Abstract
Objectives
Colorectal cancer (CRC) is the third most common and fourth most deadly cancer worldwide despite its various screening method. Thus, the search for novel and better markers is continuous. This study aimed to assess the combined expression levels of miR-133a, miR-574-3p, and miR-27a in early diagnosis of colorectal cancer in comparison to traditional tumor markers (CEA and CA19.9).
Methods
miR-133a, miR-574-3p, and miR-27a were assessed in sera of 120 participants categorized into healthy control group (n = 20), benign group (n = 30) and malignant group (n = 70) using real-time PCR.
Results
miR-133a, miR-574-3p, and miR-27a expressions showed significant difference among different staging, grading and tumor size of CRC. The sensitivities of the three miRNAs whether combined or individually used were better than routinely used tumor markers (CEA and CA19.9) leading to more accurate and faster diagnosis of CRC.
Conclusion
Synergetic detection of miRNA-133a, miRNA-574-3p, and miRNA-27a may serve as better noninvasive biomarkers with higher combined sensitivity for early diagnosis of CRC than individual detection of miRNAs.
Introduction
Colorectal cancer (CRC) is the third most common cancer as well as the fourth most lethal cancer worldwide. CRC accounts for around 10% of all new cancer cases universally, remaining the second most common cause of cancer-related deaths. 1 In Egypt, the expected percent of CRC is 6.5% of all cancers. 2
Obesity, diabetes, Irritated bowel disease, precancerous colonic lesions, and family history of CRC were the highly considerable risk factors. 2 Extensive research has recommended that nutrition may play both a causal and protective role in the development of colon cancer. 3
MicroRNAs (miRNAs) are regulatory RNAs with 18–25 endogenous non-coding nucleotides that show post-transcription gene regulation. MiRNAs regulate gene expression and contribute to development, differentiation, inflammation, and carcinogenesis. 4 Dysregulation of mature miRNA biogenesis steps can cause alterations in miRNA expression in cancer. 5
MiR-133a has been confirmed as a tumor suppressor in many cancers by suppressing cellular proliferation, migration, and invasion. 6 Human miR-133a is encoded by 2 different genes, miR-133a-1 and miR-133a-2, which are processed into matching mature sequence. miR-133a-1 and miR-133a-2 are implanted in the MIB1 gene on chromosome 18 and C20orf166 on chromosome 20, respectively. 7
MiR-133a has been shown to regulate diverse target genes in numerous cancer cells. Among them, eukaryotic translation initiation factor 4A1 (eIF4A1) acts as a serious effector of miR-133a. eIF4A1, an adenosine triphosphate (ATP)–dependent RNA helicase, is vital for assembly of the translational active ribosome. 8 MiR-133a represses the luciferase activity of Luc-eIF4A1-3′UTR by targeting the 3′UTR of eIF4A1 mRNA. The inhibitory effect of miR-133a on cell proliferation can be abolished by overexpression of eIF4A1. 9 Tumor growth inhibition by miR-133a could be attributed, at least in part, to activating p53/p21 pathway. 10
MiR-27a exerts numerus regulatory functions in different kinds of cancer. 11 MiR-27a was reported to be located on chromosome 19 and played a vital role in tumor development. 12 MiR-27a acts as a tumor suppressor. Studies have recognized sphingosine-1-phosphate phosphatase 1(SGPP1) and Smad2 as two new goals of miR-27a, which is related to signal transducer and activator of transcription 3 (STAT3) to control cancer cell proliferation, apoptosis and migration. 13
MiR-574-3p was found to act as tumor suppressor in several cancers. 14 Genes encoding miR-574-3p are located on chromosomes 4P (4p14 and 4p15) with a tumor inhibitory miRNA function. 15 It has been reported that miR-574-3p blocked the epithelial–mesenchymal transition (EMT) and promoted cisplatin sensitivity through inhibiting the activity of zinc finger E-box binding homeobox transcription factor 1 (ZEB1) in GC cells. 16 A study potentiated the role of miR-574-3p in blocking the cell proliferation of human CRC cells in vitro through targeting Cyclin D2 (CCND2), which increases cell apoptosis 17
About 15% of CRCs are diagnosed in metastatic stages (stage IV). Despite the existing screening methods and prognosis factors, there are still an excessive number of patients that are facing therapeutic failure and metastasis. 18 Thus, this study aims to evaluate the expression levels of miRs-133a, 27a and 574-3p as early diagnosis biomarkers of CRC and associate them with clinicopathological features of CRC patients and benign polyps patients.
