Abstract
The activity of the enzyme acetyl-CoA-carboxylase α (ACC-α) is rate limiting for the de novo synthesis of fatty acids. The encoding gene is expressed from three promoters in ruminants (PI-PIII). Their individual contribution to the formation of milk fat is unknown. Promoter-specific molecular probes were hybridized in situ to serial sections of mammary glands from cows and sheep to determine their developmental and spatial expression profile in the udder. We show that all three promoters are active in mammary epithelial cells (MECs) of udders from both species. This implies that, in principle, none of these promoters can be singled out as the key element controlling the ACC-α-related contribution to establishment of milk fat content, although the activity of PIII only is known to be disproportionally stimulated by lactation in MECs. We propose that all three promoters may be relevant for milk fat synthesis in cattle, whereas PII and PIII are crucial for milk fat formation in sheep. We show also that ACC-α synthesis is not strictly coupled to casein synthesis, particularly during pregnancy and involution.
Keywords
O
Activity of the ACC-α is known to be rate-limiting for the de novo formation of fatty acids (for reviews see Wakil et al. 1983; Kim 1997). Levels of regulation include allosteric interactions with metabolites and reversible phosphorylations (Hardie 1989; Kim et al. 1989), triggered extracellularly by various hormones (Iritani 1992). Long-term control is exerted at the transcriptional level (Kim 1997). The encoding gene is expressed from two distinct promoters in most mammals (Lopez-Casillas et al. 1991). PI transcribes exon 1, PII promotes exon 2, and the translation initiation codon for both moieties of transcripts is located on exon 5. Three promoters initiate transcription from this gene in sheep (Barber and Travers 1998) and cattle (Mao et al. 2001). The third promoter (PIII) resides in intron 5 and uniquely initiates expression of exon 5A (Barber and Travers 1998; Mao et al. 2001). Exon 5A harbors an alternative initiation codon for translation, resulting in the formation of an N'-terminally modified isoform of the enzyme (Barber and Travers 1998). The significance of the enzymatic variation is unknown.
Different tissue-specific restrictions are imposed on the initiation of transcription from these promoters. PI is primarily active in adipose tissue and its activation is nutritionally regulated in liver (Lopez-Casillas et al. 1991). Non-homologous elements of DNA have been recruited throughout evolution to serve as functionally homologous PI (Mao et al. 2001). However, although PI of cattle and sheep are truly homologous structures, these two evolutionary closely related species display differences in initiation from PI in the udder. PI-derived transcripts were found to be strongly expressed in the udder of the cow (Mao et al. 2001) but were not detected in sheep (Travers et al. 2001). Initiation of transcripts using the evolutionarily conserved PII is fairly ubiquitous in all species examined and hence PII is considered to represent a “housekeeping” promoter (Luo and Kim 1990; Mao and Seyfert 2002). Its mammary gland-specific activation was found to be strongly induced (∼10-fold) by lactation in rats (Ponce-Castaneda et al. 1991). In cattle, however, lactation stimulates the expression of transcripts from PII in the udder by only approximately threefold (Mao and Seyfert 2002), and hence is just proportional to the stimulation of the overall expression of the ACC-α-encoding gene (Mao et al. 2001). Activity of the Ruminantia-specific PIII is predominantly confined to the lactating mammary gland. The transcription from this promoter only is strongly induced by lactation in the udder (15–28-fold), while merely basal activities are found in virgin glands or other tissues (Barber and Travers 1998; Mao et al. 2002). In the udder, PIII is active in MECs (Mao et al. 2002). The activity of PIII only was shown to be stimulated by the lactational hormone prolactin (PRL), as signaled through activated STAT5 transcription factors (Mao et al. 2002). PRL-induced STAT5 activation of STAT5, in turn, was shown to occur exclusively in MECs of the mammary gland (Gallego et al. 2001). The cell-specific expression of the other two promoters has not been analyzed in the mammary gland thus far.
All three promoters are equipped with a unique set of cis-regulatory elements. Given the interest in understanding the hormonal and nutritional controls influencing the formation of milk fat, it is necessary to elucidate the specific role of any one of these promoters in the formation of milk fat. Therefore, we used in situ hybridization (ISH) to examine which of these promoters is used in MECs, the relevant cell type for milk fat formation.
Materials And Methods
Cloning of Probes
Probes of bovine origin were used throughout. Isolation and molecular characterization of the entire bovine promoter region have been described (Mao et al. 2001). To generate promoter-specific probes, the entire exons 1, 5, and 5A were amplified with specific primers from our initial BAC91 isolate and subcloned into the vector pGEMTeasy (Promega; Mannheim, Germany). Exon 1 served as probe for PI-derived transcripts and exon 5A as probe for PIII-derived transcripts. Exon 2 has an extraordinarily high content of G and C residues. This precludes its use as riboprobe for ISH due to crossreactivity with ribosomal RNAs, causing high background (Witkiewicz et al. 1993). Therefore, we used exon 5 as a probe with specificity for transcripts from both promoters, PI and PII. Activity of PII becomes evident by comparing signals of this probe to those obtained with the exon 1-specific probe. We used for exon 1 amplification the oligonucleotide primers Ac1ex1 (forward, 5′-GAAGTTCCCTCAGGCTCTTAATC; reverse, 5′-CTCAGAGACCTCTCTGCTTC); for exon 5 amplification the primers Acex5 (forward, 5′-CTCTGAGAGCTCATTTTGAAG; reverse, CTCATGTGTAAGGCCAAACCAT), and the primers bAc5A to amplify exon 5A (forward, 5′-AGGCGGAAGCTGCTGAGATCTAC; reverse, 5′-TCTTCAGCTGTCGGCCTTG). The lengths of the resulting probes were 244 bp, 253 bp, and 474 bp for exons 1, 5, and 5A, respectively. All template clones were linearized 3′ of the insert with either NcoI or SalI before in vitro translation. Linearized templates were purified twice with phenol/chloroform extraction and then precipitated with ethanol before in vitro transcription.
