Abstract
Matrix metalloproteinases (MMPs) 8 and 13 comprise the collagenase subfamily in rats and mice, and only MMP13 has been implicated in degradation of the collagenous matrices during development of bone and cartilage. On the hypothesis that MMP8 is also involved in bone and cartilage development, the present study was designed to investigate gene expression of MMP8 in rat embryonic mandibles and hind limbs. Expression of MMP8 was examined with in situ hybridization and RT-PCR and was compared with that of MMP13. Osteoblastic and chondrocytic cells expressing collagenous matrix molecules were identified using in situ hybridization for collagen Types I and II. The results demonstrated that MMP8 is expressed by osteoblastic progenitors, differentiated osteoblasts, osteocytes, and chondrocytes in the growth plate for the first time. Furthermore, the expression of MMP8 is much broader than that of MMP13, for which expression is confined to differentiated phenotypes of osteoblastic and chondrocytic lineage.
Keywords
The Major Constituents of bone and cartilage extracellular matrices are collagens, i.e., Type I for bone and Type II for cartilage (von der Mark et al. 1976; Kosher et al. 1986; Devlin et al. 1988; Sasano et al. 1992). Osteoblasts and chondrocytes synthesize the collagens and other extracellular matrix (ECM) molecules during development of bone and cartilage (Up-holt and Olsen 1991; Stein et al. 1996; Zhu et al. 2001). It has been suggested that these bone and cartilage ECM molecules are degraded and turned over during embryonic development (Partridge and Winchester 1996; Chin and Werb 1997; Sasano et al. 2000a,b). However, little information is available regarding enzymes that remodel collagenous matrices in developing bone and cartilage.
The matrix metalloproteinases (MMPs) are believed to play a central role in the breakdown of ECM, which is essential for embryonic development, morphogenesis, and tissue remodeling (Birkedal-Hansen et al. 1993; Nagase and Woessner 1999). Collagenases are the only members of the MMP family that degrade native fibrillar collagens of Types I, II, and III (Gack et al. 1995; Franchimont et al. 1997; Johansson et al. 1997; Woessner and Nagase 2000). MMPs 1, 8, and 13 comprise the collagenase subfamily in humans, but only two of them, MMPs 8 and 13 have been identified in rats and mice (Jeffrey 1998; Woessner 1998; Woessner and Nagase 2000).
MMP13 is the only collagenase that has been implicated in degradation of the collagenous matrices during development of bone and cartilage (Gack et al. 1995; Mattot et al. 1995; Johansson et al. 1997; Winchester et al. 1999). MMP8 (neutrophil collagenase) was originally believed to be confined to neutrophils (Tschesche and Pieper 1998; Woessner and Nagase 2000), but recent studies indicate that it may be expressed in other cell types, such as osteoarthritic chondrocytes (Shlopov et al. 1997), articular chondrocytes (Cole et al. 1996), synovial fibroblasts and endothelial cells (Hanemaaijer et al. 1997), and odontoblasts and dental pulp cells (Palosaari et al. 2000). However, it is not known whether MMP8 is expressed during osteogenesis and chondrogenesis. We hypothesized that MMP8 is another collagenase involved in development of bone and cartilage. To test the hypothesis, the present study was designed to investigate gene expression of MMP8 in rat developing mandibles and hind limbs. Expression of MMP8 was examined using ISH and RT-PCR and was compared with that of MMP13. Osteoblastic and chondrocytic cells expressing collagenous matrix molecules were identified using ISH for collagen Types I and II.
Materials and Methods
Preparation of Tissue
The principles of laboratory animal care (NIH publication no. 86–23, revised 1985) were followed as well as specific national laws. The preparation was performed according to animal protocols that were institutionally approved by the Tohoku University. Wistar rat embryos 14 to 20 days post coitum and Wistar rats 1 week post natum were used. At least three embryos or postnatal animals at each stage were examined. Heads and hind limbs were resected and immediately fixed by immersion in 4% paraformaldehyde with 0.5% glutaraldehyde (GA) in 0.1 M phosphate buffer, pH 7.4, at 4C overnight. Some of the fixed specimens were decalcified in 10% EDTA in 0.01 M phosphate buffer, pH 7.4, for 3–4 days for embryos and 2 weeks for 1-week-old rats at 4C. The EDTA solution was autoclaved before use. After dehydration through a graded series of ethanol solutions, the tissues were embedded in paraffin (Sasano et al. 1996; Zhu et al. 2001). Serial sections 5 μm thick were cut and the adjacent sections were stained with hematoxylin-eosin or processed for ISH.
Preparation of Riboprobes
Digoxigenin (DIG)-labeled single-strand riboprobes were prepared using the DIG RNA labeling kit (Roche; Mannheim, Germany) according to the manufacturer's instruction.
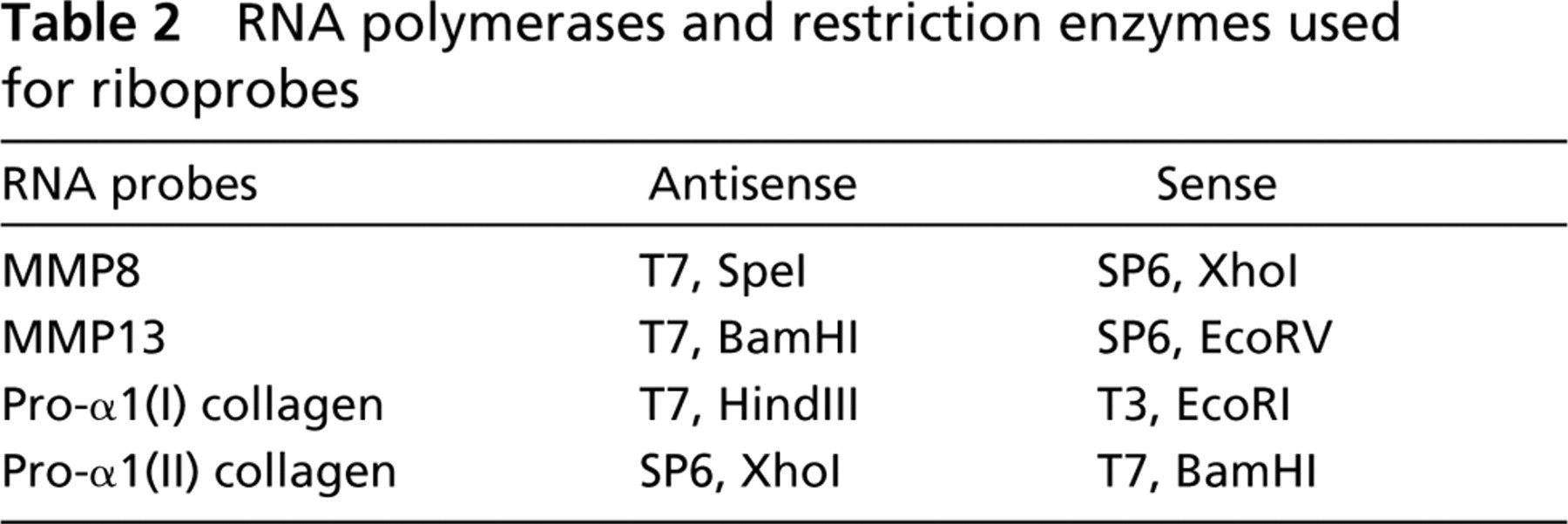
Fragments encoding rat MMP8 (171–1253 bp:GenBank AJ007288) and rat MMP13 (1547–2411 bp:GenBank M60616, M36452) were obtained from the total RNA of embryonic rat limbs using reverse transcription followed by polymerase chain reaction (RT-PCR) and subcloned into the PCR II TOPO (Invitrogen; Carlsbad, CA). A fragment encoding rat pro-α1(I) collagen (2838–4329 bp:GenBank Z78279) was obtained from the total RNA of rat skin using RT-PCR and subcloned into the pT7/T3–18 plasmid (Life Technologies; Grand Island, NY). A 545-bp fragment encoding mouse pro-α1(II) collagen was obtained from the total RNA of embryonic mouse limbs using RT-PCR as described previously (Takahashi et al. 1998). Oligonucleotide primers used for the RT-PCR are shown in Table 1. The cDNA was verified by digestion with restriction enzymes and was confirmed by dideoxynucleotide sequencing. Riboprobes were generated as indicated in Table 2.
