Abstract
Purpose
We previously attempted to develop quantitative enzyme-linked immunosorbent assay (ELISA) systems for the PDA039/044/071 peptides, potential serum disease biomarkers (DBMs) of pregnancy-induced hypertension (PIH), primarily identified by a peptidomic approach (BLOTCHIP®-mass spectrometry (MS)). However, our methodology did not extend to PDA071 (cysteinyl α2-HS-glycoprotein341–367), due to difficulty to produce a specific antibody against the peptide. The aim of the present study was to establish an alternative PDA071 quantitation system using liquid chromatography-multiple reaction monitoring (LC-MRM)/MS, to explore the potential utility of PDA071 as a DBM for PIH.
Methods
We tested heat/acid denaturation methods in efforts to purify serum PDA071 and developed an LC-MRM/MS method allowing for specific quantitation thereof. We measured serum PDA071 concentrations, and these results were validated including by three-dimensional (3D) plotting against PDA039 (kininogen-1439–456)/044 (kininogen-1438–456) concentrations, followed by discriminant analysis.
Results
PDA071 was successfully extracted from serum using a heat denaturation method. Optimum conditions for quantitation via LC-MRM/MS were developed; the assayed serum PDA071 correlated well with the BLOTCHIP® assay values. Although the PDA071 alone did not significantly differ between patients and controls, 3D plotting of PDA039/044/071 peptide concentrations and construction of a Jackknife classification matrix were satisfactory in terms of PIH diagnostic precision.
Conclusions
Combination analysis using both PDA071 and PDA039/044 concentrations allowed PIH diagnostic accuracy to be attained, and our method will be valuable in future pathophysiological studies of hypertensive disorders of pregnancy.
Keywords
Introduction
It is clinically important to understand pregnancy-induced hypertension (PIH) in more detail to improve feto–maternal safety during pregnancy.1,2 Presently, monitoring of blood pressure and quantitation of urinary proteins are common forms of clinical care, but fail to predict prognosis or the risk of PIH complication onset.
Previously, we used a peptidomic analytical method to identify several candidate serum peptide fragments as disease biomarkers (DBMs) for PIH. 3 Among them, we focused on three peptide fragments (PDA039/044 both derived from kininogen-1, and PDA071 from α2-HS-glycoprotein) to confirm their utilities as DBMs. 4 Using established sandwich enzyme-linked immunosorbent assay (ELISA) methods, PDA039 (kininogen-1439–456), but not PDA044 (kininogen-1438–456), was validated as a DBM of PIH, but we could not develop an ELISA for PDA071 (cysteinyl α2-HS-glycoprotein341–367) because we encountered problems with the antibody specificity towards the native form of the serum peptide. 4 Concerning the pathological classification, the relationship between gestational hypertension (GH) and preeclampsia (PE) is controversial among the hypertensive disorders of pregnancy at present, the Japan Society for the Study of Hypertension in Pregnancy provides the clinical classification of PIH syndrome that includes both GH and PE. 4
Several studies have reported pathophysiological monitoring of disease progression using combination of DBMs for accurate diagnosis and prediction.5–7 We felt that a similar approach using the concentrations of PDA039/044 in combination with that of PDA071 would be of potential of clinical utility. Therefore, it was necessary to establish a serum PDA071 assay system using a non-immunological method.
Our primary aim of this study was thus to develop a serum PDA071 assay featuring targeted proteomics. We developed a new method for PDA071 quantitation using liquid chromatography (LC) coupled with multiple reaction monitoring (MRM)/mass spectrometry (MS).8,9 This method showed higher quantitative accuracy and can be widely applicable without the specific BLOTCHIP® system 10 used in our previous studies to screening for DBMs. 3 Using this system, we evaluated the potential utility of PDA071 as a DBM for PIH. Three-dimensional (3D) plotting of PDA071 concentrations combined with those of PDA039/044 was effective in this regard.
Materials and methods
Reagents
LC/MS-grade acetonitrile (ACN), formic acid (FA), methanol, isopropanol (IPN), and trifluoroacetic acid (TFA) were purchased from Wako Pure Chemical Industries, Ltd. (Osaka, Japan). Milli-Q water (Millipore, Billerica, MA, USA) was used as ultra-pure water. Other reagents were obtained from Sigma-Aldrich (St. Louis, MO, USA).
Peptides
PDA0714 and stable isotope-labelled PDA071 (PDA071 H28; TV(13C5,15N)VQPSV(13C5,15N) GAAAGPV (13C5,15N)VPPC(+Cys)PGRIRHF(13C9,15N)KV) were chemically synthesized (Scrum Inc., Tokyo, Japan). PDA071H28 was used for an internal standard (IS). These peptides were diluted at the concentration of 1 nmol/μL in 10% ACN/0.1% TFA solution and stored at −80℃ until use.