Probe Labeling
Probes were labeled with 20 U of either T7 or SP6 polymerases (Promega) in 20 μl reaction volumes. They contained the appropriate reaction buffer as provided by the supplier and we added 10 mM DTT, 1 U of RNasin (Promega), 10 mM each of rATP, rGTP, rCTP, and 25 μCi of [35S]-UTP (Amersham; Little Chalfont, UK).
Slide Preparation and General Procedures for ISH
Serial sections (7-μm) were cut from paraformaldehyde (4%)-fixed and wax-embedded udder samples prepared from freshly slaughtered, pregnant, lactating, or involuting sheep and cows. They were mounted adjacently on ATS (Sigma-Aldrich, St Louis, MO; cat. no. A3648)-coated slides. We followed the general procedures for slide preparation and hybridization with radioactively labeled riboprobes to such sections essentially as previously described (Molenaar et al. 1992). This reference describes also the bovine αS1-casein control probe used. 35S-labeled riboprobes were generated from the respective subclones. Approximately 40,000 cpm of the radiolabelled sense and antisense probe were added per μl of hybridization mix. Then 30 μl of hybridization mix was applied to each section. Probes were hybridized overnight at 58C. The hybridization mixes contained 0.2M DTT and washing solutions contained 20 mM of β-mercaptoethanol. The slides were washed in 50% formamide, 2 × SSC, 20 mM β-mercaptoethanol at 55C (twice for 15 min), then with several changes of 0.2 × SSC. Subsequently, all single-stranded RNA was digested away with RNase A and T1 for 30 min (10 μg/ml and 2.5 μg/ml, respectively; Roche Diagnostics, Basel, Switzerland), then rewashed in 0.2 × SCC. Slides were dehydrated in a graded series of ethanol, all containing 10 mM β-mercaptoethanol. They were coated with LM-1 nuclear emulsion RPN40 (Amersham), allowed to develop at 4C in sealed desiccant-containing boxes, and developed after an exposure of 2 weeks (αS1-casein) or 2.5 months (all ACC-α probes). They were counterstained with Gill's rapid haematoxylin and eosin.
Quantification of ISH Signal Intensities over Various Cell Types
An Olympus BH2 microscope and PM-10ADS photomicrographic equipment were used to record the relative signal intensities by setting the camera controller to spot metering (1% of the field view) and positioning the measuring area over the cell types under examination. The X40 objective was used for the cow measurements and the X100 objective used for the sheep measurements. Objective measurements were obtained by setting the illumination to full intensity and reading the exposure time displayed by the camera controller. This gave a direct measure of the signal strength, since the length of the exposure time corresponded to the extent to which the deposited silver grains blocked the light and hence gave a measure of their number. Measurements were taken of 10 representative cell types and averaged. Background was corrected by subtracting the average of six measurements from corresponding cell populations from the adjacent control sections.
RT- and Real-Time Quantification PCR
RNA for RT- and real-time PCR experiments was extracted using TRIZOL (Life Technologies; Karlsruhe, Germany) as prescribed by the manufacturer. The cDNA for these assays was primed in reverse with the oligonucleotide primer Acex8_9r (5′-CAGCCAGCCCAAACTGCTTGTTGCAC) bridging exons 8 and 9. This cDNA was used to amplify the ovine exon 1 with the primers bovine/ovine Ac1cx1f: 5′-GTCTGTCCATCTGTGAAGTATC; reverse Acex5r: 5′-CTCATGTGTAAGGCCA(G/A) ACCAT (G for bovine, A for ovine amplifications, respectively). The abundance of total and PI-derived transcripts in mammary glands from sheep was assayed with the Light Cycler device and assay kits (Roche Diagnostics), basically as described (Mao et al. 2001) but using cDNA equivalent to 25 ng of total RNA as input in these determinations and the respective ovine specific forward primer. Abundance of PIII-derived transcripts was also assayed from these cDNAs. Primers oAc_5Af (5′-AGGCAGAAGCTGCTGAGATCTAC) combined with the reverse primer Ac5A6r (5′-GCCAGACATGCTGGATCTTTG; bridging exon 5A to exon 6 in both species) served to amplify transcripts derived from ovine PIII. Relative abundances were titrated from a dilution series of the appropriate subclones containing 106 to 10 copies of the plasmids.
Results
Clear association of the signals from all of our ACC-α specific antisense probes with MECs is the most prominent feature seen in our hybridizations. The sense control probes, on the other hand, did not give rise to any specific accumulation of silver grains above any specific cell type or suborgan region. A set of serial sections from a mammary gland of the sheep at 1 week of lactation is shown in Figure 1. The low-magnification overviews (Figures 1A, 1C, 1E, and 1G) show that all alveoli expressing the αS1-casein-encoding gene are also expressing the ACC-α encoding gene, as transcribed from all three promoters. The higher magnifications (Figures 1B, 1D, 1F, and 1H) clearly show that the MECs are the site of ACC-α expression. Although in this example the lower left bundle of alveoli is devoid of αS1-casein messages, ACC-α-derived transcripts are clearly evident here when hybridized with the exon 5-derived probe detecting messages stemming from both promoters, PI and PII. This shows that αS1-casein and ACC-α gene expression are not strictly coupled, at least in the stage of early lactation.
Cells of the connective tissue and those lining the blood vessel (Figures 1B, 1D, and 1F) are clearly labeled with the probe of combined specificity for PI and PII, while the signals of the two other ACC-α specific probes appear either to be reduced (blood vessel) or absent (connective tissue).
