Abstract
Sphingosine 1-phosphate (S1P), which derives from the metabolism of sphingomyelin, is mainly synthesized, stored, and released from platelets after activation by physiological and pathophysiological events. S1P acts in cardiovascular tissues through cell surface G-protein-coupled receptors of the endothelial differentiation gene (EDG) family, i.e., EDG1, EDG3 and EDG5. The aim of the present study was to assess the precise distribution of EDG1, EDG3, and EDG5 receptors expressed in human cardiovascular tissues to investigate their respective physiological implication. When assessed by Northern blots, EDG1, EDG3, and EDG5 displayed wide expression levels in decreasing order, respectively. In particular, EDG3 was mainly detected in the aorta. Detailed analysis by in situ hybridization (ISH) and immunohistochemistry (IHC) revealed strong EDG1 expression in cardiomyocytes and in endothelial cells of cardiac vessels. In cardiomyocytes, the EDG1 receptor is likely to be co-expressed with EDG3 and EDG5, although EDG1 exhibits the most prominent expression pattern. Unlike EDG3 and EDG5, which are expressed in the smooth muscle cell layer of the human aorta, no signal corresponding to EDG1 expression could be detected in the aorta. Moreover, only EDG3 expression was also found in smooth muscle cells of cardiac vessels. The present results provide new insight into the expression pattern of S1P receptors in human cardiovascular tissues, indicating a differential pattern of expression for these receptors in human vessels.
S
Tissue expression of the EDG1, EDG3, and EDG5 receptors in mice indicates that heart and lung have the highest overall expression of these genes (Zhang et al. 1999; Ishii et al. 2001), whereas EDG6 and EDG8 receptors are mainly expressed in lymphoid (Gräler et al. 1998) and brain tissues (Im et al. 2000), respectively. The intense expression of EDG1, EDG3, and EDG5 receptors in the mouse heart, the essential role played by EDG1 receptor in vessel formation (Liu et al. 2000), the release of S1P from activated platelets (Yatomi et al. 1995), and its accumulation in certain areas of the myocardium raise the possibility that sphingolipids, and more particularly, S1P may be, through activation of seven specific transmembrane receptors, a key contributor to physiological cardiac regulation. It has recently been proposed that EDG-mediated S1P negative inotropic and cardiotoxic effects may play important roles in acute myocardial ischemia. S1P was shown to increase the influx of extracellular calcium, which then causes calcium overload and impairs cardiac function (Nakajima et al. 2000). Interestingly, it was also recently shown in rat that S1P induces cardiomyocyte hypertrophy, mainly via the EDG1 receptor (Robert et al. 2001).
Although it is believed that EDG receptor family members could mediate many S1P effects in heart cell function, the cellular mechanisms underlying these effects are still largely unknown. Cardiac tissues are composed of diverse cell types, notably including cardiomyocytes, endothelial cells, fibroblasts, and smooth muscle cells of blood vessels. EDG1, the first S1P receptor cloned from endothelial cells, was suggested to be involved in the differentiation of endothelial cells (Hla and Maciag 1990) and was shown to be expressed in rat cardiomyocytes (Nakajima et al. 2000; Robert et al. 2001). EDG3 and EDG5 mRNA have been detected in rat smooth muscle cell lines (Tamama et al. 2001). EDG1, EDG3, and EDG5 messengers were detected in human heart and atrial cardiomyocytes by Northern blotting (An et al. 1998) or RT-PCR analysis (Himmel et al. 2000), respectively. However, these expression studies used cell cultures or whole-heart homogenates and were not focused on the in situ cell type-specific localization of the different S1P receptors, especially in human cardiac tissues.
The aim of the present work was to assess the precise distribution of EDG1, EDG3, and EDG5 receptors in human cardiovascular tissues to investigate their possible physiological implication. We first compared EDG1, EDG3, and EDG5 mRNA expression patterns in different regions of the heart and in aorta by Northern blotting as well as EDG1, EDG3, and EDG5 protein detection in whole heart, aorta, and smooth muscle by Western blotting analysis. We next determined the precise localization of EDG1, EDG3, and EDG5 expression in heart, aorta, and cardiac vessels by ISH or IHC.
Materials and Methods
Northern Blotting
Human cardiovascular MTN blots (Clontech; Palo Alto, CA) containing blotted RNA from normal human cardiovascular tissues were used: 21 specimens (age 21–30 weeks; cause of death spontaneous abortion) for male fetal heart; four specimens (age 19–54 years; cause of death trauma) for male adult heart; 50 specimens (age 18–75 years; cause of death trauma) for male adult aorta; 15 specimens (age 18–88 years; cause of death trauma) for male adult apex of heart; 39 specimens (age 18–72 years; cause of death trauma) for male adult left atrium; 38 specimens (age 18–72 years; cause of death trauma) for male adult right atrium; 11 specimens (age 18–61 years; cause of death trauma) for male adult left ventricle; 20 specimens (age 18–70 years; cause of death trauma) for male adult right ventricle. The blots were pre-hybridized for 30 min at 65C in 5 ml of Express-Hyb buffer (Clontech), then hybridized for 2 hr at 65C in the same buffer containing denatured α[32P]-dCTP-labeled probe (EDG1, EDG3, or EDG5) at 2 × 106 cpm/ml. The filters were then washed for 40 min at room temperature (RT) in 50 ml of 2 × SSC, 0.1% SDS, and for 40 min at 50C in 50 ml of 0.2 × SSC, 0.1% SDS. Each blot was stripped and then re-hybridized with glyceraldehyde-3-phosphate dehydrogenase (G3PDH) probe for normalization. The relative quantification of EDG1, EDG3, and EDG5 expression was performed by densitometric analysis of the film using 1D Image Analysis Software (Kodak Digital Science; Rochester, New York). Results were normalized using the ratio of EDG probe levels to G3PDH probe levels.
EDG1-specific Peptide Synthesis and Polyclonal Antibody Production
The protein sequence of the human EDG1 receptor was analyzed for highly antigenic regions using the Jameson-Wolf antigenic index. A synthetic peptide was selected at the C-terminal intracellular tail. A homology search was performed to check for sequence redundancy with other proteins. As expected, the peptide sequences had very high homology with EDG1 receptors from different mammalian species. Homologies with other mammalian proteins were not significant, particularly with other EDG receptor family members. The following synthetic peptide (amino acids 350–367 FSRSKSDNSSHPQKDEGDN) was synthesized, purified and conjugated as described in detail elsewhere (Robert et al. 1994). After immunization of rabbits according to a standard protocol, the immunoglobulin fraction from the anti-EDG1 immune serum was obtained by affinity chromatography on protein A-Sepharose.
