Abstract
The distribution of ribosomes in mature human mast cells, a major granulated secretory cell, does not resemble that in other secretory cells, such as pancreatic acinar cells and plasma cells. By routine ultrastructural analysis, ribosomes in human mast cells are often close to, attached to, or even appear to be within secretory granules. To document better these relationships, we used multiple electron microscopic imaging methods, based on different principles, to define RNA, ribosome, and granule relationships in mature human mast cells. These methods included EDTA regressive staining, RNase digestion, immunogold labeling of ribonucleoproteins or uridine, direct binding or binding after ultrastructural in situ hybridization of various polyuridine probes to polyadenine in mRNA, and ultrastructural autoradiographic localization of [3H]-uridine incorporated into cultured human mast cells. These different labeling methods demonstrated ribosomes, RNA, U1SnRNP (a small nuclear RNP specific for alternative splicing of mRNA), mRNA, and uridine to be associated with secretory granules in human mast cells, implicating granules in a larger synthetic role in mast cell biology.
Keywords
R
We (Dvorak 1989) and others (Asboe-Hansen 1971) have noted that the regularly observed patterns of human mast cell granules (i.e., crystals, scrolls, particles, homogeneously dense, mixed) are intermittently accompanied by small numbers of irregular, electron-dense, ragged particles. These structures are often crowded close to and into apparently open ends of scrolls, which abut poorly visible membranes peripherally in granules. These structures are also intimately close to and even appear to be attached to granules and their membranes. In addition, it has long been recorded that human mast cells do not have extensive arrays of rough endoplasmic reticulum such as characterize pancreatic acinar and plasma cells (Palade and Porter 1954; Palade 1955; Palade and Siekevitz 1956; Hammel et al. 1987,1998; Dvorak 1989,1991,1992; Dvorak and Monahan-Earley 1992,1995). Recent studies, using an enzyme affinity-gold method developed to detect RNA (Bendayan 1981), were done with human mast cells (Dvorak and Morgan 1998). They showed extensive secretory granule labeling with an RNase-gold affinity reagent that was determined through many controls to reflect the presence of heparin in granules (Dvorak and Morgan 1998). A possible signal from RNA could not be unambiguously determined in or close to granules because the binding of the RNase-gold, based on the known inhibition of RNase by heparin in granules, was so extensive (Dvorak and Morgan 1998). Therefore, we pursued other avenues to examine the possible close association(s) of RNA with secretory granules in human mast cells. These include direct electron microscopic inspection, RNase digestions, regressive EDTA staining for RNA (Bernhard 1969), immunogold studies using autoimmune human sera containing antibodies to various ribonucleoproteins, a variety of ultrastructural methods to detect polyadenine in mRNA, ultrastructural autoradiographic analysis of [3H]-uridine incorporation, and immunogold detection of uridine. These efforts suggest a close association between secretory granules and RNA in human mast cells.
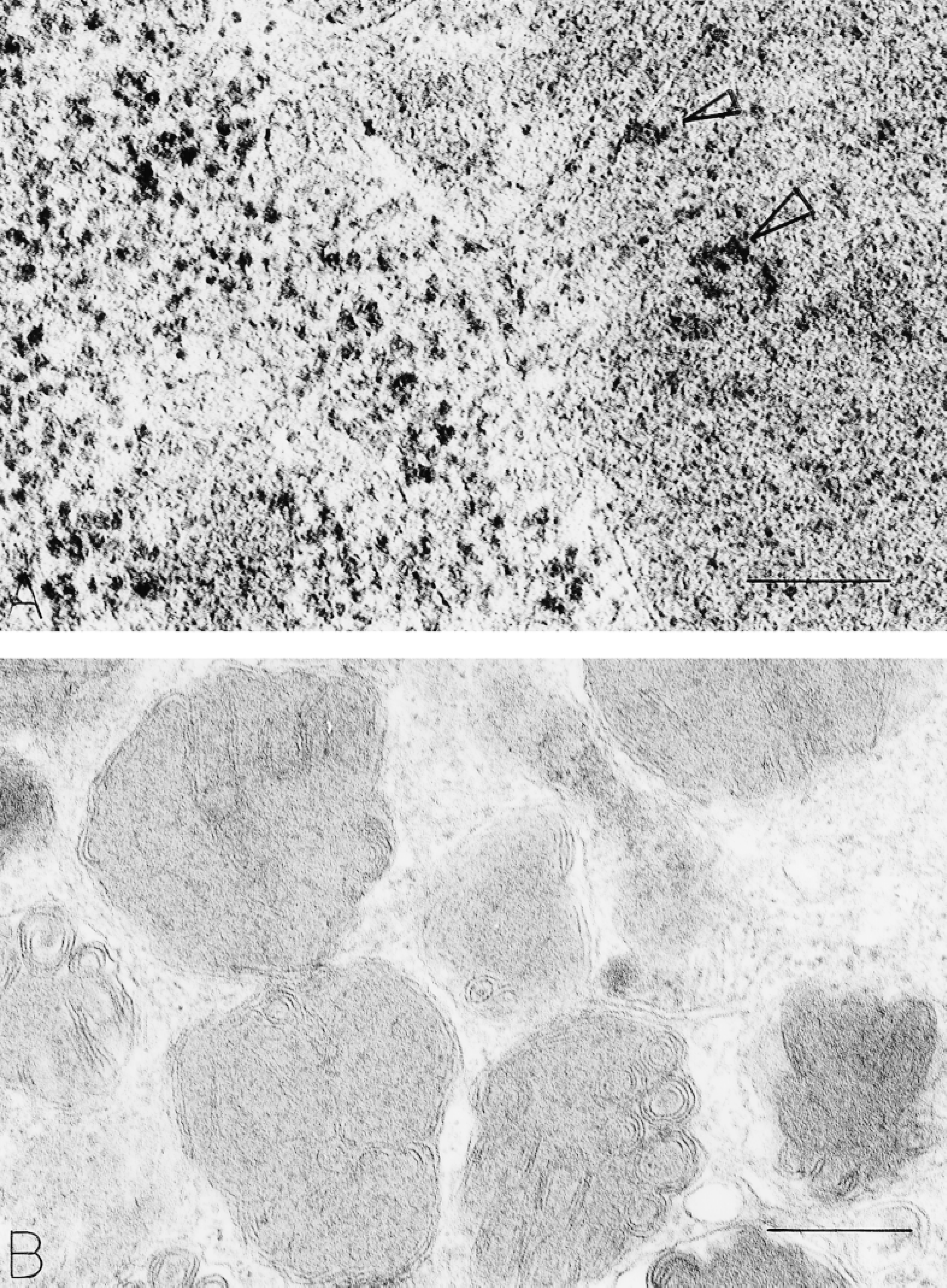
Ribosomes in human mast cells. (
Materials and Methods
