Abstract
Objectives
To explore microRNA-574-3p expression in serum of patients with hepatocellular carcinoma and investigate correlations between serum microRNA-574-3p expression and the development and prognosis of hepatocellular carcinoma.
Design and methods
Serum samples were collected from 70 patients with primary hepatocellular carcinoma, 40 patients with cirrhosis and 45 healthy controls. Serum microRNA-574-3p expression levels were detected by real-time quantitative polymerase chain reaction. The linearity, specificity and reproducibility were evaluated. In addition, the diagnostic value of microRNA-574-3p and its correlations with clinicopathologic features were assessed.
Results
The relative expression of microRNA-574-3p in hepatocellular carcinoma patients, cirrhosis patients and healthy controls was 2.306 (1.801–3.130), 1.362 (0.994–1.665) and 1.263 (0.765–1.723), respectively, indicating that it was significantly higher in hepatocellular carcinoma patients than that in the other two groups (U = 439.5, 514.5, both P < 0.0001) and was significantly correlated with hepatitis B virus DNA copies (U = 383.0, P = 0.018). In hepatitis B virus-positive hepatocellular carcinoma patients, the relative expression of microRNA-574-3p was significantly correlated with hepatitis B virus DNA concentration (r = 0.348, P = 0.022). Compared with healthy control group, AUCROC of serum microRNA-574-3p in hepatocellular carcinoma group was 0.837 with 95% CI: 0.763–0.910. Combining microRNA-574-3p, AFU and alpha-fetoprotein together, the sensitivity was highest compared with other markers alone or combined.
Conclusions
The relative expression of serum microRNA-574-3p in hepatocellular carcinoma patients was significantly higher than that in cirrhosis patients and healthy controls, and it may be an important biomarker in the auxiliary diagnosis of hepatocellular carcinoma.
Keywords
Introduction
Hepatocellular carcinoma (HCC) is the most common human malignancy in China, which has a poor prognosis and low survival rate. 1 The high-risk factors of HCC are hepatitis B virus (HBV) or hepatitis C virus (HCV) infection, haemochromatosis psychosis, alcohol abuse and so on. 2 About 350 million people were infected with HBV, of which there were about 120 million infected people in China. 3 Currently, the combination of serum alpha-fetoprotein (AFP) ultrasound surveillance and computed tomography is the common strategy for the screening and detection of HCC in the high-risk group. 4 AFP is an unsatisfactory marker due to its low sensitivity and high false-positive rate, which was also influenced by the tumour size and cancer stage. 5 Consequently, there is an urgent need for effective and reliable tools for an early diagnosis to improve the survival rate of patients with HCC.
MicroRNAs (miRNAs) are small non-coding RNA molecules (18–25 nucleotides), which could bind to the 3′-untranslated region (3′-UTR) of target mRNA and silence the expression levels of target genes. 6 Aberrant expression of microRNA (miRNA) has been detected in a variety of human malignancies, suggesting that they serve essential roles in tumourigenesis and tumour development. 7 Previous studies indicated different miRNA expression in HCC pathogenesis. miR-21, miR-122 and miR-223 were upregulated in HCC tissues compared with that in normal tissues, 8 whereas miR-122a, miR-22 and miR-152 were downregulated. 9 In addition, differential expression of some serum miRNAs, such as miRNA-375, miRNA-25, miRNA-21, let-7f and miRNA-16 in patients with chronic hepatitis B as well as HBV-associated HCC, 10 might serve as biomarkers.
Previous studies have reported that miR-574-3p expression was significantly upregulated and may serve as a tumour promoter in human osteosarcoma, 11 gastric cancer, 12 breast cancer 13 and type 2 diabetes. 14 However, the clinical significance of miR-574-3p in HCC patients remains unclear. In the present study, the expression of miR-574-3p in HCC was investigated using real-time quantitative polymerase chain reaction (PCR) (RT-qPCR). Furthermore, the association between the expression of miR-574-3p and the clinicopathological characteristics of patients with HCC was analysed.
Materials and methods
Characteristics of patients
Seventy patients with primary HCC, 40 patients with cirrhosis and 45 healthy controls were admitted to Nantong Tumor Hospital from March 2015 to November 2016. All of the recruited liver cancer patients have been pathologically diagnosed as HCC, including 24 females and 46 males ranging in age from 42 to 72 years with a mean age of 35 years. Cirrhosis patients included 16 females and 24 males ranging in age from 37 to 69 years with a mean age of 48 years. The age of the healthy group ranged from 40 to 68 years with a mean age of 49 years. This study was approved by the local ethics committee, and written informed consent was obtained from all of these participants.