Western Blotting
For Western blotting, 50 μg of total protein (Protein Medley; Clontech) was used from normal human heart, aorta, and smooth muscle from small intestine. Proteins were extracted by the supplier from whole aortas pooled from 49 males/females (ages 15–65; cause of death trauma or sudden death), small intestine smooth muscle pooled from 16 males/females (ages 16–50; cause of death trauma or sudden death) and whole heart from a 44-year-old white man (cause of death trauma). Western blots were subjected to SDS-PAGE after polyvinylidene difluoride (PVDF) membrane blotting (Biotrace PVDF; Pall-Gelman, Ann Arbor, MI). EDG1 receptor protein was detected using a purified polyclonal antibody (Ab729; 0.5 μg/ml) and an immunogenic synthetic peptide (10 μg/ml) as competitor. EDG3 and EDG5 receptor proteins were analyzed using commercial monoclonal antibodies (MAbs) with high interspecies homology (0.1 μg/ml) (Antibody Solutions; Palo Alto, CA). The molecular weights were estimated by using a prestained protein standard mixture (Biolabs; Beverly, MA). For EDG1 detection, the membrane was developed for only 15 sec using an ECL(+) kit (Amersham-Pharmacia Biotech; Poole, UK), whereas 45 min was necessary for EDG3 and EDG5.
In Situ Hybridization
Probe Synthesis. RNA probe synthesis was carried out with an RNA transcription Kit (Stratagene; La Jolla, CA). Plasmids (pBluescript; Stratagene) containing EDG1, EDG3, and EDG5 cDNA were linearized to give rise to the anti-sense and sense probes. One μg of the linearized DNA template was incubated for 2 hr at 37C in a solution containing 1 × transcription buffer, dithiothreitol (30 mM), rATP (0.4 mM), rGTP (0.4 mM), rCTP (0.4 mM), α[35S]-UTP (5 μCi/μl), RNase inhibitor (1.6 U/μl), and RNA polymerase (0.4 U/μl) T7 (sense) and T3 (antisense). The DNA templates were then digested with RQ-1 DNase (10 U) for 15 min at 37C. After incubation, 10 μg yeast tRNA was added to the sample. Probe isolation was achieved on a Sephadex G50 column. After precipitation the probe was dissolved in hybridization mix (50% formamide, 0.3 M NaCl, 20 mM Tris-HCl, pH 8.5, 5 mM EDTA, 10% dextran sulfate, 1 × Denhardt's solution, 0.5 μg/μl yeast tRNA, and 10 mM DTT) at a final concentration of 2.104 cpm/μl and stored at — 70C until hybridization.
Slide Treatment. Paraffin sections were obtained from Novagen (Hybrid Ready Tissues; Madison, WI). Human adult normal tissue sections originated from male donor aorta and heart tissues, ages 26–36 (cause of death trauma). Tissues were removed, fixed in 4% paraformaldehyde, embedded in paraffin, sectioned at 5 μm (manufacturer's protocol), and stored at RT. Radioactive ISH was performed as previously described (Mazurais et al. 1999). Comparisons between EDG1, EDG3, and EDG5 were performed on tissues from the same donor. After treatment with xylene (three times for 5 min) to remove paraffin and rehydration through an ethanol series, sections were rinsed in 0.85% NaCl and phosphate buffer (PBS 0.1 M, pH 7.4), postfixed in 4% paraformaldehyde, and then treated with hydrochloric acid (0.2 M) and proteinase K (40 μg/ml) diluted in TE buffer (50 mM Tris-HCl, 5 mM EDTA, pH 8). After a rinse in PBS, the sections were refixed in 4% paraformaldehyde to stop proteinase K activity and then rinsed in PBS before acetylation with acetic anhydride (0.25% in triethanolamine 0.1 M, pH 8). Finally, sections were rinsed in distilled water before dehydration through an ethanol series. After air-drying, tissue sections were hybridized under coverslips with radioactive probe diluted in hybridization buffer (2.104 cpm/μl) overnight at 55C in a humid chamber. After hybridization, coverslips were removed in 5 × standard saline citrate (1 × SSC = trisodium citrate 15 mM, NaCl 150 mM, pH 7), 10 mM DTT at 55C for 15 min and the slides were washed (30 min) with 2 × SSC, 50% formamide, 10 mM DTT at 65C. After a rinse (10 min) in NTE buffer (10 mM Tris-HCl, 0.5 mM NaCl, 5 mM EDTA, pH 8), sections were treated with RNase A (20 μg/ml in NTE) for 30 min at 37C. Slides were than washed in NTE for 15 min at RT and incubated for 30 min in 2 × SSC, 50% formamide, 10 mM DTT at 65C. Before autoradiography, the tissues were rinsed in 2 × SSC and 0.1 × SSC at RT and dehydrated in an ethanol series containing 0.3 M ammonium acetate. Autoradiography was performed by dipping the slides in Ilford K5 nuclear track emulsion and exposing the slides in the dark for 14 days at 4C. After development, the sections were counterstained with toluidine blue and mounted in Pertex (Microm; Francheville, France). The photographs presented in Figures 3–7 are representative of three independent experiments.
Immunohistochemistry
Human paraffin sections were obtained from Novagen (Hybrid-Ready Tissues). Normal human adult tissue sections originated from donor heart tissues, ages 20–65 (cause of death trauma). Tissues were removed, fixed in 4% paraformaldehyde, embedded in paraffin, sectioned at 5 μm (manufacturer's protocol), and stored at RT. After treatment with xylene and rehydratation through an ethanol series and PBS, the specimens were heated in target retrieval solution, pH 6 (Dako; Glastrup, Denmark), 40 min at 95C and then cooled for 20 min on the bench. After two washes of 2 min in TBS solution (Dako), sections were incubated for 1 hr at RT with anti-EDG1 primary antibody at 2 μg/μl. Use of commercially available EDG3 and EDG5 antibodies (Antibody Solutions) did not obtain suitable results. For control slides, competition was performed by overnight incubation of the primary antibody with a related peptide used to generate the anti-EDG1 rabbit polyclonal antibody. After washes in PBS, the goat anti-rabbit secondary antibody and the tyramide signal amplification peroxidase IHC detection kit (Dako) were used according to the manufacturer's instructions with diaminobenzidine as a substrate. After hematoxylin counter-staining, sections were analyzed on a Nikon (Eclipse E800, Sony camera DXC-950P) microscope. The photographs in Figures 3 and 5 are representative of three independent experiments.
