Abstract
Background
A number of immunoassays have been developed to measure antibodies specific to SARS-CoV-2. More data is required on their comparability, particularly among those with milder infections and in the general practice population. The aim of this study was to compare four high-throughput automated anti-SARS-CoV-2 assays using samples collected from hospitalized patients and healthcare workers with confirmed SARS-CoV-2 infection. In addition, we collected general practice samples to compare antibody results and determine seroprevalence.
Methods
Samples were collected from 57 hospitalized patients and nine healthcare workers at 14 days and at 28 days following confirmed SARS-CoV-2 infection. Samples were also collected from 225 patients presenting to general practice. Four assays were used: Abbott Architect IgG, Beckman Coulter DxI 800 IgG, Roche Cobas e801 total antibody and Siemens Advia Centaur XPT total antibody.
Results
All four assays showed concordance at 14 days in 83.9% of hospitalized patients and in 66.7% of healthcare workers. All four assays showed concordance at 28 days in 88.4% of hospitalized patients and 77.8% of healthcare workers. The sensitivity to detect recent infection was higher for the IgG assays than the total assays. All four assays showed concordance of 95.1% in the general practice population. Seroprevalence ranged from 4.9 to 5.8% depending on the assay used.
Conclusions
All four assays showed excellent comparability, but it may be possible to obtain a negative result for any of the anti-SARS-CoV-2 assays in patients with confirmed previous SARS-CoV-2 infection. An equivocal range would be useful for all anti-SARS-CoV-2 assays.
Introduction
The severe acute respiratory syndrome – coronavirus 2 (SARS-CoV-2) causing COVID-19 (Coronavirus infectious disease 2019) was first reported in China at the end of 2019. Following extensive human-to-human transmission of the virus, it became a major worldwide concern with the World Health Organization (WHO) declaring a pandemic on 11 March 2020. 1 The diagnosis of current SARS-CoV-2 infection is carried out using real-time polymerase chain reaction (RT-PCR) by detecting viral RNA in respiratory samples.
Coronaviruses are enveloped RNA viruses, and the 3′ part of the genome encodes four major structural proteins: spike (S), membrane (M), envelope (E) and nucleocapsid (N). 1 Antibody responses to SARS-CoV-2 have been shown to occur 8 to 10 days following infection with IgG and IgM rising either simultaneously or sequentially.2,3 It is still unclear whether the presence of anti-SARS-CoV-2 antibodies in the serum confers that individual any immunity. 4 However, these assays are useful to determine previous exposure to SARS-CoV-2 infection and are being used as tools for determining seroprevalence in different populations.
The major assay manufacturers have rapidly developed serological assays to detect antibodies to SARS-CoV-2 (targeted to the nucleocapsid or spike protein) using high-throughput immunoassay analysers that are already available in routine clinical laboratories.5,6 As the Roche Elecsys (total) and Abbott (IgG) assays were early to market, a number of studies have been published with these assays, generally using serum samples from patients who have been hospitalized with severe COVID-19.7,8 However, there are limited published data and assay comparison data using the more recently released high-throughput assays (such as the Beckman Coulter IgG assay and the Siemens total antibody assay). It is therefore important to see how these assays compare against one another particularly, as they are being used routinely to determine previous exposure to SARS-CoV-2. In the UK, large-scale sero-surveillance studies using samples collected from blood donors are being performed by Public Health England using the EUROIMMUN anti-SARS-CoV-2 assay. Recent data from the North West England region (incorporating our Lancashire and South Cumbria network) show that the adjusted seroprevalence has decreased to 6.8% in August 2020 (from above 10% in May 2020). 9 There has been some recent debate about the suitability of the serological assays to detect mild infection with a suggestion that assays that target the spike protein may be most suitable for this purpose. 10
As in the rest of the UK, anti-SARS-CoV-2 analysis was introduced in our region in Summer 2020. This test has been available for both healthcare workers and also for patients attending general practice to determine previous exposure to SARS-CoV-2. Within Lancashire and South Cumbria, we have four different high-throughput immunoassay analysers within our Biochemistry and Virology departments (Abbott Architect, Beckman Coulter DxI 800, Roche Cobas e801 and the Siemens Advia Centaur XPT). Anti-SARS-CoV-2 assays are available on all the analysers and we have three of these assays in routine clinical use. There has been no published comparison data of the anti-SARS-CoV-2 assays on these four high-throughput immunoassay analysers.
We wanted to directly compare the performance of the assays to understand the impact of using different assays within our network in individuals with confirmed previous SARS-CoV-2 and within the general practice population. The first aim was to directly compare the assays in hospitalized patients and healthcare workers who had tested positive for SARS-CoV-2 by RT-PCR in serum samples collected at 14 days and at 28 days later. The second aim was to determine how the assays compared by randomly selecting a subset of serum samples received from general practice and determining the seroprevalence.
