Abstract
Cancer is a complex and heterogeneous disease that poses a significant threat to global health. Early diagnosis and treatment are critical for improving patient outcomes, and the use of liquid biopsies has emerged as a promising approach for cancer detection and monitoring. Traditionally, cancer diagnosis has relied on invasive tissue biopsies, the collection of which can prove challenging for patients and the results of which may not always provide accurate results due to tumor heterogeneity. Liquid biopsies have gained increasing attention as they provide a non-invasive and accessible source of cancer biomarkers, which can be used to diagnose cancer, monitor treatment response, and detect relapse. The integration of -omics technologies, such as proteomics, genomics, and metabolomics, has further enhanced the capabilities of liquid biopsies by introducing precision oncology and enabling the tailoring of treatment for individual patients based on their unique tumor biology. In this review, we will discuss the challenges and advances in the field of cancer liquid biopsies and the integration of -omics technologies for different types of liquid biopsies, including blood, tear, urine, sweat, saliva, and cerebrospinal fluid.
Introduction
Despite advances in diagnosis and treatment, cancer remains a major public health threat, accounting for nearly 10 million deaths worldwide.1,2 Timely and accurate detection and diagnosis are critical to improving patient outcomes and survival. 3 The current standard of care involves surgery, radiation, and systemic therapies such as chemotherapy and immunotherapy, or targeted therapy. Appropriate treatment regimens are selected based on the patient’s cancer type and tumor staging, which in turn is determined by tumor profiling involving invasive resection/biopsies from the primary tumor.
Currently, the standard method of profiling tumors involves obtaining resected tumor samples through invasive surgeries. While necessary, obtaining both tumor quantity and quality can be a challenge. In addition, obtaining a holistic image of a tumor that has metastasized to multiple locations can be exceedingly difficult. Establishing a longitudinal tumor profile over the course of treatments would be highly useful to monitor the effectiveness of therapies and potential for recurrence, however subjecting patients to multiple invasive biopsies is less than ideal. The answer to these challenges could be found through the utilization of liquid biopsies which have emerged as a promising and more comprehensive approach in providing non-invasive biomarkers for diagnosis, prognosis, and treatment response monitoring of disease.4 -7
Liquid biopsies (LB) are minimally invasive diagnostic tools that involve the sampling and analysis of biofluids for screening, diagnostic, and prognostic purposes. The main body of research and development has been conducted in blood. 8 However, tears, saliva, urine, cerebrospinal fluid, and other biofluids serve as sources as well. Circulating biomarkers, such as circulating tumor cells (CTCs), cell-free DNA (cfDNA), and exosomes are just a few of the analytes that can be extracted from liquid biopsies for clinical evaluation and hold great promise for revealing the molecular landscape and fate of primary tumors. These biomarkers provide information on the cancer’s molecular and genetic makeup, allowing clinicians to monitor disease progression and response to treatment over the entire journey of the cancer patient. Incorporation of the isolation of these analytes from biofluids is a critical step in the liquid biopsy workflow.
CTCs are rare tumor cells shed into the bloodstream, providing insights into tumor metastasis and progression.9,10 Various techniques have been developed for CTC isolation, including immunomagnetic separation and microfluidic devices. The CellSearch system, based on immunomagnetic capture using EpCAM antibodies, is a widely used method for CTC enrichment, and is the only FDA-approved device for CTC detection and enumeration 11 Additionally, microfluidic devices offer label-free and size-based CTC isolation, enhancing the sensitivity of detection. 12
ctDNA is fragmented DNA released into circulation by apoptotic and necrotic cells. 13 Isolation methods for ctDNA involve plasma or serum extraction followed by DNA purification. Commercial kits, such as QIAamp Circulating Nucleic Acid Kit, provide efficient extraction, although it is a manual platform. 14 To analyze ctDNA mutations, platforms such as digital droplet PCR (ddPCR) and high-throughput next-generation sequencing (NGS) are most commonly used. 15
Exosomes are small membranous vesicles secreted by cells that contain a wide array of biomolecules, such as proteins, nucleic acids, and lipids.16,17 Ultracentrifugation, size-exclusion chromatography, and immunoaffinity capture are common methods for exosome isolation. ExoQuick and exoEasy are 2 of the most frequently used and commercially available platforms. 18 Further characterization of exosomes involves techniques such as nanoparticle tracking analysis (NTA) and electron microscopy.19,20
The integration of -omics technologies, genomics, proteomics, transcriptomics, and metabolomics into liquid biopsies has enabled a more comprehensive analysis of the tumor’s molecular and genetic profile, giving way to a more personalized approach to cancer treatment by identifying specific targets for therapy.21 -26 Genomics is the hallmark of biomarker research and aids researchers to study the gene expression and regulation providing a deeper understanding of the molecular mechanisms involved in a disease at a genome level.27 -29 Similarly, proteomics has long served as a powerful technology to identify new protein biomarkers. By measuring the abundance of proteins, proteomics sheds light on a level above gene expression and the integration of these 2 fields provides a comprehensive view of cellular responses from transcription to translation. Recent advances have shown the importance of metabolomics and transcriptomics in their relevance to understanding cancer progression and monitoring. Metabolomics provides a comprehensive understanding of the metabolic state of a biological system which can be used to identify disease markers.30 -32 Transcriptomics, on the other hand, serves as a powerful tool for studying mRNA expression and regulation, allowing researchers to identify the mRNAs that are active in a particular biological process, and how they are regulated.33 -36 Combined, these -omics technologies serve as tools to provide complementary information and help researchers gain a better understanding of the complex biological processes that underlie health and disease. By utilizing these techniques, we can identify crucial biomarkers that can improve the accuracy and specificity of liquid biopsy tests.