Procedures for the isolation, partial purification, and stimulation of human lung mast cells (Schulman et al. 1982), standard electron microscopic specimen preparations (Dvorak 1987), RNase digestions of samples before embedding or en grid (Dvorak and Morgan 1998), Bernhard's regressive EDTA stain for RNA (Bernhard 1969), an enzyme affinity-gold method to detect RNA (Bendayan 1981), postembedding ultrastructural immunogold (Dvorak et al. 1992,1994,1997; Beil et al. 1996; Login et al. 1992,1995,1997a,b; Qu-Hong et al. 1997), ultrastructural autoradiography (Dvorak et al. 1983; Dvorak et al. 1984), and in situ hybridization (Binder et al. 1986; Capco and Jeffery 1978) have all been reported. Key materials used included [3H]-uridine (New England Nuclear; Boston, MA), human autoimmune IgG fractions specific for U1SnRNP (a-RNP) and mitochondria (a-mito) (Immunovision; Springdale, AZ), antibody to uridine (Research Plus; Bayonne, NJ), polyuridine (PU) (LKB; Piscataway, NJ), polyadenine (PA) (LKB), [3H]-polyuridine (New England Nuclear), gold tetrachloride (Sigma; St Louis, MO), photobiotinylation kit (Vector Labs; Burlingame, CA), RNase (E.C.#3.1.27.5; Worthington Biochemical, Freehold, NJ), bovine serum albumin (BSA) (Sigma), glutaraldehyde (Electron Microscopy Sciences; Fort Washington, PA), Ilford L4 emulsion (Polysciences; Warrington, PA), ELON (Kodak; Rochester, NY), sodium sulfite (Sigma), potassium thiocyanate (Sigma), and potassium bromide (Sigma).
All samples were examined by two observers in either a Philips 300 or a Philips 400 electron microscope.
Results
Evidence for Electron-dense Particles and RNA Associated with Human Mast Cell Granules
Human mast cell granules display a number of ultrastructural patterns in routinely prepared ultrastructural samples (reviewed in Dvorak 1989). Three major patterns have been termed crystal, scroll, and particle granules. Smaller numbers of granules are homogeneously dense; many granules are filled with mixtures of these components. Isolated human lung mast cells display all of these patterns. In addition, however, and in routine preparations, electron-dense ragged particles associated with granule matrix, membrane, or packed into close quarters near granules are present (Figures 1A and 1C). Rough endoplasmic reticulum is vanishingly rare in these mature mast cells. Clusters of typical nonmembrane-bound cytoplasmic ribosomes are apparent, both close to (Figures 1A and 1C) and remote from granules.
Digestion with RNase of fixed samples before embedding or of thin sections on grids before staining with lead citrate and of samples prepared either with or without osmium and uranyl en bloc showed reductions in the visibility of cytoplasmic ribosomes (Palade and Siekevitz 1956) and in granule and perigranule particulate densities (Figure 1B). Considerable inter-and intrasample variability of the RNase digestion experiments existed, however. We next used Bernhard's regressive EDTA stain (Bernhard 1969) to evaluate RNA distribution in human mast cells. This method bleaches DNA to electron lucency, thereby enhancing the electron density of RNA in samples. RNA-rich nuclear structures and cytoplasmic ribosomes were electron-dense (and nuclear chromatin electron-lucent) in human mast cells prepared with this method. In addition, secretory granules encased in dense particles and containing some were evident (Figure 2). Some ribosomes were attached to granule surfaces. Many secretory granule matrices were electron-lucent with this regressive stain for RNA.