Oligonucleotide primers to prepare for riboprobes

In Situ Hybridization
The protocol used in the present study has been reported elsewhere (Sasano et al. 1996; Zhu et al. 2001) and is only briefly described, as follows:
The sections were deparaffinized and washed in PBS.
The sections were immersed in 0.2 N HCl for 20 min and, after washing in PBS, incubated in proteinase K (20 g/ml; Roche) in PBS for 30 min at 37C.
After washing, the sections were dipped in 100% ethanol and dried in air.
The sections were incubated with the antisense or sense control probe (400 ng/ml) in a hybridization mixture for 16 hr at 45C.
After washing, the sections were treated with RNase (Type 1a, 20 μg/ml; Sigma, St Louis, MO) for 30 min at 37C.
After washing, the hybridized probes were detected immunologically using the Nucleic Acid Detection Kit (Roche), counterstained with methyl green, and mounted.
At least six sections from each of three embryos or postnatal animals at each stage were examined using the same probe. The intensity of hybridization signals was evaluated by observing at least three fields on every section.
RT-PCR
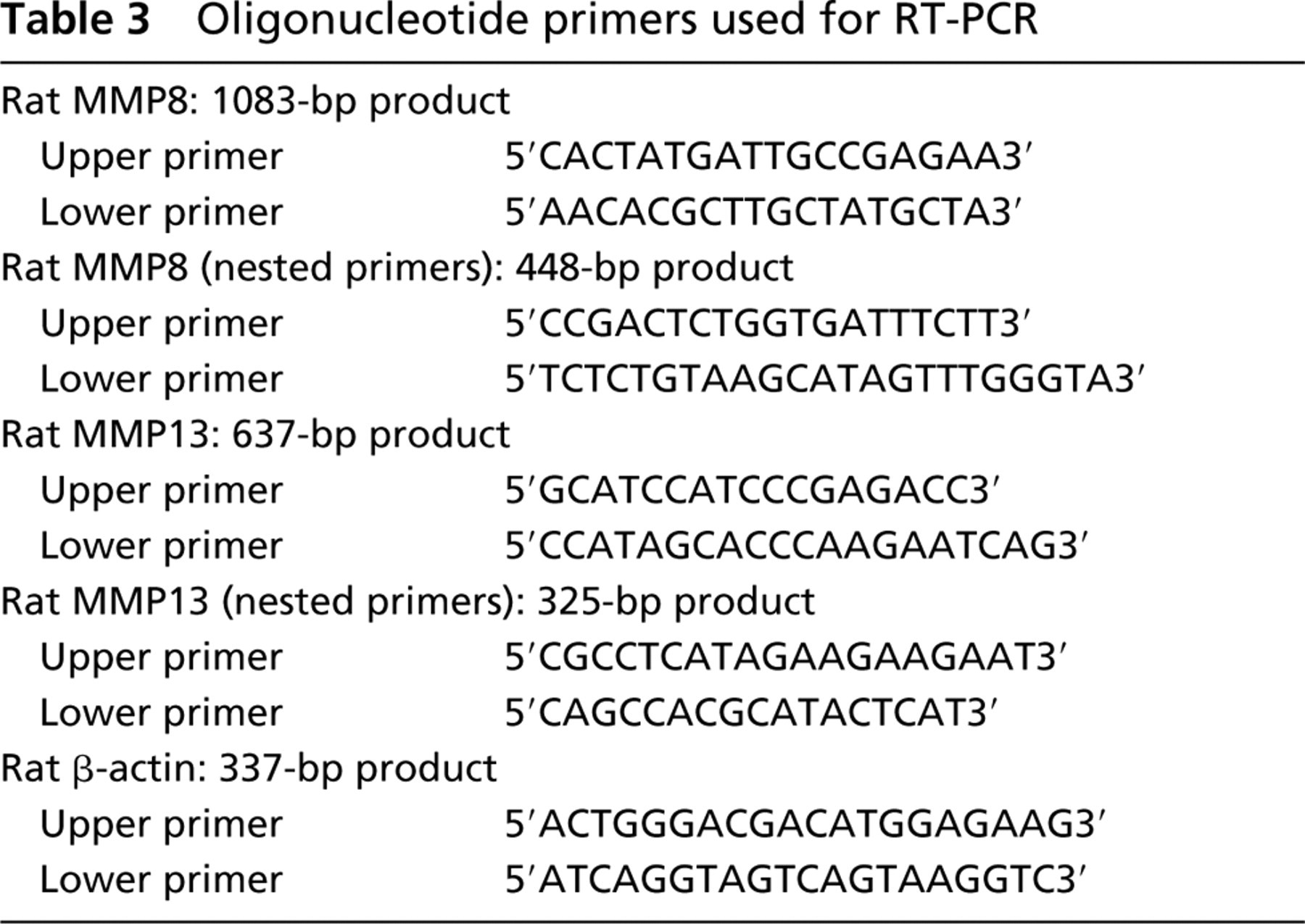
The total RNA was extracted from embryonic rat mandibles and hind limbs using the RNeasy Mini Kit (QIAGEN; Hilden, Germany) and processed as follows (Singh et al. 2000). cDNA was synthesized using 1.0 μg of total RNA primed with 1.5 μg of random primers (Invitrogen) in the presence of reverse transcriptase at 100 U/μg RNA in a reverse transcription buffer as supplied by Invitrogen. For cDNA amplification, 1.0 μl of reverse-transcription products was incubated in the presence of 20 pmol of two specific primers (Table 3), 1.5 mM MgCl2, 0.2 mM of the four dNTPs (Invitrogen), and 0.05 U of Taq polymerase in reaction buffer (Invitrogen). The reaction mixture was subjected to one cycle of denaturation for 3 min 50 sec at 95C, followed by 40 cycles of an amplification sequence that consisted of denaturation for 70 sec at 95C, annealing for 70 sec at 64C for MMP8, 63C for MMP13, and 56C for β-actin and extension for 2 min 30 sec, with an additional 7 min 30 sec for the last cycle. Subsequently, 1 μl of the amplified product was processed the same way using the nested primers for MMP8 with annealing at 64C and for MMP13 at 58C.
RNA polymerases and restriction enzymes used for riboprobes
Oligonucleotide primers used for RT-PCR

After amplification, equal amounts of PCR products were size-separated by electrophoresis through a 1.5% agarose gel and visualized by ethidium bromide staining and UV transillumination. All samples were amplified at least twice on different occasions to control for any variations in the PCR technique. Negative controls (no reverse transcription and no template RNA) did not produce bands visible on gels. Images of the stained gels were captured using the Electrophoresis Documentation and Analysis System (Kodak; Rochester, NY).