Materials and methods
Ethics
The Research and Development Departments at Blackpool Teaching Hospitals NHS Foundation Trust, East Lancashire Hospitals NHS Trust, Lancashire Teaching Hospitals NHS Foundation Trust and University Hospitals of Morecambe Bay NHS Foundation Trust reviewed the study. They all agreed that this was a method evaluation study and formal ethical approval was not required. Residual samples were used from hospitalized patients and those attending their local general practice; these samples were collected as part of their routine care and were anonymized prior to inclusion in the study, so consent was not required. For the healthcare workers, they volunteered to participate and informed consent was provided prior to sample collection.
Patients and samples
Serum samples that were collected routinely from patients with severe COVID-19 (as defined by the requirement for hospitalization) for their standard care on or near to 14 days and 28 days after the positive SARS-CoV-2 RT-PCR result (confirmed by a number of RT-PCR methods depending on reagent availability, the analytical availability within the region and the pathway entry of the individual) were identified at three hospital sites (Royal Lancaster Infirmary, Furness General Hospital and Royal Blackburn Hospital). All patient identifiable data were removed prior to inclusion. The number of days after the positive RT-PCR result was recorded, along with the patient’s sex and age. Serum samples were available from 31 patients at day 14 and from 26 patients at day 28.
In addition, nine healthcare workers who had tested positive for SARS-CoV-2 RNA by RT-PCR volunteered to assist with the method evaluation at the University Hospitals of Morecambe Bay NHS Foundation Trust. All provided consent to participate and paired samples were collected on or near to day 14 and day 28 after the positive RT-PCR result. None of the healthcare workers were hospitalized with COVID-19.
In order to further examine the differences in results and seroconversion rates using the four different assays, 225 residual serum samples routinely received in the Pathology Departments at Royal Preston Hospital (n = 75), Blackpool Victoria Hospital (n = 54), Royal Blackburn Hospital (n = 50), Royal Lancaster Hospital (n = 24) and Furness General Hospital (n = 22) were identified during August 2020. These samples were from patients who attended their local general practice and required a serum sample to be collected as part of their routine care. All patient identifiable information was removed from the samples prior to inclusion; only age and sex were recorded and we did not know if these patients had previously had a SARS-CoV-2 RNA test. All serum samples were aliquoted into four aliquots for analysis using the four different assays.
Anti-SARS-CoV-2 analysis
All serum aliquots were sent to the Royal Preston Hospital and distributed from there to the other laboratories for further analysis. Anti-SARS-CoV-2 analysis was carried out using the Abbott Architect (Abbott Diagnostics, Maidenhead, UK) at the Blackpool Victoria Hospital, the Beckman Coulter DxI 800 (Beckman Coulter, High Wycombe, UK) at the Royal Lancaster Infirmary, the Roche Cobas e801 (Roche Diagnostics Ltd., Burgess Hill, UK) at the Royal Preston Hospital and the Siemens Advia Centaur XPT (Siemens Healthcare Diagnostics, Frimley, UK) at the Royal Blackburn Hospital. All assays were performed according to manufacturer’s instructions; the assay characteristics and cut-off index recommended for the interpretation of a positive result can be seen in Table 1. The Beckman Coulter assay also has an equivocal range which is shown in Table 1. The reportable range for the Siemens assay is 0.05–10.0; for data analysis, any results <0.05 were converted to 0.05 and any results >10.0 were converted to 10.0.
The antibody detected, the target antigen and the cut-off index for the four anti-SARS-CoV-2 immunoassays.
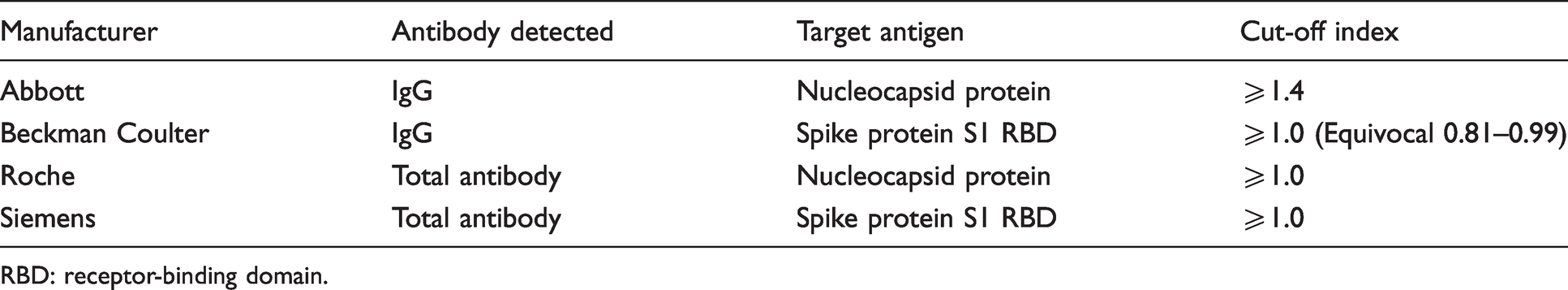
RBD: receptor-binding domain.
