Abstract
Six distinct fibroblast growth factors (FGF5) have been detected in pancreatic islets by immunohistochemistry (IHC) using commercially available antisera. We show here that these antisera are useful for Western blotting but that only two are suited for IHC. By Western blotting, these antisera detect recombinant FGFs. Detection can be eliminated by preabsorption with immunizing peptide but not with irrelevant peptide. By IHC we find specific labeling of islets with anti-FGF1 and anti-FGF2 antisera. Labeling can be abolished by preabsorption with the immunizing peptides. In contrast, prominent staining of islets by anti-FGF4, −FGF5, −FGF7, and −FGF10 antisera is unspecific because the staining cannot be competed by preabsorption with the immunizing peptides.
F
We first tested whether the antisera could detect recombinant FGFs in Western blotting experiments and, if so, whether the signal could be competed out by preabsorption with the cognate immunizing peptide. We used goat anti-FGF1 (cat RDI-FGF1Cabg), goat anti-FGF2 (cat RDI-FGFBCbg), goat anti-FGF4 (cat RDI-FGF4Cabg), goat anti-FGF5 (cat RDI-MFGF5 Cabg), goat anti-FGF7 (cat RDI-FGF7Cabg), goat anti-FGF10 (cat RDI-FGF10Cabg), rabbit anti-human FGFR1 (flg) (cat RDI-FGFR1 abr), mouse anti-human FGFR2 (bek) (cat RDI-FGFR2abm), rabbit anti-human FGFR3 (cat RDI-FGFR3abr), and rabbit anti-human FGFR4 (cat RDI-FGFR4abr). All FGF and FGFR antibodies and corresponding peptides were purchased from Research Diagnostics Inc. (Flanders, NJ). Preabsorption of antisera was performed by incubating with immunizing peptide at a fivefold excess (w/w) in PBS supplemented with 0.25% BSA and 0.3% Triton X-100 for 1–3 hr at room temperature (RT) on a rocking platform. Western blotting was performed using precast 15% SDS-polyacrylamide gels (Bio-Rad; Hercules, CA). Samples containing 50 ng recombinant human FGF protein (R&D Systems; Minneapolis, MN) were denatured for 5 min at 90C in a 20-μl volume in 1 × formamide gel loading buffer [62.5 mM Tris-HCl (pH6.8), 13% sucrose, 2% SDS, and 0.005% bromphenol blue]. After electrophoresis proteins were transferred to a Hybond nitrocellulose membrane and washed three times for 5 min in TBST [50 mM Tris-HCl (pH 7.5), 150 mM NaCl, and 0.1% Tween-20]. Thereafter the membrane was blocked in TBST with 2% skimmed milk and 2% BSA for 1 hr. Goat anti-FGF antibodies were used at a dilution of 1:200 in blocking buffer as suggested by the manufacturer. Membranes were incubated overnight at 4C, followed by three washes in TBST for 5 min and 1 hr of blocking. Secondary HRP-conjugated antibody was used at a dilution of 1:2000 in blocking buffer and was incubated with the membrane for 1 hr at RT. The membranes were washed three times in TBST for 5 min and again twice for 5 minutes in TBS (TBST without Tween-20). The blots were developed using an ECL+ Western blotting detection system (Amersham Pharmacia Biotech; Poole, UK). As shown in Figure 1, all the tested anti-FGF antisera recognized the relevant recombinant FGF and the signal could be blocked by preabsorption with the immunizing peptide but not with an irrelevant peptide. Therefore, all the tested antisera are capable of specific detection of the cognate antigen in Western blotting experiments. We also conclude that the immunizing peptides are able to block the antigen recognition sites of the antibodies.