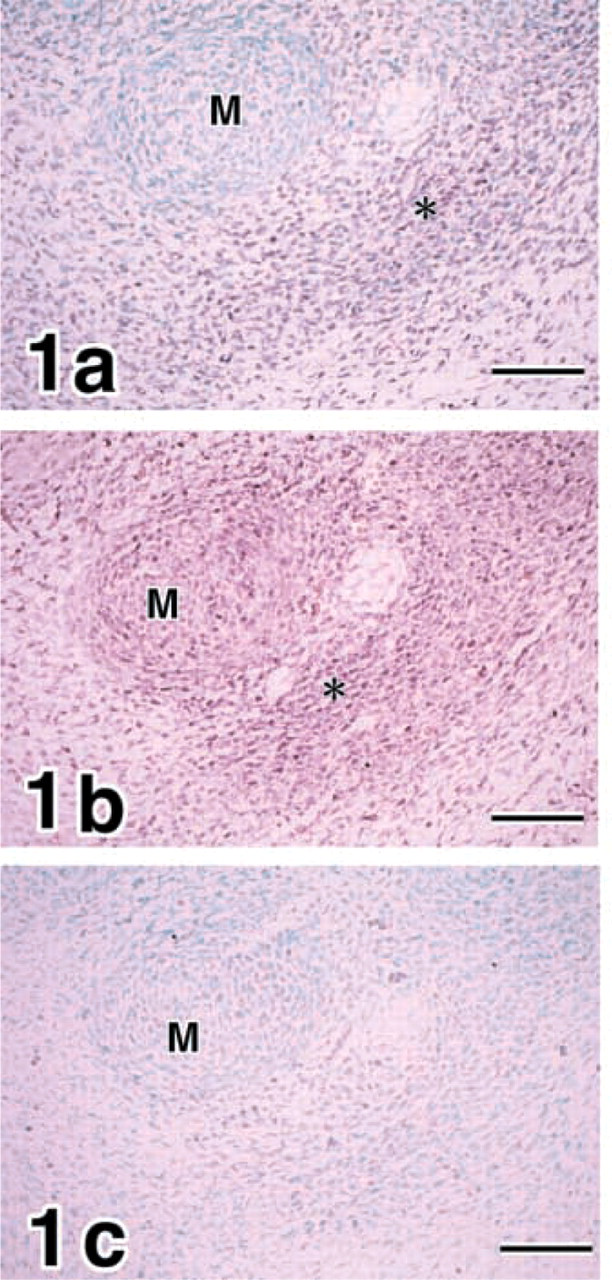
E14 mandibles. Expression of COL I is demonstrated in mesenchymal cell condensation (
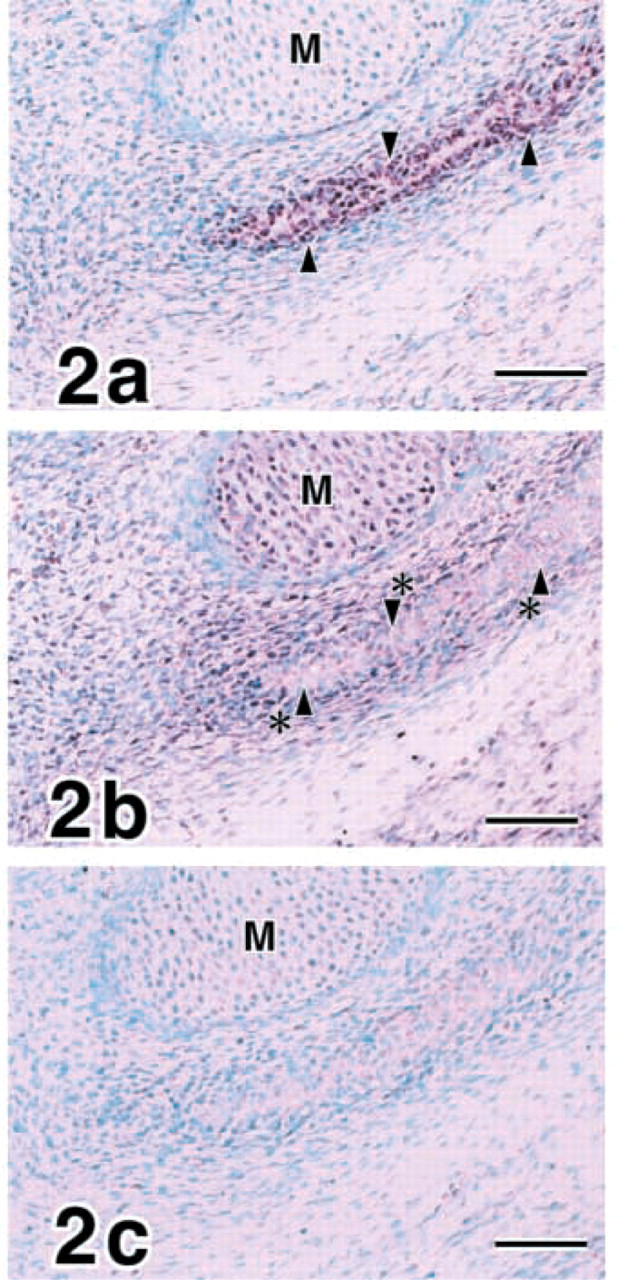
E15 mandibles. Cuboidal osteoblasts actively express COL I (
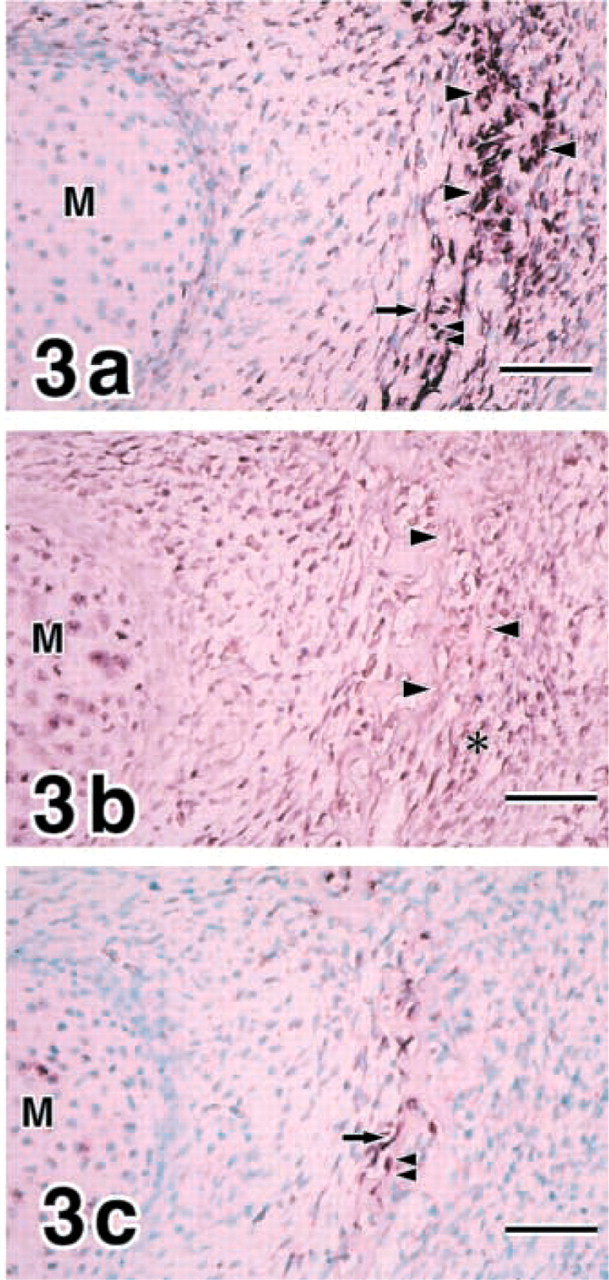
E16 mandibles. Flat osteoblasts and osteocytes as well as cuboidal osteoblasts express COL I (
The PCR products in bands corresponding to MMP8 and MMP13 were extracted using the Gel Extraction Kit (QIAGEN) and the sequence was confirmed by dideoxynucleotide sequencing.
