuse in HPV 16/18. Our findings indicate that p16 evaluation might be useful in distinguishing HPV-related lesions from other neoplastic lesions uninfected by the virus (Sano et al. 1998). Some studies have shown that inactivation of the p16INK4A gene is a common event in head and neck squamous cell carcinoma and results in the loss of positive immunostaining for the protein (Reed et al. 1996; Yamato et al. 2000; Rocco and Sidransky 2001; Shintani et al. 2001). Apparently, p16 protein expression in oral carcinogenesis may occur in relation to the functional inactivation of Rb protein by HPV infection (Sano et al. 1998). Baseline levels of p16INK4A are high in many immortal cell lines and human tumors, but apparently this occurs exclusively under conditions in which mutations in other proteins (e.g., pRB) or the presence of viral oncoproteins render it ineffective (Zhang et al. 1994; Robles and Adami 1998; Shintani et al. 2001). The E7 protein from high-risk HPV binds to Rb and Rb-related proteins (Yamato et al. 2000; Llewellyn et al. 2001). The E7 binding disrupts the interaction of Rb and E2F, allowing E2F to activate its downstream cellular target genes required for cell cycle progression. E2F also appears to activate expression of the products encoded by INK4a, leading to an accumulation of p53 by preventing its degradation (Sano et al. 1998). E6 and E7 HPV oncoproteins have been considered to be responsible for malfunction of p16INK4A, a tumor suppressor protein (Olshan et al. 1997; Zur Hausen 2002). Thus, E6/E7 oncoprotein seems to be inactivated in low-risk HPV infection (Zur Hausen 2002) and may be related to lower p16 expression (Chen et al. 1999). Therefore, p16 IHC features might be helpful to predict the presence of low- or high-risk HPV infection. However, some oral lesions with HPV 16/18 were p16-negative or sporadically positive. Possibly the presence of high-risk viral DNA alone does not give rise to E6/E7 expression, so that a normal p16 function results. In such cases the amount of E6 and E7 mRNA would need to be analyzed to determine the risk of malignancy in oral lesions infected with high-risk HPV. Detection of specific mRNA derived from integrated HPV genomes in advanced precancers can be used to identify lesions with a particularly high risk for progression to invasive cancer (Doeberitz 2002). On the other hand, several HPV-negative oral lesions gave moderate (focal) or high (diffuse) p16 expression. Intriguingly, despite the large amount of molecular genetic and biochemical data supporting a tumor suppressor role for p16, the exact biological role of this gene remains elusive (Rocco and Sidransky 2001). Previous study has suggested that alterations of the p16/pRB pathway may be implicated in betel nut- and tobacco-related oral tumorigenesis (Pande et al. 1998). Thus, p16INK4A dysfunction may not be only HPV-related, and other mechanisms and co-factors should be considered.
We can conclude that high-risk HPV types are involved in p16 overexpression, perhaps due to viral integration and malfunction of this tumor suppressor protein, and probably contribute to the multistep oral carcinogenesis process. Moreover, our results suggest that p16 immunohistochemical expression is a useful marker for high-risk HPV infection and the development of oral cancer. Further studies will attempt to uncover more details of the molecular events, involving p16 and HPV, that drive the oral carcinogenic process.
Footnotes
Acknowledgements
Supported by FUNDUNESP, a Brazilian research-funding organization.
