Abstract
The technological advancement in diagnostic techniques has immensely improved the capability of predicting disease progression. Yet, there is a great interest in developing newer biomarkers that can enhance disease risk prediction thereby minimising the associated morbidity and mortality. Circulating miRNAs, a non-coding RNA molecule, are critical regulators in the pathophysiology of various complex multifactorial diseases. In recent years, circulating miRNAs have been enormously studied and are considered as an emerging biomarker due to their easy accessibility, stability, and detection by sequence-specific amplification methods. However, there is a distinct lack of consensus regarding the preanalytical factors such as preferred sample selection, methodological aspects, etc that may independently or together influence the detection of circulating miRNAs resulting in erroneous expression profiles. Therefore, the present review makes an attempt to highlight the various pre-analytical and analytical factors that can potentially influence the circulating miRNA levels. Literature on circulating miRNA’s stability, processing and quantitation in different biofluids along with the effect of various controllable and uncontrollable factors influencing circulating miRNA expression have been summarised in the current review.
Introduction
MicroRNAs (miRNAs) are small non-coding RNAs (∼22 nucleotides long) involved in the regulation of a protein-coding gene at the post-transcriptional level by binding complementary to the 3' UTR of the target mRNA. 1 In recent years, numerous studies have explored the role of circulating miRNAs as a non-invasive diagnostic and prognostic biomarker in various complex diseases like cancer, Alzheimer’s, neurological, paediatric and cardiovascular diseases (CVD).2,3 The presence of miRNA in all body fluids made it a widely studied biomarker. However, it has been unable to overcome the barrier between bench to bedside due to the disparity within the sampling and detection methodologies as well as biological variability. Thus optimising the protocols right from sample selection, processing, and storage to efficient quantitation, detection and validation procedures is imperative for circulating miRNA to be considered as a biomarker. This review discusses the available literature on the circulating miRNAs in various biofluids and explains the methodological aspects of miRNA detection in each of these fluids.
Methodology
Sample Selection
Circulating miRNAs have been detected in various body fluids such as plasma, serum, saliva, tears, urine, amniotic fluid, breast milk, etc. 4 ; however, judicious selection of specimens is necessary to eliminate confounding results.
Circulating miRNAs in blood derivatives
Serum and plasma are frequently used biofluids, with the latter being most preferred for studying the disease-specific dysregulated miRNA due to the following reasons (i) miRNAs in platelets, WBCs & RBCs can be released during coagulation, thereby changing the repertoire of circulating miRNAs,5,6 (ii) Plasma is concentrated with miRNAs as compared to serum, 6 (iii) Serum may be contaminated with other small non-coding RNAs of 30 nucleotides long, 6 (iv) increased surface adsorption of circulating miRNA on blood clot. 7 Inspite of the known advantages of using plasma as the preferred specimen, the available literature is quite conflicting. Most recently, Mompeón et al, 2020 found varying expression patterns of selected miRNAs in plasma and serum of acute myocardial infarction (AMI) subjects, 8 whereas, Mitchell et al, 2008 on the other hand, suggested that both plasma and serum samples can be used for analysis as measurements obtained from plasma and serum showed strong correlation. 9 Studies have shown that optimising the preanalytical factors, especially specimen processing, helps to reduce the technical variability between protocols and eliminate inconsistent and incomparable results. While using serum as a starting material, Farina et al, 2014 suggested centrifuging the blood at 2000 rpm for 15 min, followed by storage at −80°C. 10 On the other hand, while using plasma, platelet-poor plasma (PPP) should be preferred in order to reduce contamination7,8 of the most abundant platelet miRNA, that is, let-7 family or miR-223-3p. 11 Additionally, plasma should be nonhaemolysed and preferably collected in Ethylenediaminetetraacetic acid (EDTA) tubes since the use of citrate or heparin as anticoagulants tends to interfere in the downstream assessment likely due to the depletion of cofactors required by DNA polymerases and also affects the miRNA integrity. 12 In contrast, Glinge et al (2017) suggested citrate can also be used as an anticoagulant along with EDTA while heparin should be avoided. 13 Furthermore, the use of whole blood for miRNA isolation is questionable, as the repertoire may have miRNAs contributed by cellular entities in whole blood. Pascut et al, (2019) observed that whole blood has almost double the miRNAs as compared to serum, most of which are still uncharacterised. 14
Circulating miRNAs in other body fluids
Other sparsely studied biofluids for circulating miRNAs.