This article highlights the advances in the field of cancer liquid biopsies using -omics technologies that were published in the last decade. We specifically focused on blood plasma as it is widely adopted and is a minimally invasive biofluid with more focus on other non-traditional biofluids such as urine, tears, saliva, sweat, and cerebrospinal fluid due to their non-invasive sample collection. We further discuss the challenges that need to be addressed moving forward to create personalized diagnostic tests that can be used in clinical testing for applications such as early cancer detection, progression, and treatment efficacy monitoring.
Liquid Biopsies: Blood
Blood-based liquid biopsies are the most extensively studied approach for cancer detection and monitoring. 37 Peripheral blood has long served as a source of oncological markers, providing insight into primary and metastatic tumor biology given that its circulatory nature allows it to easily remain in contact with primary and metastatic tumors.38 -43 The accuracy of the information derived from liquid biopsies is dependent on many factors, including the quality of pre-analytical workflow involving sample collection, processing, and storage.
The post-collection processing to obtain the desired blood fraction is a critical component of the pre-analytical workflow affecting blood-based liquid biopsies. Serum and plasma are the commonly used blood fractions used for investigation of cancer biomarkers. Plasma is obtained by adding an anticoagulant, followed by centrifugation to remove cellular material from the collected specimen. 44 However, the large proteomic dynamic range as well as potential for hemolysis during the collection process can lead to high false-positive results. 45 Based on the Plasma Proteomic Project by the Human Proteome Organization (HUPO), raw plasma is recommended over serum due to the lower degree of ex vivo degradation. 46 The timing of sample collection can also impact liquid biopsy results, with fluctuating levels of circulating tumor cells and circulating tumor DNA throughout the day and treatment course. 47 Additionally, a recent study mapped the expression variation of plasma proteins to provide insights into potential biomarkers and treatments for various age-related functional disorders. They reported that the functional analysis of the proteome revealed a significant difference in the plasma proteomic profiles between young adults and middle-aged adults or elderly adults. 48 Thus, for the purpose of biomarker discovery, it is important to establish standardized protocols for sample collection, such as using the same time of day and same collection methods across a diverse patient population.
Table 1 summarizes the recent developments in the field of cancer biomarker discovery using blood plasma. With regard to blood-based liquid biopsies, previous studies have leveraged mass spectrometry-based proteomics to identify candidate biomarkers for the detection of pancreatic cancer. They identified several proteins that were differentially expressed in patients with pancreatic cancer compared to healthy controls, including S100A6, S100A8, and S100A9. 49 Another research group employed liquid chromatography-tandem mass spectrometry to investigate novel plasma biomarkers in dissecting tumor stages and post-surgical outcomes by using albumin and immunoglobulin G-depleted plasma samples from individuals with hepatocellular carcinoma and cholangiocarcinoma. 50 The results showed an upregulation of the following proteins in hepatocellular carcinoma and a down regulation in cholangiocarcinoma-afamin, alpha-2-HS-glycoprotein, apolipoprotein B-100, clusterin, hepatocyte growth factor-like protein, and kininogen-1. Similarly, they also reported that the expression of Ig lambda chain V region 4A was upregulated in cholangiocarcinoma and had the opposite effect in hepatocellular carcinoma. Geary et al. used a sequential window acquisition of all theoretical fragment ion mass spectra (SWATH-MS) to identify proteomics signatures in late-stage cancer patients and reported a wellness score. 51 SWATH-MS technology has high precision and accuracy that allows for protein quantification without the need for labels and has been used to identify digitized proteomic signatures in other several cancer types including breast, colorectal, endometrial, and lung cancers.52 -55 Such studies provide insights into the clinical application of these biomarkers in the diagnosis and follow-up of cancers.
Recent cancer biomarker discoveries in blood.