Blood samples
Serum samples from the PIH patients (n = 34: 33.7 [mean year old] ± 5.5[SD]), healthy pregnant females at 23 (n = 15: 31.8 ± 4.7 year old) and at 33 gestational-week (n = 30: 32.8 ± 4.4 year old) were used as reported previously. 4 For establishing LC-tandem MS (MS/MS) system, serum from normal rat was used as a control serum sample. All samples were stored at −80℃ until further use and not allowed to repeat freeze/thaw cycle more than twice. The Ethic Committee board members of Juntendo University and Yamagata Saisei Hospital approved this study (registration #19-017 and #144, respectively).
Peptides extraction from serum
Heat denaturation method
The serum sample (original volume: 200 μL) was placed in 1.5 mL polypropylene centrifuge tube and diluted with an equal amount of 20% ACN/0.1% FA solution. The sample was heated for 10 min at 70℃ in a heat block or a water bath. After cooling on ice sufficiently, the sample was centrifuged at 15,000 r/min (20,400 g) for 30 min at 4℃. The supernatant (approximately 300 μL) was transferred into a new tube, and then 1500 μL of 0.1% FA solution and 10 μL of the standard solution containing 2 pmol/μL PDA071H28 were added.
TFA precipitation method
TFA precipitation was performed essentially as described previously. 3 Briefly, TFA (20 μL) was added slowly to the 200 μL of human/rat serum samples with gentle mixing. In order to precipitate serum major proteins, the samples were kept on ice for 30 min. After centrifugation at 15,000 r/min (20,400 g) for 30 min at 4℃, the supernatant (approximately 180 μL) was transferred to a new tube and diluted 10-folds with water.
Preparation of MS sample
Salts were removed from the extracted peptides sample with Sep-Pak Plus C18 solid-phase extraction cartridge (Waters, Milford, MA, USA). Peptides containing PDA071/PDA071H28 were eluted with 1 mL of 80% ACN/0.1% FA solution. Finally, the peptide solution was diluted 8-fold with 0.1% FA solution to adjust the sample condition for LC-MS/MS. In order to prevent non-specific adsorption, the peptides solution prepared from the control rat serum by the TFA precipitation method was used as a diluent for standard peptides.
MRM condition
In order to determine the conditions of MRM for detecting PDA071 and PDA071H28, the synthesized peptides were analysed by LC-MS/MS system (LC: DiNa, KYA technologies, Tokyo, Japan/MS: QSTAR Elite, SCIEX, Framingham, MA, USA).
Quantitative LC-MS/MS analysis
The peptides mixture dissolved in 10% ACN/0.1% FA was analysed with a nanoflow LC-MS/MS using a nanoLC system (Eksigent nanoLC400; SCIEX, Framingham, MA, USA) and a mass spectrometer (QTRAP 6500, SCIEX). Five microlitres of each sample was injected into reverse phase LC system (Nano cHiPLC® ChromXP C18-CL 3 μm 120 Å, trap column; 200 μm I.D. × 0.5 mm length, analytical column; 75 μm I.D. × 15 cm length, SCIEX) equilibrated 2% ACN, 0.1% FA. Gradient elution was performed using 0.1% FA (solvent A) and ACN with 0.1% FA (solvent B). The elusion conditions from the column were as follows: 2–15% solvent B/5 min, 15–25% solvent B/10 min, 25–40% solvent B/5 min, 40–90% solvent B/1 min, 90% solvent B for 10 min. Then, the column was kept at 2% B for 20 min. The total run time of the analysis was 71 min. Between each analysis, the washing method for cleaning columns was run to remove carry-over peptides with 10 μL washing solution (ACN/IPN/methanol/water, 25% each). The washing method conditions were three cycles of the following protocol: 2–90% solvent B for 10 min, maintained 90% solvent B for 5 min, dropped to 2% solvent B within 0.1 min, and kept 2% for 5 min. The column was re-equilibrated at 2% solvent B for 20 min in the end of the washing process.
The MS was operated in positive ion electrospray ionization mode. Enhanced mass scan and MRM scans were utilized to detect the analyte and IS. The MRM transitions (precursor/product) for detecting the PDA071/PDA071H28 were as follows: PDA 071: m/z 715.388/763.398, 715.388/910.992 and 572.511/763.398, PDA071H28: m/z 722.405/768.414, 722.405/919.013 and 578.125/768.414. The CE value (SCIEX) was set at 36 volt.
The MultiQuant 3.0 software (SCIEX) was used for quantitating peptide concentration. The data quantitation was performed by calculating the chromatographic peak areas of 715.388/763.398 for PDA 071 and 722.405/768.414 for PDA071H28 using the MultiQuant MQ4 algorithm. The calibration curves were built by linear regression analysis using the least-squares method with 1/x weighting to prioritize the calibration points with lower peptide concentrations.