Results
In Situ Hybridization
Mandibles. Expression of pro-α1(I) collagen (COL I) transcripts was demonstrated in mesenchymal cell condensation of the putative osteogenic region in E14 mandibles (Figure 1a. MMP8 was expressed in the mesenchymal cell condensation and chondrocytes in Meckel's cartilage (Figure 1b. In contrast, no expression of MMP13 was identified in E14 mandibles (Figure 1c.
Cuboidal osteoblasts differentiated and expressed COL I actively in E15 (Figure 2a. MMP8 transcripts were demonstrated in embryonic periosteal cells as well as chondrocytes of Meckel's cartilage but not in cuboidal osteoblasts (Figure 2b. No expression of MMP13 was identified in E15 mandibles (Figure 2c.
Flat osteoblasts and osteocytes appeared in addition to cuboidal osteoblasts, and all of these cell types expressed COL I at E16 (Figure 3a. MMP8 was continuously shown in chondrocytes in Meckel's cartilage and periosteal cells around cuboidal osteoblasts (Figure 3b. MMP13 expression was demonstrated in flat osteoblasts and osteocytes strongly at E16 (Figure 3c. Chondrocytes in Meckel's cartilage expressed MMP13 weakly.
E18 mandibles. Cuboidal osteoblasts, flat osteoblasts, and osteocytes express COL I (
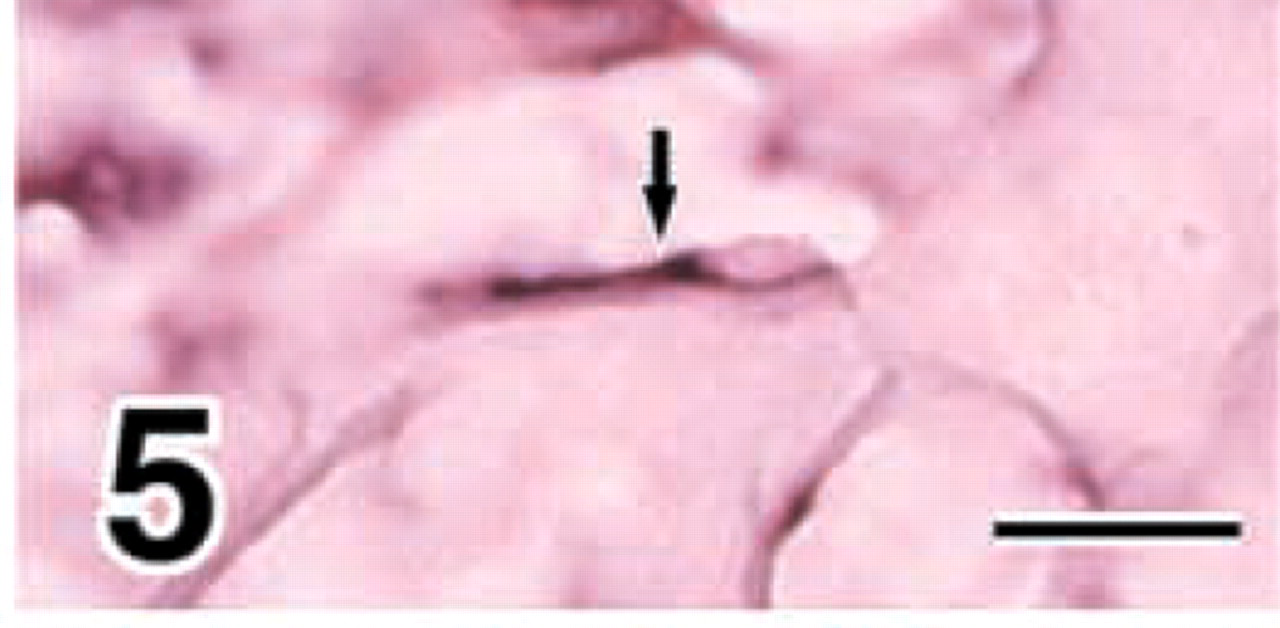
Flat osteoblasts (arrow) express MMP8 in E18 mandibles. Bar = 25 μm.
Mandibles at 1-week postnatum. Osteocytes (arrowheads) express COL I (
Gene expression patterns of COL I, MMP8, and MMP13 in rat embryonic bone development of mandibles and hind limbsa
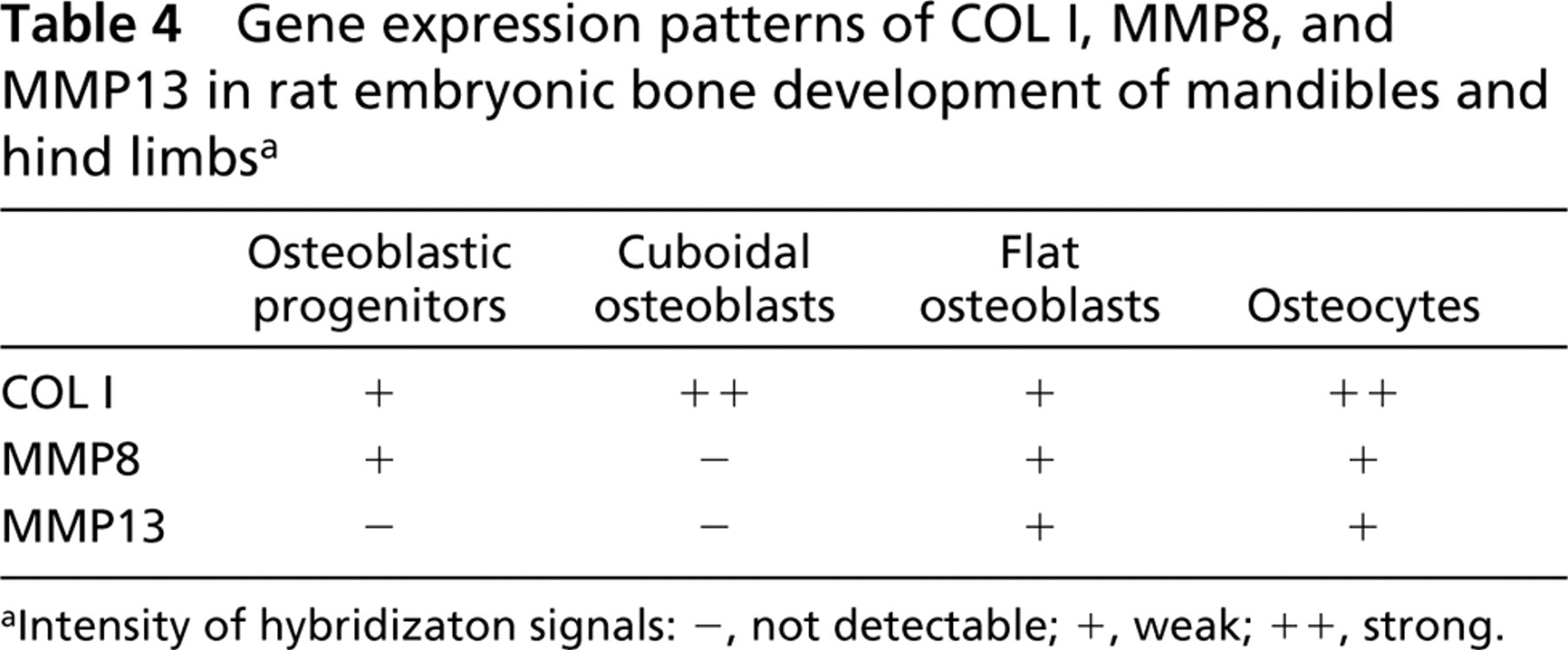
Intensity of hybridizaton signals: -, not detectable; +, weak; ++, strong.
