Abstract
Background
It is difficult for clinical laboratories to identify samples that are labelled with the details of an incorrect patient. Many laboratories screen for these errors with delta checks, with final decision-making based on manual review of results by laboratory staff. Machine learning models have been shown to outperform delta checks for identifying these errors. However, a comparison of machine learning models to human-level performance has not yet been made.
Methods
Deidentified data for current and previous (within seven days) electrolytes, urea and creatinine results was used in the computer simulation of mislabelled samples. Eight different machine learning models were developed on 127,256 sets of results using different algorithms: artificial neural network, extreme gradient boosting, support vector machine, random forest, logistic regression, k-nearest neighbours and two decision trees (one complex and one simple). A separate test data-set (n = 14,140) was used to evaluate the performance of these models as well as laboratory staff volunteers, who manually reviewed a random subset of this data (n = 500).
Results
The best performing machine learning model was the artificial neural network (92.1% accuracy), with the simple decision tree demonstrating the poorest accuracy (86.5%). The accuracy of laboratory staff for identifying mislabelled samples was 77.8%.
Conclusions
The results of this preliminary investigation suggest that even relatively simple machine learning models can exceed human performance for identifying mislabelled samples. Machine learning techniques should be considered for implementation in clinical laboratories to assist with error identification.
Keywords
Introduction
The preanalytical phase is a major source of error in clinical laboratory testing. Mislabelling of a sample with the details of an incorrect patient is particularly difficult to identify. Current laboratory practice is to use delta checks to screen for these errors and have results failing delta checks manually reviewed by laboratory staff. 1 Preliminary investigations have demonstrated that machine learning (ML) models are superior to delta checks in identifying mislabelled samples. 2 , 3 The relative performance of different ML algorithms has not been established, however. Moreover, it is unknown how ML models compare to human-level performance. In the following short report, the performance of eight different ML models and laboratory staff for the identification of simulated mislabelled samples was compared.
Methods
De-identified data was extracted from the laboratory information system (LIS) of a public hospital in Sydney, Australia for the period 1 June 2019 to 26 October 2020. Patient age, sex, current and previous results for sodium, potassium, chloride, bicarbonate, urea and creatinine, as well as the date and time of collections were extracted. Samples without previous results within seven days were excluded and 141,396 samples remained.
Mislabelled samples were simulated at a rate of 50% by random switching of patients’ current results with results from other patients, with a record stored of which results were switched. This data was then randomly split into three subsets: 80% (n = 113,116) for ML model training, 10% (n = 14,140) for ML hyperparameter tuning and 10% (n = 14,140) for testing final ML models and volunteer performance. These proportions, commonly used for ML problems, were selected to maximize training set size while still allowing adequate sample numbers for hyperparameter selection and testing of model performance.
ML algorithms were trained, tuned and tested in the R environment using the packages ‘class’, ‘rpart’, ‘randomForest’, ‘e1071’, ‘xgboost’, ‘keras’ and ‘caret’. 4 Two decision tree models were developed, one complex and one simple. The complex model was selected to maximize accuracy without limitations on complexity, while the simple model was limited to fewer than 10 decision nodes to allow it to be easily deployed within standard LISs. Further detail regarding the methods used is provided as supplementary material.
Human performance on the test data was assessed via a purpose-written web application (app). Staff with biochemistry roles in the network of public hospital laboratories were invited to access the app. De-identified information was collected regarding the volunteer’s position, years’ of experience validating biochemistry results and proportion of work time spent validating. For each volunteer, the app randomly selected a set of results, which were presented along with the age and sex of the patient, and the date and time of collection of the current and previous sample. The volunteer was then asked whether or not they thought the current results were from a mislabelled sample. Ten sets of results were presented to each volunteer.
Ethics review was not required because the project met criteria for exemption from such review, as a quality improvement activity, as per National Health and Medical Research Council criteria. 5 Data analysis was performed in the R environment. 95% Confidence intervals (CI) of parameter estimates were calculated as twice the standard error. Associations between volunteer characteristics and accuracy were assessed by t-tests of linear regression coefficients.
Results
Fifty laboratory staff volunteered to participate. Twenty-eight held the position of ‘Technical Officer’, 15 ‘Scientific Officer’ and five ‘Senior Scientific Officer’. Two recorded their position as ‘Other’ (not pathologist). Volunteers had a median of 4.5 years’ experience validating biochemistry results (range 0–40 years).