Statistical analyses
Statistical analysis was performed with GraphPad Prism version 5.0 (GraphPad Software Inc., La Jolla, CA, USA). The differences in peptide concentrations among groups were evaluated by the Mann-Whitney U-test for non-parametric data. Discriminant analysis was employed for PDA039, 044 and 071. Wilks’ lambda 11 or Jackknifed12,13 classification was performed in order to ascertain two-group classification success per domain and across domains. Percent correct classification is based on the overall success of the total samples’ cumulative classification in this manner. A probability of P < 0.05 was considered statistically significant.
Results
Conditions of MRM-mediated detection: Standard PDA071
PDA071 exhibited protonated precursor ions at m/z values of 715.388 (4+) and 572.511 (5+) upon electrospray ionization MS. Fragmentation of these ions yielded two major product ions at m/z values of 763.398 (2+) and 911.992 (2+). The peptide sequence and mass confirmed that these ions were generated from PDA071. We then selected three MRM transitions to detect PDA071: m/z 715.388/763.398, 715.388/911.992 and 572.511/763.398.
MRM transitions for detection of isotope-labelled PDA071 (PDA071H28: IS) were also determined, and were m/z 722.405/768.414, 722.405/919.013 and 578.125/768.414. We applied these MRM parameters in triple quadrupole mass spectrometry (using a QTRAP6500 instrument) and constructed an LC-MRM/MS assay system that detected both synthetic PDA071 and PDA071H28 diluted with a control rat serum peptide solution at the same retention times.
Of the three transitions of each peptide, we selected the most intense MRM signals for quantitation; these were m/z 715.388/763.398 for PDA071 and m/z 722.405/768.414 for PDA071H28.
Calibration curve for quantitation: Standard PDA071
Chemically synthesized PDA071 was diluted from 0.977 to 1000 fmol/μL with a control rat serum peptide solution. PDA071H28 (the IS) was added to each sample, to 5 fmol/μL and the samples were then analysed via LC-MRM/MS. To construct a calibration curve, the peak areas of the MRM signals of PDA071 (715.388/763.398) and the IS (722.405/768.414) were calculated. The PDA071 concentration was calculated by reference to the ratio of the signal peak area of PDA071 and PDA071H28. We applied linear fitting to the log–log data plot of the calculated and actual amounts. The calibration curve was linear from 4.88 to 2500 fmol (r = 0.999) (Supplementary Figure S1), corresponding to PDA071 concentrations of 12.2 to 6250 fmol/μL in the initial serum samples. Although quantitation of the peptide was possible at 2.44 fmol, analytical precision is not good when the lower limit of quantitation (LOQ) was set at 2.44 fmol. Therefore, the LOQ was set at 4.88 fmol, and 2.44 fmol is indicated as limit of detection (LOD) (Supplementary Figure S1). The peak observed at the lower LOQ was much greater (over 10-fold) than the noise peaks.
Extraction of native PDA071 from human serum samples
The native PDA071 peptide was difficult to extract from serum because it adsorbed strongly to several major serum proteins. 3 Therefore, we tested two protein precipitation methods, TFA precipitation and thermal denaturation, in efforts to extract peptides quickly, in a relatively simple manner.14,15 Initially, we tested the TFA method against the human serum containing high concentrations of PDA071 peptide measured by the BLOTCHIP® method. 3 However, the peptide concentrations detectable by the LC-MRM/MS method were less than expected. This suggested that native PDA071 was adsorbed to major serum proteins and precipitated with them when TFA was added.
We next explored thermal denaturation.16,17 To completely precipitate proteins, we heat-denatured serum in the presence of 10% ACN and 0.1% FA. We tested a range of temperatures (60–110℃) and denaturation times (10–120 min) to optimize the method. Our pilot experiments showed that denaturation at 70℃ for 10 min completely precipitated the major proteins and afforded the maximal yield of PDA071 (data not shown). When the samples were prepared from human serum that contained high concentrations of PDA071, the BLOTCHIP® and LC-MRM/MS methods yielded comparable peptide concentrations.
Thus, low concentrations of ACN and acid were necessary when thermal denaturation was used to isolate native PDA071. However, synthetic PDA071 and PDA071H28 peptides added to control rat serum were not effectively extracted by this method; the yields were less than 10% (data not shown).