Results
Comparison of anti-SARS-CoV-2 assays in hospitalized patients with confirmed SARS-CoV-2 infection
Serum samples were obtained from 31 hospitalized patients who had tested positive for SARS-CoV-2 RNA by RT-PCR 14 days earlier (median = 14 days, range 12–18 days). The age of the patients ranged from 35 to 95 years with a median age of 81 years; 19 patients were male. Figure 1 shows the results using all four assays; 26/31 (83.9%) samples showed concordance across all four assays with 24 positive results and two negative results. The discordant results are also shown in Table 2. For two samples, the result was positive using one assay only (Abbott and Siemens). The other three discordant samples were positive using three of the assays and negative using the Roche assay or the Siemens assay or the Abbott assay in one sample. The sensitivities of the assays for samples collected at 14 days after the positive RT-PCR result were 87.1%, 87.1%, 83.9% and 83.9% for the Abbott, Beckman Coulter, Roche and Siemens anti-SARS CoV-2 assays, respectively.
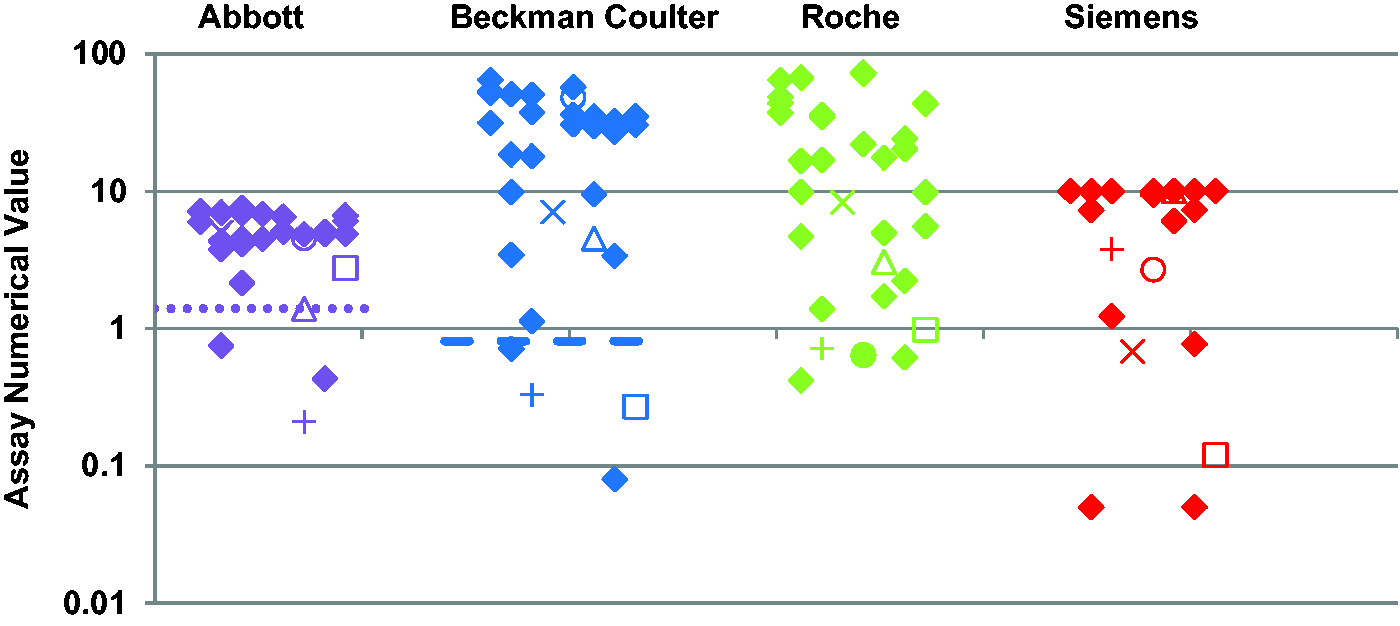
Anti-SARS-CoV-2 results using the Abbott Architect IgG, the Beckman Coulter IgG, the Roche total and the Siemens Centaur total assays (left to right) in hospitalized patients at 14 days after a confirmed positive SARS-CoV-2 RNA result. Filled diamonds show results where there was concordance between all four assays. The other symbols show the results from serum samples that were not concordant. The positive cut-off for the Beckman Coulter, Roche and Siemens assays is 1.0 (x-axis). The positive cut-off for the Abbott assay is 1.4 (shown by the dotted lines). The Beckman Coulter assay has an equivocal range between 0.81 and 0.99 (shown by a dashed line at 0.81).
The results using all assays are shown for all samples where the results were discordant.
