Abstract
Study objective:
SARS-CoV-2, which causes coronavirus disease (COVID-19), continues to cause significant morbidity and mortality. The diagnosis of acute infection relies on reverse transcription-polymerase chain reaction (RT-PCR)-based viral detection. The objective of this study was to evaluate the optimal serological testing strategy for anti-SARS-CoV-2 antibodies which provides an important indicator of prior infection and potential short-term immunity.
Methods
The sensitivity and specificity of four different ELISA assays (Euroimmun IgG, Euroimmun NCP-IgG, Fortress and DIAsource) and one CLIA assay (Roche ELECSYS) were evaluated in 423 samples; 137 patients with confirmed RT-PCR COVID-19 infection (true positives), and 100 pre-pandemic samples collected prior to October 2019 (true negatives). A further 186 samples were collected from health-care staff and analysed by all five assays.
Results
The Fortress ELISA assay demonstrated the highest sensitivity and specificity followed by the Roche ECLIA assay. The highest overall sensitivity came from the assays that measured total antibody (IgM–IgG combined) and the three assays that performed the best (Fortress, Roche, Euroimmun IgG) all have different antigens as their target proteins which suggests that antigen target does not affect assay performance. In mildly symptomatic participants with either a negative RT-PCR or no RT-PCR performed, 16.76% had detectable antibodies suggesting previous infection.
Conclusions
We recommend a combined testing strategy utilizing assays with different antigenic targets using the fully automated Roche ECLIA assay and confirming discordant samples with the Fortress Total Antibody ELISA assay. This study provides an important indicator of prior infection in symptomatic and asymptomatic individuals.
Introduction
In late December 2019, China reported the first cases of an atypical pneumonia-like illness which spread globally 1 and became known as coronavirus disease 2019 (COVID-19), caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), the seventh coronavirus known to infect humans.2–4 Until the roll-out of vaccines are escalated, attempts to contain the virus include mass quarantine, isolation of infected individuals, enhanced hygiene strategy and serial lockdown, which have significant economic implications. 5
SARS-CoV-2 is an enveloped single-stranded RNA virus of the coronaviridae family, genus β coronavirus. All coronaviruses share similarities in the organization and expression of their genome which encodes 16 non-structural proteins and the 4 structural proteins spike (S), envelope (E), membrane (M) and nucleocapsid (N). In the case of SARS-CoV-2, the receptor-binding domain is found on the S1 domain of the spike protein and binds to angiotensin converting enzyme 2 receptors, with viral loads being detected in nasopharyngeal and respiratory swabs. 6
Where there is no ‘gold standard’ for the detection of SARS-CoV-2, the most reliable method for the diagnosis of acute infection is the detection of SARS-CoV-2 nucleic acid in respiratory samples. 7 Reverse transcription-polymerase chain reaction (RT-PCR) has some recognized preanalytical and analytical deficiencies. Preanalytical issues include variability in type, site and quality of swabs taken and timing of sample in the course of infection. Analytical characteristics vary and results are generally reported as positive or negative with the absolute cycle threshold (Ct) not standardized across assay systems. 7 Further confounders include potential cross- and carry-over contamination between specimens and potential contamination of reagents. 8 Antigen tests are available which detect viral proteins at the time of infection but have been shown to have a low specificity and assay performance improvements are necessary. 9 Rapid diagnostic tests in the form of qualitative lateral flow immunoassays (LFIA) are generally POC tests, but assessment of some of these assays did not show good analytical performance to date. Overall sensitivities with LFIA were low compared with RT-PCR, 10 and studies suggest that they currently should be used in combination with other more sensitive and specific methods. 11 In clinically unwell patients, chest CT is more specific than RT-PCR in diagnosing SARS-CoV-2 infection in persons with COVID-19 pneumonia,12,13 but imaging is not necessary in the majority of patients with COVID-19 disease.
Where there are advantages, there are also shortcomings with methodologies to detect SARS-CoV-2 accurately and there is a need to develop additional testing methods.14–16 Identification of infected individuals is paramount to controlling the spread of SARS-CoV-2 and has proven difficult due to a number of factors including, the overwhelming volume of new cases, the high incidence in people with vague or no symptoms, rationing of reagents for RT-PCR testing, lack of consensus within and between countries of criteria for testing, all of which contribute to a gross underestimation of cases of infection.17–19
In this context, International Federation of Clinical Chemistry and Laboratory Medicine (IFCC) have recommended serological tests as valuable indicators of prior infection in both non-hospitalized individuals (asymptomatic and symptomatic) and when assessing community exposure via seroprevalence studies. The IFCC emphasize that antibodies against SARS-CoV-2 serve as an adjunct to molecular testing in the following circumstances; those who have symptoms and are RT-PCR negative, those with persistent positive RT-PCR in the absence of infectious virus and in multi-system inflammatory syndrome in the paediatric setting. 20
Numerous SARS-CoV-2 serological tests for IgA, IgG and IgM have been rapidly developed, including assays measuring antibodies targeted against the viral spike protein which mediates entry of SARS-CoV-2 into host cells or the nucleocapsid protein which is a highly immunogenic structural protein. The World Health Organization (WHO) have raised concerns about the accuracy of these testing measures in the absence of larger scale in-depth verification studies 21 and emphasized the critical need for comprehensive evaluations of these different assays and technologies in clinical samples in order to identify the optimal testing strategies for the COVID-19 pandemic.
