Abstract
Introduction:
Chronic pain and depression are highly comorbid and difficult-to-treat disorders. We previously showed this comorbidity is associated with higher depression severity, lower antidepressant treatment effectiveness and poorer prognosis in the Australian Genetics of Depression Study.
Objective:
The current study aimed to assess whether a genetic liability to chronic pain is associated with antidepressant effectiveness over and above the effect of genetic factors for depression in a sample of 12,863 Australian Genetics of Depression Study participants.
Methods:
Polygenic risk scores were calculated using summary statistics from genome-wide association studies of multisite chronic pain and major depression. Cumulative linked regressions were employed to assess the association between polygenic risk scores and antidepressant treatment effectiveness across 10 different medications.
Results:
Mixed-effects logistic regressions showed that individual genetic propensity for chronic pain, but not major depression, was significantly associated with patient-reported chronic pain (PainPRS OR = 1.17 [1.12, 1.22]; MDPRS OR = 1.01 [0.98, 1.06]). Significant associations were also found between lower antidepressant effectiveness and genetic risk for chronic pain or for major depression. However, a fully adjusted model showed the effect of PainPRS (adjOR = 0.93 [0.90, 0.96]) was independent of MDPRS (adjOR = 0.96 [0.93, 0.99]). Sensitivity analyses were performed to assess the robustness of these results. After adjusting for depression severity measures (i.e. age of onset; number of depressive episodes; interval between age at study participation and at depression onset), the associations between PainPRS and patient-reported chronic pain with lower antidepressant effectiveness remained significant (0.95 [0.92, 0.98] and 0.84 [0.78, 0.90], respectively).
Conclusion:
These results suggest genetic risk for chronic pain accounted for poorer antidepressant effectiveness, independent of the genetic risk for major depression. Our results, along with independent converging evidence from other studies, point towards a difficult-to-treat depression subtype characterised by comorbid chronic pain. This finding warrants further investigation into the implications for biologically based nosology frameworks in pain medicine and psychiatry.
Introduction
Over the past three decades, chronic pain and depression have remained in the top three leading causes of global disability (GBD 2017 Disease and Injury Incidence and Prevalence Collaborators, 2018). Both disorders are highly comorbid – having one condition substantially increases the risk of developing the other (Viana et al., 2018), with around one-third of depression patients presenting with severe pain (Stubbs et al., 2017), and 23–60% of chronic pain patients manifesting depression (Orhurhu et al., 2019). The presence of pain is also strongly associated with depression onset and relapse (DeVeaugh-Geiss et al., 2010; Gerrits et al., 2014a, 2014b; Means-Christensen et al., 2008). Furthermore, comorbid chronic pain and depression is typically more difficult to treat than either primary condition alone. However, virtually all clinical guidelines and recommendations for either condition (Bates et al., 2019; Brueckle et al., 2020; Haibach et al., 2014; IsHak et al., 2018; Moore et al., 2015; Urits et al., 2019) have failed to adequately address treatment options for this common comorbidity. This major evidence gap can be attributed (in part) to the reliance on pharmacotherapeutic and other interventional trials excluding specific comorbidities (Baune et al., 2019; Cipriani et al., 2018; Kendrick et al., 2019), and are thus not representative of the inherent clinical heterogeneity across psychiatric populations.
We recently reported on a large pharmacoepidemiological study of comorbid chronic pain and depression (Roughan et al., 2021). This data-driven methodological approach (Bates et al., 2019; IsHak et al., 2018; Moore et al., 2015; Urits et al., 2019) took into account the wide individual variation to examine comorbid chronic pain and depression risk factors and antidepressant treatment effectiveness in the Australian Genetics of Depression Study (AGDS). The main findings concerning disease severity and treatment response were as follows: (1) higher pain severity (intensity) was associated with longer durations of pain and an increased number of depressive episodes; (2) patient-reported effectiveness from taking particular antidepressants (i.e. sertraline, escitalopram, venlafaxine) were significantly lower compared to those without chronic pain; (3) individuals prescribed particular antidepressants (i.e. sertraline, escitalopram, citalopram) for chronic pain (among other indications) reported significantly lower effectiveness compared to those taking the same medications but for a different indication; (4) the vast majority of participants with chronic pain (i.e. 75–98% across antidepressants) reported their antidepressant prescription was not for chronic pain; and (5) individuals with comorbid chronic pain reported fewer positive functional benefits from taking antidepressants (e.g. ‘relief of depressive symptoms’, ‘return of normal emotions’, ‘getting back to normal daily activities’). We therefore provided independent converging evidence that comorbid pain is a strong predictor of non-remission with antidepressant medication treatment as found in previous clinical trials and observational studies (Bair et al., 2004; Fishbain et al., 2014)).
Here, we continued this work by leveraging genotype data in AGDS to investigate whether the phenotypic association between chronic pain and a poorer antidepressant effectiveness is genetically mediated. The relationship between depression severity and lower antidepressant effectiveness is established (Gueorguieva et al., 2011). We previously reported an association between chronic pain and lower patient-reported antidepressant effectiveness (Roughan et al., 2021). However, this negative association could be confounded by the (dose-dependent) correlation between chronic pain and depression severity (DeVeaugh-Geiss et al., 2010), and by the expected relationship between depression severity and antidepressant effectiveness. Using genome-wide association study (GWAS) summary statistics for multi-site chronic pain (Johnston et al., 2019) and major depression (MD) (Wray et al., 2018), polygenic risk scores (PRS) (Wray et al., 2020) were generated for each of these conditions. We examined whether PainPRS and MDPRS were associated with patient-reported chronic pain, and antidepressant treatment response. We further examined whether any associations between patient-reported chronic pain and PainPRS with lower antidepressant effectiveness are robust to measures of MD severity and liability. These analyses aim to disentangle the confounding associations and determine whether genetic risk for chronic pain predicts antidepressant treatment response independently of the genetic risk for depression.