Results
EDG1, EDG3, and EDG5 Receptor Expression in Heart and Aorta
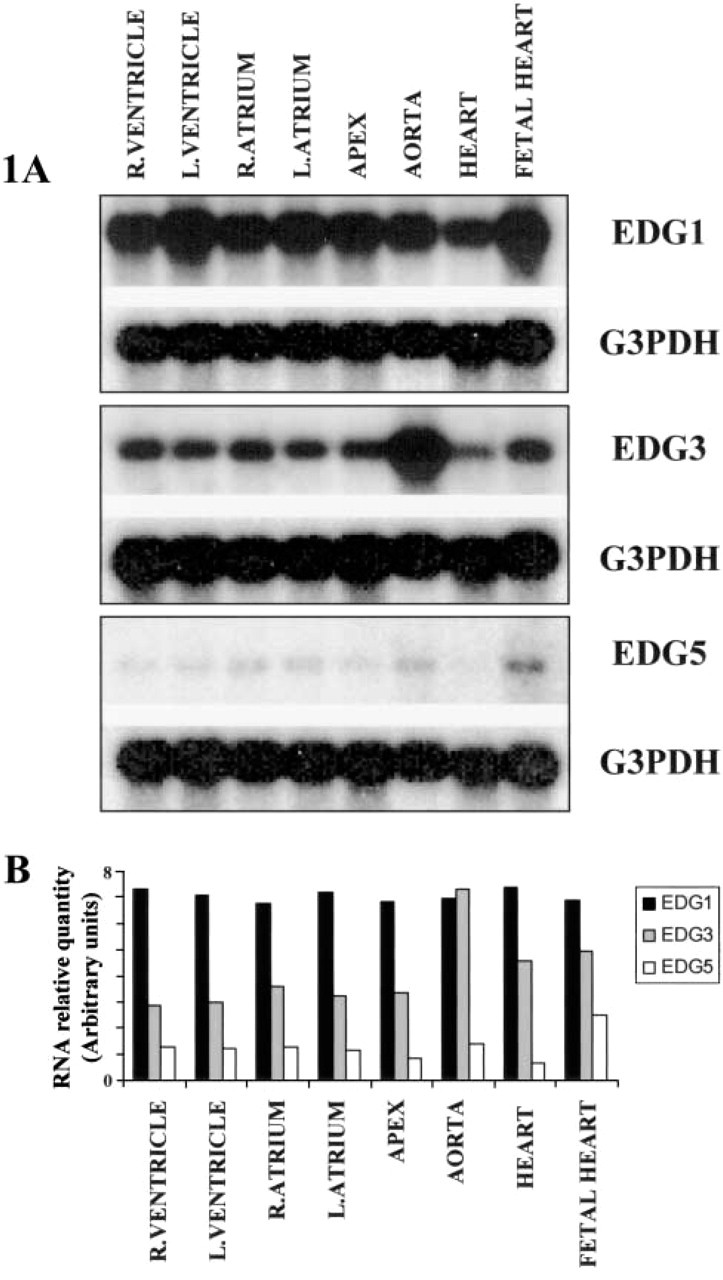
To determine the comparative mRNA expression patterns of EDG1, EDG3, and EDG5 receptor genes in different regions of human heart, Northern blots were probed at high stringency with EDG1, EDG3, and EDG5 cDNA probes corresponding to their respective coding regions (Figure 1). The 3.0-kb EDG1 mRNA was strongly and equally expressed in the different compartments (right and left ventricle, right and left atrium, apex) of the adult human heart, in the whole fetal heart, and in aorta. In contrast, the ~4.6-kb EDG3 mRNA was more abundant in the aorta than in heart regions (Figure 1B). Finally, EDG5 receptor transcript is weakly and equally expressed in the different regions of the heart and in aorta.
Western blotting experiments were performed to detect EDG1, EDG3, and EDG5 proteins in adult human heart, aorta, and smooth muscle of small intestine (Figure 2). It is worth noting that a long film exposure of 45 min was necessary to visualize EDG3 and EDG5 signals from total proteins loaded. Conversely, only 15 sec was sufficient to detect the EDG1 receptor. Considering that each antibody exhibits proper affinity, it is difficult to compare relative abundance at the protein level of each receptor in different tissues. The 38-kD EDG1 protein was detected at a high level in aorta and heart as well as in smooth muscle of small intestine. Similarly to results obtained by Northern blotting, the EDG3 45-kD protein was abundant in the aorta and smooth muscle compared to adult human heart. EDG5 receptor protein was equally present in whole heart, aorta, and smooth muscle.
(
Localization of EDG1, EDG3, and EDG5 Transcripts and Proteins in Human Heart
Full-length riboprobes of human EDG1, EDG3, and EDG5 receptors were used for ISH on normal human tissue sections. In parallel, IHC using an anti-EDG1 antibody was performed. Commercially available anti-EDG3 and anti-EDG5 antibodies (Antibody Solutions) did not allow successful IHC studies.
In the left ventricle, EDG1 mRNA was expressed in cardiomyocytes (Figures 3A and 3B) which also homogeneously displayed EDG1-immunoreactive signal (Figure 3E). It is interesting to note that the strong density of labeling (Figure 3B) was detected around cardiomyocyte nuclei because of the larger amounts of mRNA in this area. The expression pattern was remarkably consistent from one experiment to the other, and the specificity of the signal was confirmed by the absence of signal on the control slides, either with sense probe (Figures 3C and 3D) or polyclonal antibody preincubated with immunogenic EDG1 peptide as competitor (Figure 3F). ISH experiments using EDG3 (Figures 4A and 4B) and EDG5 (Figures 4E and 4F) probes indicated lower expression of the respective messengers in cardiomyocytes, compared to EDG1. Absence of any significant signal on the control slides demonstrated the specificity of this labeling (Figures 4C, 4D, 4G, and 4H). Because cardiomyocytes are densely packed, it was difficult to ascertain whether all cells were labeled, but a large majority displayed silver grains with the three different probes. As a consequence, EDG1, EDG3, and EDG5 are most likely co-expressed in a large number of cardiomyocytes. Both in atrium and septum, expression of EDG1, EDG3, and EDG5 could also be detected in cardiomyocytes (data not shown). Although it is difficult to estimate a precise distribution between different areas of the myocardium, such as inner, middle, outer myocardium, interstitium, or endocardium, by ISH or immunocytochemistry, no differences in gene expression or protein detection were identified for each receptor independently. In addition, localization of EDG1, EDG3, and EDG5 expression by ISH and IHC were revealed at the base of a growing tricuspid valve. These experiments revealed the expression of EDG1, EDG3, and EDG5 mRNA in the myocardium but absence of labeling in the fibrosa (data not shown).