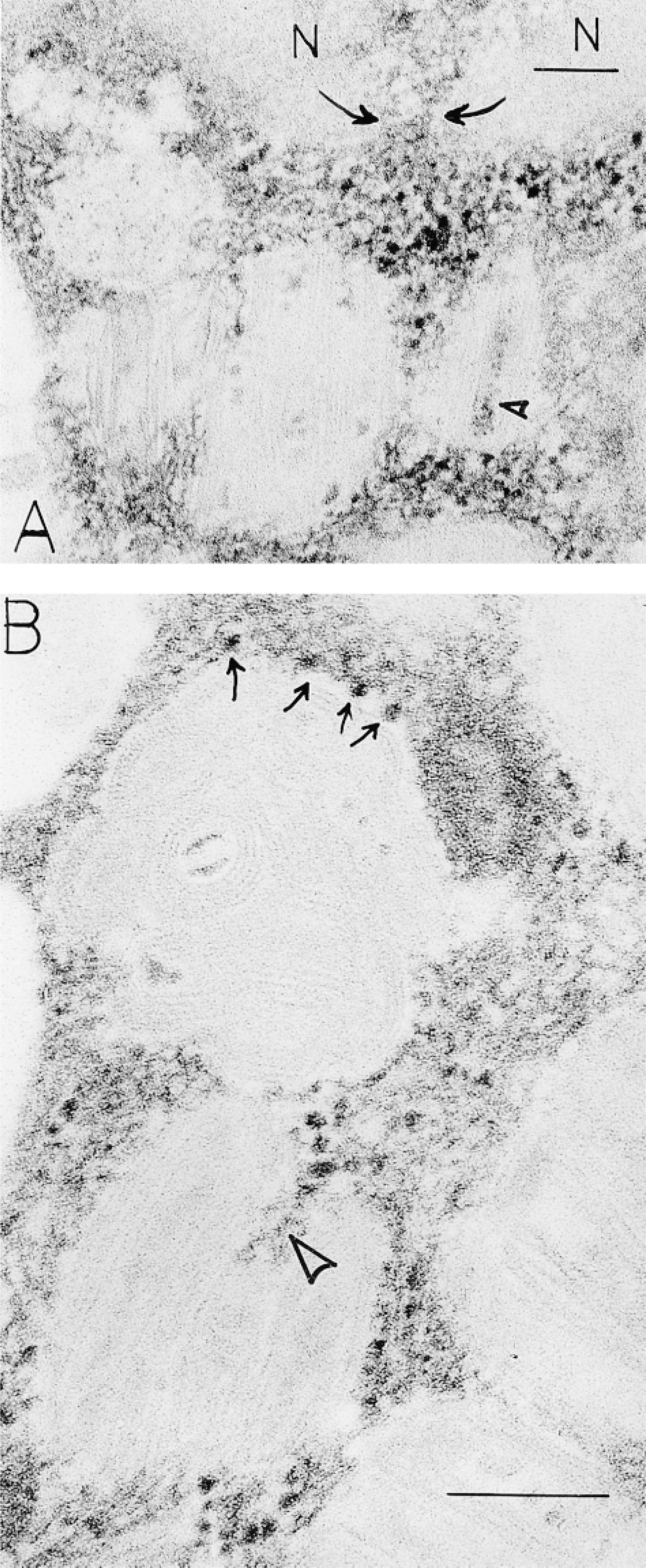
Human mast cells, stained with the Bernhard EDTA regressive method for RNA, show ribosomes as electron-dense ragged particles surrounding cytoplasmic granules. Nuclear (N) chromatin is electron-lucent and electron-dense RNA fills the nuclear pore (
Multiple Mechanistic Methods Define RNA Distribution in Human Mast Cells
An enzyme affinity-gold ultrastructural method (Bendayan 1981), based on the affinity of the enzyme RNase for its substrate RNA, showed extensive binding of RNase-gold to human mast cell granules as reported earlier (Dvorak and Morgan 1998). Binding was reduced when osmium was omitted but was still extensive. As in previous work (Dvorak and Morgan 1998), virtually all of this label was demonstrated to be bound to granule heparin, a known RNase inhibitor (Lindahl and Höök 1978; Salmivirta et al. 1996), and was present in human mast cell granules in great quantities (Dvorak and Morgan 1998) (Figure 3A). Reductions in binding occurred with RNA blocking but these qualitative reductions were not statistically significant (Dvorak and Morgan 1998). The RNase-gold method (Bendayan 1981) for defining a relationship of RNA to secretory granules, although suggestive, could not be used for this purpose.