MECs express transcripts initiated from all ACC-α-encoding promoters also at full lactation (Figure 2). The example shown was taken from bovine sections and is also representative of the images seen from sheep tissue at full lactation. There appears to be considerable variation in the intensity of ACC-α expression among even closely adjacent alveoli. In addition to the heterogeneity of milk protein gene expression associated with histologically obvious changes in alveolar morphology, hybridization for αS1-casein and α-lactalbumin reveals such substantial variation in the status of alveolar activity for these mRNAs also in cows and sheep alveoli that appear otherwise histologically similar (unpublished observations; and Molenaar et al. 1992, 1995). From the example shown, it appears that cells lining those ducts do not express the ACC-α (e.g., Figure 2, running from upper left to bottom). However, ACC-α expression of such cells may be readily seen in other ducts (not shown).
ACC-α expression may be uncoupled from αS1-casein gene expression also in stages of mammary gland differentiation (Figure 3). This can be clearly seen in sections taken from either pregnant or involution animals. The examples shown in Figure 3 were taken from cows but, again, are representative for sheep as well. It appears from the sections of the pregnant animal that some groups of alveoli are not yet prepared for αS1-casein gene expression (Figure 3D, arrow), while already well engaged in ACC-α gene expression (Figure 3A-3C). Likewise, we observe that, during involution ACC-α gene expression may eventually persist longer in residual alveolar regions than does the expression of the αS1-casein-encoding gene (Figures 3E-3H).
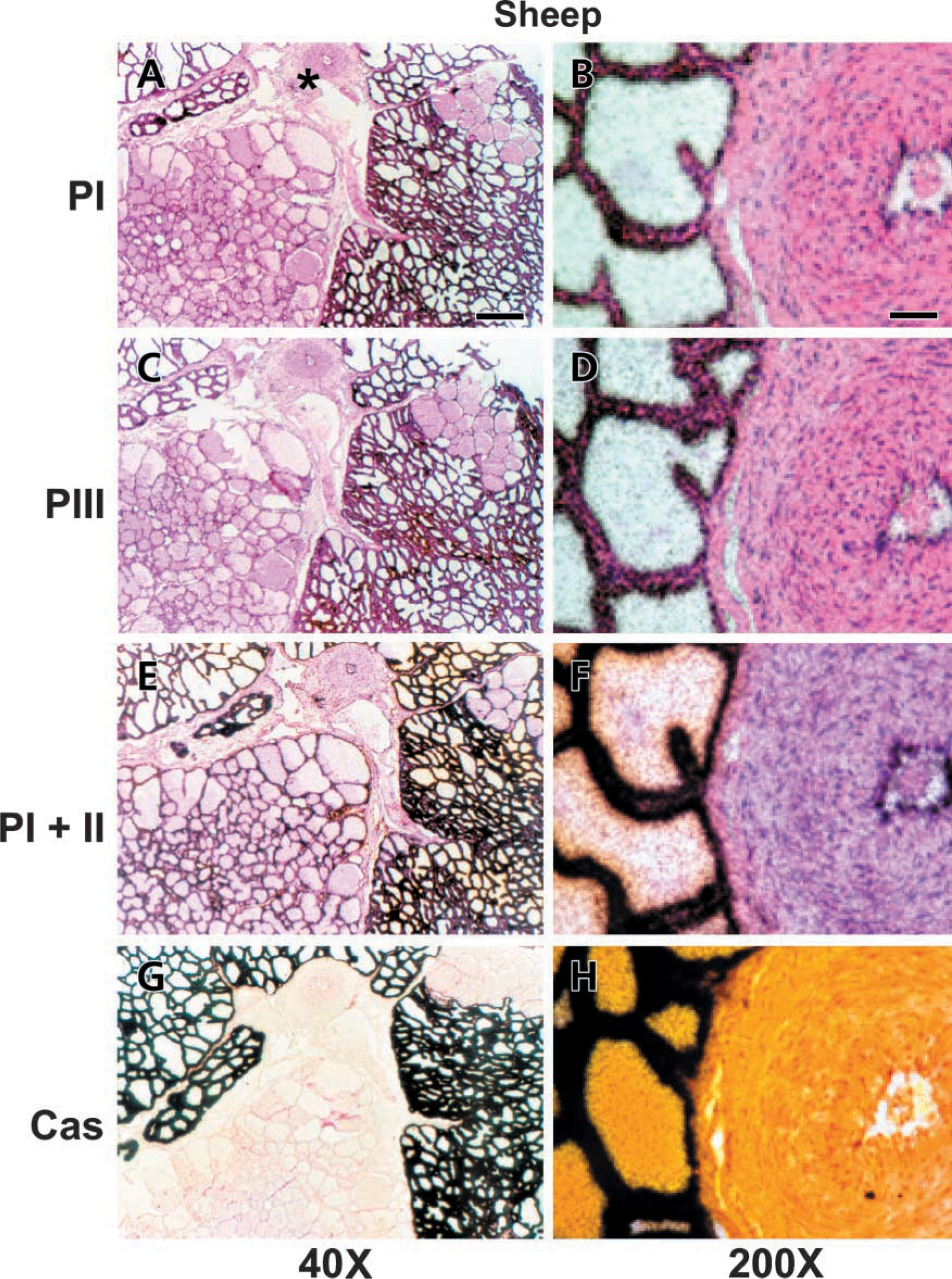
All three promoters are expressed in MECs of the lactating mammary gland of the sheep; Serial sections of the udder from an early lactating ewe were hybridized with antisense probes specific for transcripts expressed from PI (A,B), PIII (C,D), or from both PI and PII (E,F). The probe specific for αS1casein (G,H) reveals that the lower left bundle of alveoli was not actively synthesizing casein. The left panel (A,C,E,G) shows low-magnification overviews. Bar = 2.5 mm. The panel at right shows details photographed at higher magnification (B,D,F,H) around the central blood vessel (cf. A, at asterisk). MECs are clearly labeled by all probes. Bar = 0.6 mm. The three ACC-α-specific probes also give rise to silver grains above the endothelial cells lining the vessel, unlike the αS1-casein-specific probe.