Another rapidly growing area within the vast field of genomics is the detection of specific mutations in ctDNA using digital droplet PCR or next-generation sequencing. For example, a study used targeted amplicon sequencing to detect EGFR mutations in ctDNA from non-small cell lung cancer patients. Results suggested that ctDNA analysis was concordant with tissue biopsy analysis in 85% of cases. 64 In addition to proteomics and genomics, metabolomics has been used to identify metabolic alterations that are associated with cancer. A study used untargeted metabolomics to identify metabolic changes in pre-diagnostic serum (up to 24 years prior to diagnosis) from pancreatic ductal adenocarcinoma patients. They identified over 30 metabolites that were significantly altered related to subclinical disease, including glycylvaline, tryptophylgutamate, and aspartate. 63
The integration of transcriptomic analysis has enabled longitudinal monitoring of response to treatment as well as the prediction of treatment response to neoadjuvant therapies. A study by Di Cosimo et al. analyzed expression levels of ct-miRNA in plasma samples from HER2-positive breast cancer patients during treatment with trastuzumab and revealed increases in ct-miR-148a-3p and ct-miR-37a-5p as early as 2 weeks following treatment commencement for patients attaining pathological complete response (pCR). 61 Another study highlighted the diagnostic and prognostic capabilities of miRNA in NSCLC by isolating exosomes from NSCLC patient serum to analyze miR-382 expression. They found a significant drop in miR-382 levels in NSCLC cases compared to healthy controls, as well as improved sensitivity and specificity when combining the miR-382 expression with carcinoembryonic antigen. 62 On the immunotherapy front, miRNA profiling is also enabling the development of prognostic scores that predict overall survival to immunotherapy in advanced-stage NSCLC. Moreover, Rajakumar et al. demonstrated how the 5-microRNA risk score they defined (miRisk) proved superior to tissue-based PDL1 scoring/staining currently used in the clinic. 65
However, several challenges need to be addressed for the clinical implementation of these biomarkers, including the low abundance of CTCs and tumor-derived exosomes (TDEs) in the blood, the heterogeneity of ctDNA, and the lack of standardized collection and analysis methods.
Liquid Biopsies: Urine
Following blood, urine is one of the most commonly used biofluids for isolating markers for urine and nonurinary tract diseases. Urine-based liquid biopsies have gained significant attention in recent years as a non-invasive method for the diagnosis and monitoring of cancer, given its less invasive sampling nature.
Urine samples can be collected using various methods, including midstream voided urine, catheterization, and urine bags. 66 Midstream (clean-voided) urine is the most common method used for urine collection, which involves collecting the urine in a clean container after discarding the initial stream to avoid contamination. 67 Catheterization is a more invasive method that involves inserting a catheter into the bladder to collect urine. 68 Urine bags are also used for urine collection in infants and young children. 69 Regardless of the collection method used, it is essential to follow standard operating procedures to avoid contamination and ensure the accuracy and reliability of results. Urine samples should be collected in sterile containers and processed as soon as possible to avoid degradation of the biomarkers of interest.
While nucleic acids and metabolites can also be analyzed, proteins are the most common biomarkers analyzed in urine. Given that following ultrafiltration of plasma, less than 1% is excreted as urine, the protein concentration remaining in urine is significantly lower than that of plasma, containing up to 150 mg of protein in approximately 1.5 L of urine output a day.70,71 The main protein found in urine is albumin, however, low abundant proteins can also be identified, such as prostate-specific antigen (PSA) for prostate cancer and urokinase plasminogen activator receptor (uPAR) for pancreatic cancer.72,73 Potential biomarkers for ovarian, breast, and pancreatic cancer have been identified through proteomic analysis of urine as well. Stockley et al. found MCM5 (mini chromosome maintenance 5) levels to be significantly increased in urine samples of both ovarian and endometrial cancer patients compared to controls, with overall sensitivities of 61.5% and 87.8%, respectively. 74 Similarly, in breast cancer patients, investigators found 59 urinary proteins to be differentially expressed compared to healthy control subjects. 75 Tomiyama et al. isolated extracellular vesicles (EVs) in urine samples and following proteomic analysis, they found 6 EV proteins to be significantly upregulated in bladder cancer patients compared to healthy individuals. 76
Nucleic acids, including DNA and RNA, are also analytes sourced from urine-based liquid biopsies. These markers provide valuable insight into mutations and alterations at the DNA and RNA levels, which can indicate the presence of cancer. For instance, urine-based liquid biopsies have been used to detect mutations in the telomerase reverse transcriptase (TERT) promoter region in bladder cancer patients. Specifically, Hosen et al. investigated pre-diagnostic urine samples from asymptomatic patients who subsequently developed bladder cancer and were able to identify TERT promoter mutations at a sensitivity of 46.67% and specificity of 100%. 77
Amino acids, organic acids, and lipids are of additional interest to investigate in urine. A recent study identified over 20 different urinary volatile metabolites to be altered in bladder cancer patients. Specifically, they reported 11 metabolites to be down-regulated in bladder cancer patients, and 12 metabolites to be up-regulated, including 2-furaldehyde and 4-metholoctane. 78 Table 2 summarizes the recent developments in cancer biomarker discovery using urine as a non-invasive biofluid.
Recent developments in cancer biomarker discovery using urine as a biofluid source.