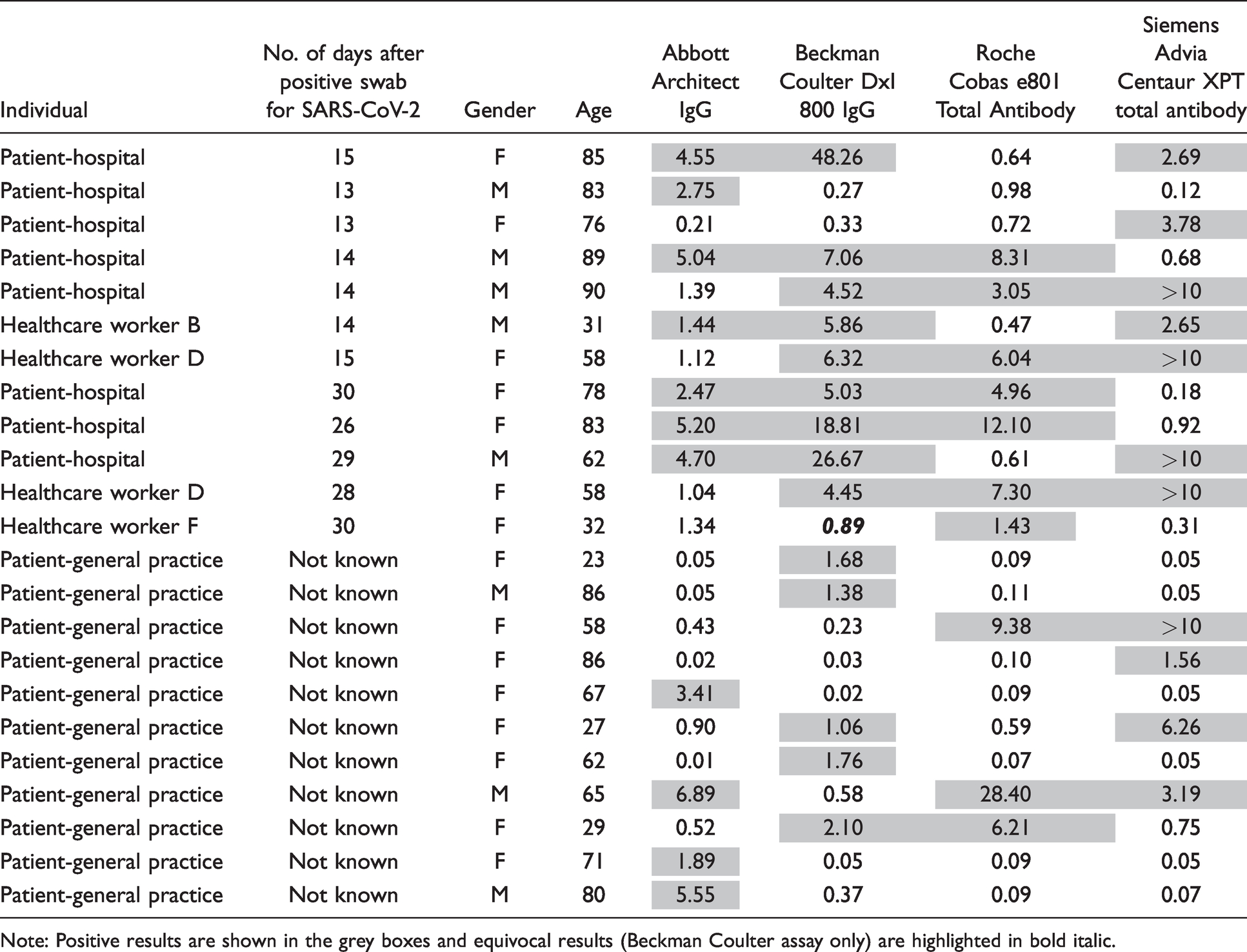
Note: Positive results are shown in the grey boxes and equivocal results (Beckman Coulter assay only) are highlighted in bold italic.
Serum samples were obtained from 26 hospitalized patients who had tested positive for SARS-CoV-2 RNA by RT-PCR 28 days earlier (median = 28 days, range 26–35 days). The age of the patients ranged from 42 to 88 years with a median age of 76 years; 18 patients were male. Figure 2 shows the results using all four assays; 23/26 (88.4%) samples showed concordance across all four assays with 23 positive results. The discordant results are also shown in Table 2; all three were positive using all assays except Roche for one sample and Siemens for two samples. The sensitivities of the assays for samples collected at 28 days after the positive RT-PCR result were 100.0%, 100.0%, 96.2% and 92.3% for the Abbott, Beckman Coulter, Roche and Siemens anti-SARS CoV-2 assays, respectively.