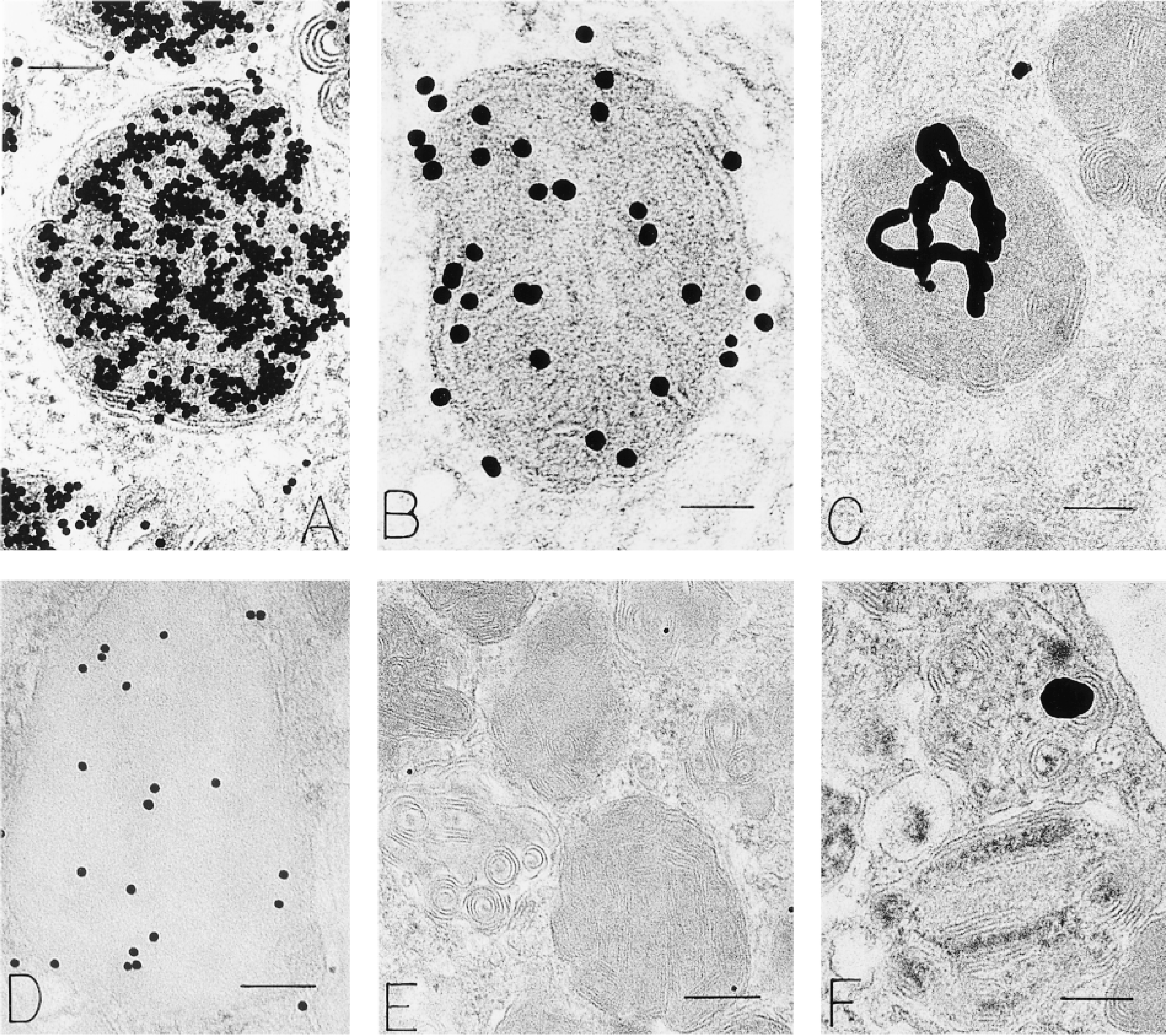
Human mast cell granules labeled with RNase-gold (
Autoantibodies circulate in humans during specific diseases (Conner et al. 1982; Tan 1991; Galperin et al. 1996; Steiner et al. 1996). Some of these are specific for different types of ribonucleoproteins and occur regularly in patients with systemic lupus erythematosus and related diseases (Farris et al. 1997). We did postembedding immunogold staining with several of these and found granule, cytoplasmic and nuclear labeling (Figure 3D; and unpublished data). The nuclear and cytoplasmic labeling for anti-RNP was similar to that previously reported (Farris et al. 1997); binding to secretory granules has not been noted for human mast cells or secretory cells generally. The antibody to RNP localized to granules, lipid bodies, and nucleus in human mast cells. Omission of primary antibodies and substitution with irrelevant primary antibodies or human serum (Figure 3E) did not label human mast cells.
[3H]-Uridine has been used for years to label RNA (Ornelles et al. 1986; Davis et al. 1987). We used this isotope combined with electron microscopic autoradiography to localize RNA in isolated, purified human lung mast cells (Figures 3C, 3F, and 4). In these studies, multiple recovery times in culture after stimulation with anti-IgE (to degranulate human mast cells) (Dvorak 1991) were also examined for isotope incorporation. These studies showed that nuclear, granular and cytoplasmic compartments of human mast cells were labeled with [3H]-uridine (Figure 4). Of 5812 autoradiographic grains counted in 121 cells (cultured for 18 hr with [3H]-uridine), the relative distribution of label associated with granules (20%) exceeded that overlying cytoplasm (10%), mitochondria (1%), and Epon (6%) (p<0.001); 63% of the autoradiographic grains were, as expected, in the nuclear compartment. The preparations of human lung mast cells also contained small numbers of eosinophils. Granules in these secretory cells, as reported (Wickramasinghe and Hughes 1978), also incorporated [3H]-uridine (Figure 5).