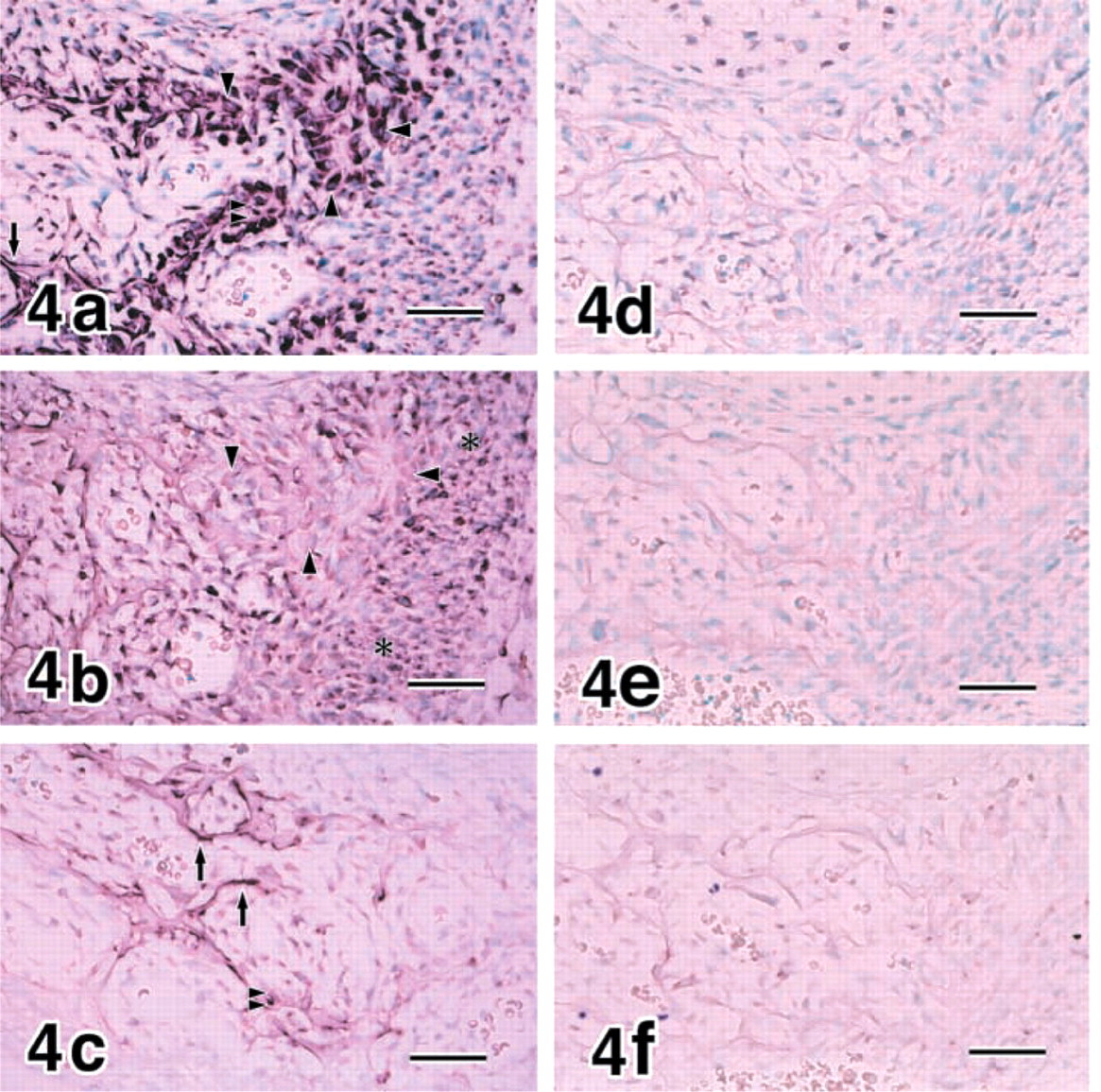
Cuboidal osteoblasts, flat osteoblasts, and osteocytes expressed COL I at E18 (Figure 4a and E20 (data not shown). MMP8 was demonstrated in chondrocytes, periosteal cells around cuboidal osteoblasts, and flat osteoblasts at E18 (Figures 4b, and 5) and E20 (data not shown). The strong expression of MMP13 was confined to flat osteoblasts and osteocytes at E18 (Figure 4c and E20 (data not shown).
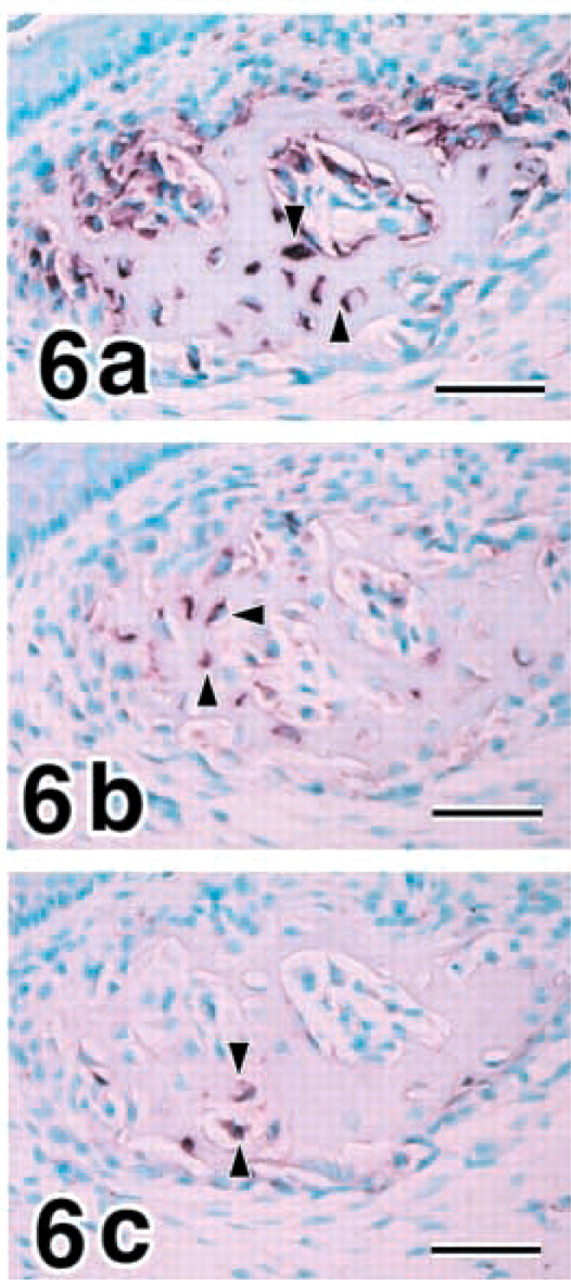
Osteocytes in 1-week postnatal mandibles expressed COL I, MMP8, and MMP13 (Figures 6a-6c). The pattern of gene expressions for COL I, MMP8, and MMP13 in osteogenic cells is summarized in Table 4, where the hybridization signal is graded as not detectable, weak, or strong according to the intensity.