Measurement of native PDA071 extracted from serum by the LC-MRM/MS method
Native PDA071 peptide concentrations were quantified by LC-MRM/MS using PDA071H28 as an
IS. To avoid destruction of the spiked PDA071H28 standard during heat denaturation and
precipitation, PDA071H28 was added to native PDA071 samples only after thermal
denaturation (Figure 1). This
allowed us to correctly measure PDA071 concentrations in serum samples. LC-MS/MS spectra of peptides. A typical total ion chromatogram of peptide solution
purified from human serum (a), and MRM ion chromatogram of PDA071 (b) and PDA071H28
(c) by LC-MS/MS. The three most intense precursor/product ions of PDA071
(m/z 715.388/763.398, 715.388/910.992, and 572.511/763.398), and
PDA071H28 (m/z 722.405/768.414, 722.405/919.013,
and 578.125/768.414) were traced. PDA071H28 was used as IS for measurement. The IS
peptide was added at the concentration of 5 fmol/μL after heat
denaturation. LC-MS/MS: liquid chromatography coupled with tandem mass spectrometry;
IS: internal standard; CPS: count per second.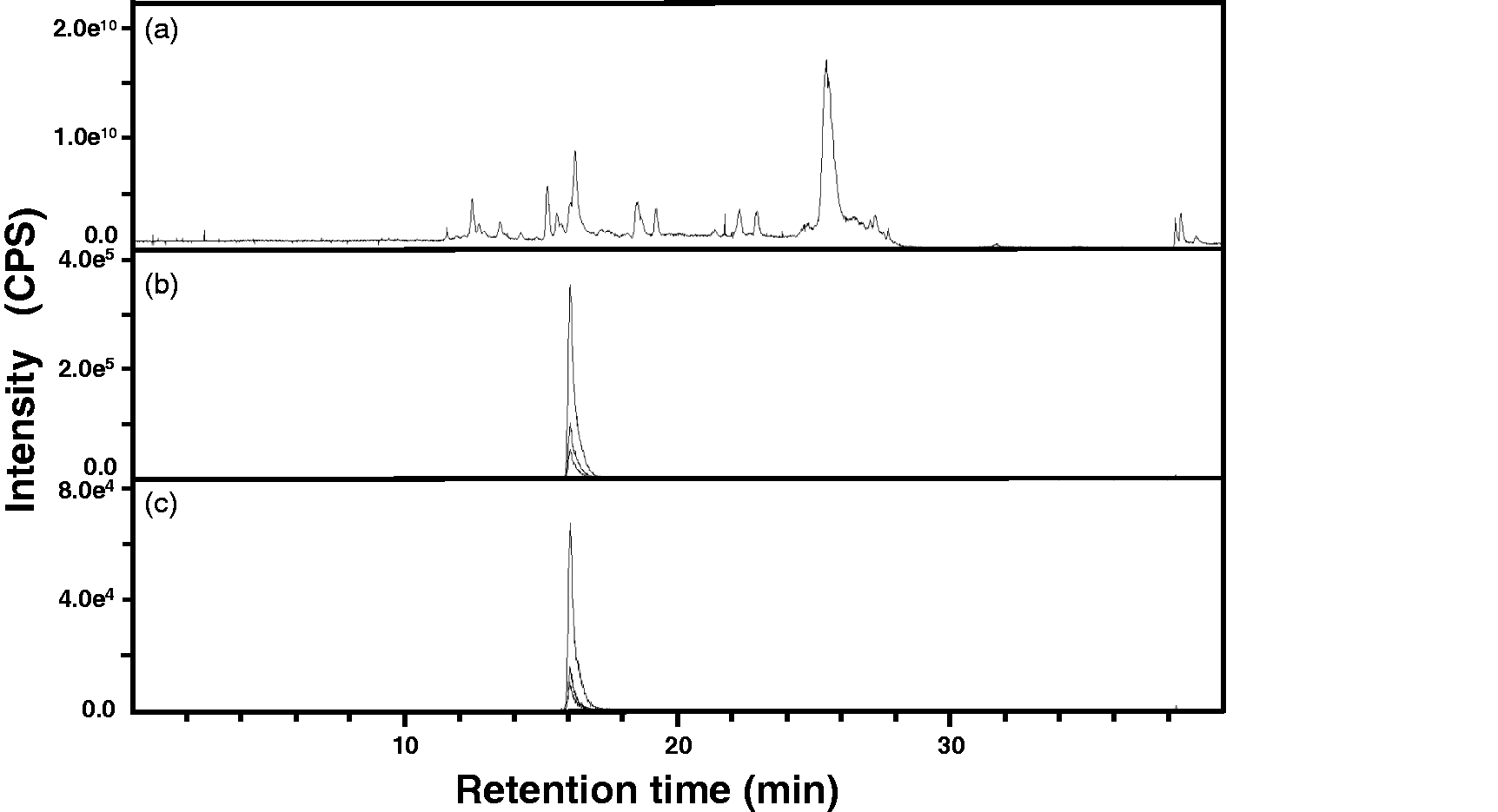
To confirm the reproducibility of the method, a serum sample from a healthy male was processed seven times and the PDA071 peptide concentrations measured. The concentrations were near-identical, confirming reproducibility (Supplementary Figure S2). We evaluated the difference in serum and plasma PDA071 concentrations (three samples) of the same subjects; the differences were less than 10% (data not shown).
Comparison with peptide concentrations measured by the quantitative LC-MRM/MS and BLOTCHIP® methods
In our previous study, the concentrations of PDA071 in 19 serum samples were measured by
the BLOTCHIP® method.