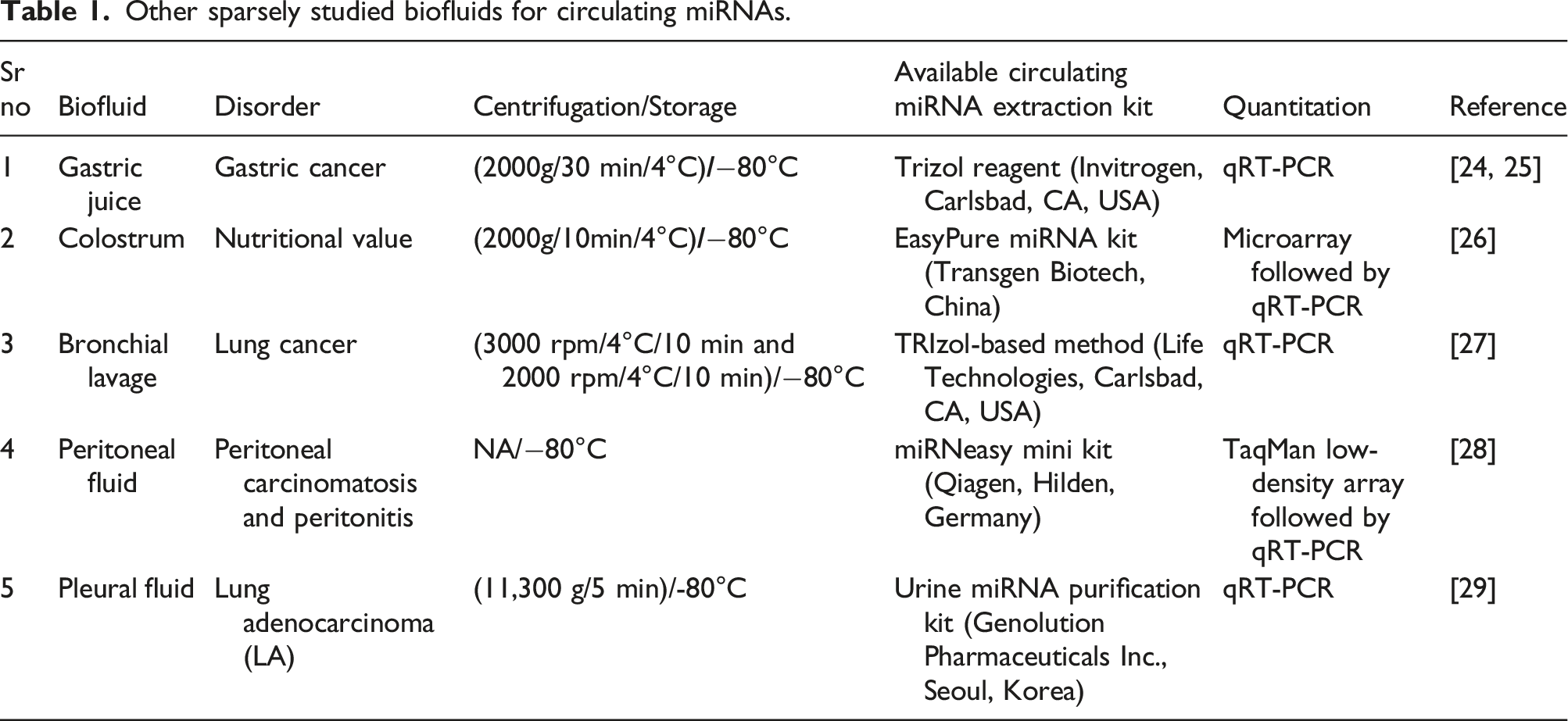
Stability of circulating miRNAs
Stability of circulating miRNAs in biofluids
Circulating miRNAs serve all the properties of an ideal biomarker; one of them is their stability in biofluids. Studies have determined the stability of limited circulating miRNAs since each miRNA behaves differently in different biofluids. The stability of circulating miRNAs is dependent on many factors, particularly the duration between sample procurement and processing, or centrifugation and the temperature during the holding period. In case of blood derivatives, it is necessary to process the plasma and serum within 2 h 30 to up to 4 h of collection. 5 The most accurate representation of miRNAs in the urine milieu is best measured within the first 24 h and can be preferably stored at 4°C to have the least degradation, 31 while Fuessel et al 2018, suggest that urine can be either be stored at 4–8°C for up to 4 h or should be processed immediately. 32 Salivary miRNAs are stable at room temperature (RT) for almost 48 h 33 . miRNAs in the seminal fluid are stable for up to 24 h at RT. 34 miRNA stability in most of the aforementioned biofluids has not been studied. Therefore, a detailed stability study should be carried out to determine stability in subsequent studies. However, maintaining minimal time between sample procurement to centrifugation will have the least impact on miRNAs; if not attainable, one can store the samples at a lower temperature (4–8°C) until processing. Physical disturbance is another important factor that may affect the stability of miRNAs. Excessive movement during transportation should be avoided, and in case of prolonged mechanical disturbance, a non-centrifuged whole sample should be preferred compared to centrifuged counterparts. 13
Stability of circulating miRNAs in archived biofluids
Biofluids, when stored at lower temperatures (−20 to −80°C), show extended stability; however, there exists a conundrum about utilising irreplaceable archived samples. A 2015 study evaluated the miRNA quantity, quality and variability in serum samples that were stored for 40 years, from a prospective biorepository. The study found the miRNAs to be stably present in archived serum samples that were frozen at −25°C for at least 40 years and that approximately 500 μL of serum can provide sufficient miRNA for small RNA sequencing and quantitative reverse transcriptase-polymerase chain reaction (qRT-PCR). 35 Similarly, for plasma specimens, studies have shown that miRNAs can be detected with satisfactory intra- and inter-array reproducibility from plasma stored for almost 20 years. 36 With no available literature for other biofluids, one should keep in mind that before using any archived samples, one must ensure that all the samples were processed using a standardised protocol, the number of freeze-thaw cycles (≤4 cycles) is noted, the volume is sufficient for extraction, and in the case of blood derivatives, they are non-haemolysed. This may help to avoid confounding results arising due to technical variability.