The mean (±95% CI) accuracy of volunteers was 77.8% (±4.6%). There was no association between volunteer accuracy and their position or number of years’ experience (P > 0.1). There was a non-statistically significant trend (P = 0.08) to higher accuracy among volunteers who spent ‘almost all’ of their work time validating results.
All ML algorithms exceeded human-level performance (Figure 1). The poorest performing model, the simple decision tree (Supplementary Figure 1), achieved an accuracy on the test data of 86.5% (±0.25%). The best performing algorithm was the artificial neural network (ANN) with an accuracy of 92.1% (±0.44%). Area under the receiver operating characteristic curve varied from 0.901 (±0.005) for the simple decision tree to 0.977 (±0.002) for the ANN. Details of the performance of all ML algorithms are given in Supplementary Table 1.
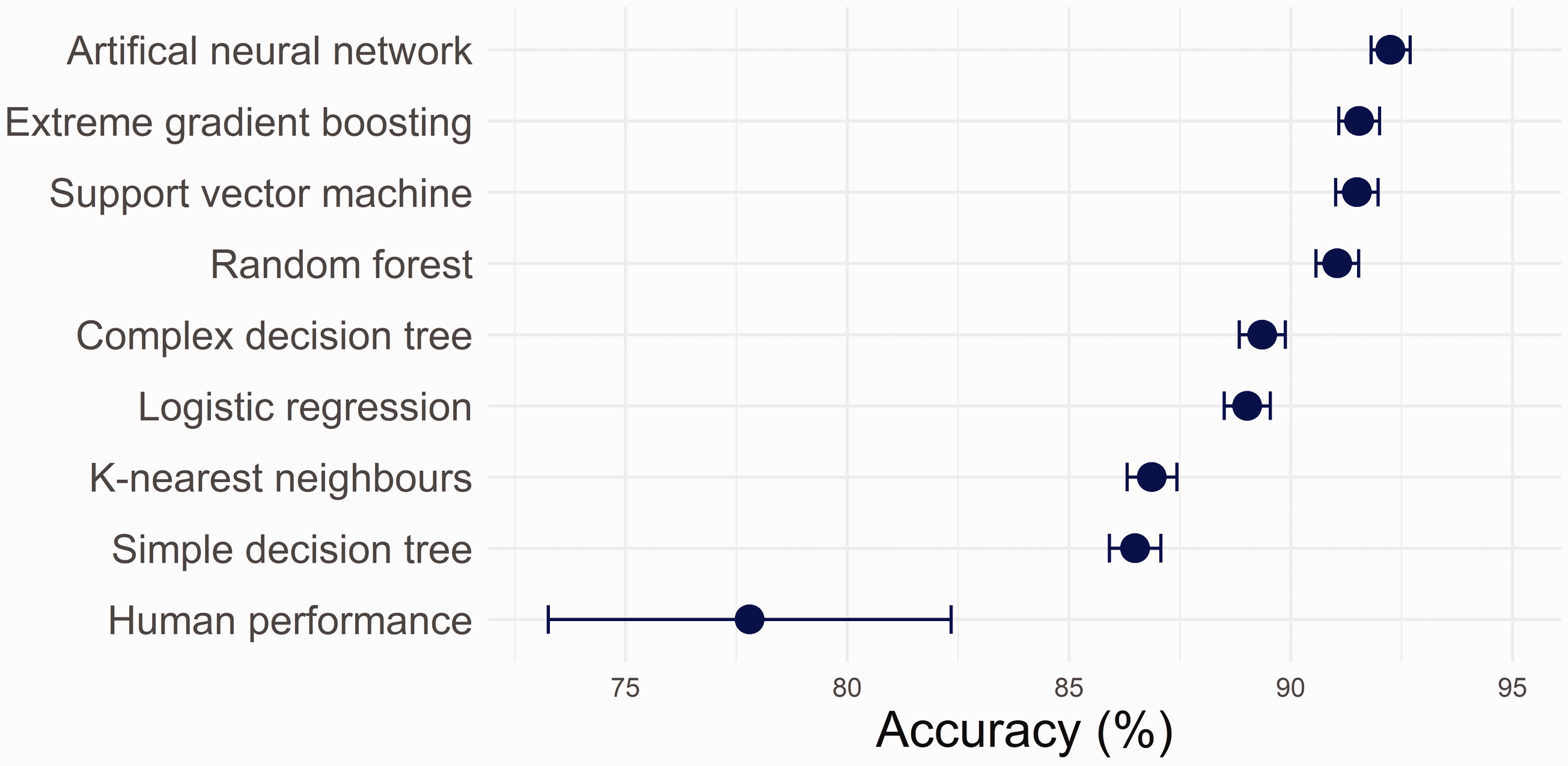
Accuracy of machine learning algorithms in identifying simulated mislabelled samples on a test data-set (n = 14,140) of current and previous (within seven days) electrolytes, urea and creatinine results. Also presented is the accuracy of laboratory staff volunteers in manually identifying mislabelled samples on a random subset of this data (n = 500).