3
Using the same serum samples, we measured serum PDA071 concentrations by
the LC-MRM/MS method and compared them with the earlier data. When the LC-MRM/MS PDA071
concentrations were plotted against the peak intensities of the BLOTCHIP® data, the values
correlated significantly (data not shown). Experimental data as shown in Figure 2(a) are the correlation
between serum concentration of PDA071 analysed by LC-MRM/MS and the values from relative
intensity of the peptide in the serum by BLOTCHIP®-MS. We previously demonstrated that
accuracy of peptide quantitative analysis from PIH patient serum using BLOTCHIP®-MS in
general.
3
Accordingly, the values of relative signal intensities obtained from BLOTCHIP®-MS can be
regarded as serum concentration (the intensities have an equilateral association with
serum peptide concentration, and these are expressed in a linear function). Based on above
evidence, a Bland-Altman plot to compare the two methods is represented in Figure 2(b). Therefore, the LC-MRM/MS
method may be used to quantitate PDA071 in serum. Comparison of serum PDA071 concentrations analysed by LC-MRM/MS and BLOTCHIP®-MS.
The serum PDA071 peptide concentrations (n = 19) analysed by
LC-MRM/MS were plotted against the values by BLOTCHIP®-MS (a) and a Bland-Altman
plots comparing data obtained from both assay systems (b). Asterisk: These values
were calculated from quantitative BLOTCHIP-MS.
3
PDA071: mean difference (solid line)
0.522, +LOA/−LOA (dotted lines) 0.106/0.881.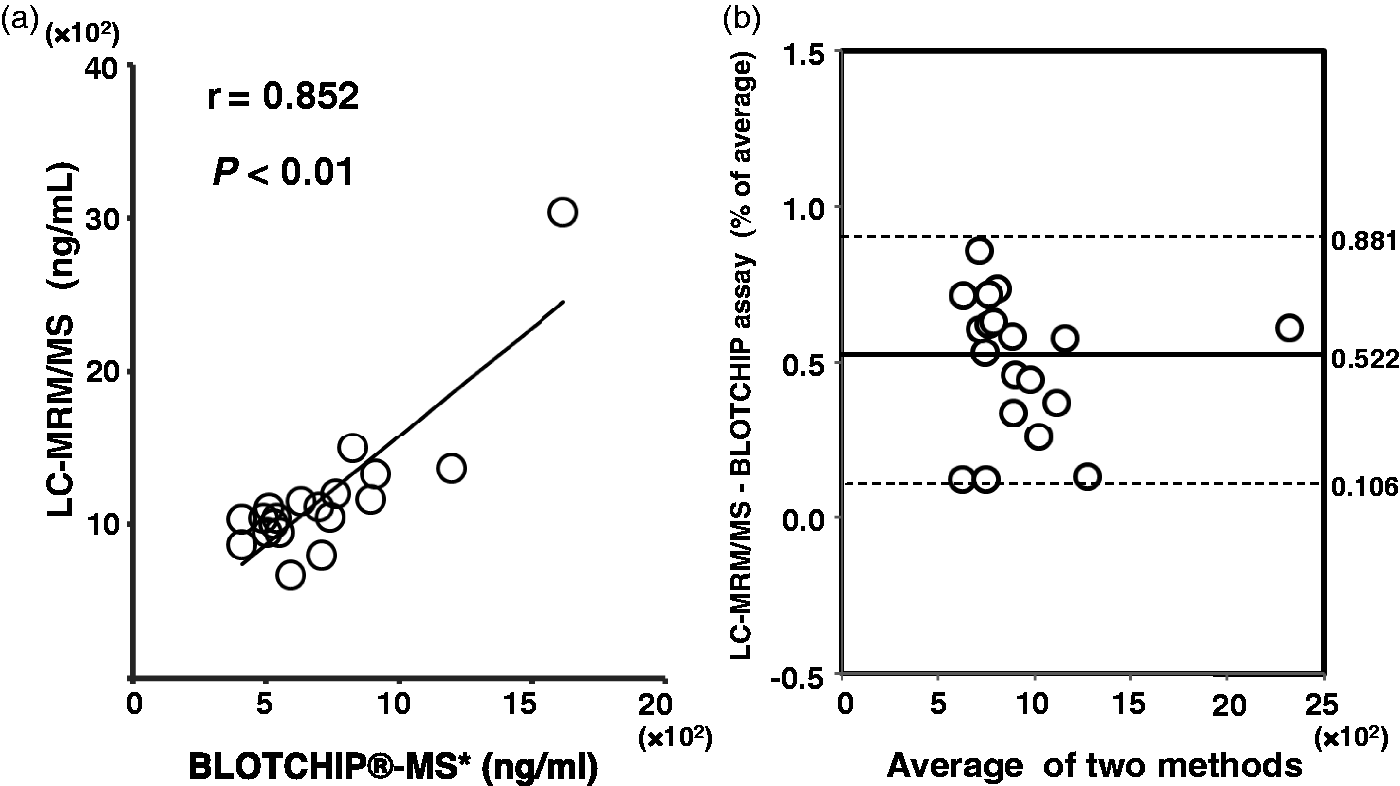
Quantitation of PDA071 concentrations in clinical samples using MRM
PDA071 concentrations in the serum samples of PIH patients and normal 33-week pregnant