Processing of biofluids
Biofluid collection and centrifugation
Enrichment of circulating miRNAs from biofluids requires efficient elimination of the cellular counterparts in order to avoid confounding results. The lack of clarity in the existing centrifugation speed, time and temperature in various studies makes it imperative to have a standardised centrifugation protocol.
As mentioned earlier, platelets should be essentially removed from plasma to obtain the true miRNA repertoire present in the circulation. The intracellular miRNAs in platelets are substantially higher than the levels found in the extracellular environment, which contribute largely to the original pool of circulating miRNAs. Also, if platelets are not removed before freezing plasma, the miRNAs in residual platelets will be released during subsequent freeze-thaw, 37 thereby making it necessary to obtain platelet-poor plasma. A two-step centrifugation of blood at 3000g for 15 min each, helps to reduce the platelets to 0–1×109/L. 38 Furthermore, Cheng et al (2013) suggested an additional filtration (0.22 µm) step after the two-step centrifugation of plasma (3400g x 10 min + 1940g x 10 min) and serum (clot 30 min + 3400g x 10 min) to effectively eliminate microparticles and platelets. The additional filtration could remove 99–100% of residual platelets relative to standard centrifugation (3400g × 10 min) in plasma while in serum; the platelet count was almost unaltered, but the microparticles were eliminated. Platelet contamination in archived samples (6 years) can also be eliminated by an additional centrifugation step at 1940 g × 10 min 39
Urine is one of the most readily accessible biological fluids, and it harbours numerous biologically significant miRNAs. Up to 50 mL of urine can be collected for miRNA profiling, although there is ambiguity about which urine voids should be collected. Studies have mentioned that either morning midstream urine 40 or spontaneous voids41–43 can be collected in sterile containers. The collected urine can be stored directly at −20°C to −80°C41,43 or can be centrifuged at a speed varying from 1000-4000 rpm or 1500 g for 10–15 min at temperatures lower than 10°C.32,42,44
Saliva, another easily obtained biofluid, should be centrifuged to remove any contaminant present. However, before collecting saliva, one should be refrained from eating, drinking or oral hygiene procedures for at least 2 hours before the collection. In addition, subjects should rinse their mouth with water before saliva collection to minimise contamination. 45 Yet centrifugation is an inescapable step, as whole saliva may include microbial origin RNA as contaminants. Centrifugation speed may range between 2500 and 3000 g at 4°C for 10–15 min 18
Amniotic fluid is procured by an invasive procedure amniocentesis, performed percutaneously under transabdominal ultrasound guidance in sterile conditions. Procured amniotic fluid can either be immediately stored at −80°C19,46 or centrifuged at 3000 rpm for 20 min at 25°C before storing at −80°C. 47
Tears can be obtained via various methods, although should be performed atraumatically. Micropipettes can be used to obtain tears from the inferior tear meniscus of eyes or after instillation of 2.5 μL of distilled water in case of few tears. 48 Also, one can collect tears by adding 60 μL of 0.9% saline dripped into the conjunctival sac and mixing it with tears. 49 Schirmer’s paper strips used to determine tear production can also be used for tear collection. The strip is placed in Eppendorf tubes containing 500 μL of water with RNase inhibitors, vortexed for 5 min, and stored at −80°C. 50 No centrifugation step is mentioned; therefore, tear samples can be stored directly for further use.