One of the main challenges with urine analysis is the variabilities observed in the inter- and intra-individual samples which is dictated by several factors such as sex, age, and lifestyle.84 -86 Additionally, urine samples have relatively low proteins but high salt concentrations thus necessitating the need for preprocessing samples to lower the salt concentrations via dialysis or precipitation.
Liquid Biopsies: Tears
Tears are an easily and continuously accessible biological fluid that holds great promise for the identification of biomarkers. 87 Tears are clear, extracellular fluids produced by the lacrimal gland and are composed of 3 distinct layers: the inner mucin layer, middle aqueous layer, and the outer lipid layer. They are ultrafiltrate of blood plasma and contain a diverse mixture of molecules, including proteins, microRNAs, lipids, mucins, and small molecule metabolites. 88 They serve as an affordable source and require minimal storage requirements as tear samples can be preserved for years without undergoing any degradation when frozen and stored at low temperatures (−20°C to −80°C). 89 Moreover, basal, open-eye tears have a high concentration of proteins, usually ranging from 6 to 11 mg/mL.87,90 Additionally, tears are highly sensitive to changes in systemic and ocular conditions, which makes them a valuable diagnostic tool for monitoring and evaluating various health conditions. 87
Microcapillary tubes, polyester/polyvinyl wicks, and Schirmer strips are some of the methods employed for tear collection. 91 Schirmer strips are more common in clinical settings as they are used in standard ophthalmologic testing for dry eye disease. Compared to microcapillary tubes, Schirmer strips are easier to use, quicker, and more reliable. 92 One of the main difficulties in tear analysis is the low volume of tears (~3-10 μL) collected and researchers typically pool the samples to address this issue. Sample pooling offers the advantage of increased volumes for analysis while reducing the inter- and intra-subject variability, but this method does not provide individual variability. With the recent technological advancements of high-speed, high-resolution mass spectrometers, several new studies perform analysis on individually collected tear samples and have reported single samples with 500 to 1500 identified proteins involved in multiple signaling pathways.93 -96 Additionally, a recent study by our group conducted a comparative analysis of tears and blood plasma collected from the same individual and evaluated the samples by protein microarrays and small RNA sequencing. We reported that the Explorer protein microarray identified 71 proteins and RNAseq identified 250 microRNAs that were commonly expressed across both samples. 97
As these biofluids offer a plethora of biomarkers, they have been studied in breast and colon cancers. Lebrecht et al. reported the differences between healthy and breast cancer patients by the tear proteome profiling utilizing mass spectrometry.98,99 Further studies by the same group noted that the proteins were involved in immune-response pathways, such as C1Q1 and S100A8, and metabolic pathways, such as ALDH3. 100 Our group reported similar results with the upregulation of S100A8 and S100A9 proteins among 9 other proteins that could serve as potential biomarker candidates (sensitivity of 84.8% and specificity of 86.4% with an AUC of 0.902) for differentiating healthy and breast cancer patients. 101 Additionally, a subsequent study was conducted with a large single-tear analysis of over 800 samples on the calgranulin-A and -B proteins to validate these biomarkers along with clinical covariates. 102 We showed the successful early classification of samples using 2 proteins and minimal clinical covariates by proposing 3 models with sensitivity ranging between 52% and 90% and specificity ranging from 31% to 79%. Another recent study by Kaufmann et al. reported 18 upregulated and 62 downregulated proteins when they analyzed tears from healthy and colorectal cancer patients. 103
Besides using proteins as biomarkers for early diagnosis and screening of cancer, microRNAs have also been reported as potential biomarkers. The study evaluated the expression of breast cancer-specific miR-21 and miR-200c and found them to be significantly higher in tear exosomes collected from metastatic breast cancer patients compared to healthy volunteers. 104 This study also revealed that the tear exosomal markers, CD9 and CD63, were significantly expressed in higher quantities compared to serum exosomes, which could be due to the loss of exosomes during the serum sample preparation. Furthermore, the same group explored the possibility of developing an antibody-conjugated nanocavity-based platform which was fabricated using molecular imprinting approach. These platforms can potentially serve as a sensor for detecting cancers using tear exosomes. 105 Table 3 summarizes the recent advancements in using tears for cancer biomarker discovery.
Recent cancer biomarker discovery studies involving tears as a non-invasive biofluid source.

Tears are advantageous due to their ease of collection, non-invasive nature, and require minimal sample preprocessing while providing crucial genetic and proteomic information. However, similar to the challenges discussed in the previous section, standardized sampling collection and handling methods along with large-scale single tear analysis are required to increase reproducibility, which can address the gaps in the field.