Subjects and methods
Enrolled participants
Upon obtaining ethical approval from the Research Ethics Committee (faculty of medicine Ain shams university, R173/2022), this case-control study enrolled 120 subjects and categorized into three groups, healthy control (20 subjects), benign group (30 subjects), and malignant group (70 subjects). All participants have signed informed consent.
Patients’ inclusion criteria were newly diagnosed CRC patients before having colorectal operations, receiving radio- or chemotherapy and have no other malignancies or distant metastasis, while patients did not achieve these criteria were excluded from the study. The clinicopathological data were collected from patients’ clinical sheets.
Sample size calculation
Sample size is 94 or more measurements/surveys are needed to have a confidence level of 95% that the real value is within ±5% of the measured/surveyed value
Samples collection and processing
Patients attending oncology unit at faculty of medicine Ain Shams University during period from February 2018 to December 2019 were enrolled. Blood samples collected from enrolled individuals were allowed for clotting 30 min at room temperature. Centrifugation at 10,000 g for 10 min at 4°C were followed by aliquoting and storing at −80°C for tumor markers assessment and miRNA expression analysis.
Determination of serum tumor markers
The quantitative measurement of human carcinoembryonic antigen (CEA) and human cancer antigen 19.9 (CA19.9) in serum was performed by the solid phase enzyme-linked immunosorbent assay using available commercial ELISA kit (Catalog No.C29-333) (Immunospec Corporation, Netherland).
Reverse transcription (RT-q) PCR
- MiRNA extraction: Circulating miRNAs were extracted from the serum samples using miRNeasy Mini kit (Catalogue # 217004, Qiagen, Germantown, MD), as recommended by the manufacturer’s instructions: miRNA extraction was carried out using 100 μL of serum sample lysed in 500 μL of QIAzol Lysis Reagent. The RNA was eluted using RNase-free water in a final volume 30 μL. The purity and the concentration of the purified miRNA were detected using nano-drop spectrophotometer (Quawell, Q-500, Scribner, Staten Island, NY) and stored at −80°C till further assessments. - Reverse transcription and cDNA preparation: As recommended in the manufacturer’s instructions using MiScript II reverse transcription kit (CAT number # 218161, Qiagen, USA) cDNA was synthesized at 37°C for 60 min in total volume of 20 μL of reverse transcription reaction components (4 μL MiScript HiFlex buffer, 2 μL nucleic mixture, 2 μL MiScript RT mixture, RNase-free water (variable depending on the volume of the added template miRNA) and template of purified miRNA with adjusted concentration 100 ng/mL). Complementary DNA concentration and purity were detected using nano-drop spectrophotometer (Quawell, Q-500, Scribner, Staten Island, NY) and stored at −20°C till performing qPCR. - Quantitative real-time PCR (qPCR): RT-qPCR was achieved by MiScript primer assay (Cat number 218300, Qiagen, USA) for miR-133a (Hs_miR_133a_2miScript Primer Assay, MS00031423), miR-27a (Hs_miR_27a_2miScript Primer Assay, MS00003241), miR-574-3p (Hs_miR_574-3p miScript Primer Assay, MS00032025), and reaction was performed via MiScript SYBR Green PCR kit (Cat number 218073, Qiagen, USA). The reaction for miScript primer assays was carried out by using cDNA with concentration adjusted to 2 ng/mL and a total volume of 20 μL. PCR protocol was initial activation for 15 min at 95°C and 40 cycles of 94°C for 15s followed by annealing at 55°C for 30s and extension for 30s at 70°C. All of reactions run in triplet. Also, an endogenous control, that is, RNU6-2 (Hs_RNU6-2_11 miScript Primer Assay, MS00033740) was used to normalize the investigated miRNAs expression level. Fluorescence detected by Strata gene real-time PCR system (Max3005P QPCR system, Strata gene, Agilent biotechnology, Santa Clara, CA). Relative expression was normalized to an internal control (miRNA U6) and calculated according to Livak and Schmittgen
19
using the (2)-ΔΔCT method.