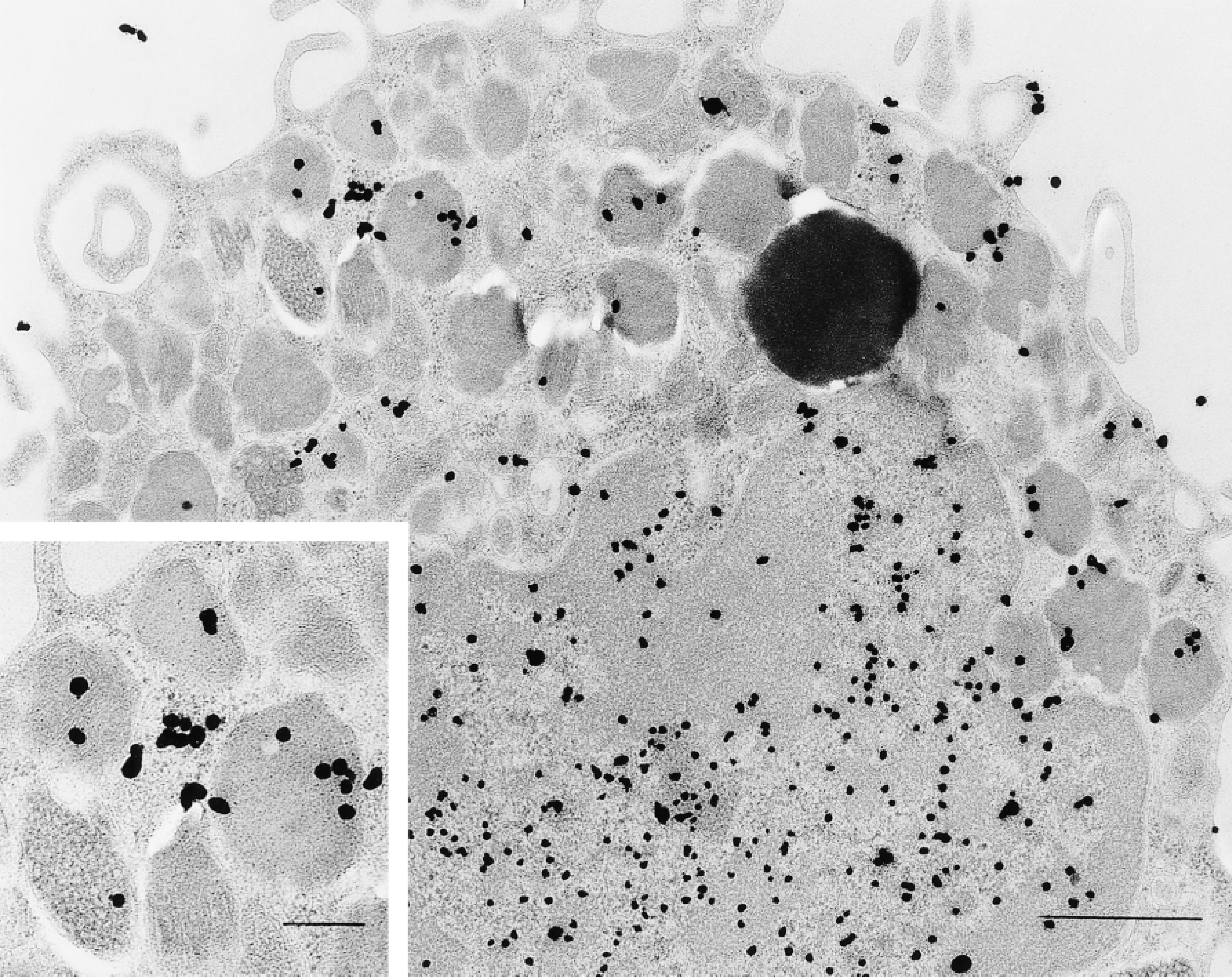
Isolated human lung mast cell incubated for 18 hr in [3H]-uridine and prepared for autoradiography was developed for small silver grains and shows extensive nuclear and secretory granule label. Additional label was associated with clusters of ribosomes adjacent to granules (
We used a series of affinity probes to bind directly to thin sections of human mast cells en grid, as well as in several in situ hybridization (ISH) protocols (Capco and Jeffery 1978; Binder et al. 1986), to localize messenger RNA based on the affinity of polyuridine (PU) for the polyadenine (PA) tail of mRNA (Ornelles et al. 1986) (Figure 6). The methods so examined included the following affinity probes for postembedding detection of mRNA: [3H]-polyuridine ([3H]-PU) (Figure 6G); polyuridine-gold (PUG) (Figures 6A–6F); polyuridine-biotin (PUB) (Figure 6H); polyuridine-BSA-gold (PUBG) (Figure 6J). Several reporter systems were used to image the photobiotinylated PU probe. [3H]-PU, after an ISH protocol, was imaged by photographic development analogous to that used for autoradiographic samples and gold-labeled probes (PUG, PUBG) were imaged directly with and without an ISH protocol. With all of these methods, granule label was observed (Figures 6A–6D, 6G, 6H, and 6J). Expected nuclear and cytoplasmic sites were also labeled. Appropriate positive (Figure 6D) and negative (Figures 6E, 6F, 6I, and 6K) controls for these methods were done.