Liquid Biopsies: Saliva
Saliva holds great potential as a non-invasive source of new biomarkers for the diagnosis and prognosis of cancer. It is primarily produced by 3 major salivary glands—the parotid, submandibular, and sublingual glands, along with 300 to 400 minor salivary glands located within the oral cavity. 106 Saliva is a complex fluid that contains proteins, DNA, mRNA, microRNAs, metabolites, and microbiota, making it a promising source of biomarkers and offers several advantages over blood and tissue, including non-invasiveness, easy storage, cost-effectiveness, and dynamic availability for monitoring.107,108 These biofluids can reflect the physiological state of the body and allow for the monitoring of oral and systemic health and diseases. Additionally, about a third of the proteins found in saliva are also detected in plasma. 109 Salivaomics is a widely used term that refers to a diverse range of technologies utilized to investigate the various types of molecules present in saliva. Several saliva-based diagnostic kits for detection of drugs, COVID, and pregnancy are commercially available and several others evaluating the cardiac risk and malaria are in development. 110
Proteomic biomarkers have been evaluated in saliva for various cancer types including oral, breast, pancreatic, and lung. Chu et al. created an innovative and cost-effective method by integrating salivary immunoglobulin A purification and affinity mass spectrometry to identify biomarker candidates that differentiated healthy individuals from those who had oral cavity squamous cell carcinoma. 111 They reported that 7 salivary autoantibodies had the potential to serve as biomarker candidates and developed a 4-autoantibody diagnostic panel with high sensitivity and specificity consisting of vesicular integral-membrane protein VIP36, prostaglandin reductase 1, Ras-related protein Rab-13, and Cytochrome b-c1 complex subunit 2, mitochondrial. More recent studies have also showed an altered expression level in the salivary metabolites such as lactate, proline, glycine, citrulline, inositol trisphosphate, 2-oxoarginine, and glycerate-2-phosphate in oral cancer patients.112,113
A recent meta-analysis conducted by Koopaie et al. reported that unstimulated saliva-derived biomarkers for breast cancer, including c-erb-B2 and sialic acid, had sensitivity and specificity comparable to that of serum. 114 Another interesting finding in the study was that the levels of autoantibodies against MUC1, CA 15-3, and adiponectin in saliva demonstrated a stronger correlation with breast cancer than serum levels. With the advancement of microarray panels for genomic analysis, several miRNAs have been studied for their importance in salivary diagnostics. Xie et al. reported a model with reasonable sensitivity and specificity, which involved 2 salivary miRNAs—miR-3679-5p and miR-940, that can detect pancreatic cancer. 115 Another group quantified salivary miRNAs using qRT-PCR and reported that hsa-miR-21, hsa-miR-23a, hsa-miR-23b and miR-29c had significant expression in the saliva of pancreatic cancer patients. 116 Table 4 summarizes the fairly new developments in cancer biomarker discovery using saliva as a biofluid source.
Recent development in cancer biomarker discovery using saliva as a non-invasive biofluid source.
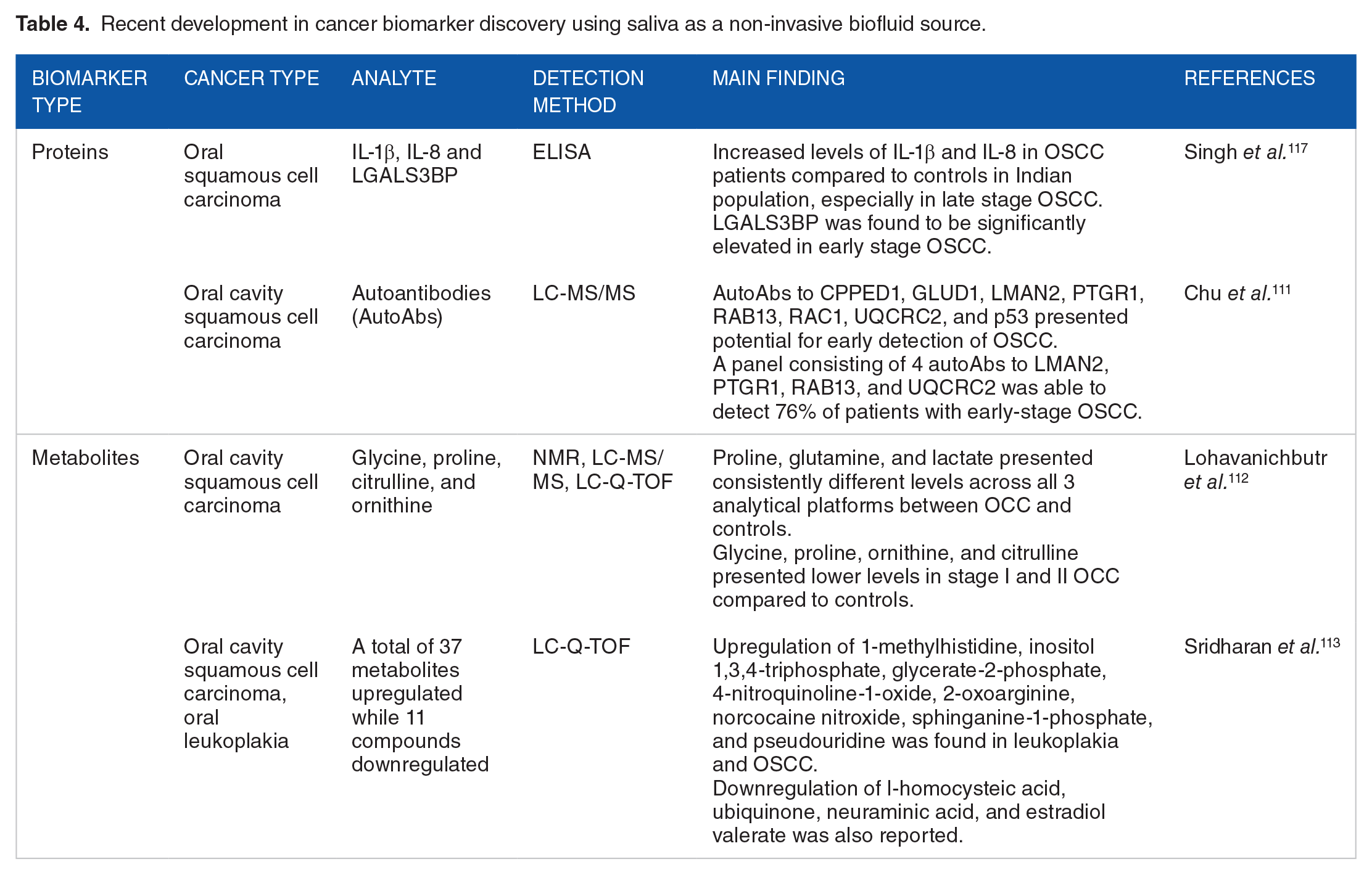