Collection of CSF should be performed by experienced physicians using the standard lumbar puncture (vertebral L3-L5), external ventricular drain or cisternal aspiration at the time of craniotomy. 51 Cerebrospinal fluid-containing blood cells should be excluded. Cerebrospinal fluid should be processed within 1 h of collection. It is usually centrifuged at a lower speed, ranging between 300 and 500 g for 10–15 min either at RT or 4°C,51,52 or one may even filter the collected samples using a 0.8 μm pore size membrane filter. 53 This ensures removal of any cellular contaminants.
Seminal fluid should be collected after 3–5 days of sexual abstinence, and should be kept for liquefaction at 37°C within 30 min of collection. Seminal fluid can be centrifuged by one-step centrifugation at 13,400 g for 5 min54,55 or two-step centrifugation either 800 g/15 min or 1600 g/10 min at 4°C followed by 16,000g for 10 min at 4°C.22,34,56 Inappropriate centrifugation speed and time may result in residual cells that may contaminate cell-free miRNAs; thus, it should be given more attention.
Haemolysis
Utmost care should be taken while working with blood derivatives as miRNA released from erythrocytes during haemolysis can dramatically alter the miRNA profile in the circulation. A study demonstrated that blood cells can be a major contributor to circulating miRNA and that perturbations in blood cell counts and haemolysis can alter plasma miRNA biomarker levels by up to 50-fold. 57 Haemolysis can be simply avoided by following good phlebotomy practices. Visual detection of haemolysis is less reliable due to its low accuracy. The extent of haemolysis can be checked spectrophotometrically by detecting oxyhaemoglobin’s absorbance peak at 414 nm, 541 nm and 576 nm with the additional peaks at 541 and 576 nm indicative of high levels of haemolysis. Samples with absorbance less than 0.2 at 414 nm can be processed for extraction. 58 Although lipaemia invariably interferes with optical density, absorbance at 414 nm and 385 nm (as a lipaemia indicator) can detect haemolysis in the presence of lipaemia. A lipaemia-independent haemolysis score (HS) can be calculated based on the mean A414 and A385 values: HS = ∆(A414−A385) + 0.16 ∗A385, with HS threshold ≤0.057, since samples with HS > 0.057 contain increased levels of free haemoglobin. 59 A qRT-PCR based method can sensitively detect haemolysis by assessing the levels of two erythrocyte-specific miRNAs miR-451a and miR-23a-3p [delta Cq (miR-23a-3p—miR-451a)]. It is observed that the miRNA ratio can detect haemolysis down to approximately 0.001%, whereas the spectrophotometric method could detect down to 0.004% haemolysis. 60 Not only blood derivatives but also biofluids like urine and CSF may have RBCs leading to haemolysis.
Aliquoting
Besides functioning as a technique merely to separate a larger sample into smaller parts, aliquoting plays a very important role in studies conducting extraction procedures. One must systematically plan for the number and volume of aliquots to be made, as storing a higher volume than required may result in frequent freeze-thaw, while storing less sample volume may require more than one aliquot for extraction. Moreover, samples like CSF, tear, saliva, amniotic fluid and seminal fluid are obtained in lower volumes, therefore they should be judiciously aliquotted without wasting the valuable sample. One can choose the volume and number of aliquots depending upon the kit intended for extraction in the future.
Storage and freeze-thaw
Stability of circulating miRNA in whole blood, plasma, serum and urine.
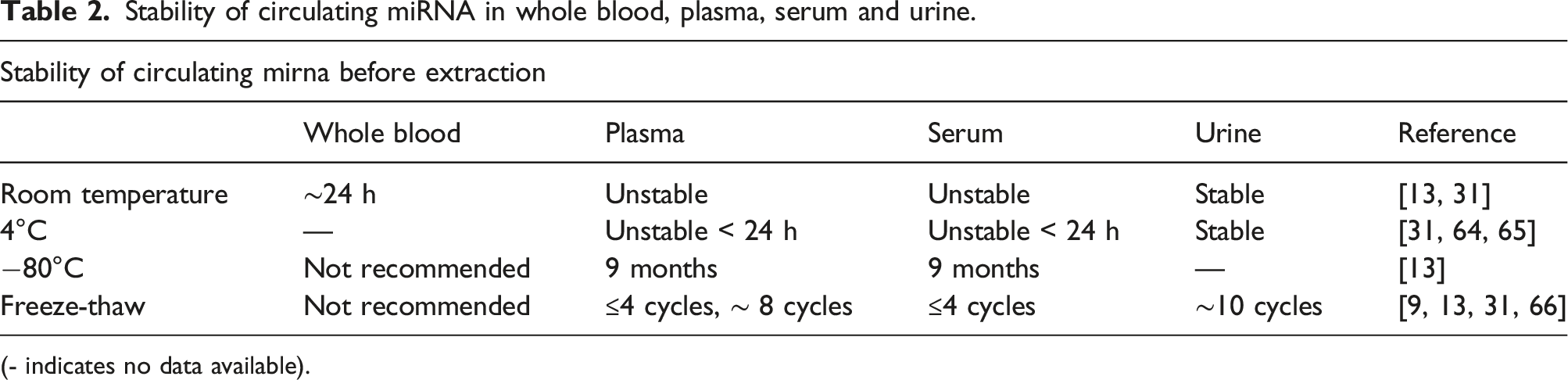
(- indicates no data available).