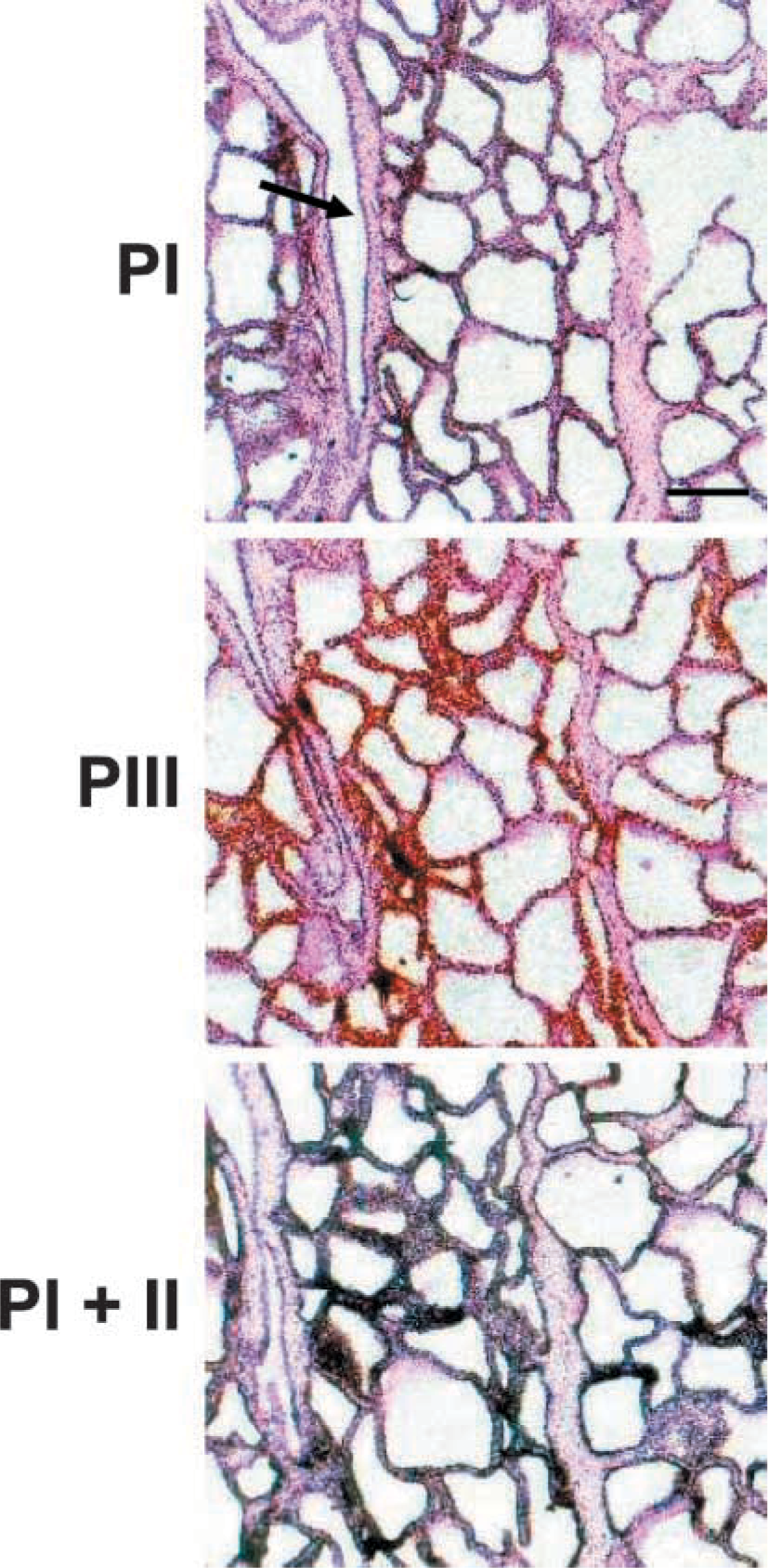
MECs express the transcripts from all three promoters at full lactation of the cow. Serial sections from the fully lactating udder of a cow, hybridized with the indicated antisense probes. Silver grains are always associated with the MECs, whereas the cells lining the small ducts (arrow) are devoid of label. Bar = 1 mm.
The images from the pregnant cow (Figures 3A-3D) show similar findings to those from the sheep at 1 week of lactation (cf. Figure 1). Specifically, as seen in sheep, it appears that adipocytes are most prominently labeled by the exon 5-derived probe, detecting messages initiated from either PI or PII (Figure 3C, inset). Neither the PI- nor the PIII-specific probe seems to give rise to specific signals in this area. The qualitative evaluation of the hybridization signals therefore augments the conclusion drawn already from sheep (Figure 1) that PII might be the relevant promoter for fatty acid synthesis in the adipocytes of the mammary gland. Likewise, the exon 5-specific probe gives rise to quite abundant silver grains over areas of connective tissue containing stromal cells during involution. Neither one of the other two promoter-specific probes reveals substantial mRNAs there.