Methods
Sample recruitment
The AGDS sample was recruited through two main streams: (1) a mailout by the Australian Department of Human Services (DHS) on our behalf, targeted at people who had been prescribed antidepressants at least four times in the past 4.5 years; and (2) a national media publicity campaign. Interested participants were directed to a website where they provided informed consent prior to participating through online questionnaires. Upon completion of the core questionnaire, participants were mailed a GeneFix GFX-02 2 mL saliva DNA extraction kit (Isohelix plc) to use at home and then returned through the post for subsequent genotyping. The package included a consent form specific to genetic information to be signed by participants. Sample recruitment has been extensively described elsewhere (Byrne et al., 2020). All protocols and questionnaires were approved by the QIMR Berghofer Medical Research Institute’s Human Research Ethics Committee.
Phenotype ascertainment
This study focuses on patient-reported antidepressant effectiveness (N = 12,863) (Byrne et al., 2020). Participants were first asked to confirm whether they had been prescribed any of the 10 most commonly prescribed antidepressants in Australia (i.e. sertraline, escitalopram, venlafaxine, fluoxetine, citalopram, desvenlafaxine, duloxetine, mirtazapine, amitriptyline, paroxetine). For each antidepressant participants had taken, they were asked to rate its effectiveness using the item: ‘How well does/did [name of the antidepressant] work for you?’. The possible responses were as follows: ‘not at all well’, ‘moderately well’ and ‘very well’. Participants failing to self-report a diagnosis of major depressive disorder (MDD) were excluded from the current study. Data on chronic pain were gathered through the general health and lifestyle module of our online survey (N = 11,370) (Roughan et al., 2021). Participants were asked whether they experienced pain in their daily life and to rank its intensity on a scale from 0 to 10. Only those reporting pain (i.e. rating > 0) progressed to the remainder of the pain module, which included questions about pain duration and location. We defined chronic pain according to the International Association for the Study of Pain (IASP) as pain persisting or recurring for longer than 3 months (Treede et al.,2015, 2019). We have previously reported on the comorbidity between chronic pain, depression severity and their phenotypic association with antidepressant treatment response in this sample (Roughan et al., 2021).
Genotyping and QC
Genotyping was performed using the Illumina Global Screening Array (GSA V.2.0.) across three genotyping centres. A common set of high-quality markers between the different genotyping centres was identified prior to joint imputation. Pre-imputation marker exclusion criteria consisted of unknown or ambiguous map position, strand alignment in a BLAST search, missingness > 5%, p(HWE test) < 10−6, MAF < 1% and GenTrain score < 0.6. The Michigan imputation server was used to impute the genotypes using the HRCr1.1 reference panel. Individuals were excluded based on high missingness (missing rate > 3%), inconsistent (and unresolvable) sex, or if they were deemed ancestry outliers from the European population (i.e. > 6 SD from the first two genetic principal components).
Polygenic risk scoring
We employed a recently developed method, SBayesR (Lloyd-Jones et al., 2019; Ni et al., 2020) to estimate conditional (i.e. simultaneously fitting all genetic variants) GWAS summary statistics for both multisite chronic pain (Johnston et al., 2019) and MD (Wray et al., 2018). This process prevents inflation arising from using correlated single-nucleotide polymorphisms (SNPs) due to linkage disequilibrium (LD) (Lloyd-Jones et al., 2019). Prior to PRS calculations, we excluded insertion/deletion variants, those with strand ambiguity or low-imputation quality (R2 < 0.6). We then used the resulting GWAS effect sizes to estimate multisite chronic pain PRS (PainPRS) and MD PRS (MDPRS) with PLINK 1.9 (Chang et al., 2015) on the AGDS imputed genetic dosages.
Statistical analyses
All analyses were performed on a subset of unrelated participants with European Ancestry. Mixed-effects logistic regressions were used to examine the association between patient-reported chronic pain and PainPRS or MDPRS. The regressions were adjusted for sex, age at study enrolment, and the first 20 genetic principal components to adjust for potential population stratification. Cumulative linked mixed models (CLMM; also known as mixed ordered logistic regressions) were used to assess the association between the main variables of interest (i.e. PainPRS, MDPRS) and overall treatment effectiveness. This more complex modelling was warranted given the ordinal nature in which antidepressant effectiveness was measured. A fully adjusted regression with both PainPRS and MDPRS was used to assess whether any association between MDPRS and antidepressant effectiveness is mediated by a genetic correlation between MD and chronic pain. To assess overall treatment response, repeated measures from participants taking more than one antidepressant were adjusted by including a random effect intercept for each participant. CLMM were also used to assess the association between the effectiveness of specific antidepressants with PainPRS and MDPRS. Statistical significance was defined with Bonferroni multiple-testing correction per PRS of interest, adjusting for the repeated testing of 10 antidepressants (p < 0.005). These analyses were performed in R using the base and ordinal libraries.