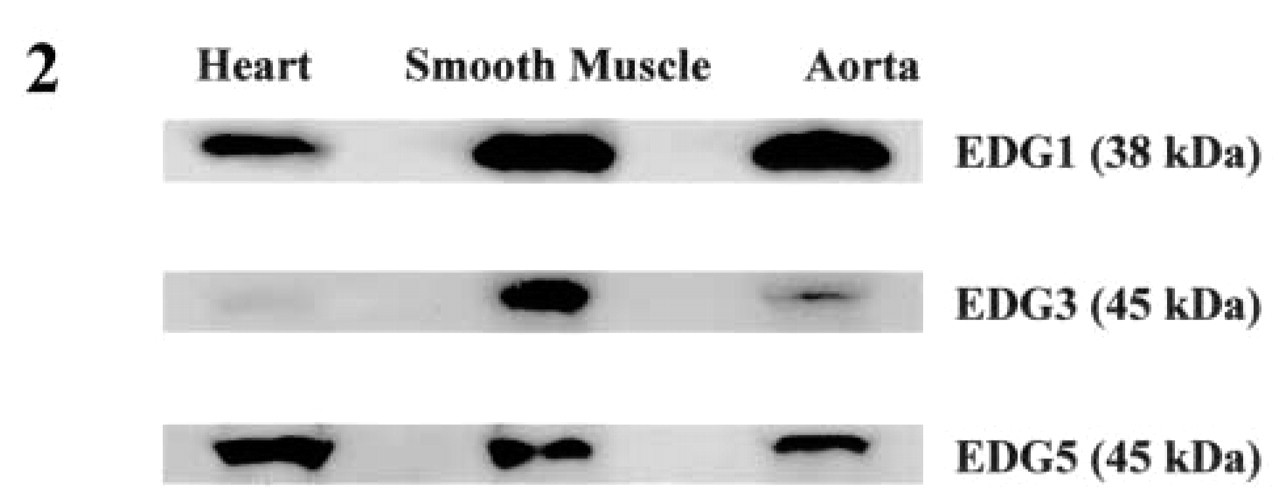
Detection of EDG1, EDG3, and EDG5 proteins in human whole heart, aorta and smooth muscle by Western blotting. Analysis of EDG1, EDG3, and EDG5 was performed using 50 μg of total proteins (Protein Medley; Clontech) of human heart, aorta, and smooth muscle. For EDG1 detection, the membrane was developed for only 15 sec using an ECL(+) kit (Amersham-Pharmacia Biotech), whereas 45 min was necessary for EDG3 and EDG5.
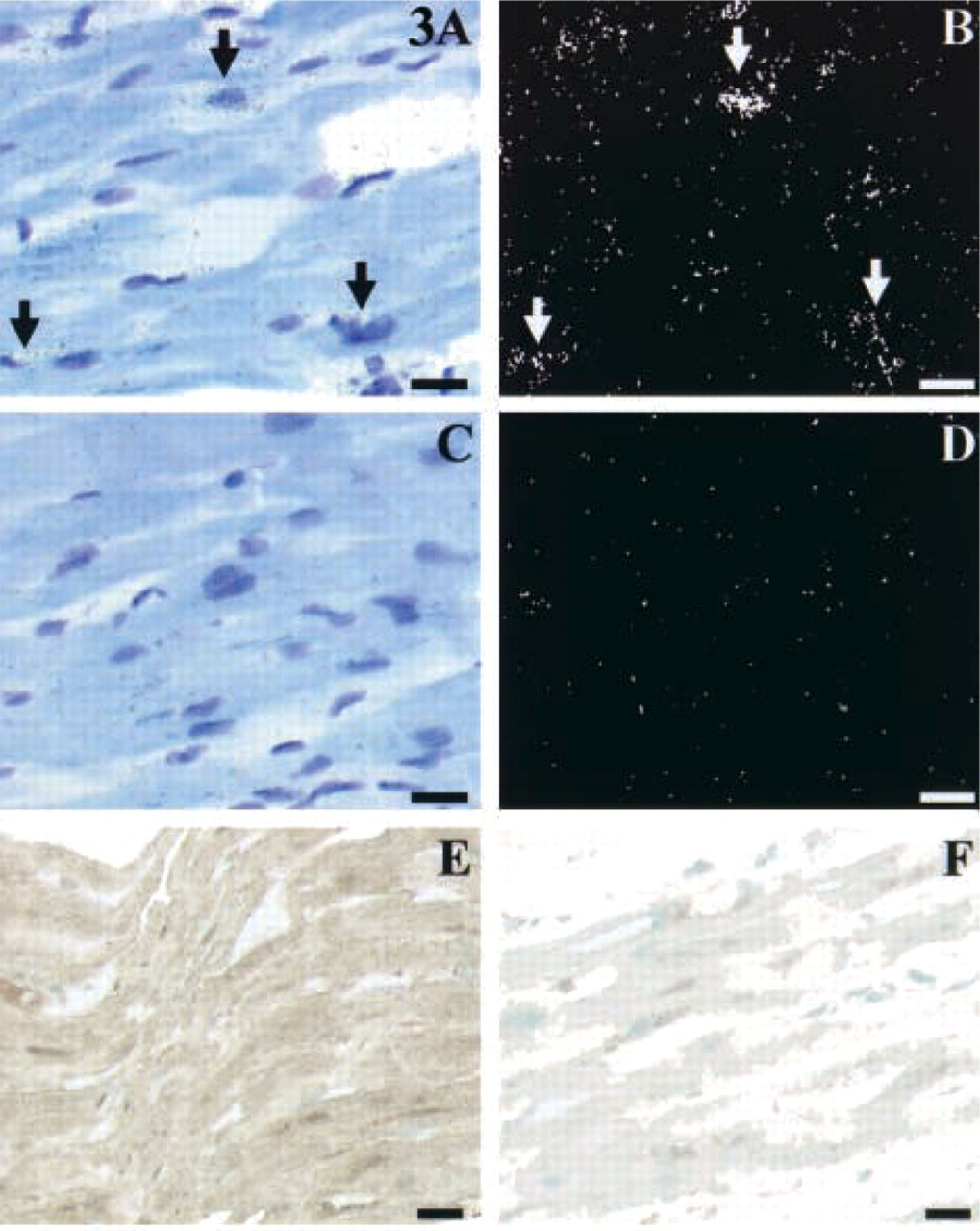
EDG1 expression in cardiomyocytes of the left ventricle. (
Strong radioactive labeling corresponding to EDG1 mRNA localization was detected in the endothelial cells of coronary arteries in the left ventricle (Figures 5A and 5B). The nature of the endothelial cell layer was confirmed by IHC analysis using an antibody raised against factor VIII as an endothelial cell lineage marker (data not shown). IHC study confirmed the localization of EDG1 protein limited to the endothelial cell layer of vessels (Figure 5E). No specific signal could be detected in the media and adventitia of vessels. The signal specificity was confirmed by the absence of signal on the control slides, either with the sense probe (Figures 5C and 5D) or with the anti-EDG1 antibody preincubated with immunogenic EDG1 peptide as competitor (Figure 5F). Endothelial cells of atrial and septal vessels were shown by ICC and ISH to express the EDG1 receptor (data not shown). Special attention was focused on the differential expression of EDG1, EDG3, and EDG5 mRNA in aorta and in cardiac vessels. In agreement with the Northern blotting experiment, EDG3 and EDG5 were expressed in the smooth muscle cells of the aortic media layer (Figures 6A–6F), whereas expression of EDG1 was not detected in smooth muscle cells of human aorta. In addition, microscopic analysis of EDG1, EDG3, and EDG5 expression was performed on serial tissue sections at the level of a small left ventricular coronary (Figure 7). This study showed that, unlike EDG1, which is expressed in the endothelial cells (Figures 7A–7C), EDG3 mRNA was largely distributed in the smooth muscle cells of the coronary vessel (Figures 7D–7F). No specific EDG3 signal could be detected in the endothelial cells. The specificity of the signal was confirmed by the absence of labeling on the control slides using the sense probe (data not shown). Regarding EDG5, in contrast to the smooth muscle cell layer of human aorta, no significantly positive cells could be detected in the different layers of the coronary vessel walls (Figures 7G–7I).