The overall aim of this study was to compare and evaluate the clinical and analytical performance and the functional sensitivity and specificity of a number of available anti-SARS-CoV-2 serological tests and to establish their performance characteristics and limitations in line with IFCC recommendations. Where the majority of previous studies have evaluated serological tests in hospitalized patients with severe SARS-CoV-2 infection, this study addresses the evaluation of antibody status in non-hospitalized SARS-CoV-2-positive patients, the majority of whom were health-care workers.
Methods
The study of the diagnostic utility was carried out in the Immunology/Serology laboratory of the Mater Private Dublin and was approved by the National Research Ethics committee Ireland (20-NREC-CoV-058). Informed consent was obtained from each participant who completed a detailed questionnaire prior to sample collection.
Patient recruitment
In total, 323 samples were collected primarily from health-care workers and their close contacts in two hospitals, a private and a state-funded acute hospital between June and September 2020 and a further 100 bio-banked samples collected prior to October 2019 were analysed. The samples were classified as follows; participants with RT-PCR confirmed positive COVID-19 disease (n = 137) which were considered ‘true positive’ samples and were used to assess the sensitivity of all the assays. These samples were collected >14 days post positive RT-PCR test or post onset of symptoms. The specificity of the assays was evaluated using only the samples collected prior to October 2019 (n = 100).
The remaining 186 samples that were collected were classified as follows: healthy asymptomatic participants (n = 109), and symptomatic participants (n = 77) with at least one or more of the following symptoms (fever, headache, sore throat, runny/stuffed nose, diarrhoea, cough/short of breath, myalgia, fatigue and loss of taste or smell). Of the 109 healthy asymptomatic participants, 13 had a SARS-CoV-2 RT-PCR test performed due to being close contacts of COVID-19-positive patients, all of which were negative. Of the 77 participants who had one or more of the COVID-19-related symptoms listed above, 31 of these participants had an RT-PCR test performed based on their symptoms and the remaining 46 participants did not have an RT-PCR test performed as their symptoms did not meet the criteria for testing at that time. Patient characteristics are summarized in Table 1.
Study groups including those who were RT-PCR positive, negative/no RT-PCR result and bio-banked samples collected prior to October 2019.
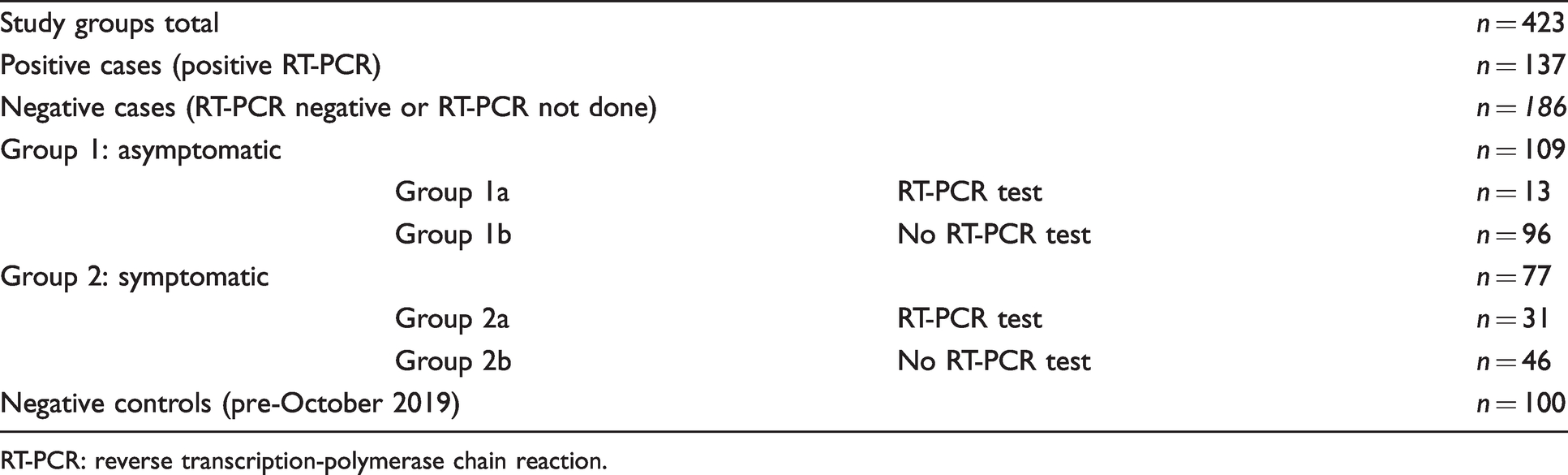

RT-PCR: reverse transcription-polymerase chain reaction.