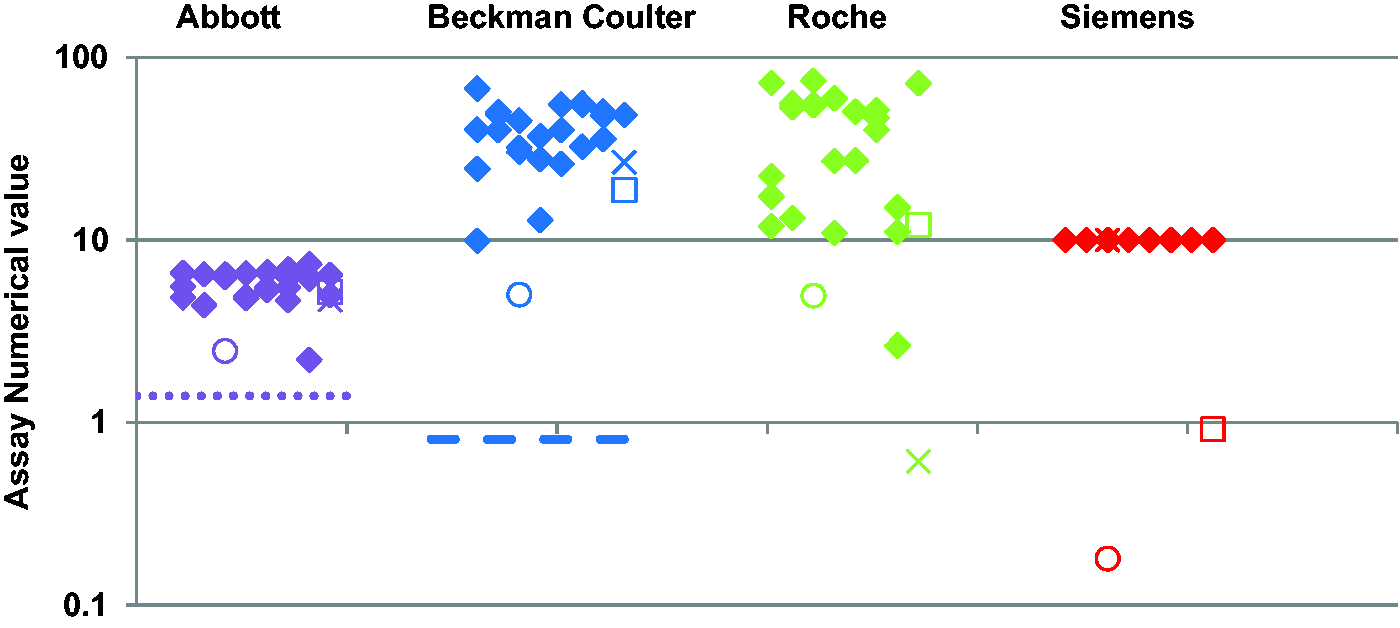
Anti-SARS-CoV-2 results using the Abbott Architect IgG, the Beckman Coulter IgG, the Roche total and the Siemens Centaur total assays (left to right) in hospitalized patients at 28 days after a confirmed positive SARS-CoV-2 RNA result. Filled diamonds show results where there was concordance between all four assays. The other symbols show the results from serum samples that were not concordant. The positive cut-off for the Beckman Coulter, Roche and Siemens assays is 1.0 (x-axis). The positive cut-off for the Abbott assay is 1.4 (shown by the dotted lines). The Beckman Coulter assay has an equivocal range between 0.81 and 0.99 (shown by a dashed line at 0.81).
Comparison of anti-SARS-CoV-2 assays in healthcare workers with confirmed SARS-CoV-2 infection
Nine healthcare workers provided serum samples at 14–20 days (median = 14 days) and at 28–33 days (median = 29 days) after testing positive for SARS-CoV-2 RNA by RT-PCR. The age of the healthcare workers ranged from 31 to 58 years with a median age of 49 years; eight were female. None were hospitalized with COVID-19. Figure 3(a) and (b) shows the results using all four assays at 14 days and 29 days, respectively. Five healthcare workers showed positive or equivocal results using all four assays on both serum samples at 14 days and 28 days. One healthcare worker was negative using all four assays on both samples (Healthcare worker C in Figure 3). The concordance between the four assays for either a negative result or a positive/equivocal result was 66.7% and 77.8% for serum samples collected a median of 14 days and 29 days after a positive SARS-CoV-2 RNA result, respectively. The discordant results are shown in Table 2.