The three different ACC-α-specific probes gave rise to hybridization signals of quite different intensities. Specifically, the signals from the exon 5-derived probe with the combined specificity for transcripts derived from PI and PII appeared to be much stronger than those from the other two probes. This makes it difficult to evaluate the relative abundances of transcripts across hybridizations. Therefore, we measured the density of silver grains at 10 different locations over selected cell types on any given slide to obtain an unbiased estimate of the relative abundance of the respective transcripts in different cell types. The analysis shows (Figure 4) that highly casein-expressing alveoli contain the highest concentrations of transcripts complementary to all three ACC-α-specific probes. Moreover, it becomes very clear that the expression of PIII is not restricted to MECs, but is also quite prominent in other cell types, particularly in the lactating udder of the cow (Figure 4A). Expression of PI, on the other hand, is clearly not restricted to adipose tissue, but is also quite prominent in blood vessels or cells lining the ducts. Finally, we note that the relative abundance of silver grains caused by the exon 5-specific probe above adipose tissue is not stronger than that caused by the other probes (Figure 4C and 4D, filled columns). The mere visual inspection of the hybridizations had provoked a different interpretation (cf. insets in Figures 3A-3C).
Recently, a lack of PI activity in the mammary gland of the sheep was reported (Travers et al. 2001). We conducted two experiments to examine the obvious discrepancy between this report and our very clear in situ images demonstrating that ovine PI initiated expression in udders. First, we conducted RT-PCR amplifications of PI-derived transcripts, using RNA from three lactating sheep. PI-derived transcripts are readily detected with this technique (Figure 5). Two amplification products were obtained, one comprising exons 1, 4, and 5 and the other one exons 1 and 5, as indicated by their sizes. This result confirms the 5′-UTR heterogeneity of PI-derived transcripts as known from all species examined thus far, including sheep (Travers et al. 2001) and cow (Mao et al. 2001). However, we noted a quantitative difference of the abundance of PI-derived transcripts between sheep and cow in this more qualitative assay.
We used real-time PCR to compare more precisely the abundance of PI-derived transcripts in lactating and pregnant udders of the sheep (Table 1). We assayed RNA preparations from lactating or pregnant sheep (three each) for the abundances of transcripts derived from PI and PIII. Moreover, we measured the relative abundance of all ACC-α-encoding transcripts from lactating ewes to approximately evaluate the contribution of PI-derived transcripts to all ACC-α-encoding messages in the lactating mammary gland of the ewe. We included a set of four samples from lactating cows to compare this experiment with previous reports (Mao et al. 2001). The data show that PI in sheep is only poorly stimulated