Sample collection
Serum samples were collected from each participant and stored at –20 °C. Samples were thawed and mixed thoroughly prior to analysis, kept at 4°C for up to seven days and re-frozen with freeze thaw cycles minimized (fewer than five cycles).
Antibody analysis
Five different commercially available immunoassays were evaluated as follows: Elecsys anti-SARS-CoV-2 Total Ab (Roche, Germany)(electrochemiluminescence [ECLIA]), anti-SARS-CoV-2 IgG ELISA (Euroimmun, Germany, anti-SARS-CoV-2 NCP IgG ELISA (Euroimmun Germany), anti-SARS-CoV-2 Total Ab ELISA (Fortress Diagnostics, Antrim, United Kingdom/Wantai BioPharm, Beijing, China) and anti-SARS-CoV-2 IgG ELISA (DIAsource, Belgium/Epitope Diagnostics, USA).
Assays were performed in accordance with the manufacturer’s instructions by experienced laboratory scientists in an accredited laboratory (ISO 15189). Assay-specific controls and calibrants were used to ensure correct assay performance. The assay characteristics, protein targets, manufacturer’s suggested cut-offs and mode of analyses are summarized in Table 2.
Characteristics of the five different manufacturers’ SARS-CoV-2 antibody assays.
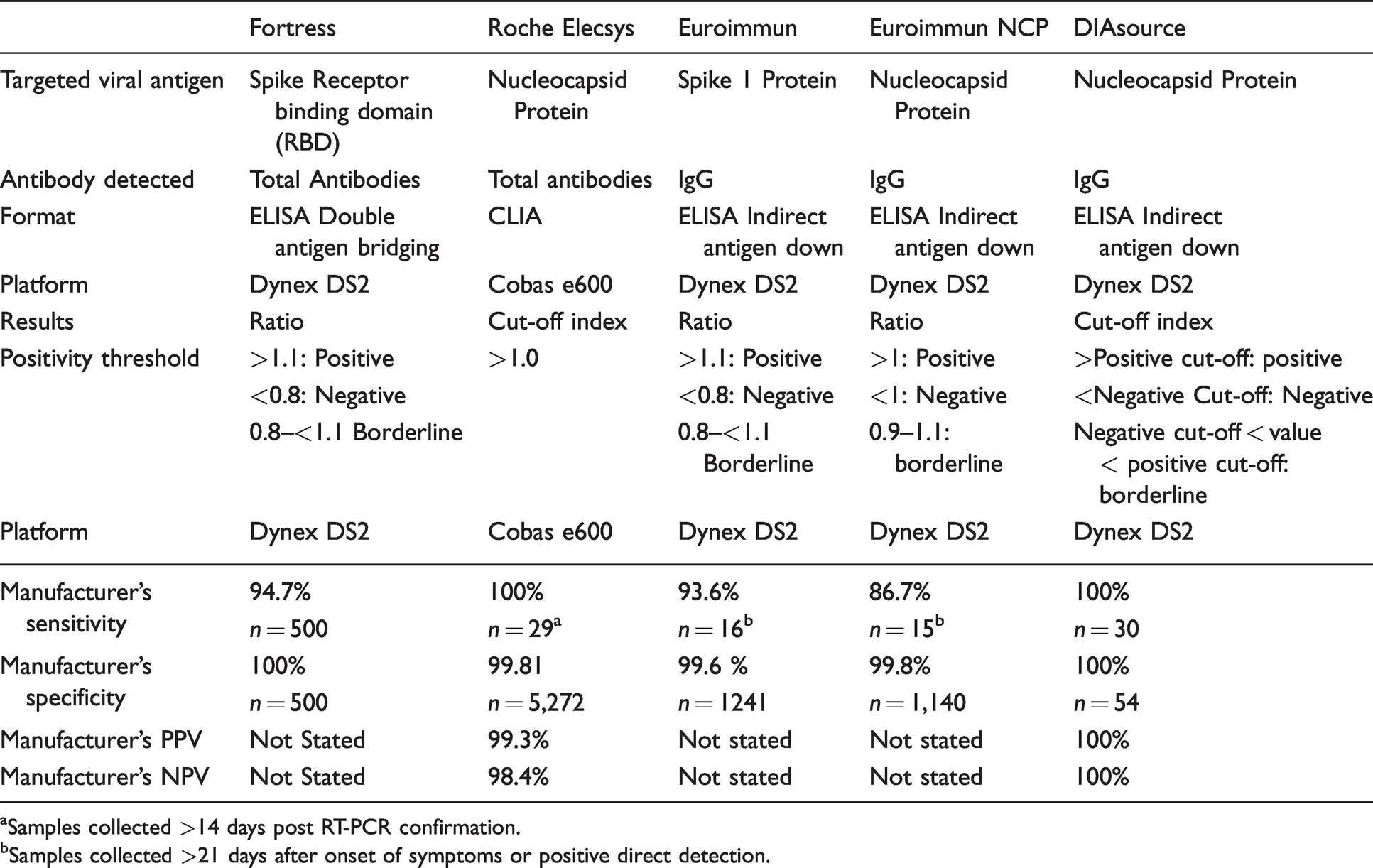
aSamples collected >14 days post RT-PCR confirmation.
bSamples collected >21 days after onset of symptoms or positive direct detection.