Despite the advantages offered by saliva-based biomarkers, most of the published literature involves smaller sample sizes, which poses a limitation. Several studies use healthy controls for comparison with a disease with limited validation of inflammatory conditions that might be associated with the disease studied. Such complex interactions need to be included in the study design to develop accurate tests that can reduce the false positive rates. Additionally, saliva protein composition databases that support the investigation and comparison of these biofluids are currently available, which can serve to advance the field. 118
Liquid Biopsies: Sweat
Sweat is a less commonly used biofluid for liquid biopsy compared to those previously discussed for cancer detection. It is secreted by the eccrine and apocrine glands and comprised mainly of water and electrolytes, metabolites, cytokines, cortisol, ammonia, serum components, and other organic compounds.119,120 Sweat specimens can be collected by non-invasive methodologies that vary from simple and direct collection of sweat off the skin and into testing tubes, to more involved and specifically designed methodologies involving commercially available products, such as Pharmcheck® patches and the Macroduct® Sweat collection system. The Pharmcheck patches are non-occlusive and absorbent bandages composed of a cellulose layer that adheres to the skin and in which larger molecules are deposited. The patch is covered by an external polyurethane layer that protects the patch from external contaminants, allowing it to be worn during normal activities and for longer periods of time. 121 Other non-commercial techniques involve hydrogel micropatches, glass rollers used on the arms, or glass pipettes used on areas such as the forehead, chest, or back.122,123 A wide variety of sweat collection methods have been used across several studies, depending on the analyte under investigation.
Several studies have shown the promising potential of sweat as a non-invasive liquid biopsy for cancer diagnosis. Calderón-Santiago et al. performed metabolic analysis of sweat from patients with lung cancer and found that trisaccharide phosphate best discriminated between lung cancer patients and controls (sensitivity 72.7%, specificity: 76.5%). Moreover, they developed a panel of 5 different metabolites, including suberic acid, tetrahexose, trihexose, nonanedioic acid and monoglyceride (22:2), that achieved a specificity of 80% and sensitivity of 79%. 124 More recently, Monedeiro et al. leveraged headspace-gas chromatography coupled to mass spectrometry (HS-GC-MS) to investigate the volatile fraction of sweat collected from healthy controls and individuals with different types of cancer including lung, prostate, gastric, kidney, head and neck, pancreas, colorectal, and lymphoma. 125 Sweat was isolated from PharmChek patches that had previously been applied to patients. Using a machine learning approach, sweat VOC profiles allowed the classification of healthy and diseased patients with 100% accuracy, when considering a panel consisting of 2-ethyl-1hexanol, octanal, and hexanal. 125 On the breast cancer front, Zadák et al. report on the diagnostic capabilities of metabolite markers isolated from apocrine sweat. Using liquid chromatography with tandem mass spectrometry (LC-MS/MS) and mathematical-statistical methods, they identified 20 metabolites that were able to discriminate between breast cancer patients and healthy controls with a sensitivity of 97% and specificity of 72%. 126 Table 5 summarizes the recent developments using sweat as a source for identifying cancer biomarkers.
Recent development in cancer biomarker discovery using sweat as a biofluid source.