Results
The final sample consisted of 12,863 participants with genotype and patient-reported antidepressant effectiveness data. Of these, 5809 (45%) participants reported having chronic pain; 3784 (30%) did not report chronic pain; and 3270 (25%) had missing data from the pain module survey. Participants with missing data from the pain module had similar demographics to the other subgroups (Table 1) and were used for polygenic risk scoring as they had genotype data available. Mean age was 43.7 (SD = 15.1) years and approximately 76% were female. Of the participants with available data on pain, 60% fulfilled the current IASP definition (Treede et al., 2019) used for chronic pain. We found PainPRS was significantly associated with chronic pain (OR = 1.17 [1.12, 1.22]), whereas MDPRS was not associated with chronic pain (OR = 1.01 [0.98, 1.06]).
AGDS sample composition.
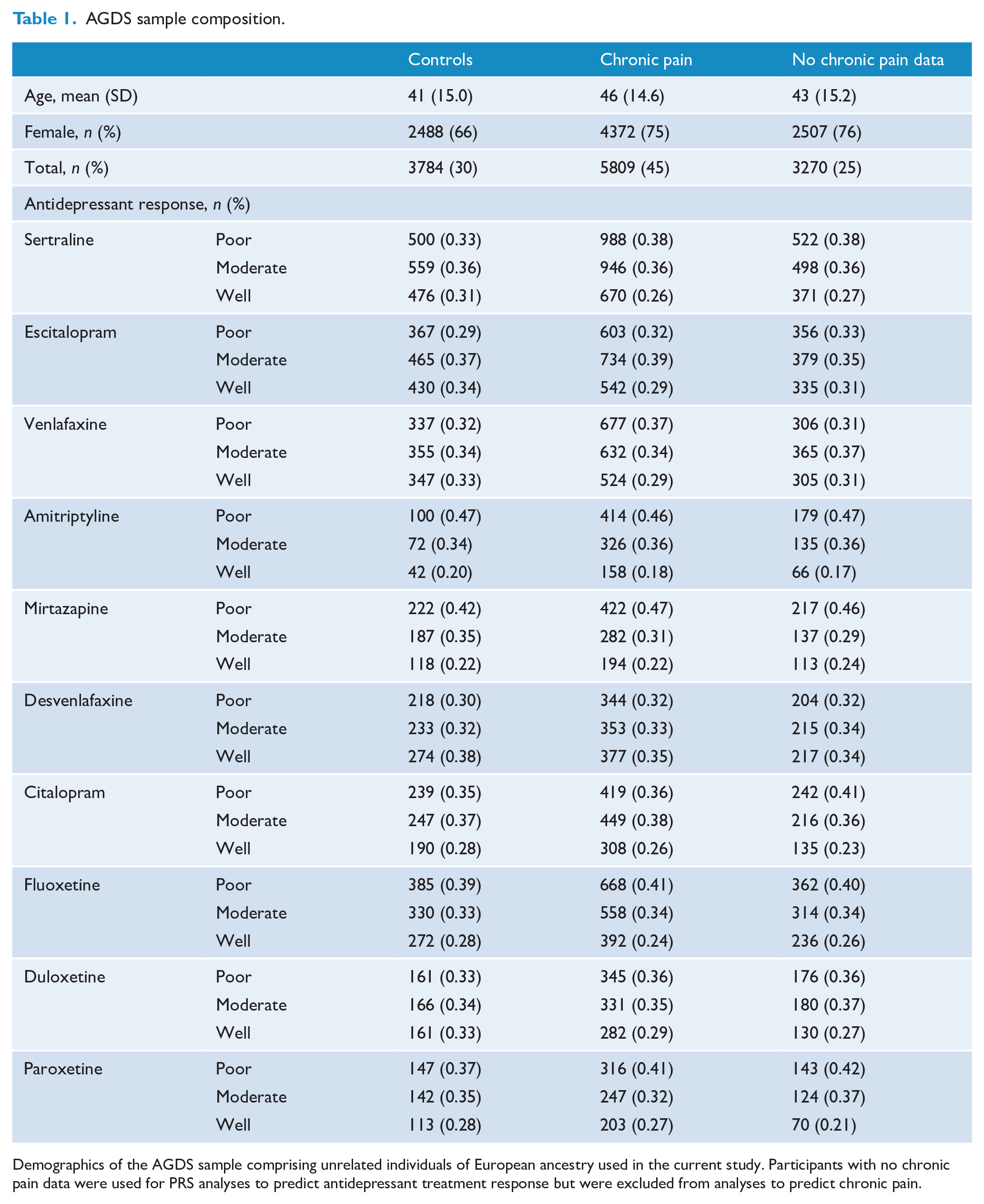
Demographics of the AGDS sample comprising unrelated individuals of European ancestry used in the current study. Participants with no chronic pain data were used for PRS analyses to predict antidepressant treatment response but were excluded from analyses to predict chronic pain.