Samples were assayed by all five methods and where more than one sample was available for the same participant, samples were de-duplicated and the last sample obtained was used for analysis.
Analytical performance
Method verification studies were performed for all five assays with within- and between-run imprecision determined using quality control samples provided by the assay manufacturers and pooled antibody positive and negative patient samples. To assess within-run imprecision, a pooled negative patient sample and a pooled positive patient sample near the positive cut-off assay were analysed in 20 replicates. Between-run imprecision for each individual assay method was calculated using the assay kit negative and positive controls where provided. Due to limited reagent availability, between-run imprecision was calculated over a minimum of 10 days and a maximum of 20 days. For the Roche ECLIA assay, between-run imprecision was calculated using the same pooled positive and negative samples that were used for the within-run imprecision, as no controls were provided with the kit.
Statistical analysis
Results of antibody measurements on each assay were evaluated according to the manufacturer’s stated cut-off indices or ratios. Diagnostic sensitivity and specificity were calculated according to the following criteria; all samples collected pre-pandemic (prior to October 2019) were considered ‘true negative’ (n = 100), and all samples with a confirmed positive RT-PCR result were considered ‘true positive’ (n = 137). Optimal test cut-off values were determined based on Receiver operator curves (ROC).
Results
Assay performance
The Roche ECLIA, Fortress ELISA and Euroimmun ELISA assays showed acceptable assay performance for both within- and between-run assay imprecision. When analysing the within-run assay performance, the Euroimmun NCP IgG ELISA assay and the DIAsource IgG ELISA assay failed to yield any positive results when the pooled positive sample was analysed in replicates of 20. Within-run assay precision using a pooled positive sample in replicates of 20 was attempted on three separate assay runs for the DIAsource assay and due to limited reagent availability on one assay run for the Euroimmun NCP assay. Due to the failure of the assays to yield any positive results for this sample, within-run imprecision for a positive control could not be calculated. In addition, the DIAsource kit positive control provided by the manufacturer failed on a significant number of assay runs; thus, between-run imprecision could not be calculated for the positive control on this assay. Limited kit availability prevented any further imprecision studies being carried out on the DIAsource and Euroimmun NCP assays. Larger CV values were observed for the between-run data on the negative control for both Euroimmun assays which was likely due to analyses being performed on different lot numbers because of limited reagent availability. Within-run analytical CVs were 10.6%, 6.4% and 9.6% and between-run CVs were 6.7%, 5.4% and 3.96% for the Fortress Total Ab, Roche Total Ab and Euroimmun IgG, respectively.
Method evaluation
The Fortress ELISA assay was the best performing assay demonstrating 98.5% sensitivity (RT-PCR positive, n = 137). The Roche assay demonstrated a diagnostic sensitivity of 97.1% followed by the Euroimmun spike protein IgG ELISA assay with a sensitivity of 92.7%. The Euroimmun NCP ELISA assay and the DIAsource IgG ELISA assay showed poorer performances with sensitivities of 83.9% and 78.8%, respectively. Analysis of assay specificity was performed on 100 pre-pandemic samples. Apart from the Euroimmun NCP assay, all of the assays showed 100% specificity with no positive antibody results on these samples. The Euroimmun NCP assay demonstrated a specificity of 96% with four of the samples collected prior to October 2019, yielding a positive antibody result.
ROC curve analysis demonstrated area under the curve (AUC) ranging from 0.998 (Fortress) to 0.938 (DIAsource) (Figure 1). We derived method-specific cut-offs where the sensitivity corresponded to the specificity, which ranged from 99% (Fortress) to 88% (DIAsource) all with positive likelihood ratios (Table 3). The derived method decision cut-off index was lower than the manufacturer’s stated cut-off index.
Overall diagnostic performance of five assays and most appropriate positivity value based on ROC analyses.