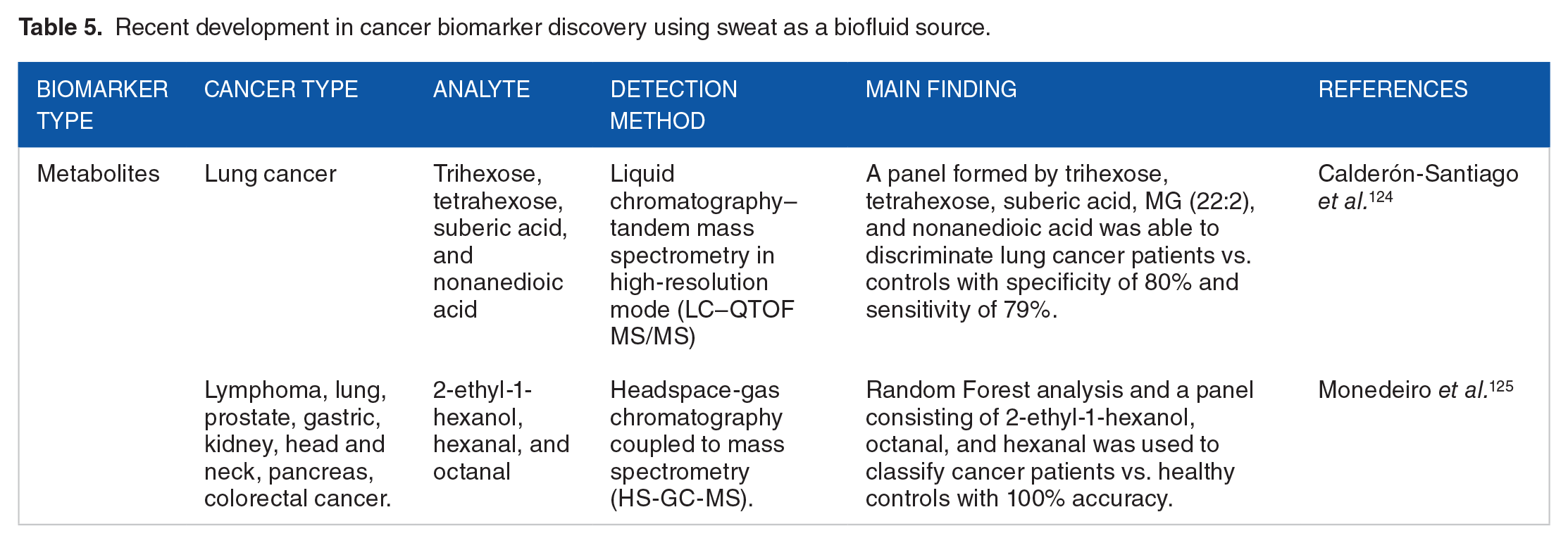
Several challenges need to be addressed to make sweat a more reliable and clinically useful liquid biopsy for cancer diagnosis. One of the main challenges is the low concentration of cancer biomarkers in sweat, which makes their detection difficult. 127 Furthermore, sweat is easily influenced by environmental factors such as temperature, humidity, and physical activity, which may affect the accuracy and reproducibility of the results and can vary greatly from skin to skin. However, with the development of more sensitive and specific analytical methods and the standardization of sample collection and processing protocols, sweat may become a promising non-invasive liquid biopsy for cancer diagnosis.
Liquid Biopsies: Cerebrospinal Fluid
Cerebrospinal fluid (CSF) is a clear, colorless fluid that safeguards the central nervous system (CNS) by providing mechanical protection and immunological buffering, as well as enabling nutrient and waste transport within the CNS.128,129 Secreted primarily by the choroid plexi in the brain’s ventricles, CSF circulates within the subarachnoid space surrounding the brain and spinal cord, presenting unique access to CNS-specific biomarkers. 130 CSF is made up of approximately 99% water, with the remaining part consisting of a wide array of biomolecules, including proteins, glucose, ions, nucleic acids, metabolites, vitamins, and neurotransmitters, all of which can be indicative of underlying pathophysiological processes, including cancer. 131 The identification of cancer-related biomarkers in CSF offers the potential for monitoring treatment response and assessment of disease progression in CNS-involved malignancies. 132
Collection of CSF most commonly involves a lumbar puncture, also referred to as a spinal tap. During this procedure, a puncture needle is carefully inserted into the subarachnoid space of the spinal column, allowing for the withdrawal of CSF. 133 The collected fluid is then subjected to a number of analyses to identify disease-specific markers. CSF analysis is particularly relevant in CNS-related malignancies such as glioblastoma, medulloblastoma, and leptomeningeal metastasis brought about by other cancers. Previous studies have reported on the reliability of CSF to act as a source of cell-free ctDNA, presenting levels of ctDNA higher than those reported in plasma. 134 In the case of brain cancer, De Mattos-Arruda et al. investigated the genetic landscape of brain tumors through the analysis of cfDNA isolated from CSF. 134 Their results underscored the importance of CSF-derived ctDNA in accurately reflecting genomic alterations present in CNS tumors. Furthermore, the work of Ballester et al. leveraged metabolomics to analyze 129 distinct metabolites in CSF samples from patients with a wide variety of CNS tumor types and revealed higher levels of D-2-hydroxyglutarate in the CSF of IDHG-mutant tumors compared to patients of other tumor types or controls (Table 6). 135
Recent development in cancer biomarker discovery using cerebrospinal fluid as a biofluid source.