Specimen collection and serum RNA extraction
Blood samples were collected using plastic gel vacuum blood collection tubes and centrifuged at 1000 × g for 10 min. The separated serum was placed in RNase-free centrifuge tubes and stored at −80℃ for use. Total RNA was extracted from 300 μL serum using the serum RNA extraction kit (Life Technologies, US) and stored at −80℃ after confirmation of the concentration and the purity by ultraviolet spectrophotometry.
cDNA synthesis
The extracted RNA was reverse transcribed into cDNA using the reverse transcription kit (Thermo Fisher Scientific, USA), miR-574-3p and U6 stem-loop RT reverse primer (Shanghai Generay Biotech Co., Ltd, Shanghai, China). The reverse transcription system was as follows: RNA 300 ng, 5 × reaction buffer solution 4 μL, RT primer 1 μL, DNA 2 μl, RNA inhibitor 1 μL, RTase 1 μL, and addition of nucleic acid-free pure water to 20 μL. The mix was incubated at 42℃ for 60 min and 70℃ for 5 min. The synthesized cDNA was kept at −20℃ until further use.
Real-time quantitative PCR
Real-time PCR was performed using an ABI 7500 PCR Detection System (ABI, USA). The miR-574-3p primer sequences are as follows: forward primer: 5′-ACACTCCAGCTGGG CACGCTCATGCACACA-3′ and backward primer: 5′-TGGTGTCGTGGAGTCG-3′; U6 primer sequences: forward primer: 5′-CTCGCTTCGGCAGCACA-3′ and backward primer: 5′- AACGCTTCACGAATTTGCGT-3′. Twenty microlitres of PCR reaction mix included 10 μL SYBR Green I mix, 3 μL cDNA, 1 μL forward primer, 1 μL reverse primer and 5 μL RNase-free H2O. Reactions were incubated in optical tubes at 95℃ for 10 min, followed by 40 cycles at 95℃ for 15 s and 60℃ for 31 s. All experiments were done in triplicate and data were calculated using the comparative Ct (2−ΔΔCt) method. U6 was used as an internal control to normalize RNA input in the real-time PCR assay.
Detection of CEA and CA199 concentrations
The concentrations of AFP and AFU in the serum samples from HCC patients were detected in the Laboratory Medicine Center of Nantong Tumor Hospital by MODULAR ANALYTICS E170 (Roche, Germany) and 7600 automated biochemistry analyser (HITACHI, Japan).
Statistical analysis
Statistical analyses were performed using the SPSS 17.0 statistical software (SPSS, Chicago, IL, USA) and GraphPad Prism 5 (GraphPad Software Inc., CA, USA). Data were expressed as medians (lower quartile and upper quartile). Differences of the relative expression level of serum miR-574-3p between the HCC, cirrhosis and healthy groups were calculated using Mann-Whitney U test. Paired comparison between the miR-574-3p concentration and the clinicopathologic features in HCC group was evaluated through Mann-Whitney U test. Kruskal-Wallis H test was used for multiple comparisons between the groups. Correlations between miR-574-3p and HBV-DNA concentrations in HCC patients were analysed by Spearman correlation analysis. Their diagnostic efficacy as a diagnostic marker was assessed by receiver-operating characteristics (ROC) and the area under the curve (AUC) (95% CI). P values of < 0.05 were considered statistically significant.
Results
Linearity
RT-qPCR was performed to detect serial dilutions (1:1, 1:10, 1:100, 1:1000 and 1:10,000) of miR-574-3p cDNA. The R2 of the miR-574-3p standard curve was 0.997, and the regression line was y = −3.40x+22.20 (Figure 1). The amplification efficiency of target gene was 0.97 according to the formula: E = 10−1/slope−1, indicating that RT-qPCR could be used to detect different concentrations of serum miR-574-3p.