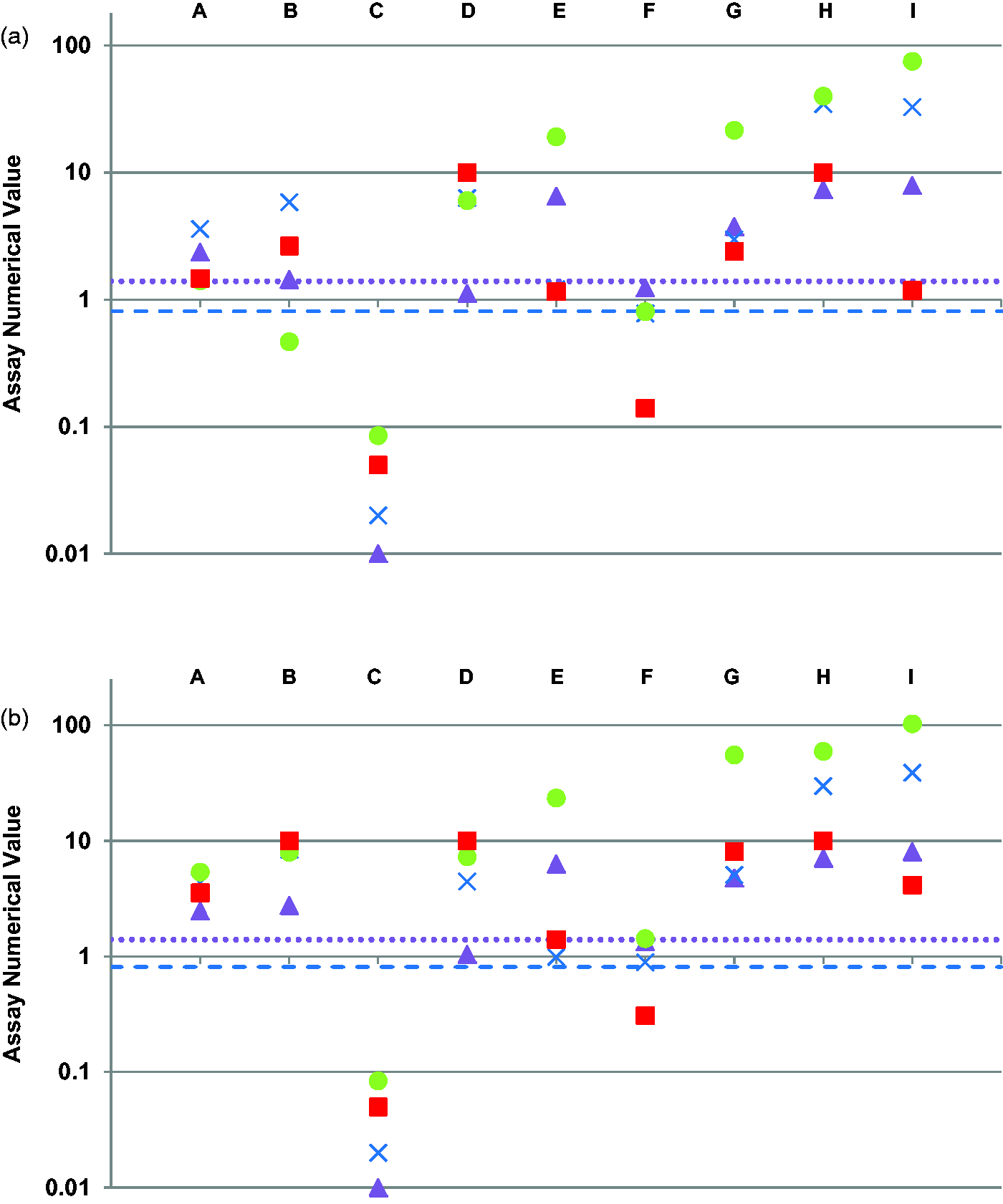
SARS-CoV-2 antibody results using the Abbott Architect IgG (triangle), the Beckman Coulter IgG (cross), the Roche total (circle) and the Siemens Centaur total (square) assays in nine healthcare workers (A-I) at (a) 14 days after a confirmed positive SARS-CoV-2 RNA result and (b) 29 days after a confirmed positive SARS-CoV-2 RNA result. The positive cut-off for the Beckman Coulter, Roche and Siemens assays is 1.0 (x-axis). The positive cut-off for the Abbott assay is 1.4 (shown by the dotted lines). The Beckman Coulter assay has an equivocal range between 0.81 and 0.99 (shown by a dashed line at 0.81).
Comparison of anti-SARS-CoV-2 assays in general practice samples and seroprevalence
There were a total of 225 serum samples collected from patients who had attended general practice. All were anonymized and previous exposure to SARS-CoV-2 was not known. The age of the patients ranged from 13 to 94 years with a median age of 62 years; 93 were male (41%). Using all four assays, the results were either consistently negative or consistently positive for 206 samples and eight samples, respectively (95.1% concordance). The results are shown in Figure 4(a)). For 11 samples, there were differing results between the four assays (see Figure 4(b)) and Table 2. Overall, the seroprevalence using the four assays in the general practice samples was 5.3%, 5.8%, 4.9% and 5.3% using the Abbott, Beckman Coulter, Roche and Siemens assays, respectively.