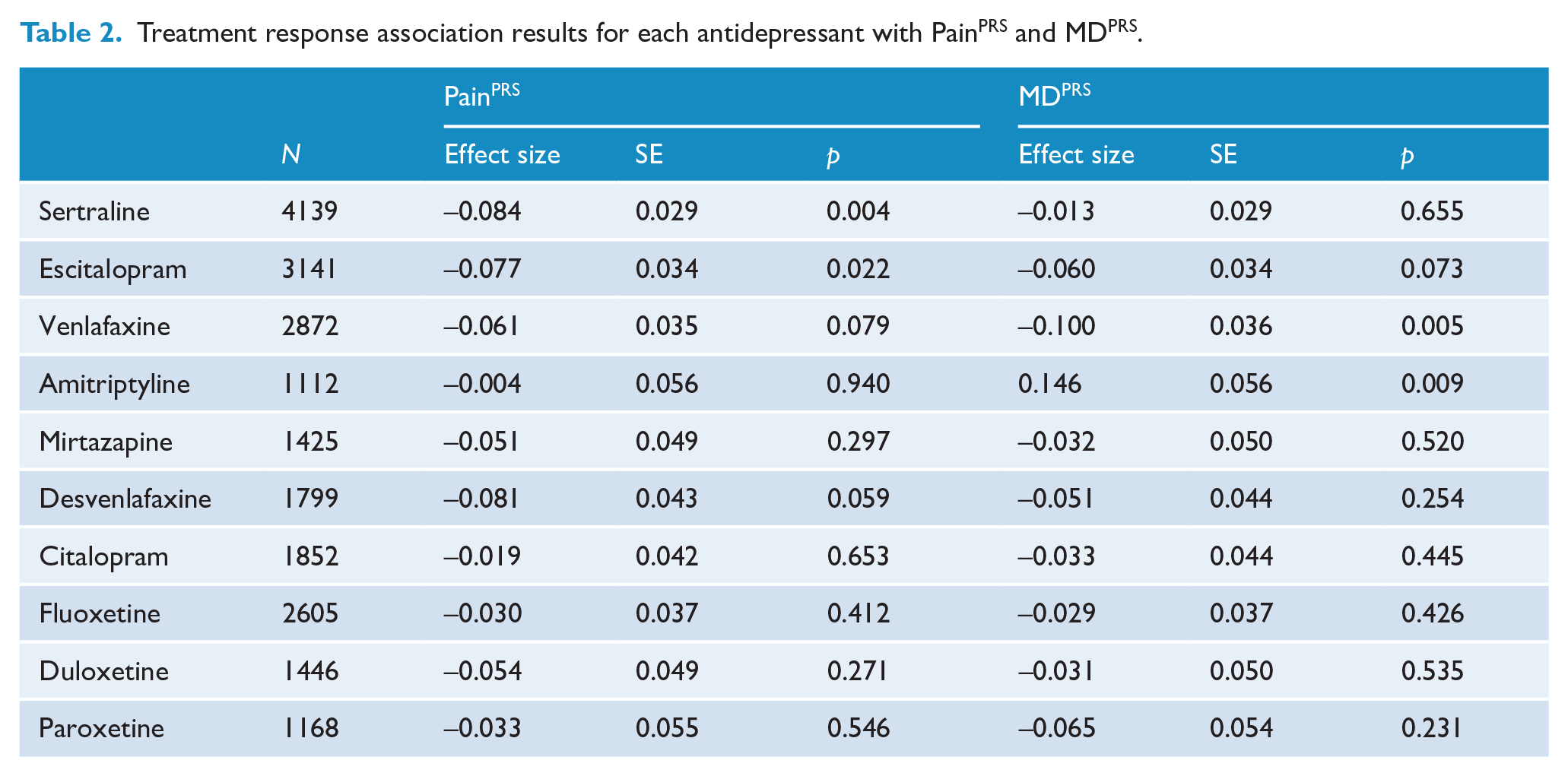
PRS for chronic pain and for MD were both associated with lower antidepressant treatment response (PainPRS OR = 0.93 [0.90, 0.96]; MDPRS OR = 0.95 [0.92, 0.99]). However, the fully adjusted model suggests the association with PainPRS (adjOR = 0.93 [0.90, 0.96]) is independent from the effect of MDPRS (adjOR = 0.96 [0.93, 0.99]) (Figure 1). Fully adjusted models stratified by antidepressant type also showed significant associations between (1) MDPRS with higher effectiveness of amitriptyline and suggestively lower effectiveness of venlafaxine; and (2) PainPRS with lower effectiveness of sertraline and suggestively lower effectiveness of escitalopram (Table 2 and Figure 2).
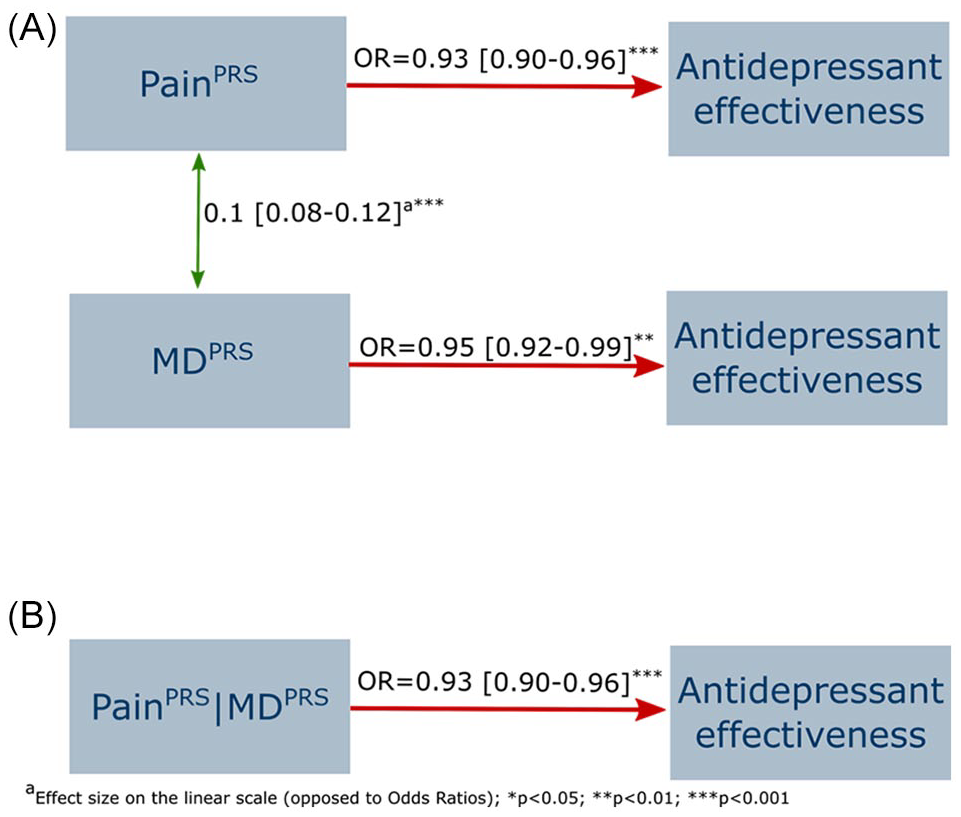

Association between MDPRS and (chronic) PainPRS with antidepressant effectiveness across all medications.