by lactation (1.4-fold) and, significantly, contributes very little (2%) to the total amount of ACC-α-encoding transcripts, unlike in cattle (30%). Although these RT-PCR quantifications are not exactly comparable across different transcripts, due to different primer efficiencies, it is clear that the activity of PI contributes much more to the total amount of Acc-α-derived transcripts in the cow than in lactating sheep. Together, these data show that PI is indeed expressed in the lactating mammary gland of the ewe, but at lower levels than in cattle.
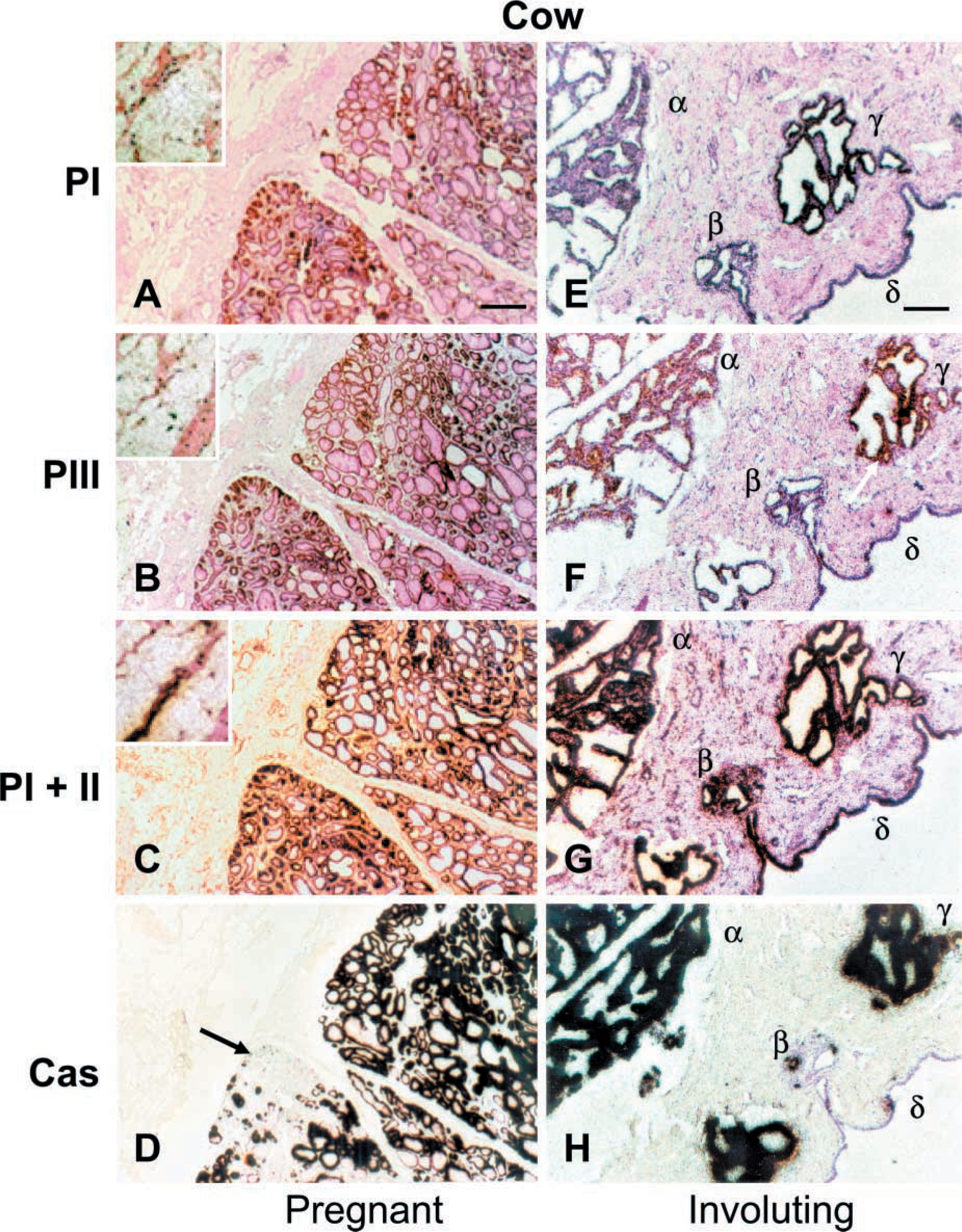
ACC-α and casein gene expression are not strictly coupled in MECs. Serial sections from a pregnant (A-D) or involuting (E-H) cow were hybridized with the different probes, as indicated. During pregnancy, some tightly packed alveoli not synthesizing αS1-casein (D, arrow) are expressing the ACC-α-encoding gene (cf. corresponding areas in A-C). Similarly, a disproportionate persistency of casein and ACC-α mRNA molecules is seen during involution (E-H). While only two (H, α, γ) from three of the denoted different areas of residual alveoli (α, β, γ) are still heavily labeled by the casein probe, all three areas are labeled by the ACCα-specific probe. However, while the cells lining the duct (δ) reveal a high abundance of transcripts hybridizing to the PI + PII-specific probe, comparable to the level seen in the alveolar area, transcripts hybridizing to either one of the other two ACC-α-specific probes are less well detected in the latter cells. Adipocytes are most prominently labeled by the PI + PII-specific probe (C, asterisk, and insets at higher magnification in A-Q, while the proportion of messages derived from PI or PIII appears greatly diminished in these areas. Likewise, the PI + PII-specific probe gives rise to an abundance of silver grains over the areas of stromal cells containing connective tissue during involution, while the other two ACC-α-specific probes reveal little or no mRNA here. Bar = 1 mm.
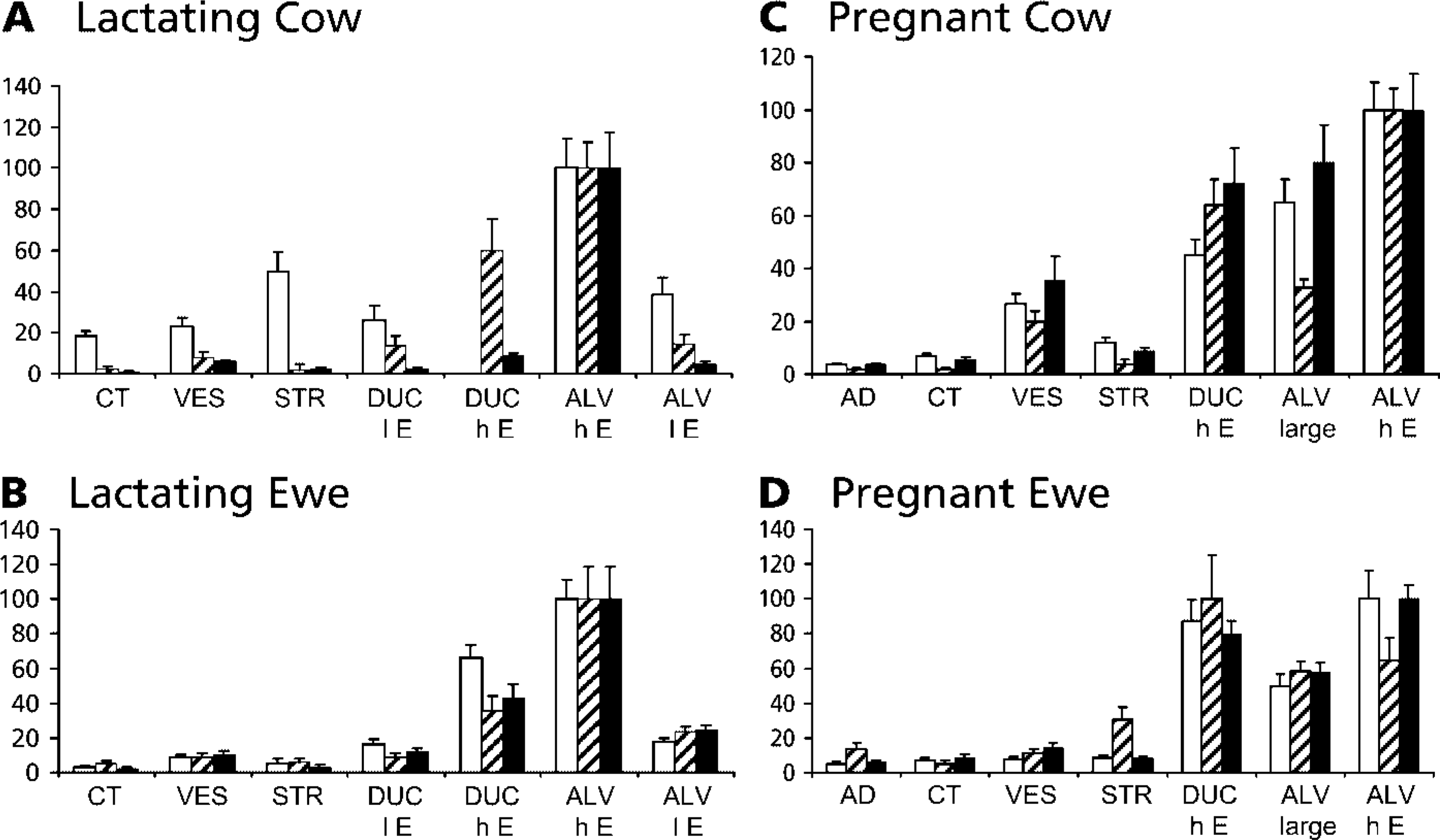
No clear cell type restriction is evident for any of the three ACC-α-specific promoters. The distribution of the relative ISH signal intensities across various tissues and cell types in the mammary gland sections was recorded from lactating (
Discussion
The significant result from our study is that all three promoters of the ACC-α-encoding gene are expressed in MECs at any developmental stage of the gland. This holds for both species examined, sheep and cattle. The data imply that none of these promoters must be neglected in analyzes aiming to disclose regulatory mechanisms governing the extent of milk fat synthesis, particularly in cattle. This conclusion could not have been drawn from previous data, which all used whole gland extracts to analyse ACC-α-derived gene expression. Previous work from other groups as well as our own suggests that milk fat synthesis might be regulated primarily either by PII in murine species (Ponce-Castaneda et al. 1991) or PIII in sheep and cattle (Barber and Travers 1998; Mao et al. 2002).