Extraction and quantitation
Commercial kits
miRNA content in peripheral blood is approximately 10 times less when compared to the tissue. 67 Thus, it is challenging to efficiently extract and accurately quantify extracted miRNAs and maintain reproducibility in miRNA-profiling studies. Most of the currently available extraction techniques are based either on phenol: chloroform extraction or combined phenol and column-based techniques, or only column-based techniques with the latter being widely used. Several commercial extraction kits are available, among which mirVana PARIS kit (Ambion, Life Technologies, Texas, USA), TRIzol-LS (Ambion, Life Technologies, Carlsbad, CA, USA), miRCURY RNA Isolation Kit – Biofluids (Exiqon, Vedbaek, Denmark) and miRNeasy Serum/Plasma Kit (Qiagen, Hilden, Germany) are favoured.68,69 Li et al. (2015) used 7 different commercial kits to compare miRNA yield as well as the amplification efficiency from plasma and observed that the miRNeasy kit from Qiagen showed better performance. 70 TRIzol reagent (Invitrogen), 41 miRNeasy (Qiagen) 43 and mirVana miRNA isolation (Thermo Fischer) 44 are used widely for the extraction of miRNA from urine. However, a specific kit, Urine microRNA purification kit from Norgen (Urine microRNA Purification Kit, Biotek) is available, which can be used for both CSF and urine (0.5 mL–1 mL). Widely used kits for miRNA extraction from CSF are mirVana PARIS Isolation Kit (Applied Biosystems)71,72 and miRCURY RNA isolation kit (Exiqon). 52 Burgos et al., compared 4 commercially available small RNA-isolation kits and found mirVana PARIS kit to give the highest yield of small RNAs, with as little as 0.5 mL of CSF starting volume to achieve reproducible results. 73 TRIzol Reagent (Invitrogen) is used for miRNA extraction from seminal fluid,22,34,54,55 whereas the most widely used kit for other samples like saliva, amniotic fluid and tears is miRNeasy (Qiagen), which is designed specifically to extract RNAs, approximately 18 nucleotides upwards. Even though the kit is intended exclusively for serum and plasma, it is also used for other biofluids, necessitating a sample-specific extraction kit. Each biofluid differs from the others in terms of protein content, nucleic acid content and cellular entities. Therefore, while using a serum/plasma kit for other biofluids, the lysis and precipitation step of extraction should be essentially standardised such that the supernatant after precipitation is devoid of all contaminants.
Quantitation techniques
Due to the low concentration of circulating miRNAs in biofluids, their quantity and purity, should be determined accurately. Various quantification techniques are available, namely Tecan InfinitePRO Nanoquant Spectrophotometer, Thermo Fisher Scientific Nanodrop 2000 Spectrophotometer, Agilent 2100 Bioanalyzer and Thermo Fisher Scientific Qubit Fluorometer with each of them requiring between 1 and 3 μL of sample volume for quantification. One of the most versatile and accurate instruments is the Agilent 2100 Bioanalyzer, which is principally a traditional gel electrophoresis on a chip format, that requires less separation time and sample consumption. The charged RNA molecules are separated by size on a voltage gradient gel, with the faster migrating smaller RNA being intercalated by fluorescent dyes and detected by laser-induced fluorescence. It also provides the RNA integrity values, but inaccessibility in small laboratories due to high cost makes it difficult for regular usage. The other common detection method is spectrophotometric analysis, viz Nanoquant and Nanodrop, based on the principle that nucleic acids absorb ultraviolet light specifically at 260 nm (for DNA and RNA) or 280 nm (for proteins). yet, Nanoquant and Nanodrop tend to overestimate the miRNA concentration since they cannot discriminate between miRNAs, other small RNAs, total RNA and free nucleotides. On the other hand, Qubit Fluorometer provides an accurate estimation as it uses fluorescent dyes and determines the concentration of small RNAs (∼20 nucleotides) even in the presence of common contaminants such as salts, free nucleotides, solvents, detergents and protein. The fluorescent dye binds specifically to miRNA thereby providing more accurate quantification. 74 The sensitivity of the Qubit can be as low as 10 pg/mL, which is far superior to the spectrophotometric assays. The latest Qubit devices (Qubit 4) can determine RNA quality, similar to that given on a 2100 Agilent Bioanalyzer, and can also determine the integrity of RNA. Dissimilarity in preanalytical techniques, miRNA extraction and handling techniques may yield imprecise and non-reproducible results. Garcia-Elias et al., 2017 mentioned that amongst the small RNAs quantified by the aforementioned methods, only 58% of the total counts are true miRNAs, thus focussing on the limitations of all current quantification techniques to estimate the miRNAs content 74 .