Statistical analysis
SPSS was used to statistically analyze data (Statistical Program for Social Science, version 26, Inc., Chicago USA). The clinicopathological and demographic factors association with investigated miRNAs were determined using ANOVA analysis. Two-tailed P value was used to discriminate positive from negative results and considered significant if < 0.05. Spearman’s correlation coefficient (CC) 20 were used to analyze correlations between investigated miRNAs. Receiver operating characteristic (ROC) curve was plotted between studied groups to detect the sensitivities and the specificities for the miRNAs and their diagnostic efficiency. 21 Area under the curve (AUC) can range from 0.5 to 1 and diagnostic tests that approach 1 designate a perfect discriminator.
Results
In this study, 120 participants were included and divided into three groups as follows: • Malignant group with mean age (48 ± 13) years • Benign group with mean age (45 ± 18) years • Healthy control group with mean age (45 ± 17) years
Clinical data of the studied patients’ groups.

Tumor markers levels
The mean, median, and rank of studied tumor markers.
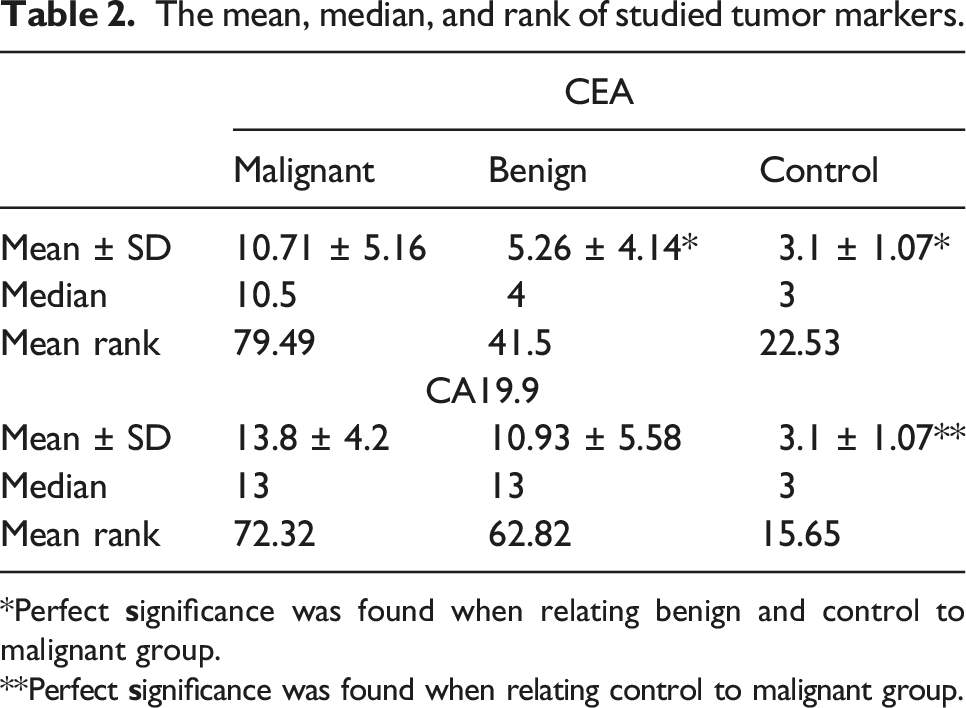
*Perfect
**Perfect

Receiver operating characteristic (ROC) curve for tumor marker (a) CA19.9 (b) CEA. The best cutoff values were 11.5 U/ml for CA19.9 and 5.5 ng/mL for CEA with AUC (95% CI) 0.736 (0.639–0.834) and 0.88 (0.814–0.946) for CA19.9 and CEA, respectively, at p < .05.
ROC curve analysis for CEA and CA19.9

miRNAs expression levels among investigated groups
Expression levels of investigated miRNAs among groups.

*Kruskal–Wallis test, P is highly significant, RQ: relative quantity.

