Abstract
We describe a versatile method for performing fluorescence in situ hybridization (FISH) in suspension instead of on a slide as usually done. This so-called suspension-FISH (S-FISH) opens new possibilities for the analysis of shape and functions of the human interphase nucleus. The procedure is described and the first results using this approach are presented.
F
Here we present for the first time a technique in which the entire FISH procedure is performed on a cell suspension and the cells are placed on a polished concave slide as the final step of the procedure, just before the evaluation. We call this procedure suspension-FISH (S-FISH) and show that it is possible to do 3D analyses on totally spherical interphase nuclei or even on 3D metaphases (Figure 2).
The S-FISH method is performed as follows:
The cell suspension is centrifuged for 10 min at 700 rpm, the supernatant is discarded, and the pellet is re-suspended in 500 μl of methanol.
Centrifuge between each step mentioned below, always using the same conditions for centrifugation as previously mentioned: 10 min, 700 rpm. Work at room temperature unless otherwise stated.
Discard 450 μl of the supernatant, add ∼450 μl 0.9% NaCl, and incubate for 3 min.
Replace the supernatant by 450 μl pepsin solution [i.e., 475 μl H2O, 25 μl 0.2 N HCl, 0.005% pepsin (Fluka 77163; Buchs, Switzerland)] and incubate for 5 min at 37C.
Replace the supernatant by ∼450 μl 1 X PBS and incubate for 90 sec to remove the pepsin.
Replace the PBS by 70% formamide/2 X SSC and incubate for 1 min.
Add 25 μl 50% formamide/2 X SSC containing the probe and 20 μg of the blocking DNA (i.e., COT1). Three times the usually applied amount of the corresponding probe is used, i.e., for centromeres 60 ng and for whole chromosome paints 3 μg. Only directly labeled probes have been used thus far. Do not centrifuge but go on with the next step.
Denature for 5–7 min at 80C and hybridize for 12–16 hr at 37C in a thermocycler. Finish the hybridization by adding 600 μl of 0.4 SSC (68C). Incubate for 2 min and centrifuge.
Replace the supernatant by 600 μl 1 X PBS at RT for 2 min.
Replace the supernatant by 600 μl 4 X SSC/0.2% Tween at RT for 2 min.
Re-suspend the pellet in 50 μl of antifade (e.g., Vectashield; Vector, Burlingame, CA); add 0.4 μg DAPI and place the solution on a slide [special slides, e.g., epoxy-covered ones (Roth L195.1) are best suited], cover with a coverslip, and evaluate with a fluorescence microscope. Before evaluation pre-cool the slide at −20C to make the antifade solution more viscous so that the nuclei do not move during image assessment.
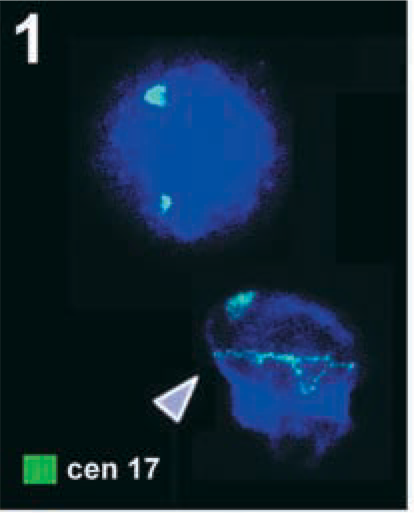
Two normal interphase nuclei from peripheral blood, added to a slide using the usual air-drying method after hybridization with a centromeric probe for chromosome 17 (cen 17, labeled in green) are depicted. Whereas the upper nucleus shows the usual hybridization pattern, the lower has one normal and one abnormal (arrowhead) green signal. The latter is not spot-like but is stretched artificially over the entire nucleus due to the flattening process. Images for Figures 1–3 were captured with the ISIS digital FISH imaging system (MetaSystems; Altlussheim, Germany) using a XC77 CCD camera with on-chip integration (Sony).
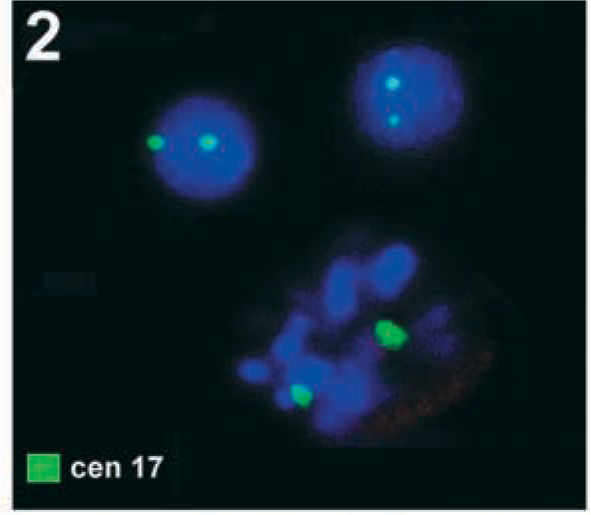
Result of S-FISH using the same centromere-specific probe for chromosome 17 (cen 17) as in Figure 1. Two interphase nuclei plus one partial metaphase showing two signals each are visible. By chance, all six signals were within the focus.
