Abstract
We studied the functional significance of marked differences in the DNA content of somatic cells and germ line nuclei by static Feulgen–DNA cytophotometry for several species of microcrustaceans that exhibit chromatin diminution during very early stages of embryogenesis. Mature females and males showed many gonadal nuclei with elevated amounts of DNA that persist until dispersal of this “extra” DNA throughout the cytoplasm as fragments and coalescing droplets of chromatin during anaphase of the diminution division.
Chromatin diminution, the fragmentation and excision of particular heterochromatic chromosome segments during early cleavage stages, occurs in some species of cyclopoid copepods and results in marked loss of DNA from the genome of presumptive somatic cells but retention of this DNA in primordial germ cell nuclei (Beermann 1977). A similar phenomenon occurs in the selective elimination of so-called “limited” DNA from somatic cell lineages for several other species of copepods (Rasch and Wyngaard 1997; Grishanin and Akif'ev 2000), in ascarid nematodes (Tobler 1986), in Japanese hagfish (Kubota et al. 1993), and in many species of ciliated protozoa (Coyne et al. 1997). These systems provide an unusual opportunity to study genomic restructuring in the context of development and differentiation.
Mature males, egg-carrying females, and juveniles at copepodid stages CIV and CV were fixed individually in a drop of 3:1 methanol/acetic acid (v/v) for 2–3 min, allowed to swell for 1–2 min in a drop of 45% acetic acid on a glass slide, squashed with moderate pressure for tissue dispersal under a siliconized coverslip, frozen on dry ice to expedite coverglass removal, thawed in two changes of absolute ethanol, and then air-dried. Each series of slides was stained simultaneously in the Feulgen reaction for DNA, using blood films of chicken, rainbow trout, and Xenopus laevis as internal reference standards of 2.5 pg DNA, 5.2 pg DNA, and 6.3 pg DNA per cell, respectively, to allow expression of integrated absorbance values for copepod nuclei in terms of pg DNA (Rasch 1985). More than 10,000 individual nuclear scans were measured at 560 nm with a Vickers M86 integrating microdensitometer, using the high-resolution scanning pattern of the instrument. It was of particular interest for this analysis to determine the DNA content of somatic cell (SC) nuclei, of putative germ cell nuclei before chromatin diminution (PD), and the DNA content of sperm as an index of the paternal contribution to the genome of M. edax embryos.
Representative distributions of the DNA content of SC nuclei and prediminuted nuclei (PD) from a mature, egg-carrying female (Figure 1A) can be compared with the DNA levels found for sperm and PD nuclei from an adult male (Figure 1B), whose SC nuclei contained 2.96 ± 0.082 pg DNA (n = 50). Sperm appeared to contain twice as much DNA as expected because half of the diploid SC genome would be about 1.5 pg DNA for this species (Wyngaard and Rasch 2000). The DNA levels of PD nuclei from both sexes appear to be equivalent. Some egg-carrying females were identified whose presumptive germ cell nuclei contain as much as 25–35 pg DNA, although a majority of the approximately 120 females analyzed to date showed PD nuclei with 15–25 pg DNA (Table 1). Juveniles at stage CIV or CV have PD nuclei with 10–20 pg DNA. There is no compelling evidence that PD nuclei adhere to regular genomic duplication, as would be shown by DNA values that would be even geometric multiples of the basic diploid genome for this species, i.e., 3 pg DNA/somatic nucleus (Rasch and Wyngaard 1997).