The standard curve of serum miR-574-3p.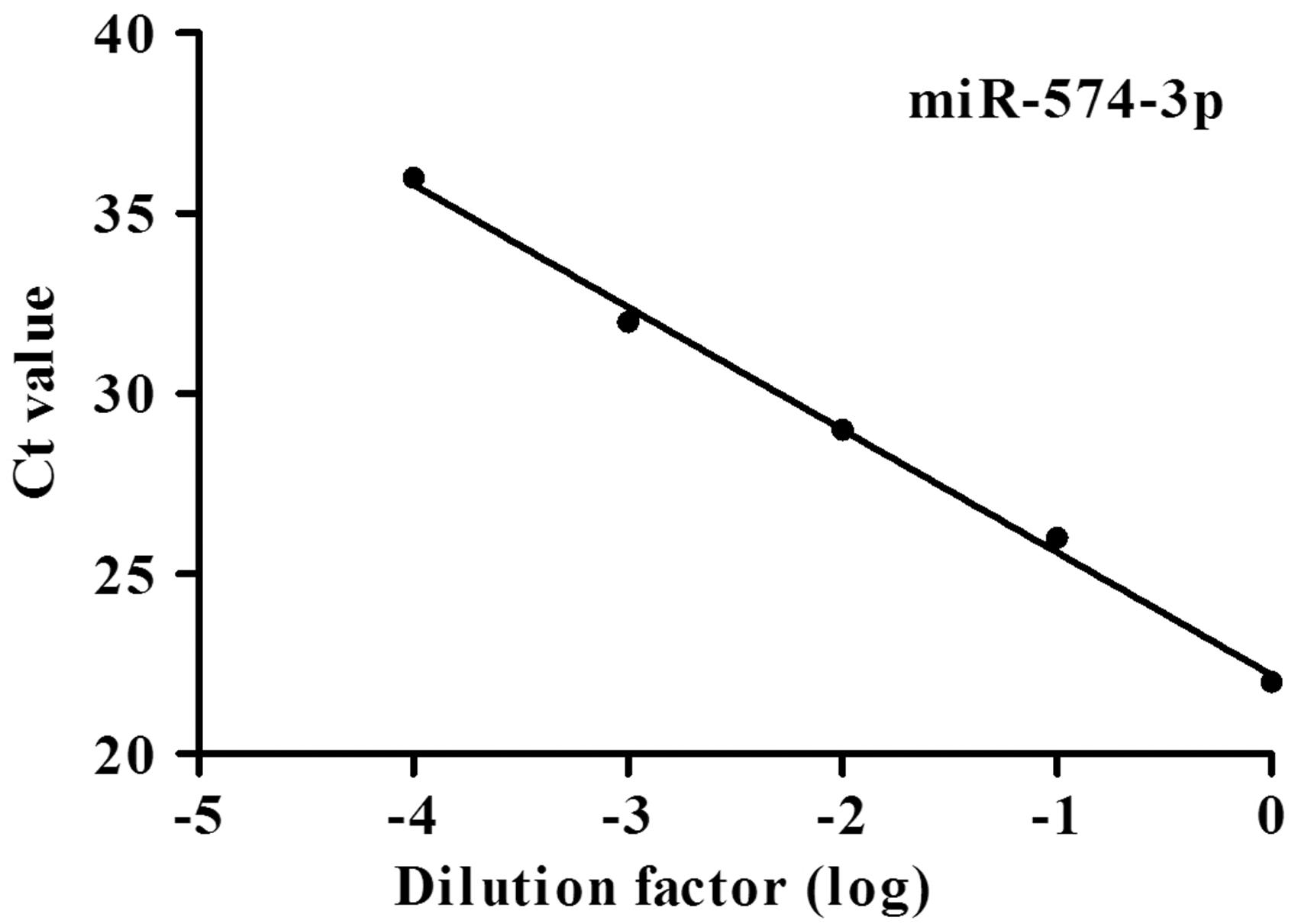
Reproducibility
For the analysis of the reproducibility of RT-qPCR, the intra-assay precision was performed in 10 replicates within one experimental process. The results are given in Table S1 (supplementary files). The intra-assay coefficient of variation (CV) was 0.851%, 0.573% for miR-574-3p and U6, respectively. Inter-assay variation was measured for 10 consecutive days. The inter-assay CV of miR-574-3p and U6 was 2.480% and 3.625%, respectively. These data proved that our study had a good repeatability.
Specificity