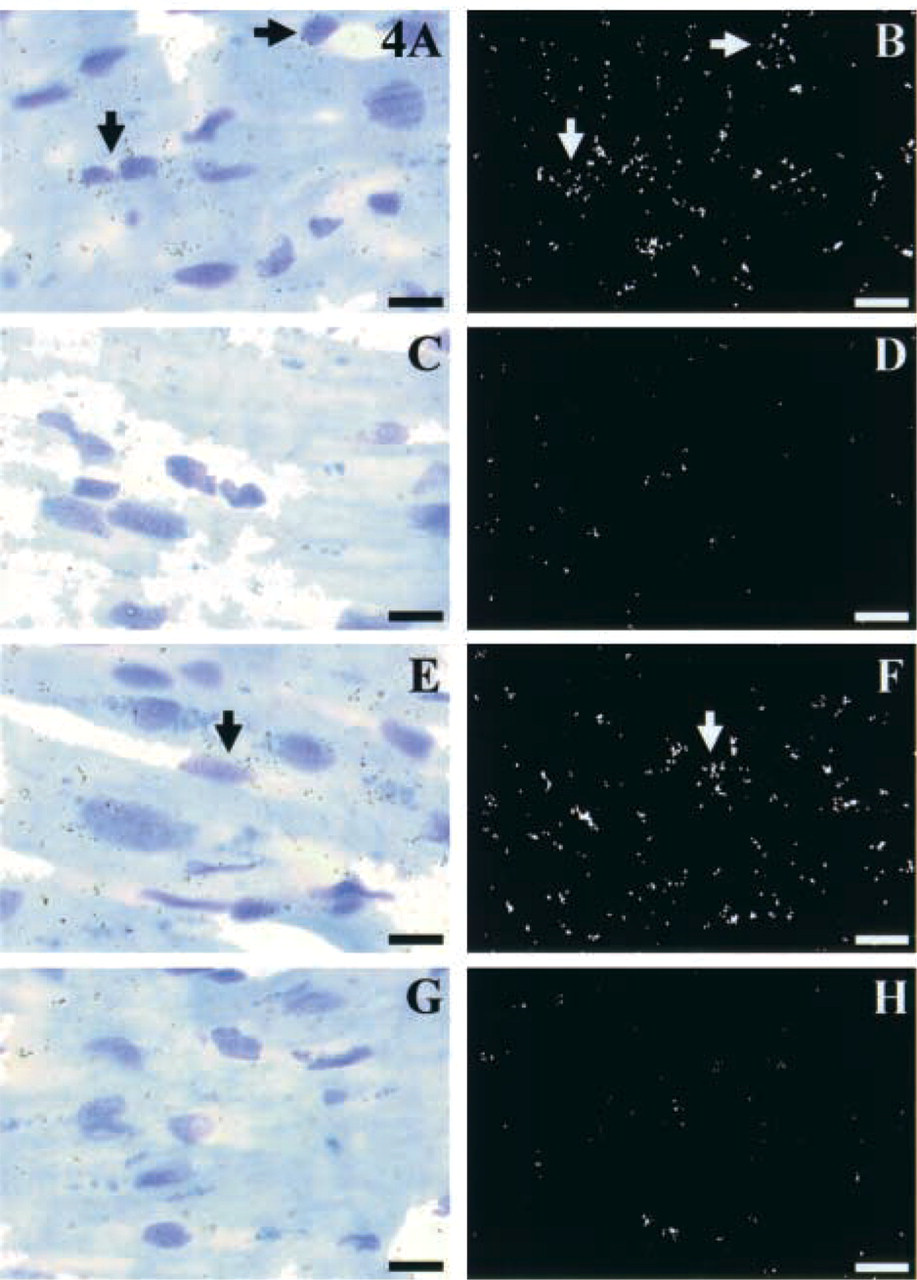
EDG3 and EDG5 expression is detected in cardiomyocytes of the left ventricle (arrows). Light microscopy of ISH with EDG3 (
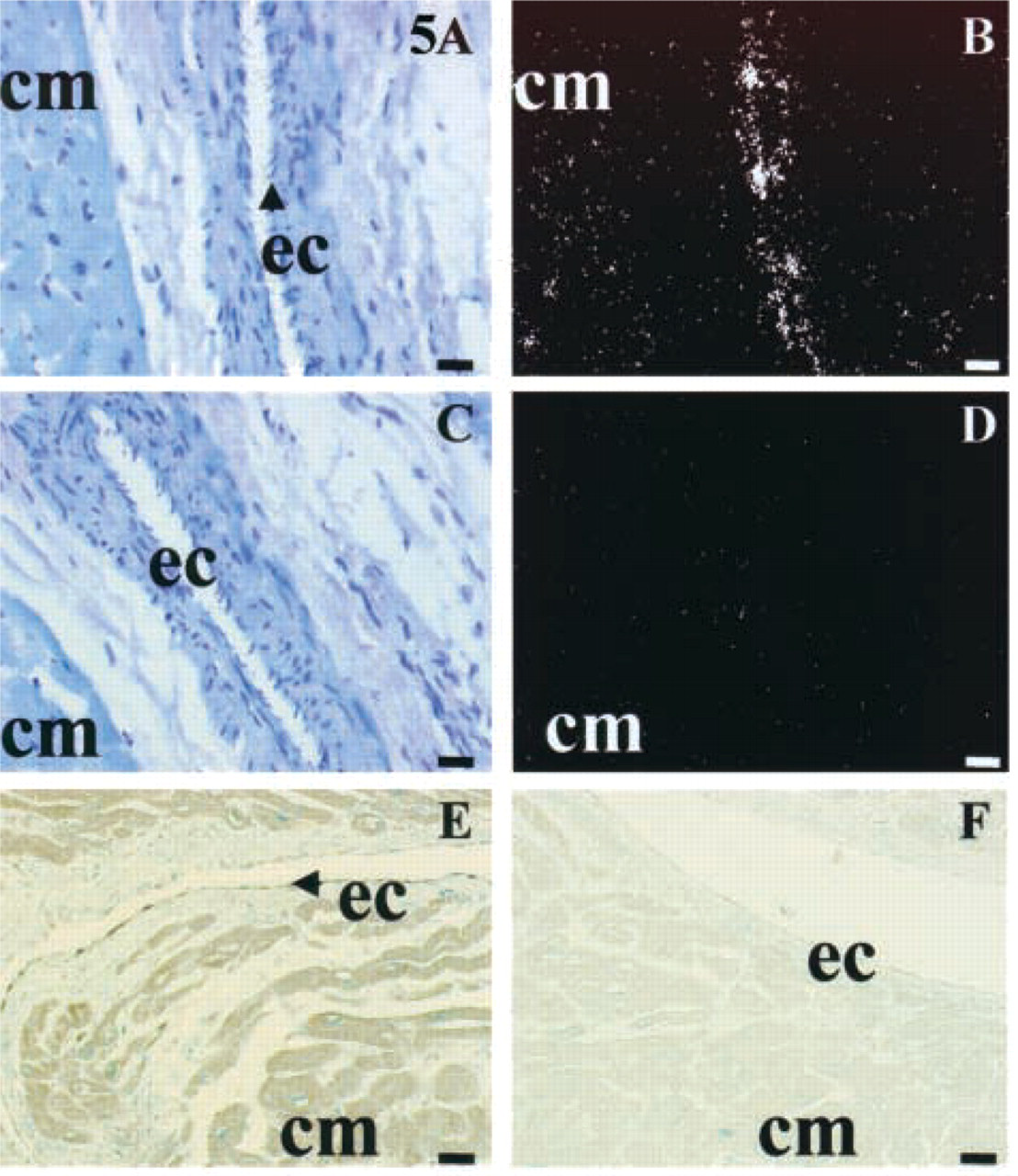
EDG1 expression in coronary arteries of the left ventricle. (
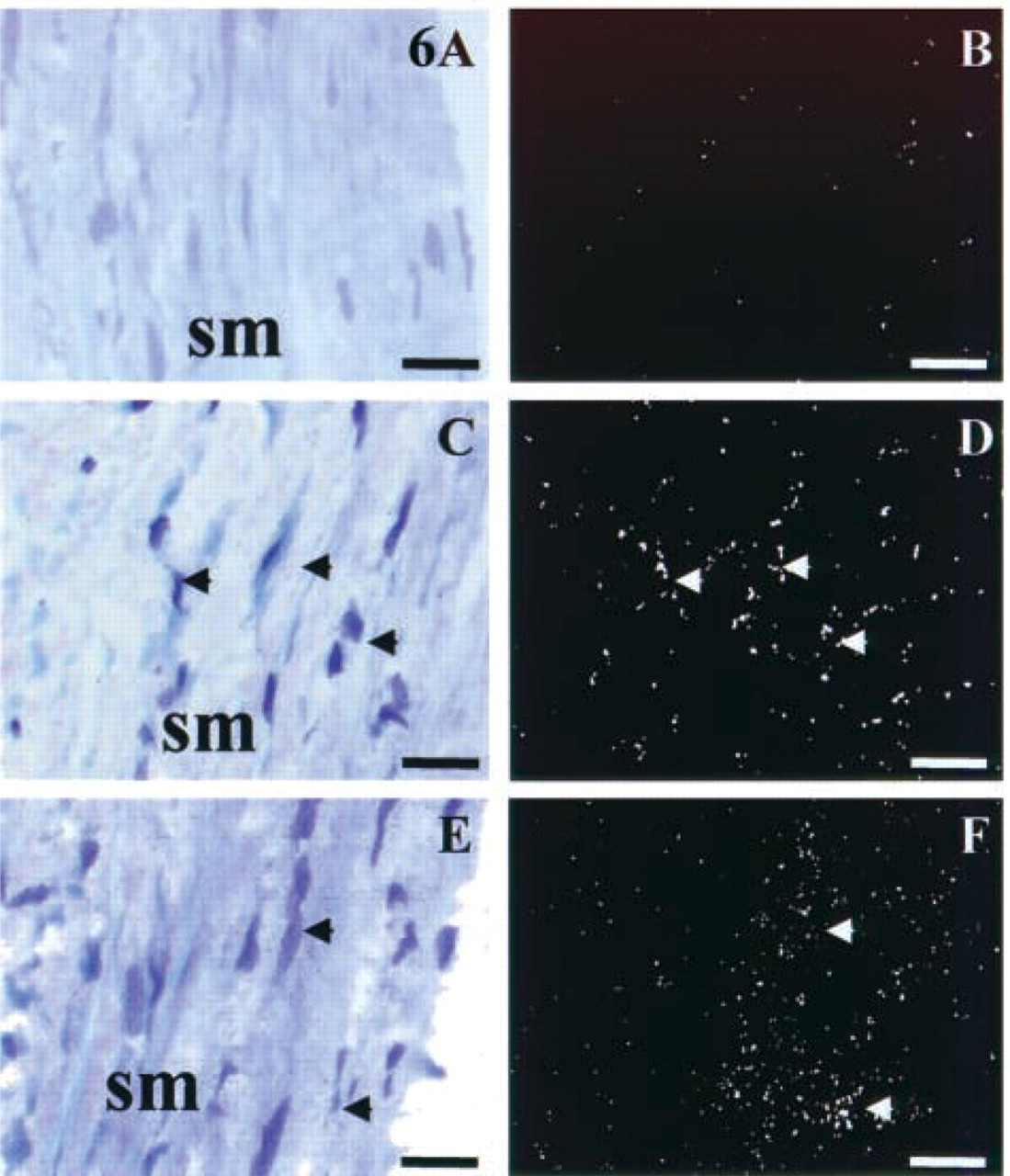
Differential patterns of S1P receptor expression in smooth muscle cells of human aorta. Light microscopy results of ISH with EDG1, EDG3, and EDG5 probes on adjacent aortic tissue sections are represented in
Discussion
This study documents the precise distribution and compares the relative expression patterns of EDG1, EDG3, and EDG5 genes in normal human cardiovascular tissues. Our data provide, for the first time, detailed information on the nature and the localization of the cardiac cells that express the S1P receptor at both the mRNA and the protein level.