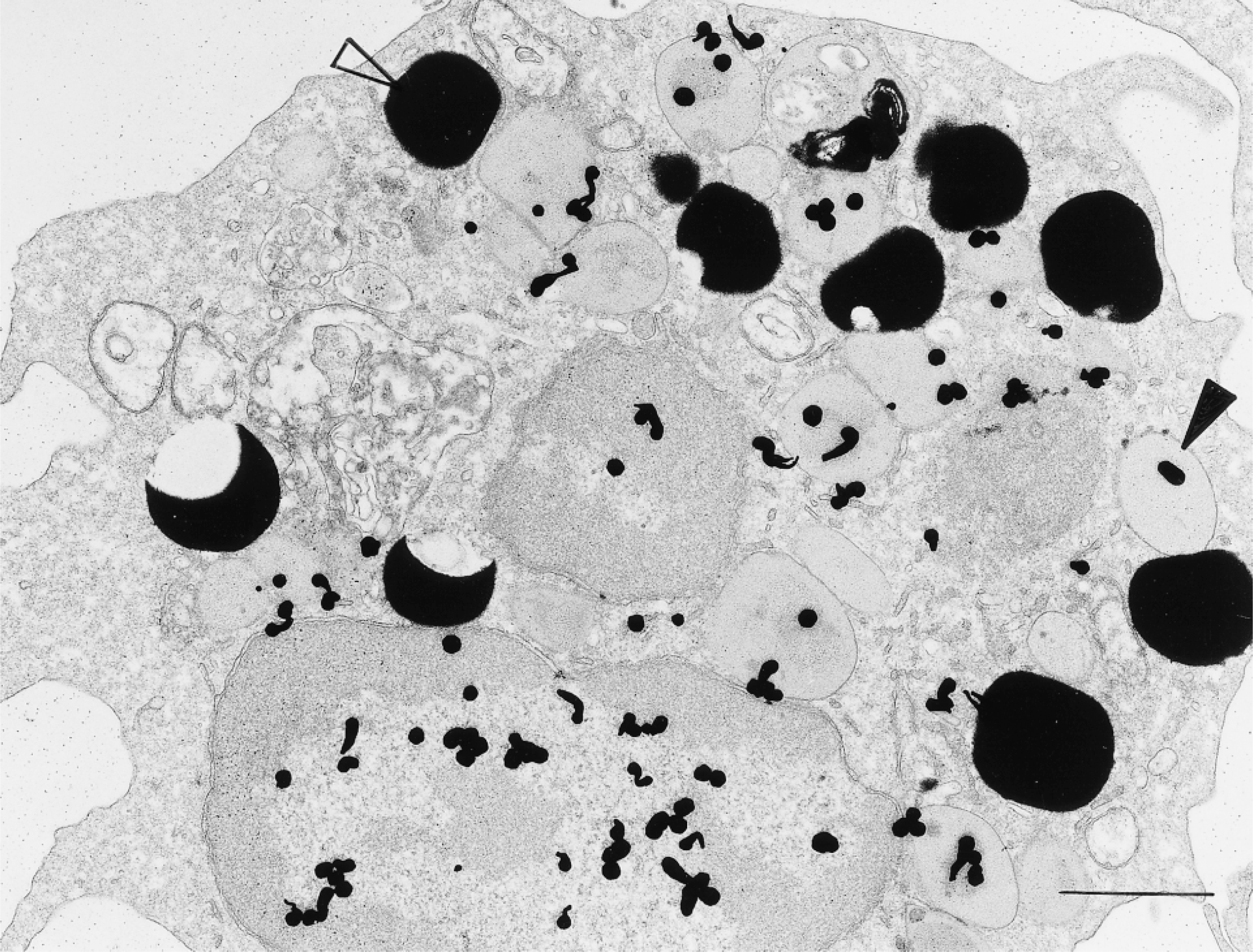
A human eosinophil present in a preparation of isolated human lung mast cells similar to Figure 4 was incubated for 2 hr in [3H]-uridine, prepared for autoradiography, and developed for small silver grains. The nucleus and secretory granules (closed arrowhead) are extensively labeled. Electron-dense lipid bodies (open arrowhead) are not labeled in this cell. The background is free of label. Bar = 1 μm.
Quantitation of the ISH procedures was done as follows. The granule gold density for PUG, imaged with an ISH protocol (Figure 6C), was 31.7/μm2, a value that decreased to 3/μm2 when absorption of the PUB reagent with PA beads preceded an ISH protocol (Figures 6E and 6F) (p<0.001). When ISH with [3H]-PU was done, 48% of a total of 370 autoradiographic silver grains were associated with granules (Figure 6G) in a total of 35 cells. ISH using a PUB probe revealed a similar proportion of gold particles (45% of a total of 455 gold particles) associated with granules (Figure 6H). This value was zero (0) when an omission control for this experiment was done (Figure 6I). In these experiments, the proportion of label associated with granules exceeded that for the Epon, cytoplasmic, and mitochondrial compartments (p<0.001) but was not significantly different from that for the nuclear compartment (38%), a good positive internal control for the procedure. Of a total of 1868 gold grains counted with the PUBG probe used with an ISH protocol, 1380 (70%) were associated with granules (Figure 6J). ISH using gold alone, a control for this procedure (Figure 6K), resulted in only six gold particles overlying granules (Figure 6K).