As shown in Figure 2(a), RT-qPCR melting curve was having a specific single peak. The melting temperature of miR-574-3p and U6 was about 78℃ and 79℃, respectively. The data aforementioned indicated our study had a high specificity. In addition, amplification curve was standard S-shaped curve, and the curve of every parallel tube was good, suggesting that the amplification efficiency of every parallel tube is similar (Figure 2(b)).
RT-qPCR melting curves and amplification curves of serum miR-574-3p and U6 in HCC patients and healthy controls.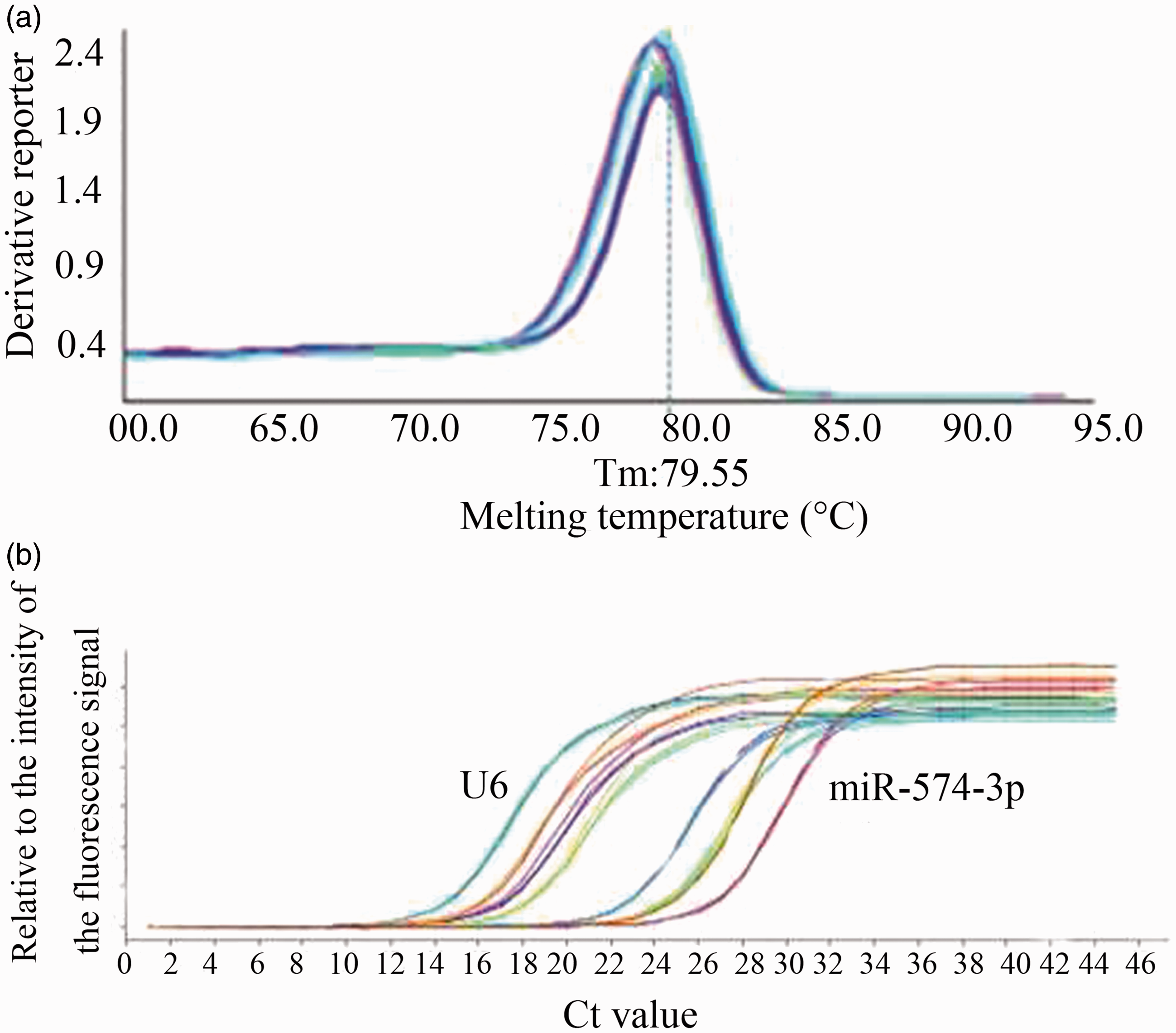
Serum miR-574-3p expression in HCC patients, cirrhosis patients and healthy controls