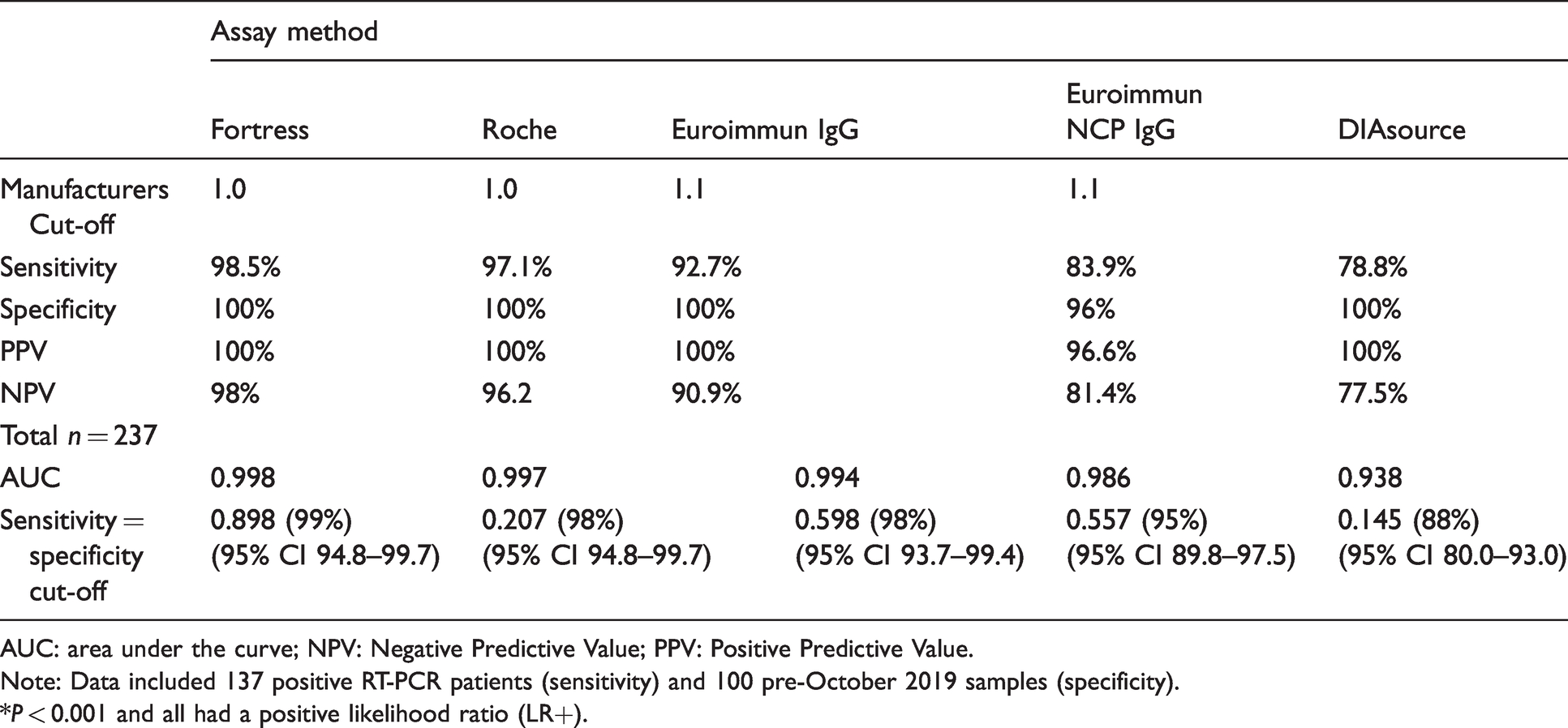
AUC: area under the curve; NPV: Negative Predictive Value; PPV: Positive Predictive Value.
Note: Data included 137 positive RT-PCR patients (sensitivity) and 100 pre-October 2019 samples (specificity).
*P < 0.001 and all had a positive likelihood ratio (LR+).
Study participant results
In the RT-PCR-positive SARS-CoV-2 antibody group (n = 137), two participant samples showed an antibody-positive response only with the Fortress ELISA assay. One of these samples was from a participant in her first trimester of pregnancy. The second sample was from a participant with a very mild illness lasting less than three days where a headache was the only symptom.
The remaining 186 study participants who did not have RT-PCR testing or had negative RT-PCR results were subdivided into those who were symptomatic (n = 77) and those who were asymptomatic (n = 109) (Table 4).
Breakdown of the 186 study participants who were RT-PCR negative or had no RT-PCR testing performed.
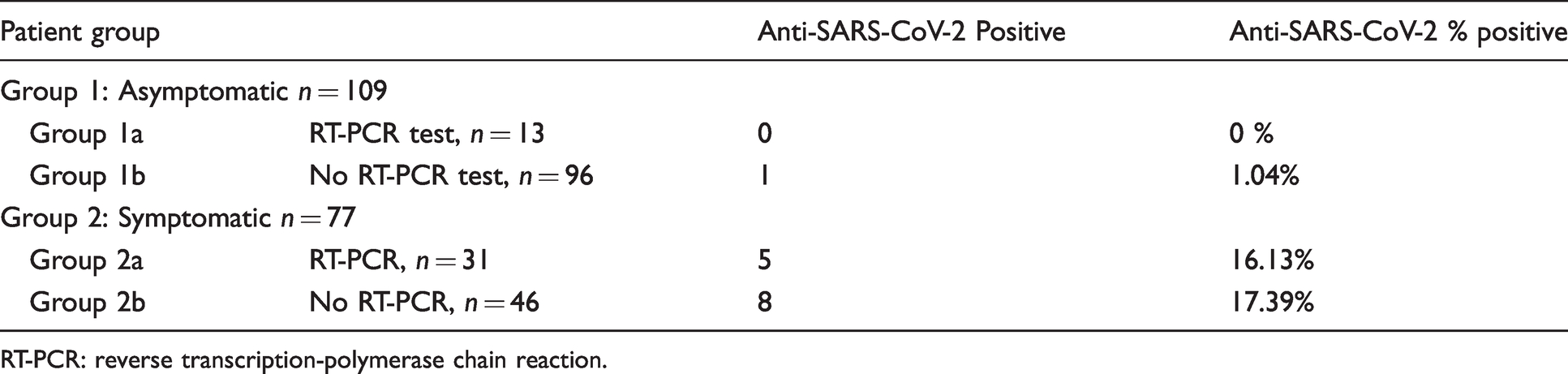
RT-PCR: reverse transcription-polymerase chain reaction.