Treatment response association results for each antidepressant with PainPRS and MDPRS.
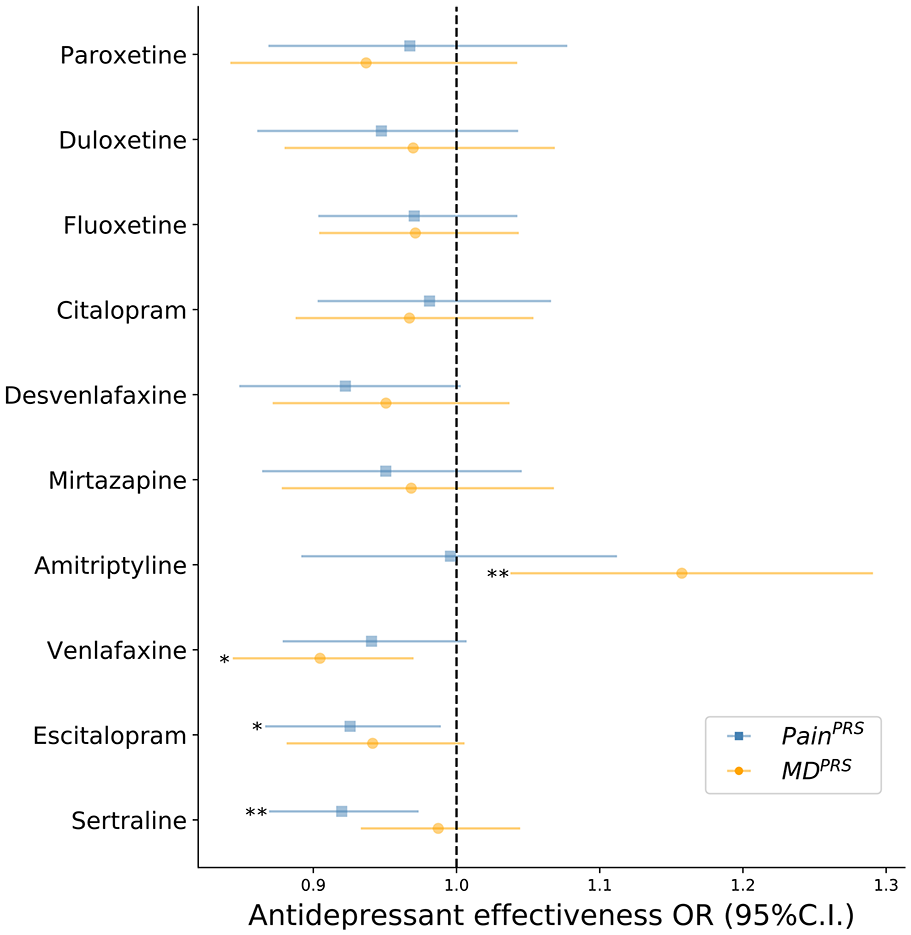
Treatment effectiveness associated with PainPRS or MDPRS stratified by specific antidepressant.
Sensitivity analyses were performed to assess the robustness of these results by adjusting for two measures of depression severity – age of depression onset and number of depressive episodes. We also adjusted for the interval between age of onset of depression and age at participation as we expect a positive relationship between this interval and number of episodes. These analyses showed that only the relationship between PainPRS and overall lower antidepressant effectiveness was sustained (OR = 0.95 [0.92, 0.98]). The same was true for patient-reported chronic pain (OR = 0.84 [0.78, 0.90]), but not for MDPRS (OR = 0.98 [0.94, 1.01]). Thus, genetic risk for chronic pain is associated with poorer antidepressant effectiveness and a mediator of treatment response that is independent of the genetic risk and severity of MD. The sensitivity analyses further revealed significant associations were also sustained between (1) PainPRS and lower effectiveness of sertraline; (2) MDPRS and higher effectiveness of amitriptyline; and (3) MDPRS and lower effectiveness of venlafaxine (Table 3 and Figure 3).
Sensitivity analysis association results for each antidepressant treatment response with PainPRS, MDPRS and patient-reported chronic pain.