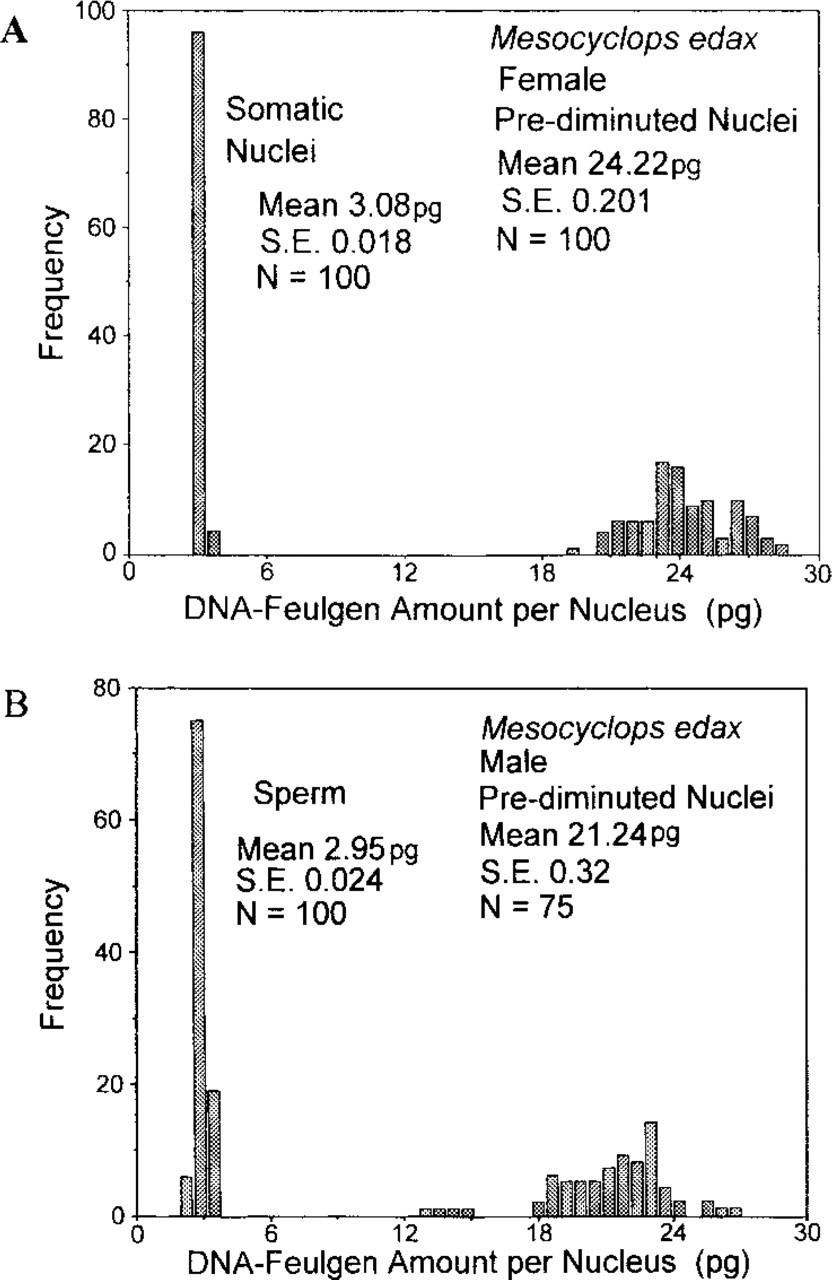
(
The heterogeneity of PD nuclear DNA levels, coupled with the absence of nurse cells and lack of mitosis preceding the appearance of large darkly stained nuclei in the gonadal regions of both males and females of M. edax, suggests that cycles of DNA endoreduplication may occur in germ cells during oogenesis (and perhaps spermatogenesis) well before egg maturation, fertilization, and the start of preparations for chromatin diminution that is routinely scheduled for an early cleavage division. The ready availability of multiple copies of DNA transcripts that can be used for protein biosynthesis during egg growth and maturation, plus a program for the providential removal of the “excess” DNA from presumptive SC nuclei by the process of chromatin diminution during a fourth or fifth cleavage division in M. edax, may constitute alternative mechanisms for more effective reproduction of this species in nature. Conversely, the elevated DNA levels of copepod germ cell nuclei may be more closely associated with nucleoskeletal DNA and/or genome evolution, as first proposed by Cavalier–Smith (1982) and later more fully explored by Gregory (in press).