Profiling of circulating miRNAs
qRT-PCR, Microarray and NGS
Several methodologies like qRT-PCR, microarray and next-generation sequencing (NGS) are available for detection of miRNAs in biofluids, with qRT-PCR considered to be the gold standard and is widely employed for validation of miRNA profiles obtained from microarray and RNA sequencing 75 . qRT-PCR involves data normalisation by using endogenous and exogenous reference genes, which can assist in identifying fundamental changes in miRNA expression levels. However, there is no consensus about an optimal normalisation strategy. Thus, judicious selection of the reference gene is important as it can affect the biological interpretation of data 76 . Donati et al., 2019 provided an overview of the reference genes currently used in various pathologies like cancer, CVD, and tuberculosis. Based on the literature, miRNAs like miR-16, miR-93, the combination of miR-221 & miR-26a, miR-191 and miR-320a were described as the potential endogenous reference genes in plasma and serum, offering researchers the opportunity to select the appropriate miRNAs for normalisation 77 . Endogenous RNAs may show variation in expression due to inter‐sample heterogeneity and pre‐isolations steps as compared to the exogenous RNAs, which may be subjected to only pre‐isolations steps rather than inter‐sample heterogeneity. Exogenous RNAs like cel‐miR‐39‐3p, cel‐miR‐54‐3p and cel‐miR‐238‐3p present less variability and can be significantly used for normalisation of both discovery and validation. 78 However, advanced technology like droplet digital PCR (ddPCR) can resolve the normalisation issue and facilitate miRNA measurement with higher throughput and precision than qRT-PCR, at a similar cost-per-sample. 79 Another limitation of qRT-PCR is primer design; currently available commercial kits make use of universal primers owing to poly-A tail addition. However, such primers allow the profiling of only existing miRNAs listed in miRBase and cannot identify novel miRNAs; 80 yet, qRT-PCR is the quickest and most reliable. Microarray is a semi-quantitative hybridisation-based detection that facilitates a parallel analysis of several miRNAs involved in a pathway. Microarrays are flexible and versatile tools due to their ability to be customised based on pathogenesis. Moreover, it is easily accessible in core laboratories as compared to highly sophisticated NGS, while the requirement of low starting material makes it of choice to researchers. However, certain limitations are inherently present, like inadequacy to detect novel microRNAs, lower sensitivity and specificity as compared to qRT-PCR, unsuitable for low input levels especially in the case of plasma/serum, inability to validate miRNA expression profile, probe designing, incapable to absolute quantify and importantly microarrays cannot distinguish mature miRNAs from precursor miRNAs or other non-coding RNAs. This may be due to the presence of similar sequences among these RNAs, thus creating a possibility of imperfect binding to probe strong enough to retain it if the chip is not washed correctly. 5 However, this can be overcome with a precise and genuine probe design. 81 Next-generation sequencing, a massively parallel sequencing technique is the most promising tool for analysing miRNA profiles that enable rapid, high-throughput analysis of the transcriptome. Unlike qRT-PCR and microarray, NGS does not require prior knowledge of miRNAs, thus facilitating the discovery of novel miRNAs. Next-generation sequencing is extremely sensitive and distinguishes between types of ncRNAs. Yet, qRT-PCR is still the only platform for absolute quantification. Liu et al. (2012) in their review elaborated on technical advancement, comparative analyses between different platforms, and software packages for analysing NGS data for profiling miRNA. 82 Several studies have performed NGS to profile miRNA expression, although most of them have a smaller sample size. The possible reason for it could be the expensive nature of technology. Moreover, small sample size makes it difficult to draw accurate statistical inferences; however, this can be overcome by pooling plasma and sera without affecting the expression profile of miRNAs.