First, Northern blots from human subjects reveal that EDG1, EDG3, and EDG5 receptors are expressed in both adult and fetal heart. More precisely, they are expressed in all anatomically distinct subregions of the adult heart, i.e., the apex, left and right ventricles, left and right atria, and the aorta. For each EDG independently, Northern blotting analysis reveals additional information on the relative amount of each receptor transcript in those cardiac regions. Whereas EDG1 mRNA is equally and strongly expressed in different parts of the heart and in aorta, the EDG3 transcript is more abundant in aorta than in the different compartments of the heart. Finally, EDG5 transcript is ubiquitously but weakly expressed in the heart and aorta in comparison with EDG3 and EDG1, except in fetal samples. These results are confirmed at the protein level by Western blotting experiments. However, it is difficult to compare the respective protein amounts because each antibody has a specific affinity and different times of exposure for detection were necessary. Taken together, Northern and Western blotting data provide new insight into previous reports indicating expression of EDG1, EDG3, and EDG5 transcripts in human (An et al. 1998) and mouse (Zhang et al. 1999) heart.
With the aim of investigating the respective roles played by EDG1, EDG3, and EDG5 receptors in cardiovascular tissues, it was important to determine the nature of the cells that express the respective messengers or proteins. For this purpose, ISH and IHC studies were performed at the cellular level. Our data indicate that EDG1 mRNA and protein are strongly expressed in ventricular, septal, and atrial cardiomyocytes as well as in the endothelial cell layer of cardiac vessels, whereas EDG3 mRNA is highly expressed in the smooth muscle cell layer of aorta and cardiac vessels and weakly expressed in cardiomyocytes from both atria and ventricles. EDG5 transcript is detected in cardiomyocytes and in smooth muscle cells of the aortic medial layer and is absent in cardiac coronary microvessels. Regarding the abundance of respective messengers in cardiomyocytes, the intensity of ISH signals obtained with each EDG-specific probe confirms the expression pattern obtained by Northern blotting described above. It was important to clarify EDG1, EDG3, and EDG5 localization in human cardiomyocytes because interspecies discrepancies between mouse (Liu and Hla 1997; Zhang et al. 1999), rat (Lado et al. 1994), and human (An et al. 1998) were previously observed for some S1P/EDG receptors in several tissues. Perinuclear labeling obtained with the EDG1 probe is much more abundant than that obtained with EDG3 and EDG5 probes. This result complements previous data obtained by RT-PCR in human atrial myocytes (Himmel et al. 2000) and by Western blotting in rat neonatal cardiac myocytes (Robert et al. 2001). Strong expression of EDG1 mRNA in human cardiomyocytes is also in good agreement with the presence of EDG1 protein in adult ventricular myocytes of rat papillary muscle (Nakajima et al. 2000). The high expression of EDG1 gene in adult and neonatal cardiomyocytes suggests its potential involvement in S1P-related cardiac functions at all stages of heart development. To support this hypothesis, developmental studies in zebrafish have recently indicated that S1P signaling via an EDG-like receptor, i.e., Mil, is essential for heart development (Kupperman et al. 2000). On the other hand, recent works have demonstrated that S1P induces cardiomyocyte hypertrophy (Robert et al. 2001) and calcium deregulation (Nakajima et al. 2000), mainly via the EDG1 receptor in rat. This result raises the hypothesis that EDG1 might be involved in cardiac pathophysiology. However, as shown by ISH, the lower expression of EDG3 and EDG5 receptors in cardiomyocytes does not rule out the possibility that these Gq-coupled receptors may also mediate S1P function, notably the cardiac calcium influx regulation as previously suggested (Nakajima et al. 2000). EDG3 expression in human cardiomyocytes is also in line with a recent study suggesting that S1P could activate the acetylcholine-stimulated inward rectifier K+ current via EDG3 receptors in human atrial myocytes and then participate in the modulation of heart rhythm (Himmel et al. 2000). Co-localization of EDG1, EDG3, and EDG5 in cardiomyocytes may result in distinct responses to S1P stimulation, because these receptors are capable of coupling differentially to one or multiple G-proteins and intracellular effectors. Whereas EDG1 was demonstrated to be coupled to Gi/o-protein and can utilize the Rho-dependent signaling pathway, EDG3 and EDG5 signal not only via Gi but also through Gαq and Gα12/α13 (Pyne and Pyne 2000).