We next tested whether the same antisera could detect a specific signal by IHC staining of sections of adult mouse pancreas. Pancreatic tissue from adult mice was fixed in 4% PFA, sucrose-protected, and 4-μm frozen sections were cut on a cryostat. FGF and FGFR antibodies were tested at a range of dilutions and were used at dilutions of 1:20 (anti-FGFs) or 1:200 (anti-FGFRs). Guinea pig α-insulin (diluted 1:500) and mouse α-glucagon (diluted 1:50) were both from Novo Nordisk (Bagsvàrd, Denmark). Cy2- or Texas Red-conjugated secondary antibodies raised in donkey were purchased from Jackson ImmunoResearch Laboratories (West Grove, PA). Preincubation of antisera with immunizing peptide was performed as described for Western blotting, except that a 20-fold excess (w/w) of peptide was used. Before addition of antibody, the sections were rinsed three times in PBS and blocked in 10% donkey serum (Jackson ImmunoResearch). Antibodies diluted in PBS supplemented with 0.25% BSA and 0.3% Triton X-100 were added to each slide and incubated overnight. Before addition of secondary antibody, the sections were washed in PBS three times for 5 min and incubated with secondary antibody for 45 min, followed by three 5-min washes in PBS. Before pictures were taken the sections were mounted in fluorescence mounting media (Kierkegaard; Gaithersburg, MD). Negative controls included stainings without primary antibody and were blank, as were stainings incubated with immunizing peptide without primary antibody. As shown in Figure 2, all six antisera labeled the islets of Langerhans but only the signal obtained with the anti-FGF1 and anti-FGF2 antisera could be competed out by preabsorption with the immunizing peptides despite the fact an amount fourfold higher than what was able to eliminate binding in Western blotting experiments was used. Therefore, whether FGF4, FGF5, FGF7, and FGF10 are truly expressed in islet β-cells cannot be determined by use of these antisera for IHC staining. We therefore conducted RT-PCR analysis of FGF expression in RNA prepared from isolated islets and used RNA isolated from E12 whole embryos and NIH3T3 cells as positive control. cDNA synthesis and RT-PCR analysis were prepared as described in Jensen et al. (1996), with the modifications described in Jensen et al. (2000). In reactions with 24 cycles of amplification, G6PDH was used as internal standard, whereas in reactions with 27 cycles TBP was used. The reactions were analyzed by polyacrylamide gel electrophoresis and visualized using a PhophorImager (Molecular Dynamics; Sunnyvale, CA). The primers used were (upstream, downstream): FGF1 (300 bp), 5′-CAACGGGGGCCACTTCTT-3′, 5′-CAGCTCCCGTTCTTCTTG-3′; FGF2 (150 bp), 5′-CCACCAGGCCACTTCAAGG-3′, 5′-AACTCCTCTCTCTTCTGC-3′; FGF4 (166 bp), 5′-ACGCGGCACGCAGAATTG-3′, 5′-CGATGCCCACGTTGCAGT-3′; FGF5 (230 bp), 5′-TCTCACTCCCGAAGGGCAA-3′; 5′-AGATCTGCAGATGGAAACC-3′; FGF7 (200 bp), 5′-GGAGCAAACGGCTACGAGT-3′, 5′-CCACGGTCCTGATTTCCAT-3′; FGF10 (190 bp), 5′-TCACCTGCCAACTCTTGGT-3′, 5′-CCGTTCTTCTCAATCGTGAG-3′; FGFR1b (278 bp), 5′-GGGCAGCAATGTGGAGTTCA-3′, 5′-TTGCCACAGGTCTGGTGACA-3′; FGFR1c (250 bp), 5′-GGGCAGCAATGTGGAGTTCA-3′, 5′-GTGATGGGAGAGTCCGATAG-3′; FGFR2b (244 bp), 5′–GGAGGGGATGTGGAGTTTGT-3′, 5′–ACTGGTTGGCCTGCCCTATA-3′; FGFR2c (271 bp), 5′-GGAGGGGATGTGGAGTTTGT-3′, 5′-CAGAACTGTCAACCATGCA-3′; FGFR3b (180 bp), 5′-TCAGTGAGAATGTGGAGGCA-3′, 5′-TGCCAGCCTCATCAGTTTCC-3′; FGFR3c (170 bp), 5′-TAACACCACCGACAAGGAGC-3′, 5′-TGCCAGCCTCATCAGTTTCC-3′; FGFR4 (190 bp), 5′-CGTGGTCGTCACTGGTACAA-3′, 5′–TGATGGAGGTTAAGGAGTC-3′. As shown in Figure 3, we could detect expression of FGF1, FGF2, FGF4, and FGF5, but not FGF7 and FGF10, in islet RNA. Identical results were obtained from two independent RNA preparations. However, because even the purest pancreatic islet isolates are contaminated by neurons, endothelial cells, fibroblasts, smooth muscle cells, blood cells, and pancreatic duct and acinar cells, we cannot conclude that the RT-PCR signal is derived from islet β-cells. We also cannot rule out that the RT-PCR signals are derived from β-cells, but high-resolution in situ hybridization or IHC detection with suitable antisera is required to determine that with certainty.