Differences in DNA content of nuclei from somatic and germ cells of Mesocyclops edax a
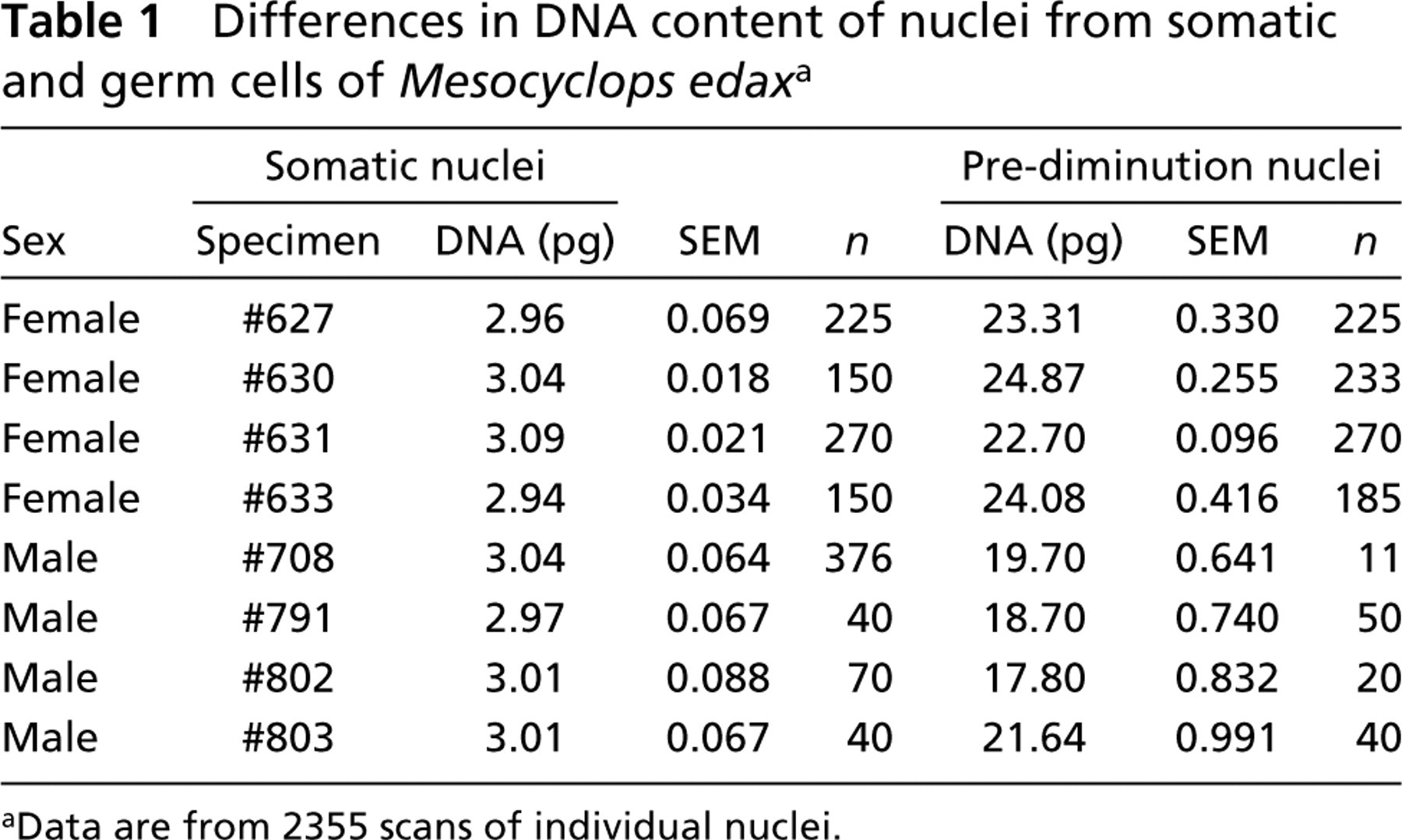
aData are from 2355 scans of individual nuclei.