Pooling samples
Pooling samples can be an economical tool for circulating miRNA profiling in high-cost RNA sequencing techniques. It allows researchers to pool samples of subjects with similar clinical conditions or demographics. Several studies have pooled plasma or sera of subjects for miRNA profiling in different diseases like cancer, 83 aging, 84 diabetes mellitus (DM), 85 etc. Comparison within these studies indicates the number of subjects included in each pool ranges from 5 to 30. A study by Niculescu et al, 2015 efficiently determined dysregulated miRNAs by creating 4 independent pools of stable angina, unstable angina, myocardial infarction and control, each pool containing 8 subjects. 86 Regardless of how successfully one implements it, Kok et al, 2018 suggest that pooling can lead to more false-positive and false-negative results as individual-specific information is lost and increases random technical variation. 87 However, pooling samples can be considered as a viable approach provided the number of samples in each pool should be representative and should not be arbitrarily high or low. 88 Anand et al, 2016 in a sequencing study containing 996 individuals, with a total of 83 pools each having 12 individuals, proposed a simple filtering guideline based on the Kolmogorov–Smirnov statistical test that removes false variants due to sequencing errors. The filters were validated by comparing pooled-sequencing to individual-sequencing data, which showed that filters provide a low rate of false-positive and false-negative variants. 89
Newer techniques
Various newly developed miRNA detection techniques make it possible to detect miRNAs even in femtomolar concentration. Nanomaterial-based detection techniques are growing due to their high surface area, excellent electrical conductivity and remarkable chemical stability. Gold, silver, copper and carbon nanostructure make ultrasensitive detection of miRNAs. Similarly, a multitude of nucleic acid amplification methods have been exploited, such as rolling circle amplification (RCA), duplex-specific nuclease (DSN)-based amplification, loop-mediated isothermal amplification (LAMP), exponential amplification reaction (EXPAR), strand-displacement amplification (SDA) and some enzyme-free amplifications. Each of these methods has its advantages and disadvantages, thus suggested to use in combinations to improve the performances of classical detection methods. 90
Factors affecting circulating miRNA
Controllable factors
Diet, lifestyle and medications
Dietary molecules directly or indirectly modulate epigenetic machinery. 91 A study demonstrated that when mice are fed high-fat diets, their plasma miRNA expression is modulated in the first 30 minutes 92 A review by Kura et al., 2019 summarised that certain foods and dietary supplements modulate expression levels of many cardiovascular-related miRNAs. Thus, foods that increase atheroprotective miRNAs can be effectively included in diet for CVD prevention and treatment. 93 miRNAs primarily involved in age-related biological processes are differentially expressed in vegetarians than in non-vegetarians. Studies showed that three vegetarian subgroups viz vegans, lacto-vegetarians and semi-vegetarians had distinctive circulating miRNA profiles when compared to non-vegetarians and suggested vegetarian dietary patterns to improve health outcomes. 94 It is still unclear how diet regulates circulating miRNAs; however, along with demographics, the diet should be considered as an influencing factor that can interfere with miRNA repertoire. Hence, most of the studies collect fasting blood samples to minimise the effect of diet on miRNAs in circulation.
Another important factor is lifestyle changes, which have become a significant factor that influences susceptibility to most diseases. miRNAs, fine regulators of transcription, are affected by the slightest change in homeostasis. A study by Fatchim et al., 2020 on individuals with impaired glucose regulation (IGR), demonstrated changes in the expression of circulating miRNA (let-7a, let-7e, miR-144 and miR-92a) when analysed before and after dietary and exercise intervention. Especially, let-7a and miR-92a were increased, among which the latter has a link with the IGF (insulin-like growth factor) system. The study also found a correlation between miRNAs and clinical variables (triglycerides, cholesterol, insulin, weight and BMI). 95 Similarly, circulating levels of miR-221 and miR-223 are modulated with exercise and diet, while miR-935 and miR-140 are differentially expressed between high and low responders to body mass loss, before and after chronic weight loss intervention. 96 Obesity or overweight in childhood also significantly affects circulating miRNA expression. 97 One cannot distinguish individuals based on lifestyle; however, it is a modifiable factor that influences various diseases. miRNAs are also modulated by medications. Russo et al., 2015 have mentioned the effect of various drugs on circulating miRNAs. 98 Certain drugs like angiotensin-converting enzyme (ACE) inhibitors or angiotensin receptor blockers (ARB), 99 statins 100 and aspirin 101 may interfere with miRNA expression. Thus, information about lifestyle and medications can be important factors to take into consideration at the time of patient recruitment.