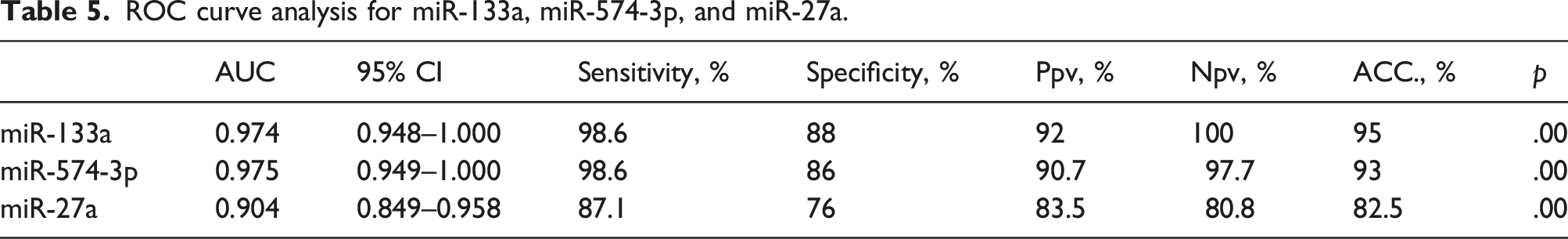
Receiver operating characteristic (ROC) curve for (a) miR-133a (b) miR-574-3p, (c) miR-27a. The best cutoff values were 36.5-, 46.5-, and 18.5-fold change for miR-133a, miR-574-3p, and miR-27a, respectively, with AUC (95% CI) 0.974 (0.948–1.000) for miR-133a, 0.975 (.948–1.000) for miR-574-3p and 0.904 (0.849–0.958) for miR-27a at p < .05.
ROC curve analysis for miR-133a, miR-574-3p, and miR-27a.
Combined sensitivities and specificities of studied miRNAs.

Diagnostic efficacy for investigated miRNAs and tumor markers
Levels and positivity rates of investigated parameters among studied groups.
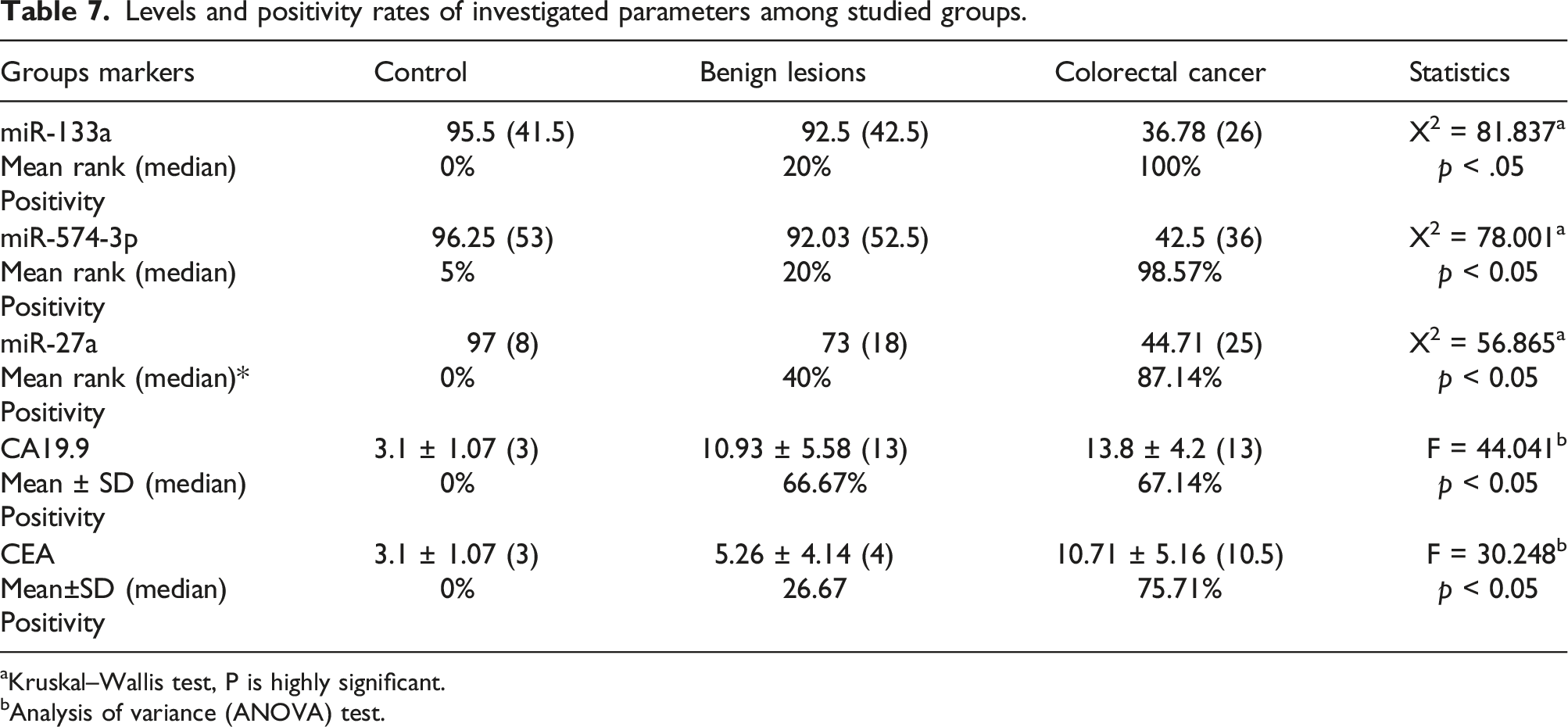
Kruskal–Wallis test, P is highly significant.