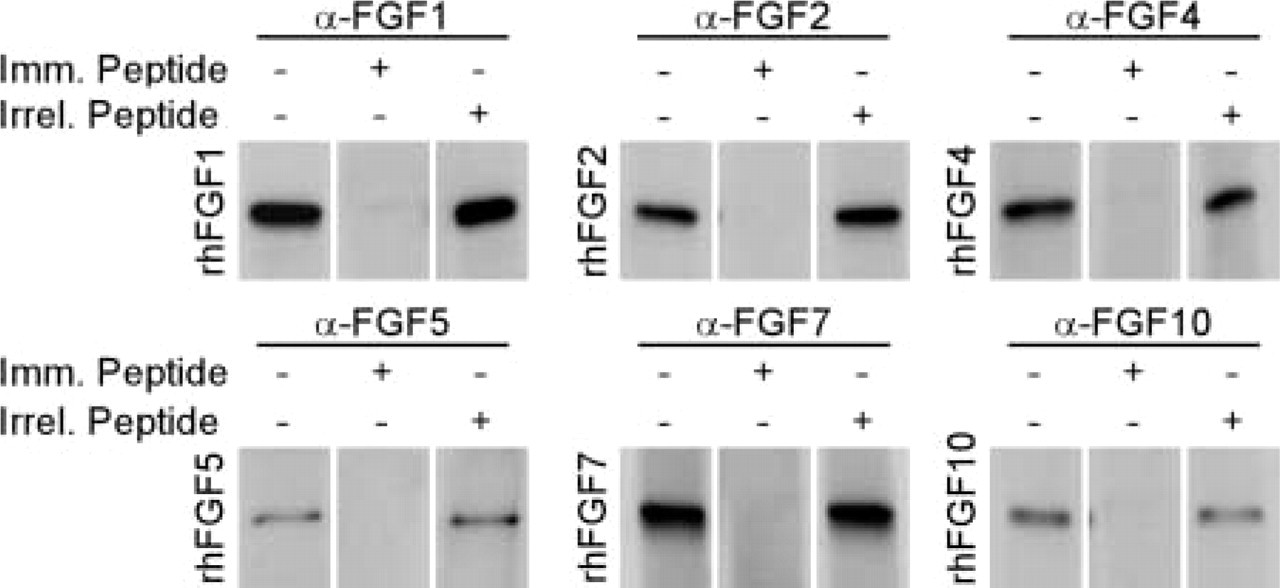
Western blotting analysis of α-FGF antisera. FGF antibodies recognize recombinant human proteins (50 ng per lane) in Western blotting analysis, and the interaction can be blocked by preincubation with a fivefold excess (w/w) of immunizing peptide (1mm. Peptide), whereas the same amount of an irrelevant peptide (Irrel. Peptide) did not influence the recognition by the antibody of the recombinant protein.