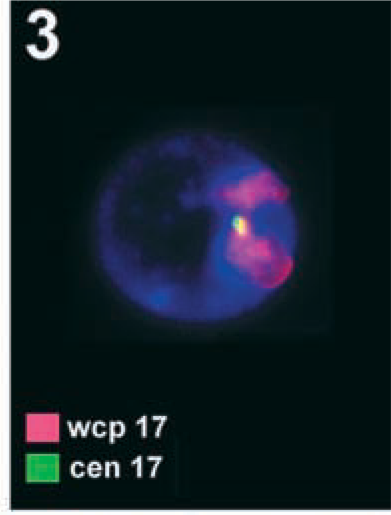
S-FISH result using two differently labeled probes. The interphase nucleus shows two red signals for the entire chromosome stained specific for chromosome 17 (wcp 17), but only one specific green signal of centromere-specific probe for chromosome 17 (cen 17) is depicted. The other was present but is out of focus.
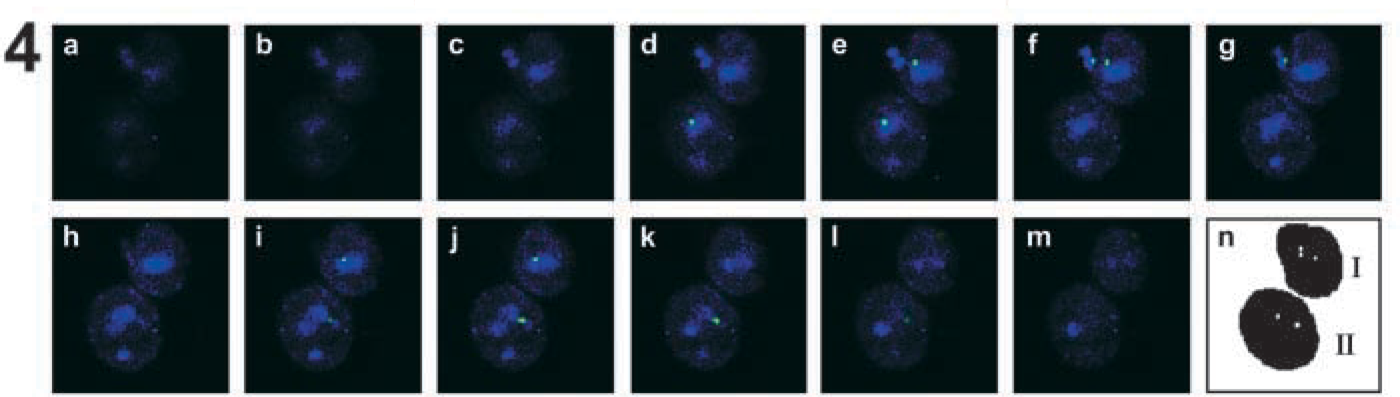
S-FISH results taken with the Zeiss Laser Scanning Microscope LSM 510 META. These are 5-μm optical sectionings of two interphase nuclei hybridized with a centromere-specific probe for chromosome 17 (cen 17) (
The artificial flattening of interphase nuclei using the air-drying method can lead to very peculiar results when interphase diagnostics are performed. Even though the single nucleus shown in Figure 1 (arrowhead) is a very extreme example, such or similar nuclei are well known to any molecular cytogeneticist. It is therefore clear that interphase analyses with the goal of learning about interphase architecture performed on such nuclei may not reflect the in vivo situation. Therefore, we developed the S-FISH approach, which makes it possible to analyze totally spherical interphase nuclei.
The S-FISH approach has been tested successfully on 10 different chromosomal suspensions using one-to five-color FISH experiments. The evaluation was done with a Zeiss Axioplan 2 microscope (Figures 2 and 3) or with a Zeiss Laser Scanning Microscope LSM 510 META (Figure 4). By these experiments it could be demonstrated (a) that is in principle possible to perform the complete FISH procedure in suspension, (b) that it is not necessary for evaluation of the interphase nuclei that they be fixed (and flattened) on the slide surface, and (c) that even metaphases can be analyzed three-dimensionally in principle by S-FISH (Figure 2). At present there is still the disadvantage that a three-fold probe volume has to be used compared to the normal FISH procedure. However, new horizons for analyses of the “real” interphase architecture are now open.
Footnotes
Acknowledgements
Supported by the Herbert Quandt Stiftung der VARTA AG, the Wilhelm Sander-Stiftung (99.105.1), and the EU (ICA2-CT-2000–10012 and QLRT-1999–31590). The continuing support of the Carl Zeiss GmbH (Jena, Germany) is gratefully acknowledged.