Analysis of variance (ANOVA) test.
On the other hand, extremely substantial raise in tumor markers levels (CEA and CA19.9) in CRC patients when compared to the other enrolled groups.
Relation between miRNAs, tumor markers and clinicopathological factors
Relation between positivity rates of investigated parameters with clinicopathological data among colorectal cancer patients.
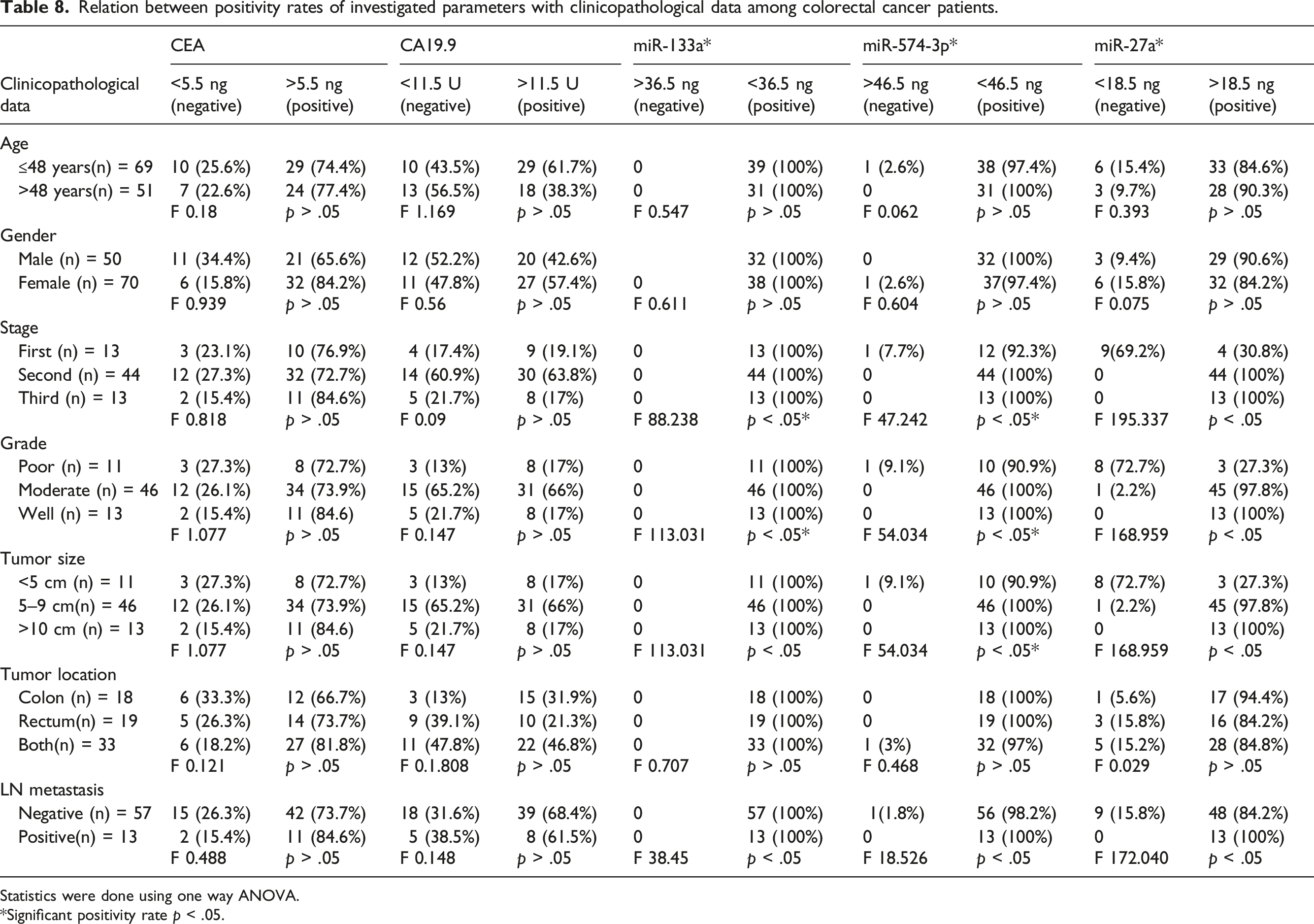
Statistics were done using one way ANOVA.