In addition to the osteogenic and chondrogenic phenotypes, muscle cells expressed MMP8 strongly (Figure 7a and MMP13 weakly (Figure 7b.
Hind Limbs. Cuboidal osteoblasts, flat osteoblasts, and osteocytes expressed COL I in growing rat hind limbs at E18 (data not shown), E20 (Figure 8a, and 1-week postnatum (data not shown), while periosteal cells showed weak expression. MMP8 transcripts were demonstrated in flat osteoblasts and osteocytes and very weakly in periosteal cells (Figure 8b. MMP13 expression was localized in flat osteoblasts and osteocytes (Figure 8c. The pattern of gene expressions for COL I, MMP8, and MMP13 in osteogenic cells was comparable to that in mandibles, as shown in Table 4.
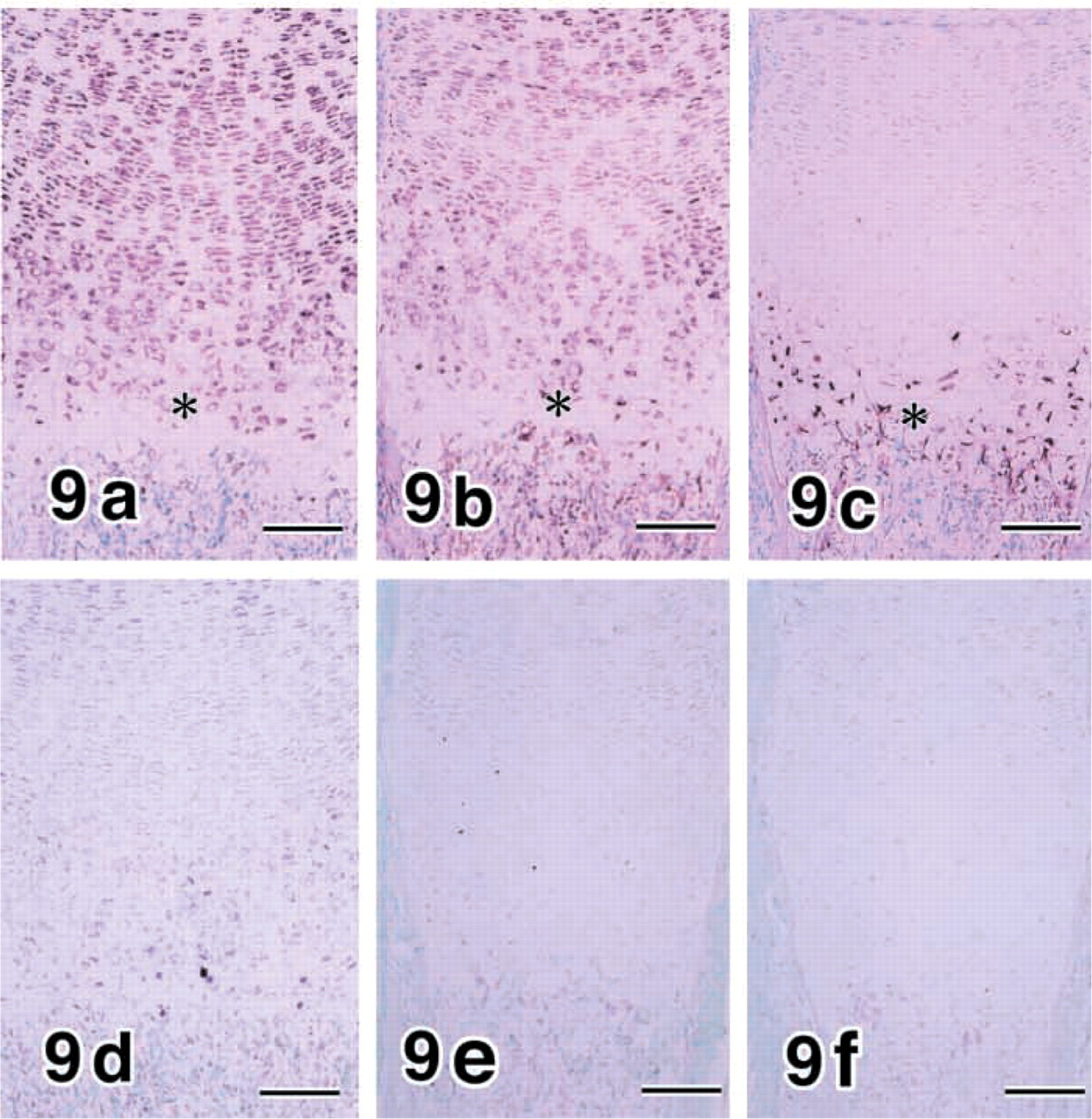
MMP8 was expressed in almost all chondrocytes (Figure 9b which expressed pro-α1(II) collagen (COL II) (Figure 9a in developing tibias and femurs, whereas MMP13 expression was localized in hypertrophic chondrocytes (Figure 9c. The pattern of gene expressions for MMP8, MMP13, and COL II in differentiating chondrocytes is summarized in Table 5, where the intensity of hybridization signals is graded as not detectable, weak, or strong.
Controls for In Situ Hybridization. No hybridization signal was identified in the adjacent sections processed with control sense riboprobes (Figures 4d-4f and 9d-9f).
RT-PCR
RT-PCR demonstrated that the gene of MMPs 8 and 13 is expressed in developing mandibles and hind limbs (Figure 10).
Discussion
Collagenases are the only members of the MMP family that degrade native fibrillar collagens of Types I, II, and III (Gack et al. 1995; Franchimont et al. 1997; Johansson et al. 1997; Woessner and Nagase 2000). Only MMP8 (neutrophil collagenase) and MMP13 (rodent interstitial collagenase) had been considered collagenases in rats and mice until a mouse orthologue of human MMP1 was recently identified (Balbín et al. 2001). MMP13 has been implicated in degradation of the collagenous matrices during development of bone and cartilage (Gack et al. 1995; Mattot et al. 1995; Johansson et al. 1997; Winchester et al. 1999). In contrast, MMP8 was originally believed to be confined to neutrophils (Tschesche and Pieper 1998; Woessner and Nagase 2000) and no information is available about its expression in bone and growth plate cartilage, although it has been suggested to cleave Types I and II collagen with high efficiency (Tschesche and Pieper 1998; Woessner and Nagase 2000).
The present study demonstrated that MMP8 is expressed in the mesenchymal cell condensation of osteoblastic progenitors (Dunlop and Hall 1995; Hall and Miyake 2000; Zhu et al. 2001), periosteal cells, flat osteoblasts, osteocytes, chondrocytes, and muscle cells in developing rat mandibles and hind limbs. This is the first report to show MMP8 expression in cells of the osteoblastic lineage, i.e., the mesenchymal condensation, periosteal cells, osteoblasts, and osteocytes. MMP8 may be involved in remodeling of collagenous ECMs during embryonic development of bone.