Discussion
The results of this investigation suggest that ML models exceed human-level performance in the identification of mislabelled samples. This finding is notable, as it would appear to be the first example in the clinical biochemistry context of ML models outperforming humans on a task traditionally performed manually.
ML models should hence be considered for routine implementation to assist with identifying labelling errors. Among the ML models, the computationally intensive algorithms performed better. LIS and middleware vendors are therefore encouraged to develop tools to facilitate the implementation of such models. Nevertheless, this study indicates that human-level performance can be exceeded with relatively simple algorithms. The simple decision tree used only seven decision nodes, and logistic regression models are computationally similar to multiple linear regression equations. Such models could be trained by laboratory or information technology staff and implemented into existing laboratory software.
This study had a number of limitations. Firstly, computer simulation randomly introduced labelling errors. This is expected to reasonably replicate errors,1–3 but it does not include non-random features that could be present in practice. Further, the specific task tested did not necessarily reflect the prior experience of laboratory staff, who encounter mislabelling errors infrequently, review results within particular workflows and have access to additional sample-related information. In contrast, the ML models were trained specifically on the parameters provided on this task. Consequently, the laboratory staff may have been at some disadvantage. Additionally, only six tests were used for comparison between current and previous samples. Both ML and human performance would likely improve as the number of tests assessed increased, but as the number of tests became large, ML algorithms might be better able to synthesize the additional information. This remains a topic for future investigation. Finally, the simulated error rate was significantly higher than sample mislabelling in practice. 6 This was necessary to assess human-level performance, as only 500 sample pairs were reviewed by volunteers. Nevertheless, by adjusting the decision threshold, ML models could be tuned to maximize the sensitivity/specificity trade-off for the rate of sample labelling errors laboratories encounter in practice.
In summary, ML algorithms exceeded human-level performance for identifying mislabelled samples. Development and deployment of ML techniques into routine practice should be considered to minimize the risk of patient harm arising from these errors.
Supplemental Material
sj-pdf-1-acb-10.1177_00045632211032991 - Supplemental material for Identifying mislabelled samples: Machine learning models exceed human performance
Supplemental material, sj-pdf-1-acb-10.1177_00045632211032991 for Identifying mislabelled samples: Machine learning models exceed human performance by Christopher-John Farrell in Annals of Clinical Biochemistry
Supplemental Material
sj-pdf-2-acb-10.1177_00045632211032991 - Supplemental material for Identifying mislabelled samples: Machine learning models exceed human performance
Supplemental material, sj-pdf-2-acb-10.1177_00045632211032991 for Identifying mislabelled samples: Machine learning models exceed human performance by Christopher-John Farrell in Annals of Clinical Biochemistry
Supplemental Material
sj-pdf-3-acb-10.1177_00045632211032991 - Supplemental material for Identifying mislabelled samples: Machine learning models exceed human performance
Supplemental material, sj-pdf-3-acb-10.1177_00045632211032991 for Identifying mislabelled samples: Machine learning models exceed human performance by Christopher-John Farrell in Annals of Clinical Biochemistry
Footnotes
Acknowledgements
The author wishes to thank Dr Adam Polkinghorne, NSW Health Pathology, with assistance in preparing this article and the volunteers who assisted with the study.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Ethical approval
Not applicable.
Guarantor
CF.
Contributorship
CF conceived the study, developed the machine learning models and web application, analysed the data and wrote the original draft of the article.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