Seventy-seven participants who were symptomatic had one or more of the documented symptoms. Of these 77 symptomatic participants, 31 had negative RT-PCR tests results and 46 did not have RT-PCR tests performed. Of the 31 symptomatic participants who had negative RT-PCR test results, five showed a positive antibody response to SARS-CoV-2 with four or more of the assay methods which represents 16.13% of this group. It is therefore likely that their RT-PCR were false-negative results. Of the 46 symptomatic participants who did not have RT-PCR tests performed, eight had a positive antibody response to SARS-CoV-2 on all five of the assays used which represents 17.39% of this group. The commonest symptoms reported in this group who were antibody positive were fatigue (66%), myalgia (47%), cough (33%) fever (31%) and headache (31%). Loss of smell and or taste was present in 40% as compared with 3% in those who were antibody negative. Where some of the symptoms are common with many viral illnesses, the latter is now accepted to be significantly associated with COVID-19 infection and its presence is indicative of the need for RT-PCR testing. 22
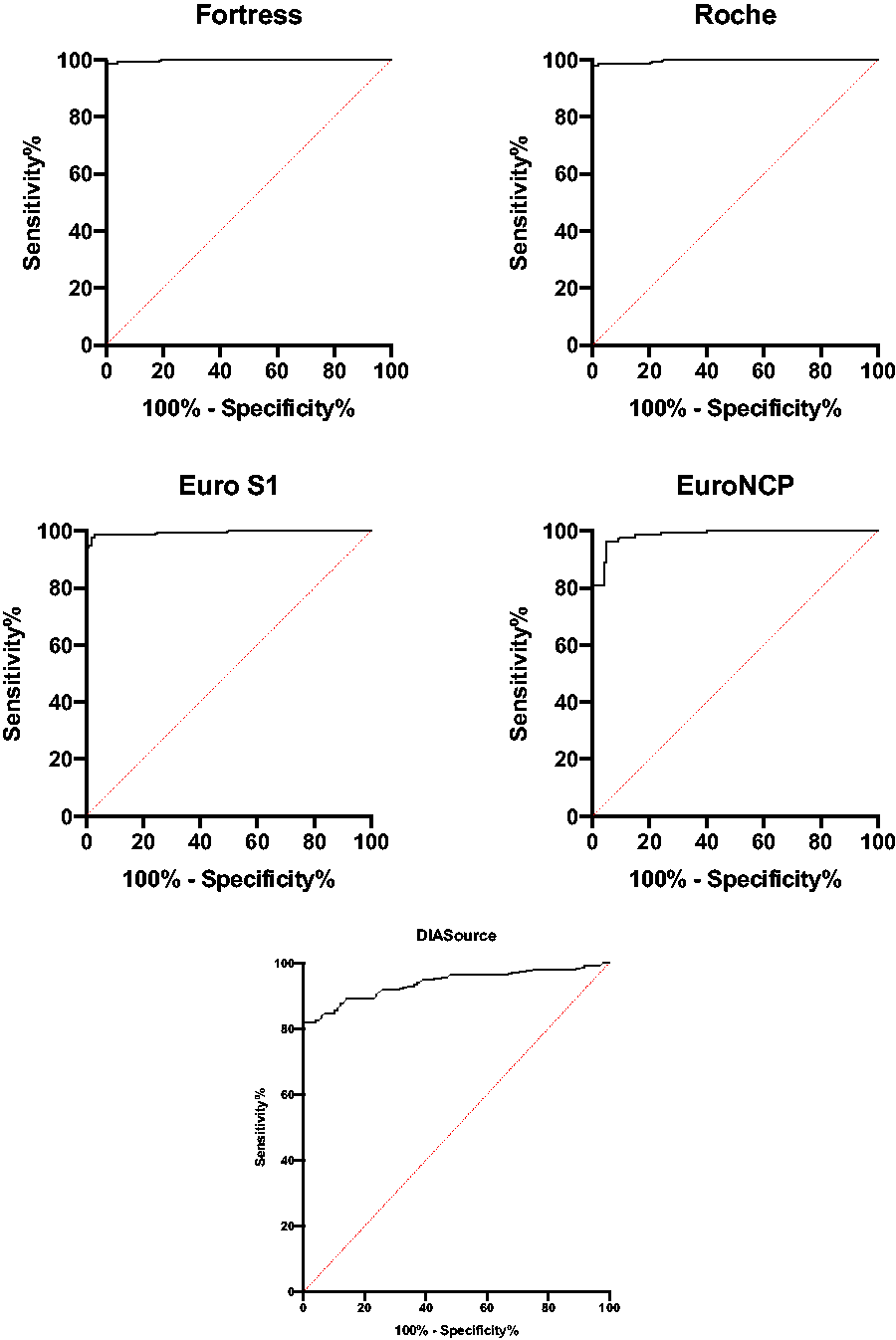
Diagnostic performance of five SARS-CoV-2 antibody assays.