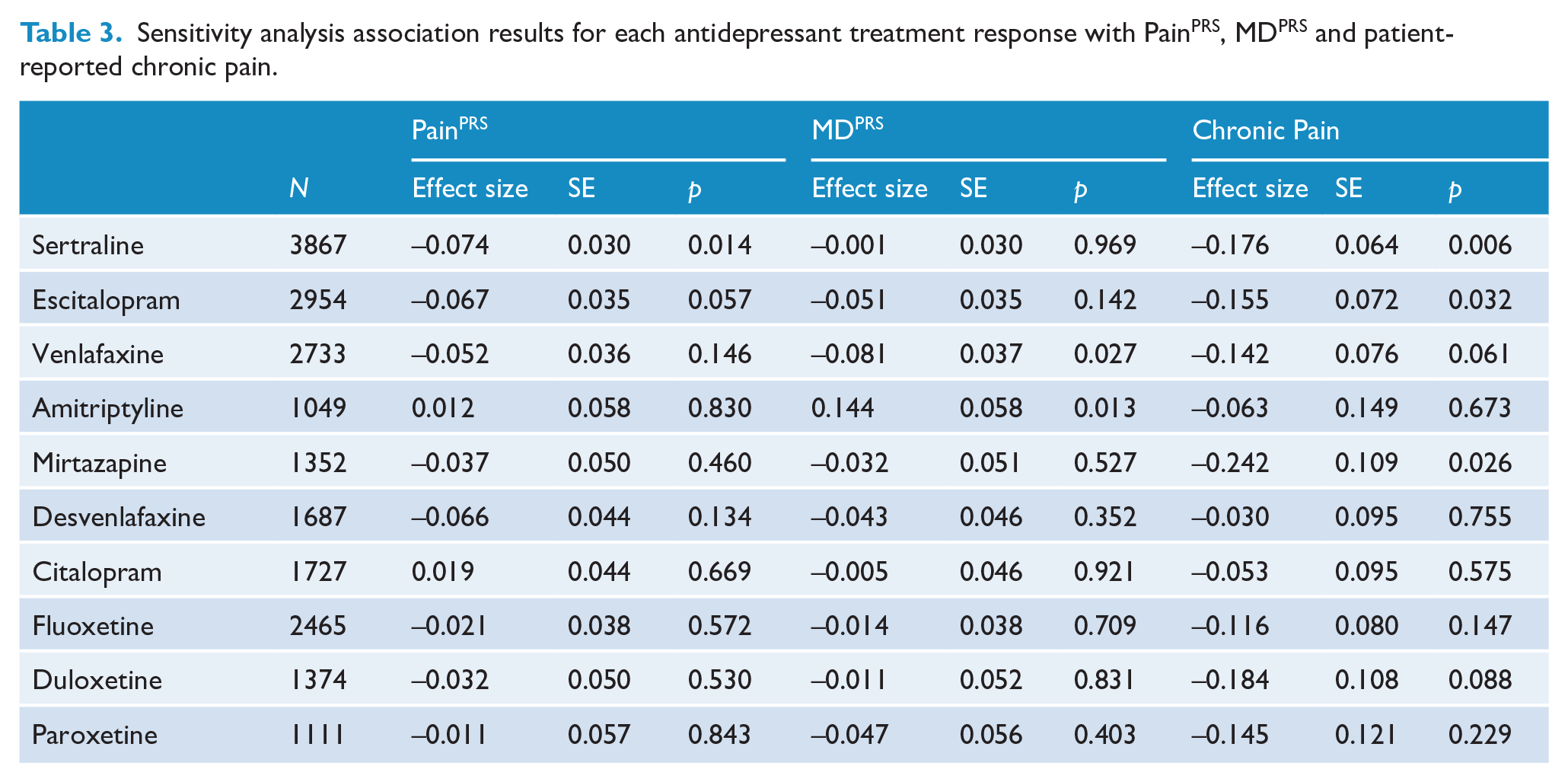
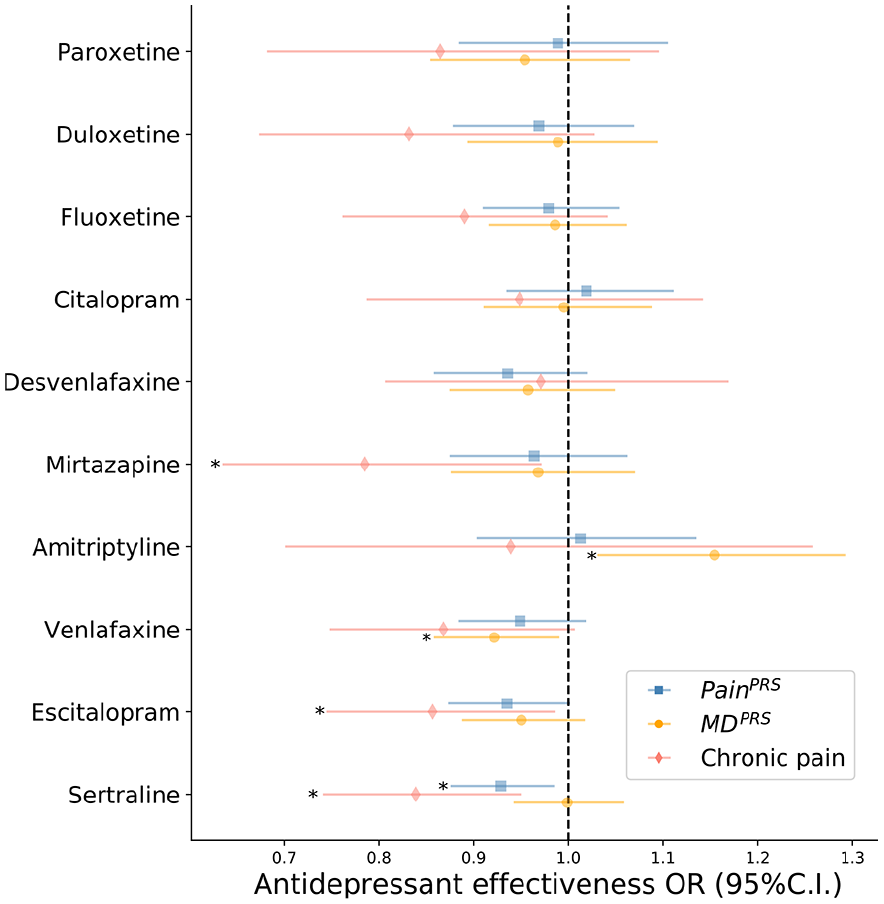
Sensitivity analyses for antidepressant effectiveness results after adjusting for age of depression onset and number of depressive episodes.