Uncontrollable factors
Age
Certain blood-based biomarker levels change with advancing age 102 including miRNAs.103,104 Thus patient recruitment in biomarker studies is crucial. Hooten et al., 2013 studied the differential serum miRNA expression using NGS and qRT-PCR in young (mean age 30 years) and old (mean age 64 years) individuals. It was observed that miR-151a-5p, miR-181a-5p and miR-1248 were significantly decreased in the elderly compared to younger individuals. These miRNAs were involved in inflammatory pathways thereby highlighting their regulatory role in aging. 104 Moreover, changes in circulating miRNAs are seen at an earlier stage in females (46–55 years) than in males (56–65 years), which can be accounted for by the hormonal changes in females as compared to males. 105 Thus, age and gender matching play a significant role in case-control studies as both can modulate miRNA expression. 103
Gender
Gender-specific differences not only play a role in type and susceptibility of disease but also in the expression of circulating miRNAs. Mostly, miRNAs show consistent expression between gender but few may be oppositely expressed in specific diseases or may show gender-specific association. 103 Individuals with metabolic syndrome show higher levels of plasma let-7g and miR-221 which are prominently expressed in women as compared to males. Moreover, it positively correlates with increased number of risk components associated with metabolic syndrome, including high blood pressure and low high-density lipoprotein (HDL) levels. 106 miRNAs regulated by female hormones or expressed from loci on the X-chromosome due to incomplete X-chromosome inactivation can also be an attributable factor for developing CVD in women. 107 Duttagupta et al., 2011 showed that a subset of plasma miRNAs (miR-548-3p, miR-1323, miR-940 and miR-1292) are significantly upregulated by 63–95% in females as compared to males, while the gene analysis showed that target genes of these miRNAs are impaired during breast cancer. 108 Thus, it is clear that gender may invariably modulate certain types of miRNAs.
Genetic heterogeneity
miRNAs are essential regulators of gene expression that are mostly encoded by noncoding genes and are subjected to genetic variation. Variants in miRNA genes can have profound effects on miRNA functionality at all levels, including miRNA transcription, maturation and target specificity, and as such, they can also contribute to disease. 109 Variants in genes coding for splicing machinery like Dicer, Ago protein 110 and the other enzymes involved in miRNA biogenesis machinery, pri-miRNA, pre-miRNA processing,111,112 as well as those that influence miRNA–target duplex formation like RNA-induced silencing complex 110 would likely make one susceptible to a particular disease which can be found in the Human MicroRNA Disease Database (HMDD). So far, the well-studied miRNA variants in Asian population that are associated with pathogenesis of CAD are miR146a (rs2910164), miR149 (rs2292832), miR196a2 (rs11614913) and miR499 (rs3746444). Similarly, Agiannitopoulos et al., 2020 studied the above-mentioned polymorphisms in the Greek population and found that only miR196a2 C>T (rs11614913) and miR499 A>G (rs3746444) were associated with increased CAD risk and myocardial infarction. 113 Thus, population-specific genetic variation could also affect the prevalence and baseline expression of miRNAs in diverse populations. 114 A study analysing five miRNAs in 106 African American and 239 non-Hispanic Caucasian patients with colorectal cancer, showed that the prognostic value of miRNAs varies with patient race/ethnicity. 115 In contrast, a multi-ethnic study in China on 932 known or suspected CAD patients from 10 different ethnicities showed no significant differences among 9 circulating miRNA in 5 main ethnicities. 116 Furthermore, Dluzen et al (2016) found differentially expressed miRNA/mRNA pairs in African American and white hypertensive women. 117 The contrasting miRNA expression observed in different populations can be attributed to SNPs that exhibit different allelic frequencies, therefore highlighting the role of genetic variants in the population. 118 Hence, one may observe high variability and poor overlap of miRNA expression in different populations.
Conclusion
Biomarkers are at the apex of translational research due to their diagnostic and therapeutic character. Circulating miRNAs, the extensively studied blood-based biomarkers, are promising diagnostic and prognostic biomarkers due to their excellent accessibility in body fluids, their long half-life and robust detection techniques. The field of circulating miRNAs to be used as a minimally invasive diagnostic and prognostic marker for various diseases is growing exponentially. Nevertheless, these studies lack consistency and are inadequate to circumvent the loopholes. To identify the repertoire of dysregulated miRNA in specific diseases, a sufficient sample size is a prerequisite as studies with small sample sizes lack statistical power, which demands large cohort and multicentre studies. Another prerequisite is optimising the sample collection and processing protocol for error-free identification and quantification. The low abundance of miRNA transcripts in biofluids makes determining its quality and quantity difficult. A standardised extraction protocol including commercial-based kits, careful selection of diseased individuals and controls for screening along with a well-established platform to sequence and quantitate may play a pivotal role in acknowledging circulating miRNA as a biomarker.
Footnotes
Acknowledgements
The authors acknowledge the National Health Education Society of P.D. Hinduja Hospital and Medical Research Centre for their support.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Ethical approval
Not applicable.
Guarantor
Dr. Swarup A. V. Shah.
Contributorship
CMM contributed to literature mining and manuscript writing. SAVS & TFA edited and critically evaluated the manuscript. All authors read and approved the manuscript.