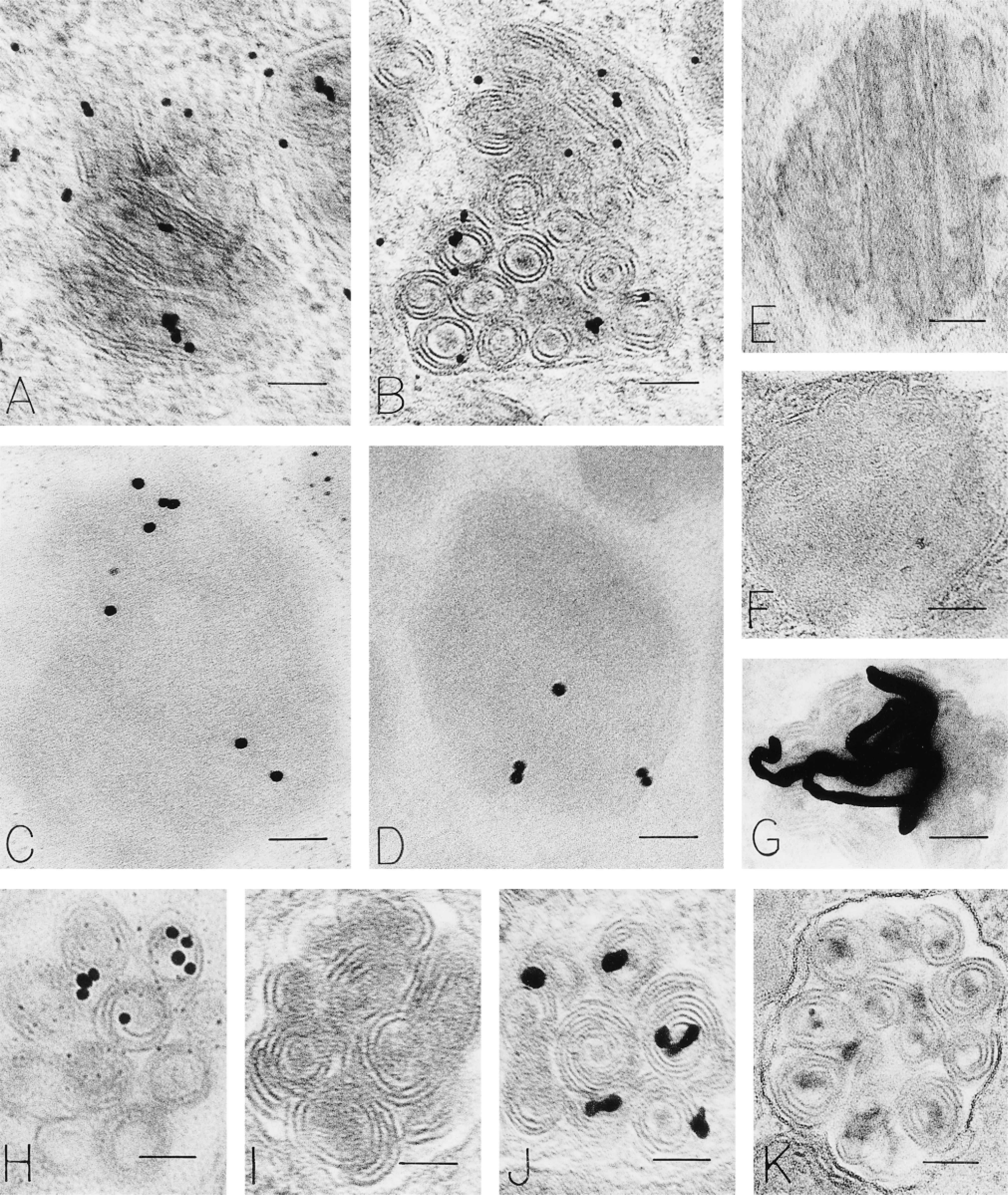
Human mast cell granules labeled with polyuridine methods to identify mRNA. Isolated, purified human lung mast cells were cultured for 6 hr, fixed, and processed for electron microscopy and prepared to detect polyadenine in mRNA by direct en grid binding of polyuridine-gold (
An antibody to uridine was used in conjunction with a postembedding immunogold method to localize RNA in human mast cells. In these experiments, nuclear, cytoplasmic, and granular compartments were labeled with gold particles, indicating the presence of uridine (Figure 3B). Omission of the primary antibody, substitution of irrelevant antibody or normal goat serum, and absorption with solid-phase PU resulted in a lack of label in human mast cells.
Discussion
Secretory granules have generally been viewed as storage organelles for synthetic products destined for export by regulated secretion from classical secretory cells (Palade 1975). Most of these cells have extensive networks of rough endoplasmic reticulum (RER) and Golgi structures. Mature human mast cells and granulocytes are also secretory cells characterized by secretory granules. Extensive networks of RER and Golgi structures are not prominently featured in these cells, however. In the work that we present here, we implicate secretory granules in synthetic processes of mast cells. The study was stimulated by observations of ribosomes within, as well as adjacent and attached to, secretory granules in cells relatively devoid of classical synthetic structures. We reasoned that these associations might be significant. Using multiple mechanistic methods for high-magnification imaging of ribosomes, RNA, and ribonucleoproteins (RNPs), we demonstrate these close associations in mature human mast cells.
Human mast cell secretory granules have variegated ultrastructural patterns (see Dvorak 1989 for review). Several patterns include scrolls and crystals, but many mast cell granules are filled with mixtures of these patterns, with homogeneously electron-dense material, particulate material, and reticular threadlike structures. Human mast cell granules are often enmeshed in cytoskeletal filaments of variable size. Microtubules are present, but particular associations with granules are not evident. Often, human mast cell granules are not completely enclosed within covering organellar membranes, and scrolls, filaments, and ribosomes crowd into these areas, often with no distinction between cytoplasm and granule interior. Ragged particles resembling ribosomes also can be found in granule interiors (Figure 1C).
Initial experiments were done using RNase digestion of samples at different stages of preparation for electron microscopy. In some preparations, dramatic reduction in granule particles was evident, suggesting the presence of granule ribosomal (Palade and Siekevitz 1956) or U1SnRNP (Conner et al. 1982) RNA; mRNA resists digestion with pancreatic RNase (Brawerman 1974). An enzyme affinity technique to identify RNA by binding RNase-gold to section surfaces (Bendayan 1981) avidly labeled mast cell granules (and cytoplasmic ribosomes) (Dvorak and Morgan 1998). Extensive controls for this technique revealed an avidity of RNase-gold for human mast cell granule heparin (Dvorak and Morgan 1998). Quantitative studies of RNA blocking showed a reduction in granule label that did not achieve statistical significance, whereas those for heparin blocking did (Dvorak and Morgan 1998). Bernhard's regressive EDTA stain for RNA (Bernhard 1969) revealed nuclear and cytoplasmic RNA sites in human mast cells. Nuclear pores were filled with RNA and perigranular cytoplasm also was darkly stained, indicating the presence of RNA. Individual ribosomes attached to granule surfaces and within granules were electron-dense.