An evaluation of the relative contribution of any one of the ACC-α-expressing promoters to control the total abundance of ACC-α-encoding transcripts in the mammary gland at lactation leads to different conclusions for different species. In rat, the activity of PII only was found to increase about 10-fold with lactation (Ponce-Castaneda et al. 1991) and may hence be the relevant promoter there. The contribution of PI may be low in sheep because the concentration of PI-derived transcripts is low at any lactational stage and does not change from pregnancy to full lactation (Travers et al. 2001). Our quantitative data regarding ovine PI expression in the mammary gland (Table 1) support these conclusions. The situation is particularly complex in cattle. PI is strongly expressed in the bovine mammary gland, accounting for ∼30% of all ACC-α-encoding transcripts (Mao et al. 2001). This previous determination is fully confirmed by the measurements reported here using samples from a different set of cows (Table 1). The concentration of PI-derived transcripts increases about three fold during lactation (Mao et al. 2001), just in balance with the increased concentrations of all ACC-α-encoding messages. The same applies for PII-derived transcripts (Mao and Seyfert 2002). Because the bovine PIII expression was shown to be basically restricted to the lactating mammary gland (Mao et al. 2002), all three bovine promoters are relevant for the synthesis of milk fat.
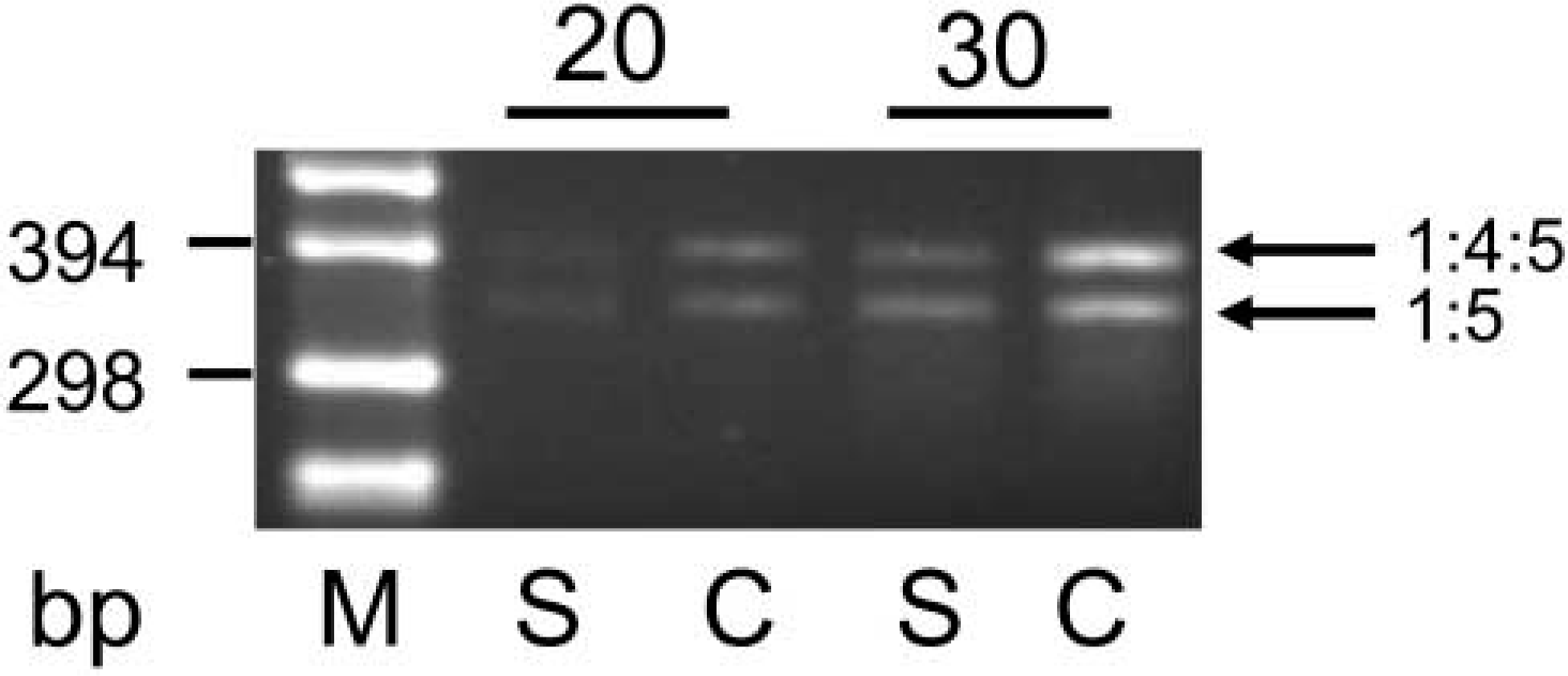
PI from sheep is expressed in the lactating mammary gland of the ewe, but not as strongly as in the cow. Equal amounts of total RNA from mammary glands of a lactating ewe or cow (Lanes S and C, respectively) were used as templates to amplify in RT-PCR transcripts derived from PI. Samples were removed after 20 or 30 amplification cycles and products resolved alongside a DNA size marker (Lane M). The product obtained from either species consists of two fragments, as expected. They represent transcripts derived from exons 1:4:5 or exons 1:5, as indicated by their different sizes (397 bp and 350 bp, as calculated from the DNA sequence files).