Of the 109 asymptomatic participants, 29 were close contacts of positive confirmed SARS-CoV-2-infected cases. Of these 29 close contacts, 13 had negative RT-PCR test results and 16 had no RT-PCR test performed. 100% of the close contact group tested antibody negative across all assay methods. Of the remaining asymptomatic participants, one sample was SARS-CoV-2-positive across all assay methods.
Discussion
Serological testing for the detection of antibodies against SARS-CoV-2 support a wide number of highly relevant applications in both the clinical management of patients infected with the SARS-CoV-2 virus and in epidemiological studies to evaluate the true rate of global infection. Results of SARS-CoV-2 antibodies are largely qualitative or semi-qualitative in design and no international standard is available for SARS-CoV-2 antibodies. The basis of introducing these tests into the accredited laboratory is that they are reliable, accurate, with good sensitivity and specificity in line with recent IFCC recommendations.19,20
This head-to-head comparison study of five commercially available qualitative SARS-CoV-2 serological assays included evaluation of 423 participants, many with workplace exposure to COVID-19 infection, facilitated the establishment of optimal assay decision cut-offs, which were below the manufacturers cut-offs in all assays.
Our results showed that the Fortress ELISA assay had the highest clinical sensitivity and specificity out of the five assays, demonstrating a sensitivity of 98.5% and a specificity of 100%. This finding correlates with a previously published study which evaluated this assay on a similar cohort of patients and observed a diagnostic sensitivity of 98.0%. 23 Other studies comparing SARS-CoV-2 antibody assays had similar findings, concluding the Fortress ELISA demonstrated the highest level of sensitivity compared with the other methods. 24 The Roche ECLIA assay also performed within levels considered acceptable for laboratory diagnostics, demonstrating a sensitivity of 97.1% and a specificity of 100%. A higher diagnostic sensitivity was observed in the Roche ECLIA assay than has been demonstrated in previous studies analysing samples collected 14 or more days after the onset of symptoms which reported sensitivities of 94%, 91.1%, 89.4% and 89.36%, respectively.23,25–27 When the ROC curve cut-off was applied, the sensitivity of the Roche assay increased to 98.5%. Similar increases in sensitivity were observed in other studies that applied optimized cut-offs when evaluating the Roche assay.25,28 The Euroimmun spike protein assay demonstrated a sensitivity of 94.1% and a specificity of 100%. The sensitivity demonstrated was higher than the sensitivity established in a previous study using a similar cohort of patient (92.86%) 23 but lower than the sensitivity reported in a study analysing samples from hospitalized COVID-19 patients (95.5%). 26 The sensitivity increased to 97.8% when optimized cut-off indices were applied. Another study did not find any increase in sensitivity when applying optimized cut-off indices to the Euroimmun spike protein assay. The Euroimmun NCP IgG and DIAsource IgG assays did not perform at a level that is considered acceptable for laboratory diagnoses in terms of sensitivity (>95%) with diagnostic sensitivities of 83.9% and 78.8%, respectively, but both demonstrated acceptable levels of specificity (>95%). The sensitivities demonstrated for both Euroimmun NCP assay and DIAsource assays were lower than levels observed in the previous studies where the Euroimmun NCP assay had a sensitivity of 88.9% 23 in a similar cohort of patients and the DIAsource assay had a sensitivity of 100%, although this study analysed far fewer confirmed positive RT-PCR samples (n = 26). 29 The sensitivities of both assays increased when the optimized cut-off indices were applied, with the sensitivity of the Euroimmun NCP assay increasing to 95.6% and the DIAsource increasing to 87.6%.