The expression of miR-574-3p was determined in the serum samples of 70 HCC patients, 40 cirrhosis patients and 45 healthy controls using RT-qPCR and normalized to the control U6. As illustrated in Figure 3, serum miR-574-3p expression in HCC, cirrhosis and healthy control groups was 2.306 (1.801–3.130), 1.362 (0.994–1.665) and 1.263 (0.765–1.723), respectively. It was significantly higher in the HCC group than that in the other two groups (U = 439.5, 514.5, both P < 0.0001), and there was no significant difference between cirrhosis and healthy control groups (U = 800, P = 0.381). These results suggested that serum miR-574-3p could be helpful in the auxiliary diagnosis of HCC.
The relative expression of serum miR-574-3p in HCC, cirrhosis and healthy control groups.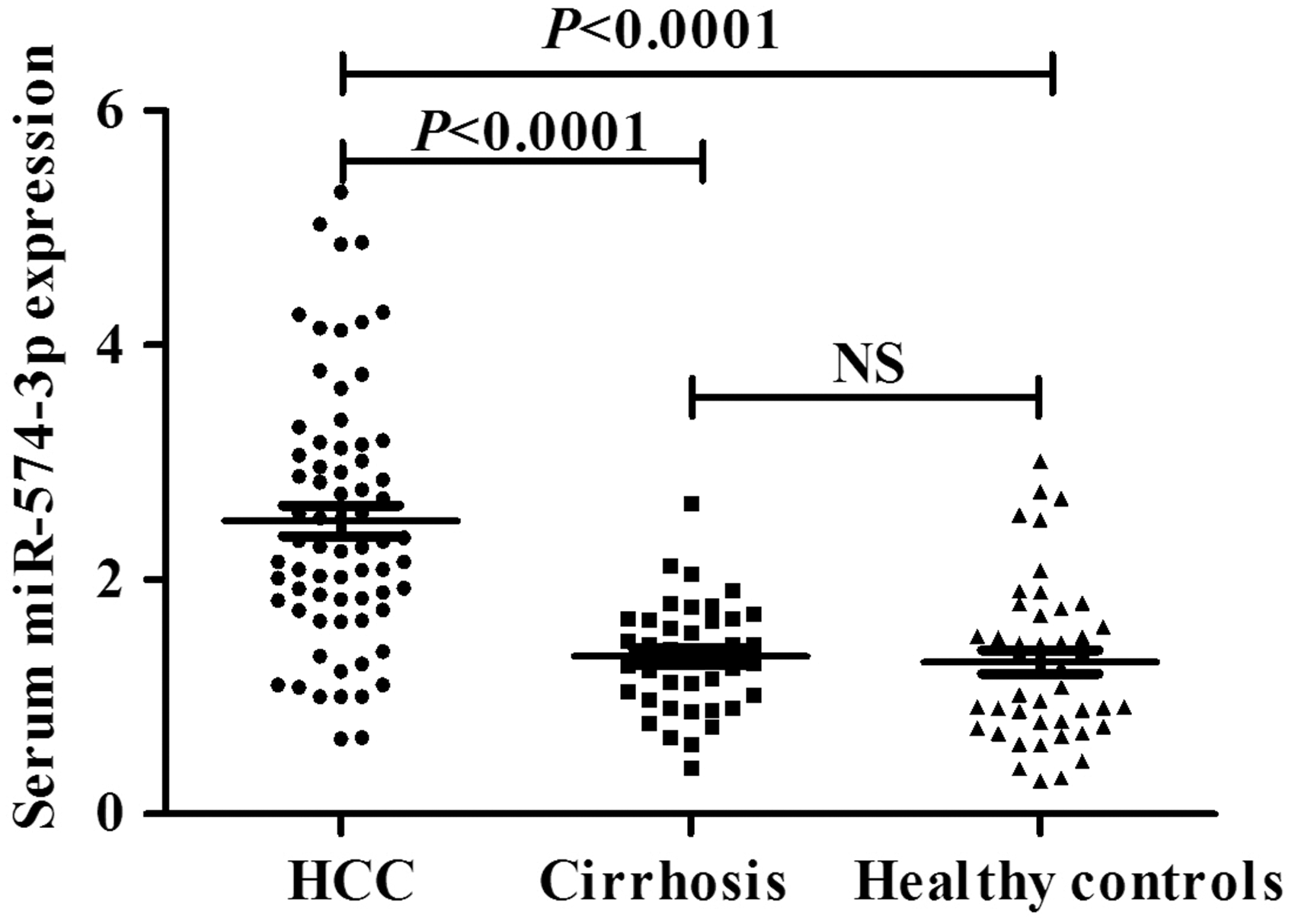
Correlations between miR-574-3p expression levels and the clinicopathologic features in HCC patients
Association between the relative expression of miR-574-3p and the clinicopathological characteristics of patients with hepatocellular carcinoma.
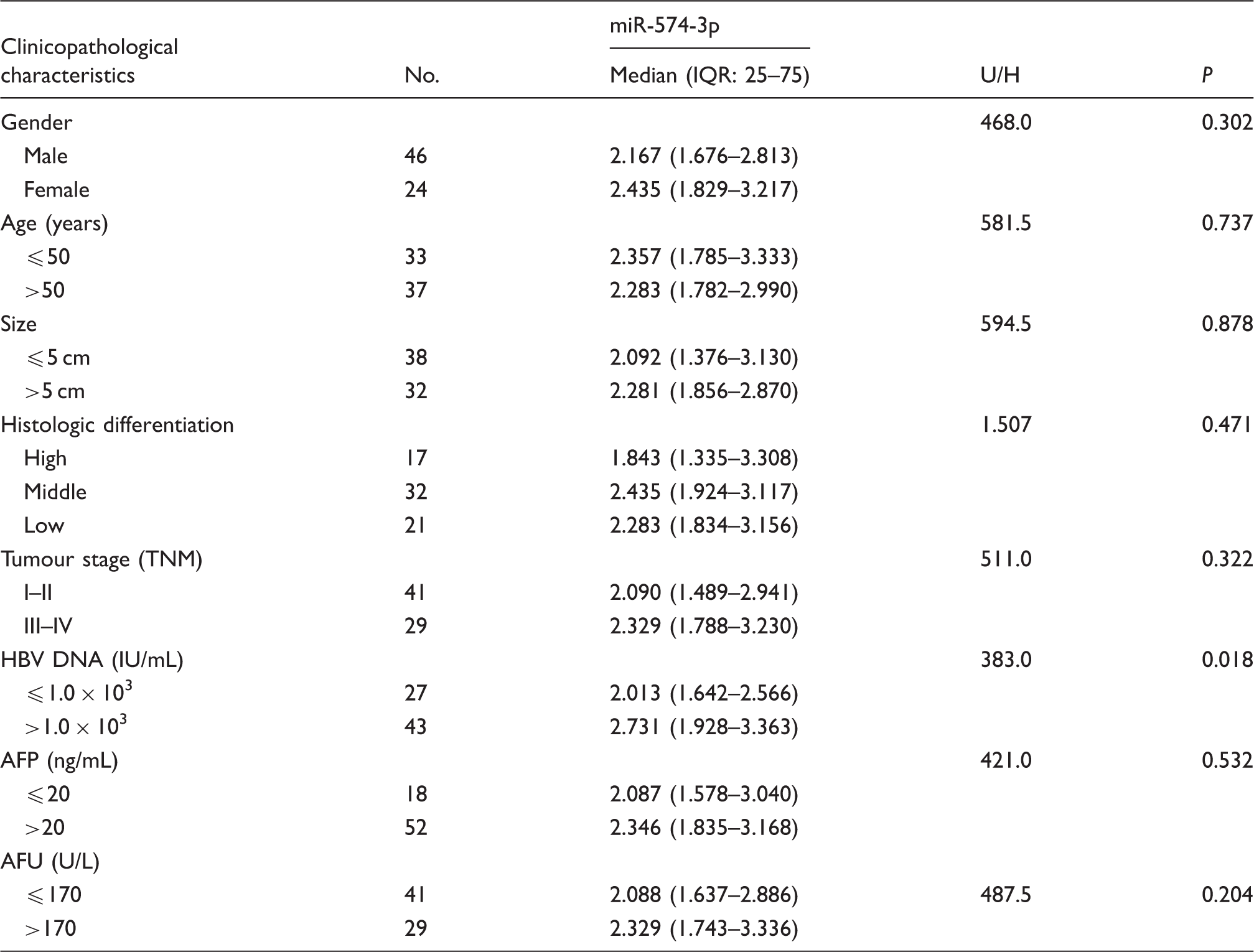
AFP: alpha-fetoprotein; AFU: α-L-fucosidase; HBV: hepatitis B virus.
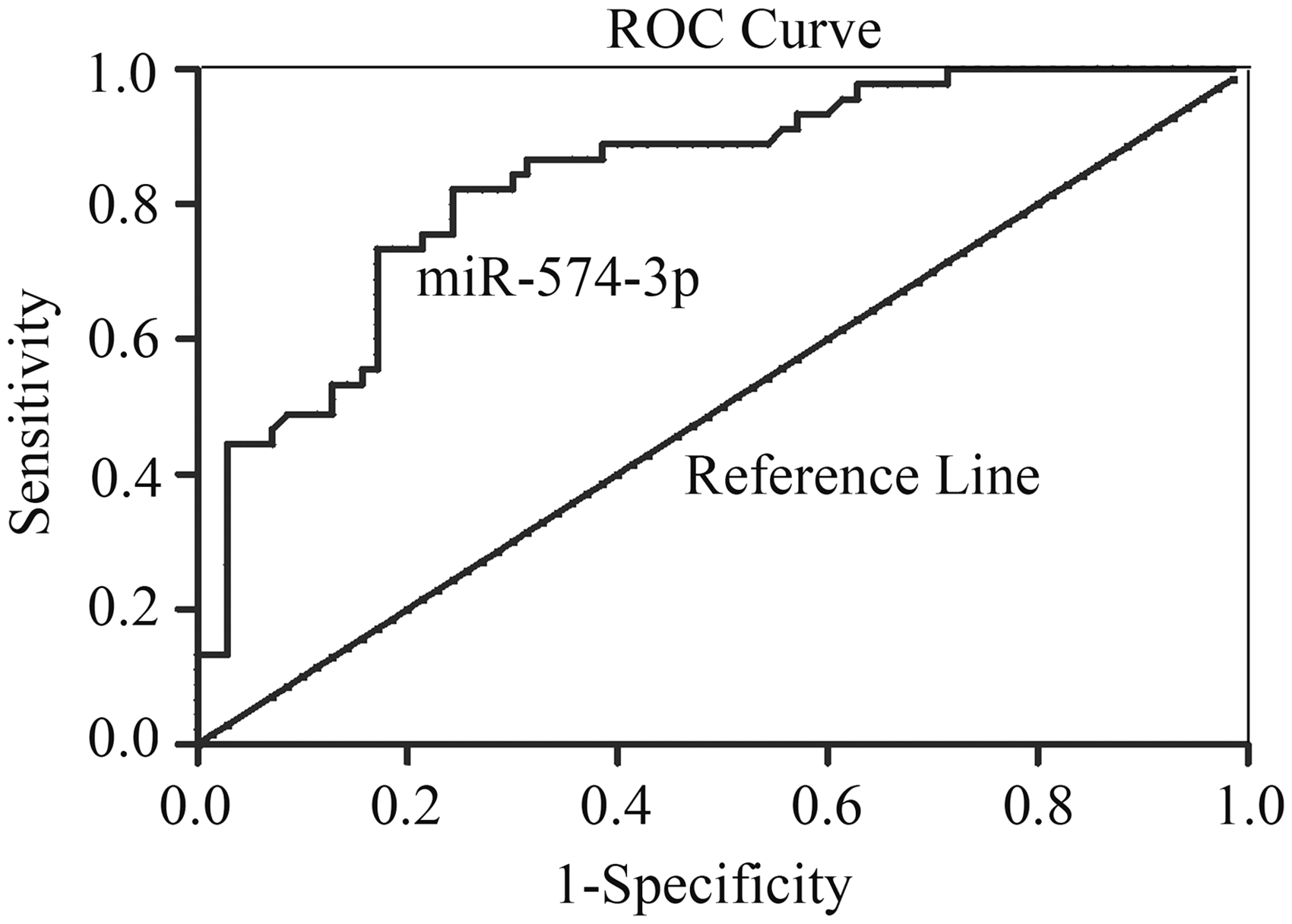
ROC curve of miR-574-3p for differentiating HCC patients from healthy control group.