*Significant positivity rate p < .05.
When the expressions of miR-133a, miR-574-3p, and miR-27a were used to detect the CRC grade as diagnostic biomarkers, they detected low grades from high grade of CRC with a positivity of 100% for high and low grades in miR-133a, 90% and 100% for low and high grade in miR-574-3p, 27.3% and 100% for low and high grade in miR-27a (Supplementary Table 1).
The expressions of miR-133a, miR-574-3p, and miR-27a could distinguish between early, moderate and late stages with positivity rate 100% for all stages in miR-133a, 92.3% for early and 100% for moderate and late stages in miR-574-3p, 30.8% for early stage and 100% for moderate and late stages in miR-27a (Supplementary Table 2).
A significant positivity rate was detected for the whole three miRNAs in distinguishing negative node metastasis from positive node metastasis in CRC patients. Sensitivity and specificity were 100% and 100% for miR-133a, 100% and 98.2% for miR-574-3p, 100% and 84.2% for miR-27a, respectively (Supplementary Table 3).
Discussion
Colorectal cancer (CRC) is the third most diagnosed cancer and second deadliest malignancy for both genders. CRC has both strong environmental associations and genetic risk factors. 22 These environmental and genetic factors cooperate to produce colon polyps that develop to colorectal cancer. The polyp to cancer development sequence is mainly motivated at the cellular level by gene alterations and epigenetic changes and is now accepted to be a heterogeneous process. 23
The routinely used tumor markers for diagnosis and prognosis of CRC aren’t significant as they are also elevated in other benign conditions. CEA is elevated in cases of hepatitis, pancreatitis, obstructive pulmonary disease and inflammatory bowel disease. 24 In this study, CEA sensitivity and specificity was 75.7% and 84%, respectively, it showed no significant difference to tumor grade and stage, which is consistent with Gao et al, 2018. 25 In addition, CA19.9 has been reported to increase in many types of cancer. 26 In this study, CA19.9 had sensitivity of 67.1% and specificity of 60% with no significant difference when related to tumor staging, grading and lymph node metastasis. The results obtained from this study showed that these tumor markers cannot be used alone for diagnosis of CRC which is consistent with Lakemeyer et al, 2021. 27 So, we aimed to find more precise markers that can be used for faster diagnosis and better prognosis such as miRNAs.
Recently miRNAs are heavily investigated to provide a new way for understanding development and progression of malignant tumors. MiRNAs are Small non-coding RNA (18–25 nt) that can stop protein expression by breaking specific target mRNAs or by inhibiting their translation. 28 They mediate gene regulation by fusing with the 3-untranslated region (3’UTR) sequences of the target mRNAs, resulting in degradation of mRNA or translational silencing. MiRNAs perform vital roles in many cellular activities, involving cell proliferation, differentiation, programmed cell death, and growth. These miRNAs are considered to control the expression of about one-third of individual protein-coding genes. 29 Also, miRNAs are stable, reproducible, and consistent. 30
Among the miRNAs, miR-133a is believed to be a tumor suppressor and a biomarker for prediction of several cancers, such as breast cancer, bladder cancer, gastric cancer, esophageal cancer (EC), osteosarcoma, and non-small cell lung cancer (NSCLC). 31 The results obtained from the current study showed that miR-133a is downregulated in CRC and that it has perfect significance when related to tumor staging, grading and lymph node metastasis with sensitivity 98.6% and specificity 88% suggesting its role as important biomarker for diagnosis and prognosis of CRC consistent with Wan et al., 2014 and Caporali et al., 2021.32,33
In the current study, miR-133a expression was considerably decreased in the serum of CRC patients with late stage and low grade, consistent with earlier reports by Luo et al., 2013, 34 which also contradicts many studies, indicating that miR-133a is not simply an oncogene as many papers reported. 31
MiR-574-3p has been verified as a tumor suppressor in several cancers such as prostate cancer, bladder cancer, gastric cancer, ovarian cancer and chronic myeloid leukemia. 14 Our results demonstrated that miR-574-3p is downregulated in CRC patients with sensitivity 98.6% and specificity 86% suggesting its role as valuable biomarker for diagnosis and prognosis of CRC.