females were examined. Although we found no significant difference between patients and
controls, the PDA071 concentration in PIH serum samples (median:
1142.9 ng/μL; interquartile range (IQR):
1043.1–1317.4 ng/μL) was slightly higher than that of normal pregnant
females (median: 1115.9 ng/μL; IQR: 1042.8–1253.8 ng/μL)
(Figure 3). No difference was
evident from 23 to 33 weeks of gestation (data not shown). PDA071 concentration in the serum of PIH patients by quantitative LC-MS. Serum
concentration of PDA071 from healthy pregnant females at 33-week of gestation (open
circles, n = 30) and PIH patients (closed circles,
n = 34) were plotted by scatter and box plot. Whiskers show
5%–95% tile.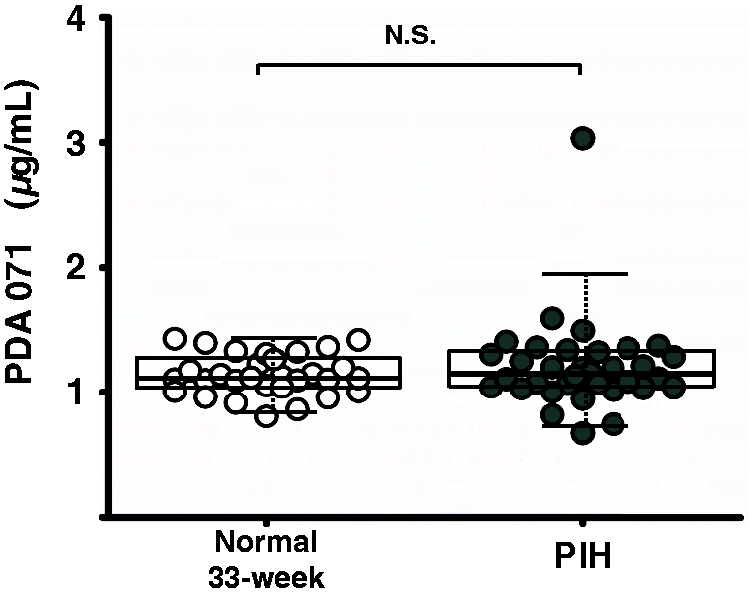
Multidimensional analyses of serum peptide concentrations for evaluation of PIH
Unexpectedly, single measures of serum PDA071 did not aid in PIH diagnosis (Figure 3). Thus, we used the PIH data
in combination with information on other DBM peptides. PDA039/044 concentrations were
examined in our previous report;
4
we thus first constructed 2D scatter plots of combinations of these
three peptides (Figure 4(a) to
(c)). The PDA071/039 serum
concentrations in PIH patients ranged rather widely (Figure 4(a)). Indeed, discriminant analysis indicated
that the serum concentrations of PDA071/039 (Figure 4(a)) and PDA039/044 (Figure 4(c)) differed significantly between normal
controls and PIH patients. Based on these data, we subjected the concentrations of the
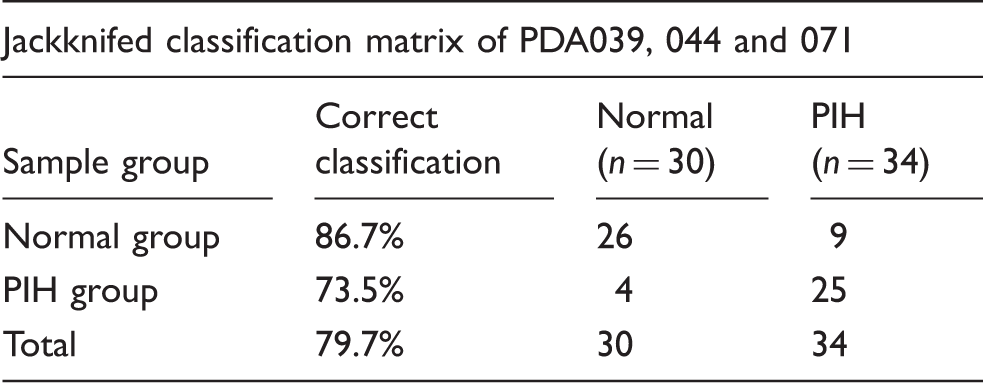
three peptides to discriminant analysis in terms of PIH diagnosis (Figure 4(d)). The controls and patients differed
significantly, and the correct diagnosis rate was 79.7% when the Jackknifed distinction
was employed (Table 1). These