Correlation between serum and HBV DNA concentrations in HBV-positive HCC patients
In our study, it was found that the relative miR-574-3p expression levels were positively correlated with HBV infection. The results showed that the relative expression level of serum miR-574-3p in HBV-positive HCC patients and HBV-negative HCC patients was 2.731 (1.928–3.363) and 2.013 (1.642–2.566), respectively. It was significantly higher in the HBV-positive group than that in the HBV-negative groups (U = 383.0, P = 0.018) (Table 1). Then, the correlations between the relative expression of serum miR-574-3p and HBV DNA copies in 43 HBV-positive HCC patients were analysed by spearman correlation analysis. It was showed that the relative expression of miR-574-3p was significantly correlated with HBV DNA concentration (r = 0.348, P = 0.022).
Combined detection of serum miR-574-3p, AFP and AFU
Diagnostic efficacy of serum, AFP and HBV DNA in HCC group as compared with control group.
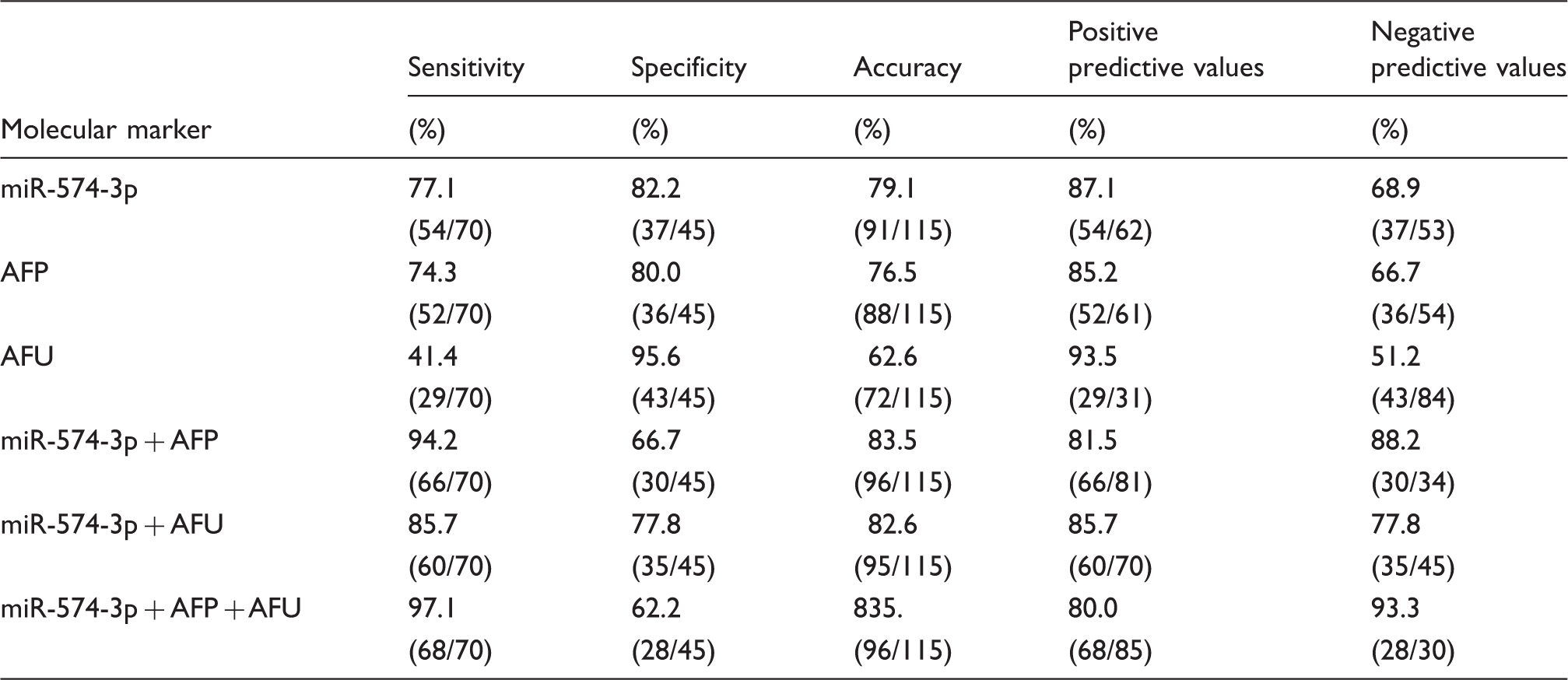
AFP: alpha-fetoprotein; AFU: α-L-fucosidase; HBV: hepatitis B virus; HCC: hepatocellular carcinoma.