Other Non-invasive Biofluids
A few studies have evaluated the potential of other biological fluids as a source for identifying cancer biomarkers. Martinez-Garcia et al. recently reported protein biomarkers in cervical fluids that can serve as a tool for early diagnosis of endometrial cancer. 137 Another group from Spain isolated exosomal vesicles from semen and reported significant changes in the miRNA signatures from these exosomes, distinguishing between prostate cancer patients and healthy controls. 138 Breast milk is another biofluid that has been reported to possess protein biomarkers that correlated with early detection of breast cancer.139,140 Though these other biofluids have been utilized to identify potential cancer biomarkers, large scale applications are still limited (Table 7).
Recent development in cancer biomarker discovery using other unconventional biofluid sources.

Challenges and Future Perspectives
Cancer liquid biopsies have emerged as a promising non-invasive diagnostic tool that can provide real-time information on cancer initiation, progression, and response to treatment. Integration of -omics technologies has provided insights into the molecular mechanisms underlying cancer and can guide personalized treatment decisions, furthering the field of precision oncology. Despite the significant advances in cancer liquid biopsies and -omics technologies, many challenges remain and need to be addressed before they can be widely implemented in routine clinical practice. One major challenge is the lack of standardization of liquid biopsy analysis. The pre-analytical, analytical, and post-analytical factors can affect the accuracy and reproducibility of liquid biopsy results. Therefore, standardized protocols and quality control measures need to be established to ensure the reliability and reproducibility of liquid biopsy analysis.
A well-designed -omics study is crucial to obtain meaningful and reliable results that can advance our understanding of biological processes and diseases. Omics technologies generate large amounts of data that require careful planning and execution to ensure that the results are meaningful and biologically relevant. Omics tools require the use of complex statistical tools; with more and more researchers being able to leverage the benefits of machine learning we are witnessing an increase in our ability to identify complex biomarker relationships with clinical factors at a tremendous rate. When designing a biomarker discovery study, researchers should prioritize the importance of maximizing the information gain and addressing the clinical needs through appropriate sample selection along with clear clinical endpoints.
Depending on the biological fluid of interest, the mode for sample collection differs and it is critical to understand the various factors that can introduce bias, such as sample collection protocols, potential contamination, and the importance of incorporating quality checks. An ideal study should involve randomization methods (simple, block, stratified, or adaptive randomizations) of the samples by allocating patients to groups that avoid any potential bias. Importance should be given to the determination of sample sizes to ensure robust results by utilizing statistical power analysis tools. Additionally, proper data sharing practices and ethical considerations need to be implemented, which promote the advancement of the field.
Another challenge is the low sensitivity of liquid biopsy analysis in some cancer types which results in high false negative rates leading to missing out on cancer diagnosis. For instance, liquid biopsy analysis has shown limited sensitivity in detecting early-stage prostate cancer which could be due to the low levels of ctDNA and the high heterogeneity of prostate tumors. 141 Similarly, tests with a low specificity result in high false positives which can lead to overscreening/diagnosis causing more burden on the healthcare system and patients. Therefore, it is crucial that researchers conduct larger cohort studies in order to develop tests with high accuracy.
The clinical implementation of -omics techniques is largely limited by the expensive equipment and technical expertise. We believe that ease of equipment use, standardized sample collection, handling, traceability, reproducibility, and automated analysis of data are required for these advanced and novel technologies to be implemented in daily clinical practice. The time and cost of developing a diagnostic test can be a major obstacle and pose a significant barrier. A collaborative effort between government agencies, academia, and the private sector is highly needed and such large multicenter trials would address these challenges. Lastly, a significant barrier to biomarker-based testing is clinician and insurer acceptance of non-traditional diagnostic tests. This can be addressed by proper education on the potential of biomarkers that provide key health information, evidenced by large clinical datasets, is required to convince physicians, clinicians, and general practitioners for the widespread adaptation of similar technologies in clinics.
As we look to the future of health and healthcare, the holy grail of laboratory medicine is often touted as personalized thresholds for disease. The work being done now to identify which markers hold potential lays the groundwork for this important perspective. To this end, each biofluid holds its own potential for breakthroughs and challenges. The most likely outcome will be that markers from each biofluid can be utilized to build a map personalized for each individual.
Conclusion
This review provides a summary of the recent advances in the field of cancer liquid biopsies that leveraged -omics technologies. We can certainly identify that the biological value of miRNAs, proteins, and other analytes obtained from liquid biopsies offers several advantages; however, they still require further validation. We believe that despite the challenges, continued research and development of cancer liquid biopsies and -omics technologies are anticipated to bring about a transformation in cancer diagnosis and tracking, consequently enhancing the well-being of patients.