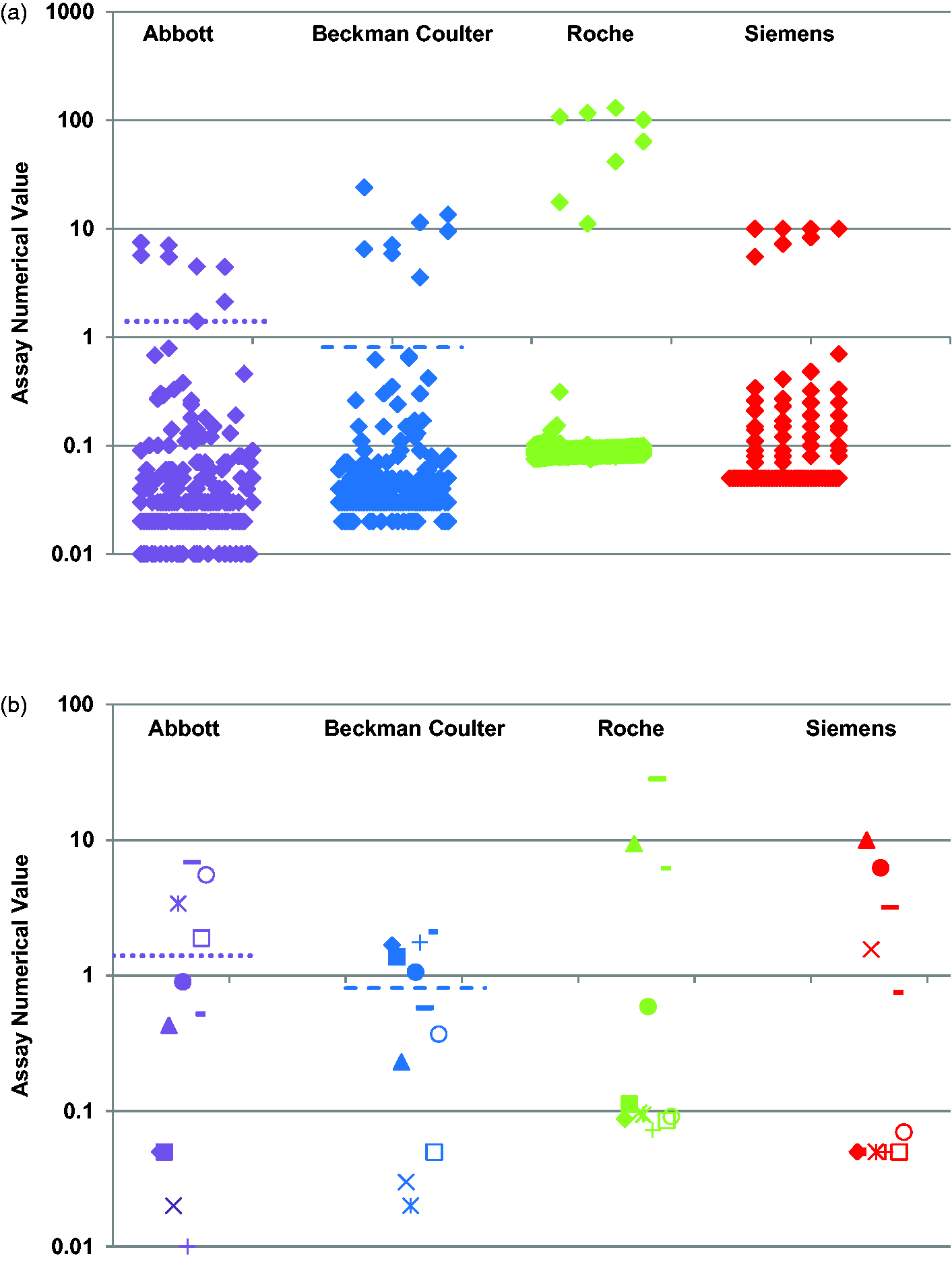
Anti-SARS-CoV-2 results using the Abbott Architect IgG, the Beckman Coulter IgG, the Roche total and the Siemens Centaur total assays (left to right) in general practice samples (n = 225). (a) The results that were concordant between the four assays (n = 215) and (b) the discordant results between the assays (n = 11). The positive cut-off for the Beckman Coulter, Roche and Siemens assays is 1.0 (x-axis). The positive cut-off for the Abbott assay is 1.4 (shown by the dotted lines). The Beckman Coulter assay has an equivocal range between 0.81 and 0.99 (shown by a dashed line at 0.81).
Discussion
This is the first study to compare all four high-throughput anti-SARS-CoV-2 immunoassays (including the Beckman Coulter assay) in both hospitalized patients and healthcare workers with confirmed SARS-CoV-2. In addition, we have determined how these assays differ when determining seroprevalence within the general practice population.
The four high-throughput anti-SARS-CoV-2 assays generally compared very well and showed good concordance, in particular for the patients hospitalized with COVID-19. However, the comparison indicated that it may be possible to obtain a negative result for any of the anti-SARS-CoV-2 assays in patients who have had a previous positive SARS-CoV-2 RNA result on RT-PCR. At 28 days, there was a higher proportion of positive results using all four assays in the hospitalized patients compared with the healthcare workers (88% vs. 67%), suggesting that these assays may not be optimized to detect antibodies in those with milder SARS-CoV-2 infection. Similar findings have been reported from other studies.11 In this study, we have compared the assays at a time period after diagnostic confirmation of SARS-CoV infection by RT-PCR rather than post onset of COVID-19 symptoms. The preincubation period after infection is now established at approximately five days.12 In our hospitalized patient group, there may have been a number of days between symptom onset and the confirmation of SARS-CoV-2 by RT-PCR on hospital admission. In our healthcare worker group, symptom onset and test date are likely to be similar. However, it is a limitation of our study that we are only able to characterize the antibody response post diagnostic confirmation by RT-PCR rather than post symptom onset.