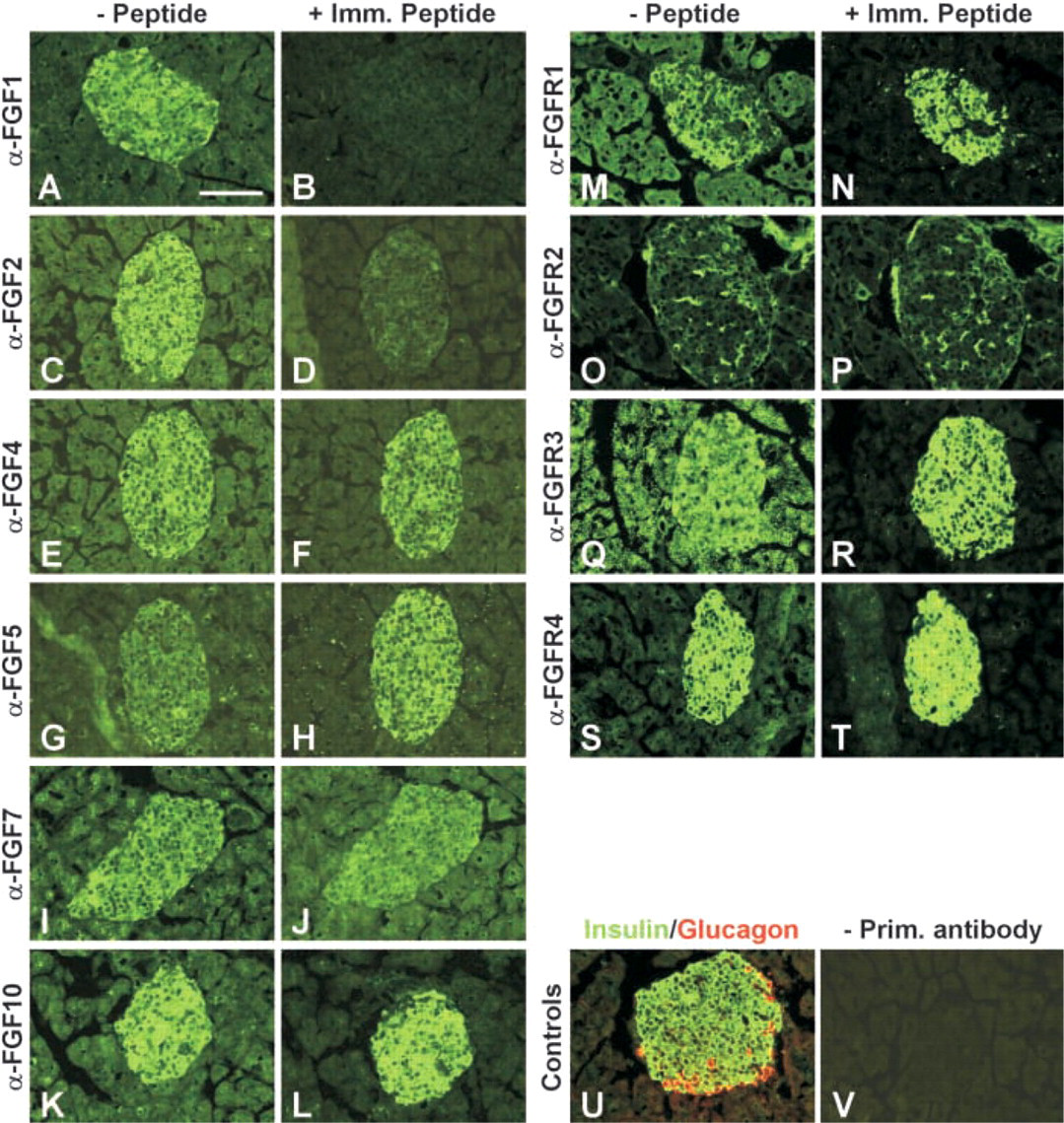
Immunostaining for FGFs and FGFRs on adult mouse pancreatic islets. IHC analysis of cryostat sections of adult mouse pancreas using FGF and FGFR antisera. Left panels (- Peptide;
We also assessed the expression of FGF receptors by RT-PCR on RNA from isolated islets. As shown in Figure 3, we could detect expression of FGFR1b, FGFR1c, FGFR2b, FGFR2c, FGFR3b, and FGFR4. Again, we cannot conclude which of the many cell types present in the islet preparations give rise to these bands. We attempted to determine if islet β-cells expressed any of the FGF receptors by IHC, but, as shown in Figure 2, even though islet β-cells were labeled by antisera against FGFR1, FGFR3, FGFR4, and endothelial cells were labeled by FGFR2 antiserum, we could not compete the labeling with the immunizing peptides. Therefore, we cannot conclude with any certainty from these data that a particular cell type in the pancreas expresses a given FGF receptor.