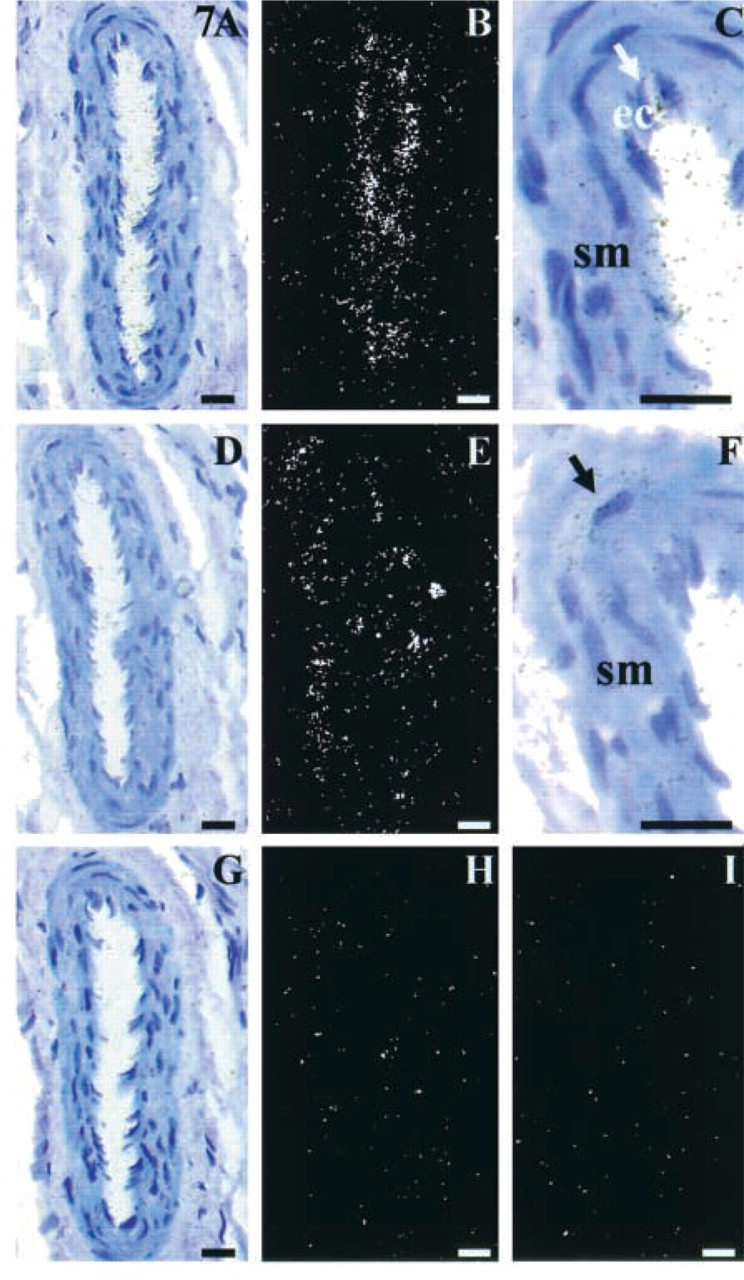
Differential patterns of S1P receptor expression in coronary vessel walls. Light microscopy results of ISH with EDG1, EDG3, and EDG5 probes on adjacent sections of ventricle are represented in
The relatively high expression of EDG3 in different compartments of the heart, as revealed by Northern blotting and the faint signal obtained by ISH in cardiomyocytes, suggested that the EDG3 receptor might be expressed in other cardiac cell types. In agreement with this hypothesis, EDG3 protein was shown to be strongly detected in whole rat heart homogenates but not in purified ventricular cardiomyocytes (Robert et al. 2001). Our ISH experiments demonstrated intense expression of EDG3 receptor gene in the medial layer of human aorta and cardiac vessels composed of smooth muscle cells. These data are in good agreement with results obtained in rat (Tamama et al. 2001) showing the expression of EDG3 receptor mRNA in isolated primary aortic smooth muscle cells. EDG3 expression in vessels is confirmed by our Western blotting analysis indicating a higher level of protein in aorta than in heart, probably due to the higher relative percentage of smooth muscle cells in the aortic tissue samples compared to heart samples. We were unable to detect significant expression of EDG5 in human cardiac microvessels, although the EDG5 transcript was strongly detected in aortic smooth muscle cells. This is in good agreement with the work of Tamama and co-workers (2001), who have demonstrated that rat aortic smooth muscle cells express EDG5 mRNA. Such discrete EDG5 expression in smooth muscle cells originating from different cardiovascular vessels might be explained by a differential and vessel size-specific gene expression. Moreover, unlike what has been previously shown in cultures of smooth muscle cells either in rodent or human (Hla et al. 1999; Rakhit et al. 1999; Tamama et al. 2001), our ISH amd IHC studies did not enable us to detect EDG1 mRNA or EDG1-immunoreactive cells in the smooth muscle cell layer of adult human aorta and cardiac vessels. This result suggests that expression of EDG1 in the smooth muscle cells of adult vessels, if any, is likely to be very weak. The use of very sensitive experiments, such as RT-PCR, to detect EDG1 expression in most of the previous studies (Hla and Maciag 1990; Rakhit et al. 1999) appears to confirm this hypothesis, which questions the physiological role played by this weak EDG1 expression.
The distinguishable spatial expression patterns of EDG1, EDG3, and EDG5 receptors suggest distinct biological roles played by each receptor in the different vessel layers. EDG3 and EDG5 expression in smooth muscle cells layers suggests that this receptor might be important for the S1P-induced mobilization of Ca2+ in human smooth muscle (Bornfeldt et al. 1995) and could then participate in regulation of the proliferation and migration of vascular smooth muscle cells, which are important events in the development of vascular disorders such as atherosclerosis and restenosis (Ross 1999). High expression of the EDG1 receptor in endothelial cells of cardiac vessels confirms its potential role in the regulation of endothelial function under physiological blood flow conditions (Takada et al. 1997). Detection of EDG1 expression in endothelial cells is also consistent with the fact that EDG1 might be involved in the phorbol ester-induced tube formation of human umbilical vein endothelial cells (HUVECs) (Hla and Maciag 1990; Lee et al. 1998). In fact, it has been demonstrated that inhibition of expression of EDG1 by specific antisense oligonucleotides abolishes the effects of S1P on capillarylike network formation (Lee et al. 1999). Moreover, the presence of functional EDG1 protein in endothelial cells is in good agreement with downstream events involving the recruitment of smooth muscle cells by S1P-stimulated endothelial cells (Liu et al. 2000). Working with EDG1 knockout mice, Liu et al. (2000) have shown in embryos that the EDG1 receptor is necessary for vascular formation via smooth muscle investment during vessel maturation. Indeed, aortic sections of EDG1 —/— mouse embryos demonstrated the lethal absence of smooth muscle cells at the dorsal surface of the aorta. The absence of EDG1 expression in smooth muscle cells of adult human cardiac vessels, unlike embryonic mouse vessels, suggests a potentially differential expression of the EDG1 receptor during development. In agreement with this hypothesis, recent results obtained in rat demonstrated that pupintimal vascular smooth muscle cells express higher levels of EDG1 mRNA than adult medial vascular smooth muscle cells (Kluk and Hla 2001). To confirm this latter hypothesis, it would be necessary to look for potential expression of the EDG1 receptor in smooth muscle cells of human embryonic cardiac vessels.
In conclusion, this study provides new insights into the expression pattern of S1P/EDG receptors in adult human cardiovascular tissues. Although further studies will be necessary to better delineate the S1P mechanisms of action in human heart and vessels, the present data suggest that the receptors of the EDG family may represent good targets for a therapeutic strategy aimed at modulating the transduction machineries involved in human cardiovascular diseases.
Footnotes
Acknowledgements
We thank Dr V. Paumier and Dr P. Noret for help in histological analysis, and Jean-Claude Camelin and Sabine Rouanet for their contribution to this project.