The highest overall sensitivity came from the assays that measured total antibody (IgM–IgG combined) which correlates with other data available on SARS-CoV-2 antibody assay studies.30–32 The three assays that performed the best (Fortress, Roche, Euroimmun IgG) all have different antigens as their target proteins (the Fortress assay targeted the RBD of the spike protein, the Roche assay targeted the NCP and the Euroimmun assay targeted the spike protein); thus, our study suggests that antigen target does not affect assay performance, although previous studies have shown that combining N and S antigens improves sensitivity. 30 The IFCC have suggested that when selecting assays, laboratories should consider employing a combined testing strategy, with assays targeting different antigenic targets and testing the same sample on an alternative platform to confirm positive results. 20
Results from this study contribute to the knowledge base of SARS-CoV-2 antibodies in asymptomatic, mildly and moderately symptomatic individuals, where there has been limited evidence to date. In our study, 5 out of 31 participants who were symptomatic but had a negative RT-PCR result had positive antibody results in four or more assays. A significant finding is the 16.1% and 17.39% of symptomatic participants with detectable antibodies but with negative or no RT-PCR results, respectively. This result is higher than other studies of seroprevalence of SARS-CoV-2 antibodies in health-care workers. A recent review estimates the overall seroprevalence of SARS-CoV-2 antibodies in health-care workers to be 8.7%. 33 The majority of these symptomatic participants did not have an RT-PCR test performed, as their symptoms did not meet the eligibility criteria for testing at the time; therefore, these results show the importance of assessing mild symptoms particularly those linked with COVID-19 such as loss of sense of smell and/or taste. This compared with 1.04% prevalence in those with no reported symptoms. This suggests that a pretest probability study type questionnaire should continue to be part of the diagnostic algorithm for SARS-CoV-2 antibody testing throughout seroprevalence assessment.
Two additional RT-PCR-positive participants, including a second pregnant patient, were antibody negative when tested with all five assays and remained negative even when applying the optimized cut-off indices devised in this study. A possible explanation for the pregnant participant is the repressed immune response after week 12 of pregnancy and the second participant’s negative results are unexplained and she was lost to follow-up. A negative antibody result in a positive RT-PCR patient could reflect a number of situations including failure to mount a measureable antibody response, sampling issues or a false-positive RT-PCR result. Three different antigen targets and detection of different antibody classes were evaluated in the study, so these factors are not a likely explanation for the conflicting results.
Importantly, our study population are primarily permanent health-care staff and their close contacts who will remain an important cohort for ongoing assessment of seroprevalence. There is evidence that antibodies confer immunity for at least six months post-COVID-19 infection 34 and further evidence that antibodies, B cell and T cell memory enhance immunity for longer. 35 There are a number of vaccines currently that have received relevant authorities in many countries and are in the process of being distributed and administered to the public to prevent COVID-19 infection. Evidence from antibody studies suggesting immunity has led to debate as to whether those who have had COVID-19 should in fact receive the vaccine in their post-COVID-19 period. Further research is needed to assess if one vaccine dose would suffice for people who have had previous COVID-19 infection, thus allowing for more targeted speedier vaccine distribution and wider population protection. This enhances the usefulness of SARS-CoV-2 antibody assessment and even longer term follow-up studies will likely define the period of immunity conferred by the presence of antibody and immunological responses. Furthermore, vaccination has resulted in symptoms in many individuals 36 and a suggestion that these are worse in those who have had COVID-19 infection in the recent past but research evidence is needed to support this. A future use of good analytical SARS-CoV-2 antibody assays is in pre-screening and monitoring of individuals receiving these vaccines and comparing the differences with the different vaccines available and between different risk groups. Combining different vaccines has been proposed as an approach to boost vaccine performance and counter anti-vector immunity. 37 A strength of our study is that we evaluated a range of immunoassay targets, which will be particularly important in vaccine assessment as most vaccines elicit responses to spike and not nucleocapsid proteins. 38 In fact, our best performing assay Fortress measures antibodies to the spike protein and will be useful in this setting.
Kontou et al. have suggested that cross-reaction with human endemic coronaviruses could lower the sensitivities of antibody and requires further investigation. 30 A limitation of our study is that due to limited sample availability, we did not evaluate cross-reactivity with other common respiratory viruses which is an important consideration. 39 However, previous studies have demonstrated good analytical specificity when analysing samples taken from patients with seasonal coronavirus infections and cross-reactivity with other circulating coronaviruses appears to be minimal.
Conclusion
In conclusion, there is undoubted clinical need for antibody status studies, and the value of robust analytical assays enhance local, national and international seroprevalence evidence and have a future and ongoing role in assessing vaccine responses. Our study concludes that the Fortress SARS-CoV-2 Total Antibody ELISA assay demonstrated the highest sensitivity and specificity, and therefore the best overall performance followed closely by the Roche ECLIA assay with the advantage of being a fully automated assay. We recommend a two-step testing strategy for screening patient samples using the Roche ECLIA assay and confirming the few discordant samples with the Fortress Total Antibody ELISA assay. This testing strategy conforms to recent IFCC recommendations.
Our study supports the benefit of recording clinical details and symptomatology in interpreting patients’ antibody results.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Ethical approval
The study was approved by the National Research Ethics Committee (NREC) Ireland (COVID study) Ref: 20-NREC-CoV-058.
Guarantor
MCF.
Contributorship
EH and MF wrote the main article. EH, LK, MMH and MF designed the research. MF gained ethical approval. NR, SD and GC provided study population and clinical information. EH and LK performed the analysis. EH, LK and MF analysed data and performed statistical analysis. All authors contributed and approved the final version of the article.