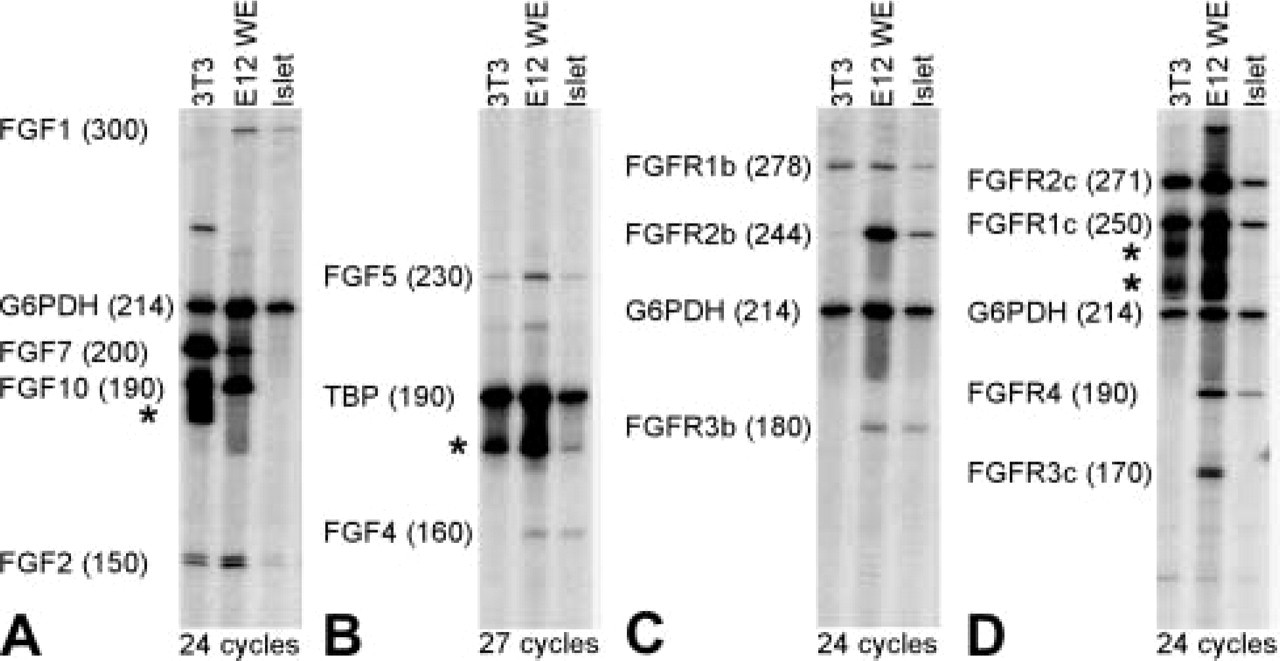
RT-PCR analysis of FGF and FGFR expression in adult mouse pancreatic islets. RT-PCR performed on cDNA isolated from NIH3T3 cells, E12 whole embryo, and isolated islet preparations. G6PDH or TBP were used as internal control. (
Our results could have implications for the autocrine FGF signaling loop among β-cells proposed by Hart et al. (2000). If an autocrine pathway exists, our results suggest that an FGF ligand, expressed in β-cells, still needs to be identified. The two FGFs that we can state are expressed in β-cells with any degree of certainty, FGF1 and FGF2, are unlikely be the responsible ligands. Both FGF1 and FGF2 can activate FGFR1c, but mice deficient for either or both of these factors have not been reported to display a diabetic phenotype (Miller et al. 2000). Closer examination of such mice might reveal a diabetic phenotype, but this remains to be shown. Evidently, other FGF ligands might be expressed in β-cells but this still needs to be demonstrated. Until such a ligand is identified, an indirect mechanism is equally plausible for the diabetic phenotype displayed by FRID1 mice.
Footnotes
Acknowledgements
Supported by funds from the Juvenile Diabetes Research Foundation and the Danish National Research Academy.
We thank Ragna Jørgensen and Helle V. Petersen for expert technical assistance.