If secretory granules have RNA activity, then RNPs (associated with RNA) may also be present within them. Autoantibodies that have selectivity for some of these RNPs routinely accompany certain human diseases (Conner et al. 1982; Tan 1991; Galperin et al. 1996; Steiner et al. 1996). We selected several of these to examine with postembedding immuogold for granule location in human mast cells (unpublished data). One of these, anti-RNP, is specific for U1RNP, a small nuclear ribonucleoprotein (U1Sn RNP), the most abundant RNP particle in the nucleus and one that is essential for splicing precursor mRNA (Hinterberger et al. 1983; Padgett et al. 1983; Krämer et al. 1984; Petterson et al. 1984). Control preparations with normal human serum or with an irrelevant antibody substituted for anti-RNP did not label granules, nor were granules labeled when omission of the primary antibody or staining with gold alone was done. The specific antibody for a-RNP was checked with reference sera (Communicable Disease Center) and was found to have a single reactivity and no reactivity to DNA (manufacturer's literature). Only patient sera that react with U1 RNP inhibit mRNA splicing in an in vitro system (Padgett et al. 1983), and control of alternative splicing is effected by differential binding of U1 small RNPs (Kuo et al. 1991). Therefore, human mast cell granules contain an RNP important for alternative splicing of mRNA, generally considered to be a nuclear function in RNA processing that creates diversity of specific individual mRNAs (Mattaj 1990).
We next adapted previously published methods (Capco and Jeffery 1978; Binder et al. 1986) for labeling mRNA to our electron microscopic samples. All of these methods were selected to detect poly(A)-positive mRNA generally and are based on the affinity of poly(U) for poly(A) (Ornelles et al. 1986). We used different reporting systems for these methods, either directly visualized or after electron microscopic ISH postembedding protocols. These methods, based on affinity of poly(U) for poly(A) in mRNA, used an autoradiographic reporter ([3H]-PU), a gold reporter directly bound to poly(U) (PUG), a biotin-anti-biotin immunogold reporter for the photoprobe poly(U)-biotin (PUB), or a gold reporter attached to a BSA-poly(U) complex (PUBG). In all of these approaches, human mast cell granules were labeled, as well as expected nuclear and cytoplasmic sites that provide positive internal controls. Appropriate ISH cytochemical controls did not label granules or nuclear or cytoplasmic sites. Therefore, all of these ISH and direct visualization protocols identify poly(A)-containing mRNA in human mast cell secretory granules.
We used two methods to detect uridine stores as a measure of RNA content in human mast cell granules. The first method, a postembedding immunogold protocol with specific immunocytochemical controls, utilized a polyclonal antibody to uridine that recognizes RNA and is known to work on aldehyde-fixed material (manufacturer's literature). Human mast cell granules were specifically labeled with this antibody; controls were negative. The second method was autoradiographic detection of [3H]-uridine incorporation by isolated living human mast cells before fixation and processing for electron microscopy. This method, regularly used for the detection of RNA (Ornelles et al. 1986; Davis et al. 1987), labeled human mast cell granules as well as expected nuclear and cytoplasmic sites, as positive internal controls.
All together, these experiments using different labeling principles document a role for secretory granules in synthetic events in human mast cells by demonstrating the existence of ribosomes, RNA, U1SnRNP (a component of spliceosomes that controls alternative splicing of mRNA), mRNA, and uridine in these secretory organelles.
The presence of ribosomes, RNA, U1SnRNP, and uridine in, around, and attached to secretory granules suggests that an important role(s) for this organellar compartment in RNA metabolism exists in human mast cells. The role might merely reflect a storage function at this site, and cellular RNA levels could be down-regulated by specific stimuli and resultant secretory granule losses. Granule stores of mRNA could also, however, reflect en site synthetic capacities that define a new role(s) for secretory granules in general. A large body of evidence has recently defined RNA transport and localization as a mechanism for facilitating site-specific synthesis at various subcellular organelles in cells of diverse origin from multiple species (Gottlieb 1990; Singer 1992,1993; Wilhelm and Vale 1993; Bassell and Singer 1997; Bassell et al. 1999; Etkin and Lipshitz 1999; Jansen 1999; Lasko 1999; Mowry and Cote 1999). Such traveling RNA has been intimately associated with all three major cytoskeletal elements, i.e., microfilaments, intermediate filaments and microtubules (Lenk et al 1977; Fulton et al. 1980; Ornelles et al. 1986; Isaacs and Fulton 1987; Gottlieb 1990; Singer 1992; Bassell 1993; Bassell and Singer 1997; Mowry and Cote 1999), and specific zip codes for localizing individual mRNAs at subcellular sites have been postulated (Singer 1992; Hill et al. 1994; Kislauskis et al. 1994; Etkin and Lipshitz 1999). Secretory granules have not been considered as such a site, but the evidence that we present suggests that they must also be included in the travels of mRNA (Wilhelm and Vale 1993; Knowles et al. 1996) in mast cell biology.
Footnotes
Acknowledgements
Supported by NIH Grant AI-33372.
We thank Peter K. Gardner for assistance in the preparation of this manuscript.