Discussion
We recently demonstrated in a sample of more than 11,000 participants that comorbid pain is a significant risk factor for poor treatment response with antidepressant medications (Roughan et al., 2021). This finding corroborated previous clinical trials and observational studies showing that pain has a significant negative impact on antidepressant treatment response (Fishbain et al., 2014). Here, we showed the increased risk of poor antidepressant response is genetically mediated using a PRS analytical approach (Wray et al., 2020). The current study has presented the first evidence for predicting antidepressant treatment response in comorbid chronic pain and depression using PRS. Our findings point towards a difficult-to-treat depression (DTD) subtype characterised by comorbid pain. The 2020 Royal Australian and New Zealand clinical practice guidelines (CPGs) for mood disorders suggests the use of ‘DTD’ to describe depression subtypes where treatment, which also could include other types of pharmacological and non-pharmacological therapies, is challenging but not unachievable (Malhi et al., 2021). This proposed subtype warrants replication in large independent samples with pain, genotype and treatment response data. We suggest that a distinct advantage of such large-scale data-driven studies – through the application of PRS analyses – is their utility in predicting medication response risk for population-level trans diagnostic stratification and disease classification (Shah et al., 2020). This work also provides further evidence for the universally recognised clinical reality that patients presenting with psychiatric and/or physical comorbidities are the rule (Gold et al., 2020; Momen et al., 2020; Viana et al., 2018), rather than the exception.
Further evidence consistent with a comorbid pain subtype was the finding of MDPRS not being associated with chronic pain, even though a phenotypic association between depression and chronic pain had been demonstrated using the same sample (Roughan et al., 2021). In the current study, we generated PRS for chronic pain based on GWAS summary statistics from an earlier report using UK Biobank pain data (Johnston et al., 2019). That report also applied Mendelian Randomisation analyses and found a causal effect of chronic pain on MD, but not vice versa. This causal direction finding is corroborated by our sensitivity analyses showing that PainPRS but not MDPRS predicts poor antidepressant response after adjusting for depression severity measures. Our results are also complemented by a recent study – notably, in a psychiatric patient cohort – which identified gene expression blood biomarkers that predicted pain state and future emergency department visits for pain, as well as new potential therapeutic targets for pain using existing antidepressant medications (e.g. amoxapine) (Niculescu et al., 2019). Furthermore, a study of medication use in the UK Biobank found depression increased the likelihood of taking analgesics (anilides) (Wu et al., 2019), thus presenting another direction to further investigate treatment-response subtypes in large genotyped samples.
A recent study that also examined PRS for depression in relation to pain and other common comorbidities used summary statistics from UK Biobank and other large population cohorts (Shen et al., 2020). They reported that MDPRS predicted recent pain along with other complex traits which showed the largest effect sizes (e.g. sleep, smoking behaviour, cardiovascular conditions). MDPRS was not reported to predict chronic pain (i.e. pain for 3+ months), which is consistent with our results. Notably, we have shown that PainPRS predicts patient-reported chronic pain. Altogether, these results support an underlying genetic difference between short-term pain and chronic pain. However, the current consensus-based IASP and ICD-11 diagnostic criteria for defining ‘chronic pain’ rests on a purely temporal and arbitrary cutoff of ‘pain persisting or recurring for longer than three months’ (Treede et al., 2019), while also failing to account for comorbidities such as depression. Furthermore, despite decades of enormous effort put into pain research, there has been no objective biological evidence found for this critical criterion. Our results on the other hand present the first evidence to support a genetic transdiagnostic subtype of depression with comorbid chronic pain. This unexpected finding has major implications, for example, on diagnostics, disease classification and early intervention approaches (e.g. screening, education, prevention) in pain medicine (Davis et al., 2020) and occupational rehabilitation (e.g. after injury/trauma) – and correspondingly – for biologically mechanisms-based nosology frameworks in psychiatry (i.e. RDoC).
In the current study, PainPRS was associated with poorer antidepressant effectiveness overall. However, when stratifying per antidepressant, we found statistically significant lower effectiveness for sertraline only. This result is likely explained by the fact that sertraline was the most commonly reported antidepressant taken and therefore had the greatest statistical power. Consistently, the effect of associations between PainPRS and other antidepressants was negative. Given the circular relationship between depression and pain, we performed fully adjusted sensitivity analyses that accounted for MDPRS and measures of depression severity (i.e. age of depression onset, number of depressive episodes; interval between age at participation and at depression onset). The association between lower overall effectiveness of antidepressants and PainPRS was robust to these factors, and a similar finding from the stratified analyses was observed.
Our results support the channelling response paradigm to treating depression. Under this paradigm, response heterogeneity is the product of different responsivities that need to be addressed for successful treatment (Malhi et al., 2021). For this subtype, one of such responsivities would be the adequate management and treatment of pain, without which depressive symptom therapies might alone cause a poor response. In a previous study (Roughan et al., 2021), we showed that, when explicitly indicated for chronic pain, duloxetine, venlafaxine and amitriptyline show a positive (but non-significant) association with better treatment response. Consistently, we found MDPRS to be associated with a higher amitriptyline response. Altogether, these results suggest that amitriptyline is effective at treating both pain and depressive symptoms, this further supports the CPGs considering it the antidepressant of choice with the highest efficacy (at the drawback of low tolerability) for depressive symptom alleviation. As such, our observations support recently published guidelines suggesting duloxetine and tricyclic antidepressants (TCAs) as preferred antidepressants for depression when pain is present as a prominent symptom (Malhi et al., 2021).