results show that the 3D analysis using the PDA039/044/071 peptides afforded a
satisfactory diagnostic precision for PIH patients. Multidimensional analyses of the three peptides in the serum. Two-dimensional
scatter diagram of PDA071 with 039 (a) or 044 (b), and PDA039 with 044 (c) were
obtained from ELISA or LC-MRM/MS analysis. Serum concentration of these peptides
from healthy pregnant females at 33-week of gestation (open circles,
n = 30), PIH patients (closed circles, n = 34)
are plotted. In 3D diagram (d), blue/red spheres indicate serum peptides
concentration from controls/PIH patients, respectively. Discriminant analysis of the three peptides. Note: Wilks’s lambda classification = 0.798, P = 0.003;
Jackknifed classification efficiency is calculated by sequentially eliminating one
case at a time, computing the discriminant function based on remaining subjects,
and using the resultant discriminant to classify the eliminated case.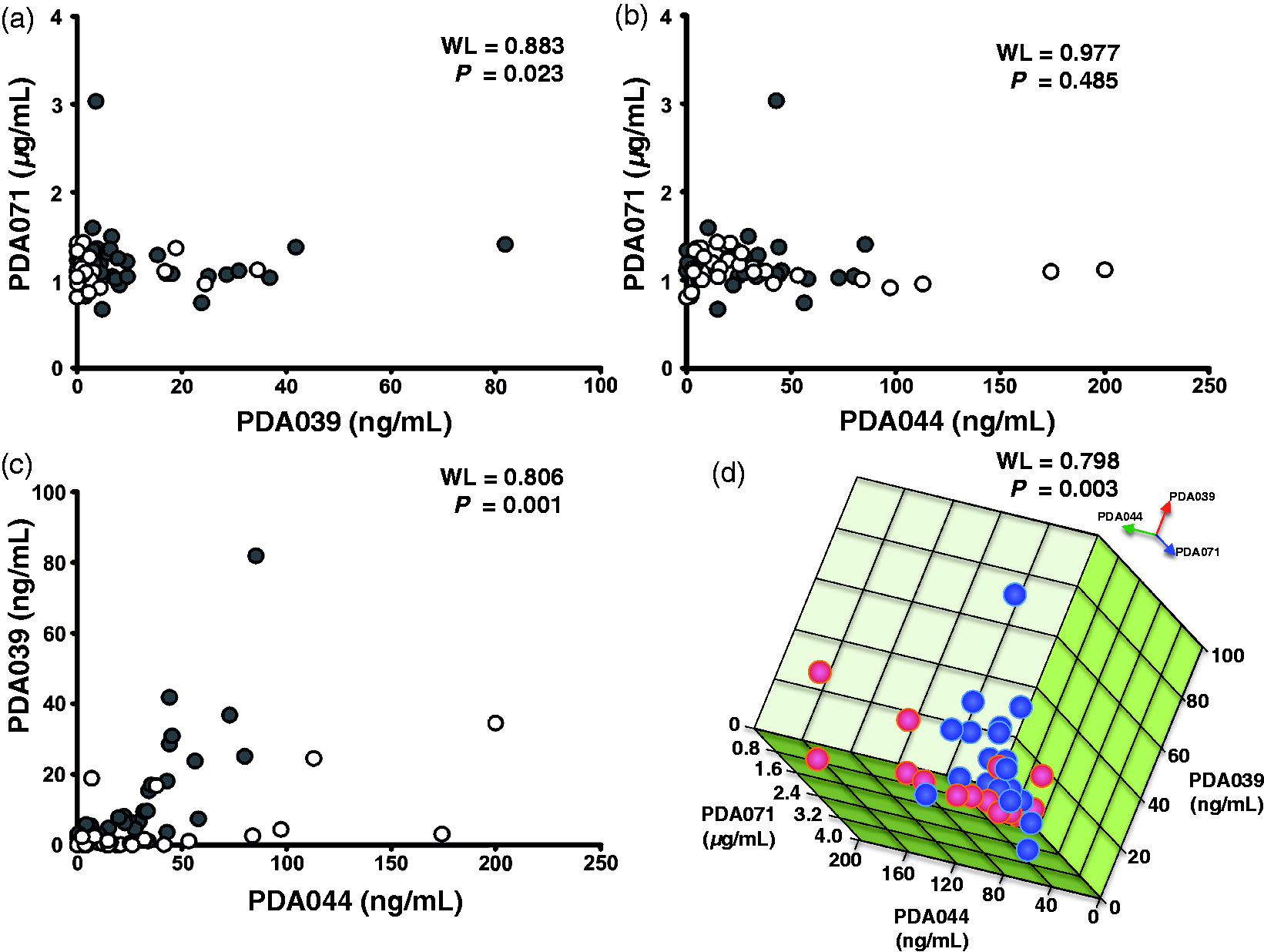
Discussion
We describe a new means of evaluating PIH pathophysiological status; we took a multidimensional approach toward this end; we employed data on the concentrations of three DBM candidate peptides, including PDA071. Unlike what was true of PDA039, the concentration of serum PDA044 or PDA071 alone did not differ between PIH patients and controls (Figure 3 in this study). 4 However, the false-positive/negative rates were reduced upon 3D analysis using the PDA039, 044 and 071 peptides (Figure 4, Table 1). We found it difficult to develop an ELISA for PDA071 because we could not create a specific antibody targeting the peptide. 4 Our new method opens up a novel approach toward PIH management using peptide data.