MMP13 is the only collagenase that has been implicated in degradation of the collagenous matrices during development of bone (Gack et al. 1995; Mattot et al. 1995; Johansson et al. 1997; Winchester et al. 1999). However, expression of MMP13 was confined to flat osteoblasts and osteocytes in the present study. Moreover, MMP13 was not expressed in any cell types of mandibles, including the mesenchymal condensation of osteoblastic progenitors and periosteal cells until E16. MMP13 may be expressed exclusively by differentiated phenotypes of the osteoblastic lineage, i.e., flat osteoblasts and osteocytes (Fuller and Chambers 1995; Zhu et al. 2001).
The present study suggested that osteocytes express both MMP8 and MMP13. Our previous studies demonstrated that osteocytes express ECM molecules, such as Type I collagen, osteonectin, bone sialoprotein, osteopontin, and osteocalcin (Zhu et al. 2001; Sasano et al. 2001). Another previous study suggested that osteocytes express MMP3 as well (Bord et al. 1998). Osteocytes may be actively involved in turnover, i.e., production and degradation, of the ECM molecules of bone.
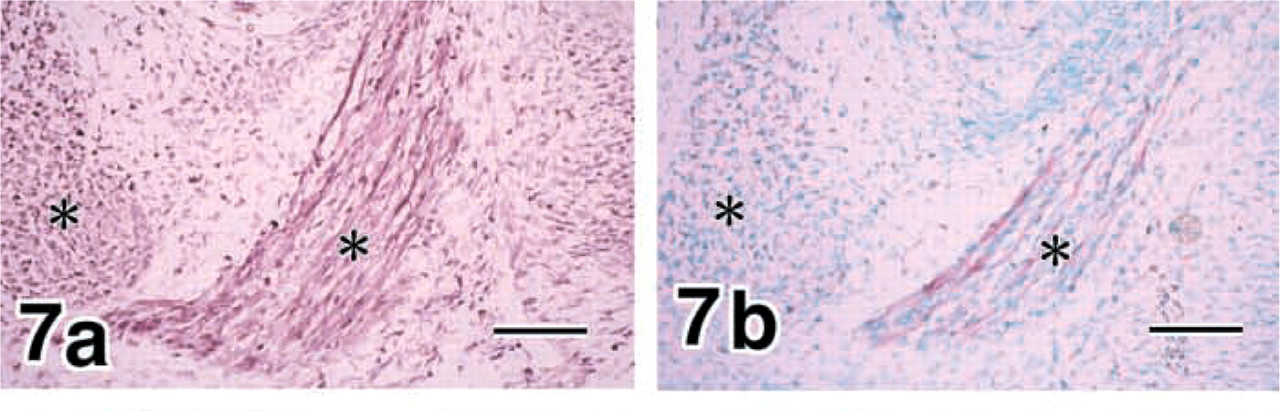
E16 mandibles. Muscle cells (asterisks) express MMP8 strongly (
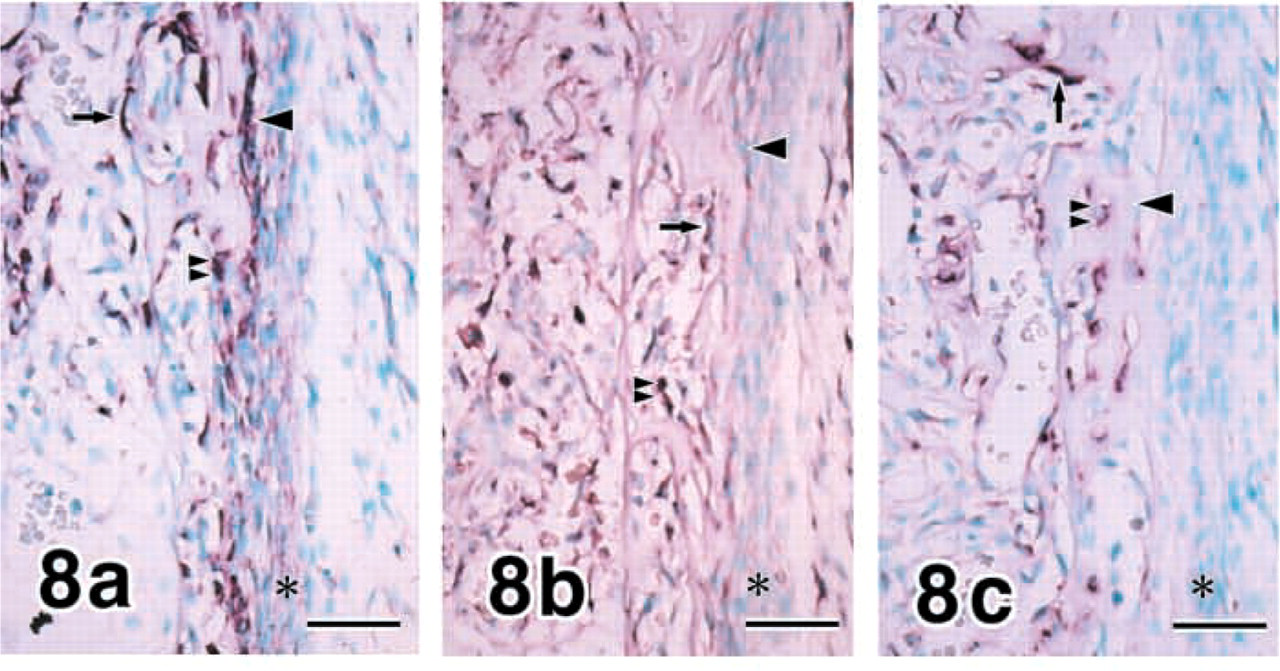
E20 hind limbs (tibiae). Cuboidal osteoblasts, flat osteoblasts, and osteocytes express COL I, while periosteal cells show weak expression (
E20 hind limbs (tibiae). MMP8 (
Gene expression patterns of COL II, MMP8, and MMP13 in rat embryonic cartilage development of hind limbsa
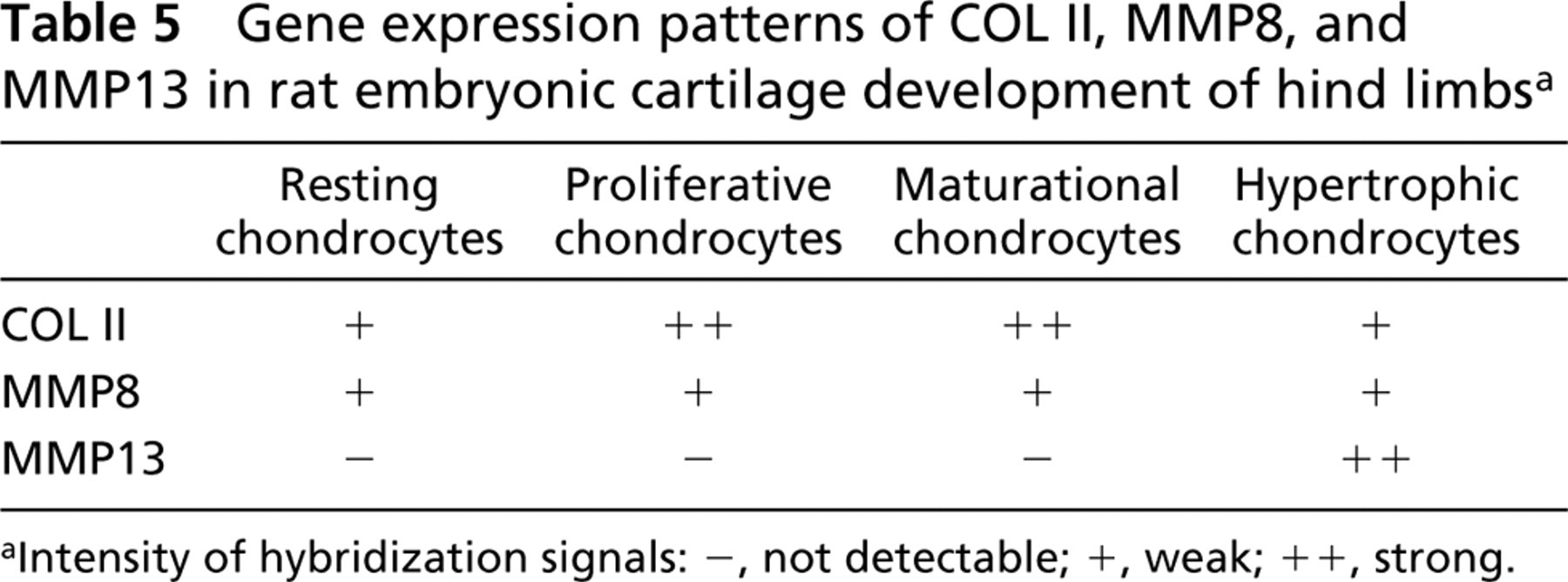
Intensity of hybridization signals: -, not detectable; +, weak; ++, strong.