Our study is the first direct assessment of whether polygenic instruments for chronic pain are associated with antidepressant treatment effectiveness. As such, there are certain limitations to be acknowledged. First, to avoid potential sources of bias that are inherent to genetic studies, analyses were restricted to a subset of unrelated individuals with European ancestry. This approach aims to reduce false-positive associations due to population stratification but limits the generalisability of the findings to other populations. Furthermore, all participants in this study had depression. As such, caution should be taken in generalising the association between chronic pain, depression and their genetic relationship across the general population. All of the studied phenotypes were also obtained through online self-administered instruments and are therefore subject to recall bias. Related to this issue is the interpretability of the antidepressant effectiveness phenotype. For example, amitriptyline is a TCA medication commonly prescribed as a treatment for chronic neuropathic pain (Moore et al., 2015). Most participants reported taking antidepressants for treating depression, as did the vast majority of participants with chronic pain (i.e. 75–98% across antidepressants) (Roughan et al., 2021). As such, those with comorbidities may have considered ‘treatment effectiveness’ as a bivariate improvement of both their depression and chronic pain. Finally, we focused only on treatment response with the 10 most commonly prescribed antidepressants in Australia. The extent to which our results may generalise to other antidepressants as well as other pharmacological and non-pharmacological therapies requires further investigation.
The current study’s findings warrant further examination by (1) performing discovery GWAS of comorbid chronic pain and depression for generating PRS in large genotyped populations to assess genetic risk prediction capacity and Generalisability; (2) replication in other DTD cohorts with primary care and genotype data (Fabbri et al., 2020); and (3) replication in a PRS-informed randomised clinical trial integrating both MD and chronic pain polygenic instruments to stratify the placebo and control groups. Further genetic epidemiology studies of treatment response can also focus on patients with chronic pain and depression of non-European ancestry, to collect additional data on medication dosages (including painkillers), the duration and magnitude of functional benefits, drug tolerability and adverse events, other prescribed and self-help therapies, pain-specific questionnaires (Roughan et al., 2021) and other common comorbidities such as sleep and cardiovascular disorders (Fabbri et al., 2020; Shen et al., 2020; Tesarz et al., 2019; van Hecke et al., 2017). Our work demonstrates a strategy for further delineating the heterogeneity across psychiatric and comorbid pain populations to potentially yield other transdiagnostic subtypes, which would pave the way for comparing particular psychotropic medications in pharmacogenomics (Bousman et al., 2019; Dorfman et al., 2020; Kaye et al., 2019; Maciukiewicz et al., 2018; Yamamoto et al., 2019) and neuroimaging trials (Schmaal et al., 2020; Thompson et al., 2020; Wang et al., 2019) specifically targeting identified subtype cohorts.
Since the advent of GWAS and associated methodologies over a decade ago, demonstrating their clinical utility in complex traits and polygenic disorders (e.g. mental health, chronic diseases) has been challenging. Here we have demonstrated the application of a commonly used genetic analysis method (PRS) to antidepressant treatment response data and showed its utility in the similarly elusive goal of identifying subtypes in psychiatry and pain medicine. We hope this approach can bring a step change in genomic medicine, pain management and multidisciplinary clinical decision-making to deliver improvements in healthcare outcomes and the identification of effective therapeutic targets.
Footnotes
Acknowledgements
The authors thank their colleagues Richard Parker, Simone Cross, Scott Gordon and Lenore Sullivan for their valuable work coordinating all the administrative and operational aspects of the AGDS project. They also thank the anonymous reviewers for their constructive comments which helped to improve the manuscript. The views expressed are those of the authors and not necessarily those of the affiliated or funding institutions.
Author Contributions
A.I.C., T.T.N. and M.E.R. designed this study and wrote the first version of the manuscript; A.I.C. performed the analyses with input from M.E.R.; N.G.M., S.E.M., N.R.W., E.M.B. and I.B.H. designed and directed the AGDS data collection efforts; and T.T.N. designed the pain module in the AGDS online survey and conceived the genetic and epidemiological investigation of comorbid pain and depression in this cohort. All authors contributed to the interpretation of the results and provided feedback on the preliminary versions of the manuscript.
Declaration of Conflicting Interests
The author(s) declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: I.B.H. has been Commissioner of Australia’s National Mental Health Commission (2012–2018); Co-director of Health & Policy at the Brain & Mind Centre, University of Sydney; leading community-based and pharmaceutical industry-supported projects (Wyeth, Eli Lilly, Servier, Pfizer, AstraZeneca) focused on the identification and better management of anxiety and depression; a member of the Medical Advisory Panel for Medibank Private until October 2017; a board member of Psychosis Australia Trust; a member of the Veterans Mental Health Clinical Reference group; and Chief Scientific Advisor to and an equity shareholder in Innowell. The remaining authors have nothing to disclose.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Data collection for AGDS was possible thanks to funding from the Australian National Health & Medical Research Council (NHMRC) to N.G.M. (GNT1086683). A.I.C. is supported by a UQ Research Training Scholarship from The University of Queensland (UQ). M.E.R. thanks the support of NHMRC and the Australian Research Council (ARC), through a NHMRC-ARC Dementia Research Development Fellowship (GNT1102821).