It was illustrated that miR-574-3p expression positivity rate was correlated with the tumor size, clinical stage and lymph node metastasis (p < .05). However, no correlation was found between the miR-574-3p expression level and age, gender or tumor location (p > .05) which was consistent with the in vitro study of Li et al., 2019. 17
Studies have shown that miR-27a showed a critical role in cancer biology, including polymorphisms, carcinogenesis, proliferation, programmed cell death, infiltration, metastasis, and angiogenesis. 35 MiR-27a acts as oncogenic miRNA that is upregulated and overexpressed in numerous cancers including gastric cancer, osteosarcoma, liver cancer, cervical cancer, ovarian cancer, pancreatic cancer, lung adenocarcinoma, breast carcinoma, and renal cell carcinoma. While it acts as tumor suppressor miRNA in esophageal squamous cell carcinoma, prostate cancer, and acute leukemia. 36 We found that miR-27a is upregulated in CRC as demonstrated by You et al., 2021 37 with sensitivity 87.1% and specificity 76%.
The present study showed elevated relative expression level of miR-27a associated with tumor stage, tumor grade and lymph node metastasis in patients with colorectal cancer, which is in line with others. 38
Statistical analysis of data obtained showed that combining both miR-133a and miR-574-3p together present perfect possible tumor markers for CRC with higher sensitivity (100%) and specificity (88%) than other markers used as illustrated in table (6). Combining the three miRNAs together suggests even better markers for early detection and prognosis of CRC with higher sensitivity (100%) than regular tumor markers used.
Limitations
The unavailability of a huge quantity of patients, non-cooperation of patients and incomplete patient records, were some of the limitations of this study. For this reason, further validation studies with larger cohorts are needed, and comparability tests are required to confirm the linkage of serum miRs-133a, 27a and 574-3p with CRC and their targeted pathway.
Conclusion
In conclusion, microRNAs are helpful in diagnosing colorectal cancer, regulating disease progression, predicting disease recurrence, and determining therapy success. Synergetic detection of miRNA-133a miR-27a and miRNA-574-3p serve as novel tumor marker with better sensitivity and specificity and help in early detection of CRC and with less invasive or surgical interference for such widely spread malignancy which could result in better prognosis and higher survival rate for CRC patients.
Also, overall, sensitivities and positive predictive values for miRNA-133a, miR-27a, and miRNA-574-3p were superior to the routinely tumor markers (CA19.9 and CEA) for early detection of CRC, especially with patients at early stage and low grade. These findings point out the value of using miRNA-133a, miR-27a, and miRNA-574-3p as early diagnostic molecular markers in CRC.
Furthermore, serum levels of miRNA-133a, miR-27a, and miRNA-574-3p can be used as a fair noninvasive method for early detection of colorectal cancer and differentiating between CRC histologic grade, stages, and lymph node metastasis.
Supplemental Material
Supplemental Material - Potential diagnostic role of circulating MiRNAs in colorectal cancer
Supplemental Material for Potential diagnostic role of circulating MiRNAs in colorectal cancer by Dina A El-Sayed, Ahmed MH Salem, Menha M Swellam and Marwa GA Hegazy in International Journal of Immunopathology and Pharmacology
Footnotes
Acknowledgments
The authors express their gratitude to Dr. Ghada R. Meckawy (Department of Clinical Oncology and Nuclear Medicine – Faculty of Medicine – Ain Shams University, Cairo, Egypt) for her kind cooperation in collecting samples from gastrointestinal oncology clinic, Ain shams university hospitals.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Ethical approval
Ethical approval for this study was obtained from the Research Ethics Committee (faculty of medicine Ain shams university, R173/2022).
Informed consent
Written informed consent was obtained from all subjects before the study.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