In one of the healthcare workers, there was no antibody response using any of the assays at either 14 days or 28 days. It is possible that their positive SARS-CoV-2 RNA result was a false-positive result. There is no gold standard assay for determining previous viral exposure. It is possible for both false-positive results to occur (e.g. due to contamination) or false-negative results to occur (e.g. due to inadequate swabbing or length of time from infection).
In terms of the sensitivity of the assays to detect antibodies following SARS-CoV-2 infection, there were slightly increased sensitivities using both the IgG assays (Abbott and Beckman Coulter) at both 14 days and 28 days in the hospitalized patients. Interestingly, these assays have different antigen targets with the Abbott assay targeting the nucleocapsid and the Beckman Coulter assay targeting the receptor-binding domain of the spike protein. This may be due to the IgG assays being more suitable for a predominantly IgG response which may be expected at 28 days following SARS-CoV-2 exposure, compared with the total assays which measure both IgG and IgM. 4
A limitation of our study is that our numbers for calculating sensitivity are low; however, they clearly show differences between the different assays. In particular, our healthcare worker group contained only nine individuals. We have included the results from this small group, as they suggest a lower proportion of positive antibody results in non-hospitalized individuals with confirmed SARS-CoV-2 infection. The lower concordance between the four assays in this group compared with the hospitalized patients suggests that the choice of assay and the testing strategy may be more important for the general population. However, further investigation of a wider group of individuals with confirmed SARS-CoV-2 infection who were not admitted to hospital is required to confirm this.
There is no clear pattern in the discordant results based on whether the assay is an IgG/total antibody assay or the antigen target. This indicates that there may be individual-specific differences in the type of antibody response and the timing of the antibody response to SARS-CoV-2 infection. This was particularly evident in our healthcare worker group where the serum samples were collected from the same individuals at different time points. Similar findings have been concluded using different assays. 7 Given that these are immunoassays, we also know that there are other factors that may affect the results of specific assays such as medications, the presence of other analytes in the serum and heterophilic antibodies. Although rare, these factors should always be considered when there is an unexpected result using an immunoassay. The recommended practice is to confirm the results using an alternative immunoassay method.13
For all groups, the results show that the Roche anti-SARS-CoV-2 total antibody assay demonstrates the clearest distinction between a ‘positive’ result and a ‘negative’ result; for example, in Figure 4, the majority of results using the Roche assay are clustered around 0.1 or >10 (compared with a cut-off of 1.0). Only the Beckman Coulter assay has an equivocal range; interestingly, some of the discordant results were due to an assay having a result very close to but not above the cut-off for that assay. For example, one hospitalized patient had a result of 0.92 (<1.0) using the Siemens assay when all other assays showed results above the respective cut-off. Similarly, a hospitalized patient had a result of 1.34 (<1.4) using the Abbott assay when all other assays were positive. In both cases, the result would be reported as ‘negative’ using one of the assays; however, if a repeat sample had been collected a few days later, it is possible that the result would have been above the cut-off. For a population study or for a person wanting to know if they have had previous exposure to SARS-CoV-2, a result close to the cut-off may be more correctly interpreted as ‘equivocal’ with a repeat sample to be collected at a later time. Alternatively, a second anti-SARS-CoV-2 assay could be used on these borderline results. Further work is needed to determine the most appropriate testing strategy and a suitable equivocal range for the Abbott, Roche and Siemens assays.
In our general practice population, the seroprevalence using the four different assays ranged from 4.9 to 5.8%. This is lower than the 6.8% reported in the sero-surveillance studies for our region by Public Health England using blood donor samples (and a different assay) at a similar time. 7 This may be due to the small sample size or the preselection of our population of patients attending general practice. It is possible that some of these individuals may have been shielding at the height of the pandemic due to other underlying health conditions or it is possible that some of these individuals only had mild infection and the assays were not able to measure the antibody response. 10 The only way to get a true snapshot of seroprevalence would be to measure antibodies in a completely unselected population.
Despite good overall conformance between the four high-throughput assays studied here, our findings do indicate that the choice of assay will impact on the results of sero-surveillance studies and re-evaluation by the manufacturers may be required for those results close to the cut-off. Additionally, in patients who present with a clear documented history of SARS-CoV-2 infection but get a negative result, analysis using a different assay may be required. Furthermore, with the roll out of SARS-CoV-2 vaccinations, the use of the assays will be changing, so a clear understanding of their application and consideration of using a second assay for confirmation of equivocal results are required.
Footnotes
Acknowledgements
We would like to thank all the laboratory staff who have been involved with analysing these samples.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Ethical approval
The Research and Development Departments at the four trusts reviewed and approved this method evaluation.
Guarantor
JO.
Contributorship
JO, SH, AB, JE and DS designed the study. JO, SH, AB, JE, BH and DS selected the samples and analysed the samples. AB and JE recruited the healthcare workers that participated in the study. JO and SH analysed the data and wrote the first draft of the article. All reviewed and agreed the final article. JO and SH contributed equally and are joint first authors.