Semiquantitative evaluation of the differential contribution of PI and PIII to the total amount of ACCα transcripts in cows and sheepa
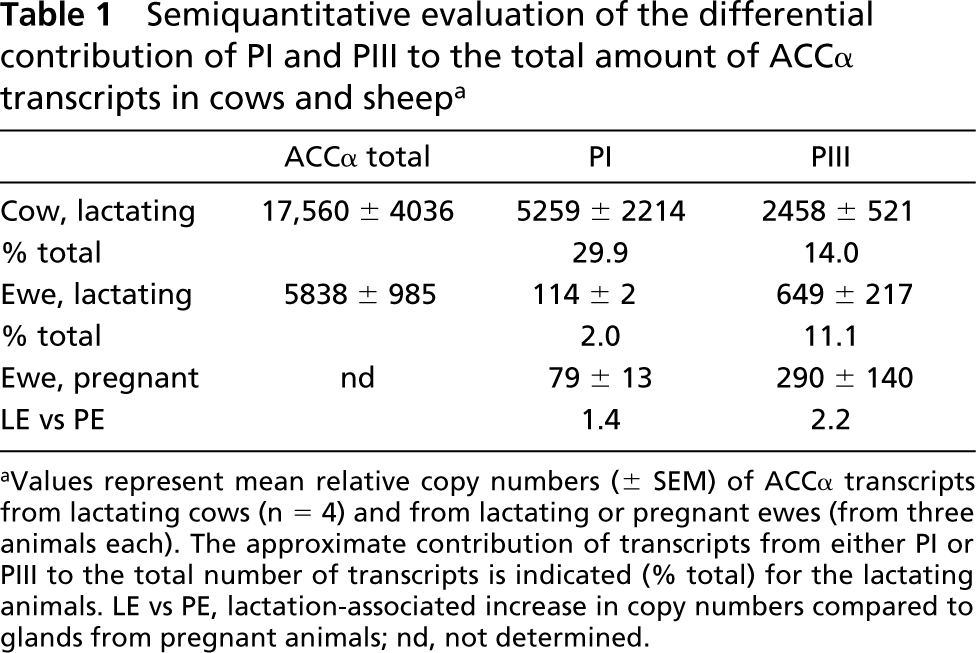
Values represent mean relative copy numbers (± SEM) of ACCα transcripts from lactating cows (n = 4) and from lactating or pregnant ewes (from three animals each). The approximate contribution of transcripts from either PI or PIII to the total number of transcripts is indicated (% total) for the lactating animals. LE vs PE, lactation-associated increase in copy numbers compared to glands from pregnant animals; nd, not determined.
Our data show that basically all promoters appear to be expressed in all cell types of the mammary gland. The visual inspection of the hybridizations had always revealed very strong signals where the exon 5 probe was applied, having specificity for transcripts from both promoters, PI and PII. Although this could have provoked the view that PII would be more widely expressed than the other two promoters, the photometrical determination of the signal intensities showed indeed that this is not the case (Figure 4). These measurements rather indicate that the ACC-α mRNA abundance, as measured from whole gland extracts, cannot be attributed to a specific cell type as single source or to the predominant activity of a specific promoter within a distinct cell type.
The ACC-α-encoding gene may be expressed in such alveoli and MECs that are not expressing the αS1-casein-encoding gene. This is evident in the sections taken at early lactation (Figure 1) but also from pregnant (Figure 3) or involuting cows or sheep (latter not shown). These qualitative data suggest that the temporal control of expression during lactational development of the gland may be tighter for the αS1-casein-encoding gene than for that of the ACC-α. Perhaps it would be more interesting to know if such an uncoupling of casein and fatty acid synthesis occurs within any one MEC at full lactation. This would have significant implications regarding the molecular control mechanisms coordinating casein and fatty acid synthesis within the MEC. However, this would require electron-microscopic analysis of ultrathin sections and cannot be resolved using our 7-μm sections as analyzed with 35S-labelled probes.
In conclusion, we have shown that all three ACC-α promoters are active in MECs of the mammary glands from cow and sheep and that ACC-α expression is not necessarily linked to that of major milk proteins. Our data together also reveal a striking difference between cattle and sheep regarding the relevance of PI for milk fat synthesis. Whereas the contribution of PI may be very limited in sheep, it contributes significantly to establishment of the final concentration of the ACC-α-encoding transcripts in the mammary gland of the lactating cow. This is a remarkable difference in the mammary gland-specific use of this promoter, which is evolutionarily truly conserved between cattle and sheep.
Footnotes
Acknowledgements
Supported by the Deutsche Forschungsgemeinschaft (DFG grant Se 326/10-2), ISTAD grant No 00-FRG-21-WHEE), and the New Zealand Foundation for Research Science and Technology.