Cuboidal osteoblasts engaged in production of ECM molecules such as Type I collagen may not be involved in expression of MMPs in rat embryos. In contrast, flat osteoblasts whose production of matrix molecules is less than the cuboidal osteoblasts could participate in degradation of the collagenous matrix with synthesis of MMP8 and MMP13 (Fuller and Chambers 1995; Zhu et al. 2001). Previous studies indicated that MMP2 (Meikle et al. 1992), MMP3 (Bord et al. 1998), and MMP10 (Bord et al. 1999) are expressed in osteoblasts. Flat osteoblasts may play a role in remodeling of ECM molecules of developing bone in concert with osteocytes.
Chondrocytes expressed MMPs 8 and 13. MMP13 was confined to hypertrophic chondrocytes, as previously reported (Johansson et al. 1997; D'Angelo et al. 2000), whereas MMP8 was expressed in almost all chondrocytes. This is the first report indicating expression of MMP8 in growth plate cartilage. MMP2 (Kawashima-Ohya et al. 1998), MMP3 (Freemont et al. 1997; Kawashima-Ohya et al. 1998), MMP9 (Freemont et al. 1997; Kawashima-Ohya et al. 1998), and MMP10 (Bord et al. 1999) are also expressed in chondrocytes. Chondrocytes may take part in turnover of ECM molecules of developing cartilage utilizing various types of MMPs.
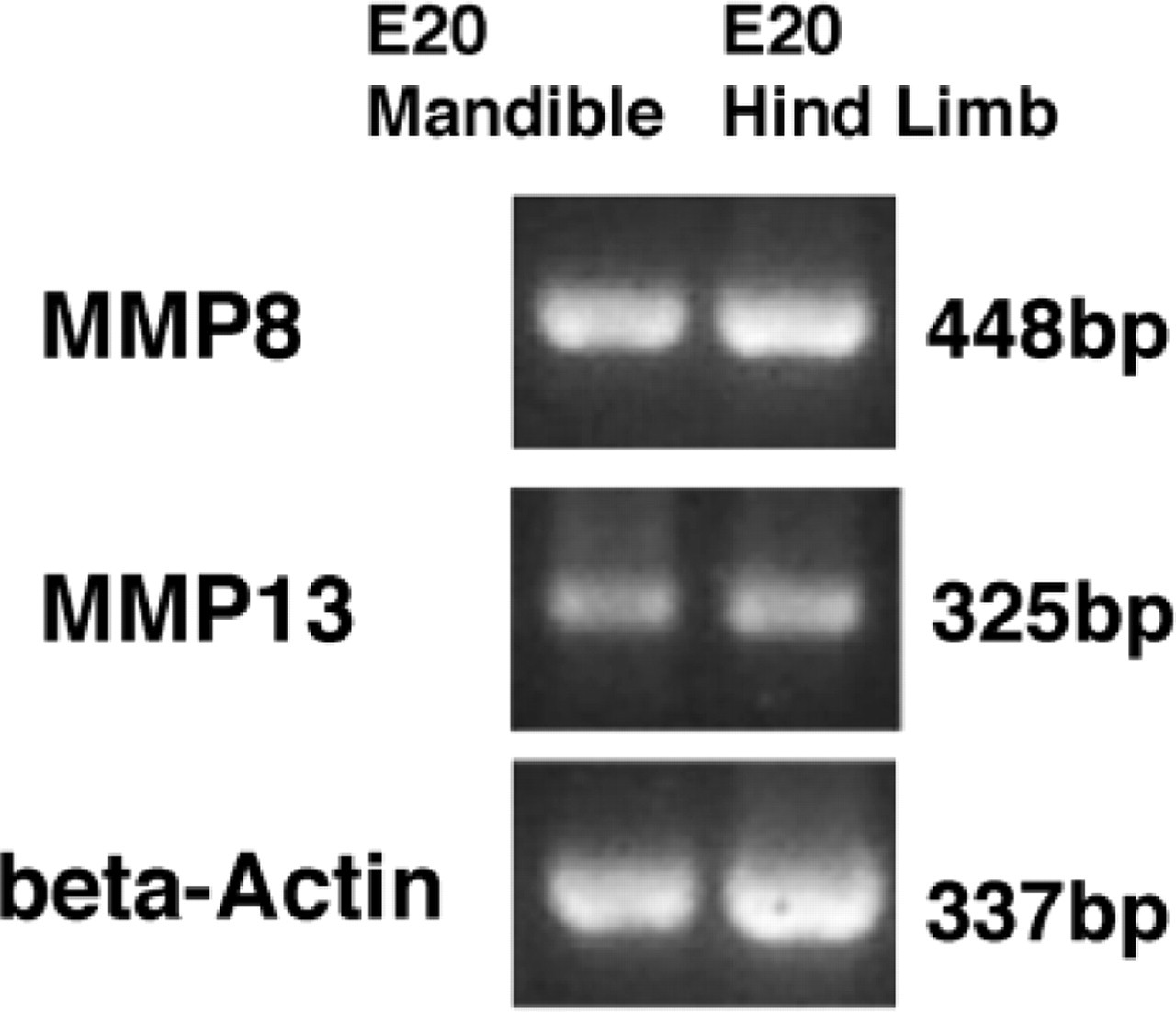
RT-PCR demonstrates that the gene of MMPs 8 and 13 is expressed in E20 mandibles and hind limbs.
Muscle cells expressed MMP8 strongly and MMP13 weakly. These collagenases may be involved in remodeling of the ECM during myogenesis, along with other MMPs, such as MMPs 2 and 9 (Guérin and Holland 1995; Kherif et al. 1999).
The previous study indicated that MMP13 is expressed during fracture healing of bone (Yamagiwa et al. 1999). MMP8 could also participate in the healing process as well as embryonic bone development. MMP8 may play an important role in remodeling of ECM molecules during bone and cartilage formation, which is distinct from that of MMP13.
Footnotes
Acknowledgements
Supported in part by grants-in-aid (12671986, 13671893, 13672076, 13470454) from the Ministry of Education, Science, Sports and Culture of Japan.
We wish to thank Mr Masami Eguchi and Mr Yasuto Mikami, Division of Oral Molecular Biology, Tohoku University Graduate School of Dentistry, for excellent assistance in this study.