Discussion
Primary HCC is the fifth popular cancer worldwide, with high invasiveness and high mortality. 1 It has been reported that the abnormal expression of molecular pathways, such as p53, RAS/MAPK, PI3K/Akt/mTOR, Wnt/β-catenin play significant roles in the pathogenesis of HCC. 15 Despite the remarkable improvements in treatment, the prognosis of HCC is poor. Therefore, it is critical for us to find more effective means or biomarkers that may provide new strategies for the early diagnosis and prognosis in HCC.
Recently, remarkable studies have revealed the vital roles of miRNA. Particularly, abnormal expression of miRNAs affects critical HCC-related pathways. It is reported that microRNA-12216 and microRNA-10117 were downregulated and microRNA-2118 and microRNA-22119 were upregulated in the occurrence of HCC. Circulating miRNAs are extremely stable and protected from RNAase-mediated degradation in body fluids, severing as candidate biomarkers for many diseases. 20 Detection of miRNAs in serum samples or plasma is thought to be a novel non-invasive diagnosis strategy with great potential for clinical application.
In this study, we detected the expression of miR-574-3p in HCC. First, 10-fold serial dilution of miR-574-3p cDNA in the serum of the same patient was detected by RT-qPCR, which indicated a good linearity and the amplification efficiency of target gene. The intra-assay and inter-assay CV of miR-574-3p were 0.851% and 2.480%, respectively, showing the method had a good repeatability. The melting curve was specific single peak, indicating our study had a high specificity. Then, the serum miR-574-3p concentrations of 70 HCC patients, 40 cirrhosis patients and 45 healthy controls were analysed. It was found that the expression of miR-574-3p was significantly higher in the HCC group than that in cirrhosis and healthy control group, suggesting it can clearly distinguish HCC and cirrhosis cases. We also investigated the association of miR-574-3p expression with clinicopathological characteristics and diagnosis value in HCC. Our results showed that serum miR-574-3p expression was positively correlated with HBV DNA concentration, indicating that serum miR-574-3p could be helpful in assessing copy number of HBV. HBV infection plays a crucial role and is a major cause of HCC in China. 21 Li et al. 22 showed that a 13 HBV-specific serum miRNA-based biomarker could clearly separate HBV-positive HCC from healthy controls. These miRNAs may be used potentially as a new tool to inhibit HBV replication and to prevent the development of HCC.
Furthermore, in terms of comparison between HCC and healthy control groups, AUC of miR-574-3p was 0.837 with 95%CI: 0.763–0.910. At the cut-off value of 1.809, the sensitivity of serum miR-574-3p was 77.1% and the specificity was 82.2%, which indicated that serum miR-574-3p had good sensitivity and high specificity. More importantly, our results showed that the combination of miR-574-3p, AFP and AFU provided optimal sensitivity for the detection of HCC. Therefore, the relative expression of serum miR-574-3p may prove to be a useful auxiliary test in the diagnosis of HCC, although there are still some limitations to generalize results mentioned above in this study. First, the samples were obtained from a single institution, and some subcategories were not included. Secondly, we evaluated only the expression of the circulating miRNAs, not the concentrations in the primary tumour or in cell lines. Further examination is necessary to evaluate the predictability of serum miR-574-3p.
In summary, our study demonstrates that serum miR-574-3p has considerable clinical value for the diagnosis of HCC, which may aid diagnosis of HCC. However, the molecular mechanisms of miR-574-3p and its value in HBV infections with high HBV DNA copy numbers in HCC need to be further studied.
Footnotes
Supplementary material
Additional supplementary information may be found with the online version of this article.
Acknowledgements
We would like to thank Nantong Tumor Hospital and the Affiliated Hospital of Nantong University.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The National Nature Science Foundation of China (81672099); Key Project of Jiangsu Province (BE2015654); the Nantong Program for Research Center of Clinical Laboratory Diagnosis (HS2015002); translational medicine project of Affiliated Hospital of Nantong University (TDF-zh201407, TDF-zh201406); Jiangsu Program for Young Medical Talents (QNRC 2016695).
Ethical approval
The study was approved by the Ethics Committee of Nantong Tumor Hospital with permit number 2015 (112).
Guarantor
SJ.
Contributorship
XS and YX researched the literature and conceived the study. HC, XW and SJ were involved in protocol development, gaining ethical approval, patient recruitment and data analysis. XS wrote the first draft of the manuscript. All authors reviewed and edited the manuscript and approved the final version of the manuscript.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