We found that the PDA071 recovery rate differed markedly when the TFA precipitation and heat denaturation methods were used. This may be because the peptide has a high affinity for major serum proteins. 3 Although TFA precipitation did extract free serum PDA071, the extent of molecular dissociation of PDA071 from major serum proteins was poor (Supplementary Figure S3). The heat denaturation method was optimal for PDA071 isolation from serum (Supplementary Figure S3). Presumably, PDA071 is heat-stable; the concentrations thereof were not affected by heat treatment (Supplementary Figure S4). Although we suppose that unexplained molecular interaction between major serum proteins and native PDA071 affect experimental results, quantitation of native PDA071 before heat treatment was quite difficult. Accordingly, it seems that there is almost no way to quantitate PDA071 before and after heat treatment. However, we have alternatively demonstrated that there is a correlation between these values analysed by LC-MRM/MS and BLOTCHIP®-MS (Figure 2), since no heat treatment is necessary to quantify peptides in the latter method.3,10 Therefore, we used this method prior to serum PDA071 quantitation with LC-MS.
Although no significant difference in serum PDA071 concentration was evident between controls and PIH patients group, one PIH case had a rather high serum PDA071 concentration (Figure 3). This patient ultimately progressed to HELLP (haemolysis, elevated liver enzymes and low platelets) syndrome. 18 Another PIH patient, also ultimately diagnosed with HELLP syndrome, had a relatively high serum PDA071 concentration (1.25 μg/mL). However, no relationship between HELLP syndrome and serum PDA071 concentration can be inferred from data based on only two cases. Although any relationship between serum α2-HS-glycoprotein (PDA071 originates from this molecule) concentration and liver function during PIH remains controversial,19,20 a recent review article suggested that the serum α2-HS-glycoprotein concentration reflects various pathophysiological states, including disorders of cardiovascular, metabolic and liver functions. 21 Thus, the evaluation of serum PDA071 derived from the glycoprotein may allow liver function to be assessed in PIH patients. On the other hand, clinical utility of the peptide DBMs for PIH, especially, excellence of the analytical methods between use of a single peptide (such as PDA039) 4 and a combination of the DBM peptides (including PDA071) is inconclusive simply at present, because additional data by multidimensional analysis from various clinical situation would be required to conclude the issue. Since we do not have enough samples from special situation of the hypertensive disorders of pregnancy (such as HELLP syndrome), therefore, the 3D analysis should be necessary to evaluate more carefully by large-scale clinical studies for the potential utility as a new inspection technique for understanding of the pathophysiological status of PIH.
To date, many proteolytic peptides associated with various health problems have been detected in human humoral fluids, including serum.22–26 However, homeostasis is maintained by permitting gradual instability within a dynamic equilibrium; it is thus difficult to understand the pathophysiology of intractable diseases (such as PIH) if it is considered that ‘a single DBM corresponds to one specific disease’. Therefore, multidimensional analyses such as those that we performed, may permit monitoring of the pathophysiological status of intractable diseases, including PIH. Our serum samples were collected from patients with varying clinical backgrounds. Large-scale studies in many hospitals/countries are necessary as stated above to describe the ‘typical’ PDA071/039/044 profile during the clinical course of PIH. Our assay system will aid in such work. Furthermore, the addition of other PIH DBM candidates to the PDA071/039/044 concentrations may predictive accuracy.
In conclusion, we have developed an assay system for PDA071, a candidate PIH DBM; we found it difficult to develop an ELISA. 4 When combined with earlier assays for other DBM candidates (PDA039/044), multiple peptide analysis allowed accurate diagnosis of PIH, and the method will be useful in future clinical studies.
Footnotes
Acknowledgements
The authors are indebted to clinical staff of Yamagata Saisei Hospital and Juntendo Urayasu Hospital for their assistance.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was funded in part by Grants-in-Aid for General Scientific Research from the Ministry of Education, Culture, Sports, Science & Technology, Japan; a grant (A-STEP) from Japan Science and Technology Agency; and Juntendo University Young Investigator Joint Project Award.
Ethical approval
The Ethic Committee board members of Juntendo University and Yamagata Saisei Hospital approved this study (registration #19-017 and #144).
Guarantor
YA.
Contributorship
KH conceived and designed the study, collected data, performed data analysis and interpretation, and wrote the manuscript. YA supervised the study, provided financial support and wrote the manuscript. MY, HI, MB, HY, DN, MS and YM collected data and performed data analysis and interpretation. KTan, KTak, MN, KY, HF and ST conceived and designed the study, provided financial support and contributed to manuscript writing. All authors have given approval to the final version of the manuscript.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
