Abstract
Background
The study aimed to evaluate a fully automated chemiluminescent immunoassay and compared it with a quantitative RNA assay and anti-HCV assay to verify the utility of this automated Ag assay as an alternative method for hepatitis C diagnosis.
Methods
A total of 229 serum samples previously tested for anti-HCV concentrations by the Architect Anti-HCV assay, were selected for HCV RNA testing by real time RT-PCR kit (Shanghai ZJ Bio-Tec Co., Ltd) and 125 specimens were tested for HCV Ag by the Architect HCV core Antigen kit.
Results
The log10 HCVAg and HCV RNA concentrations were highly correlated [r = 0.834); with HCV RNA as the comparator test, HCVAg had 100% specificity, 100% positive predictive value (PPV) and 94.8% sensitivity. We found 1 pg/mL of total HCV core Ag is equivalent to approximately 6607HCV RNA international units (IU)/mL. Receiver operator characteristic curve analysis showed that the area under the curve of HCV core Ag (0.989) was greater than HCV Ab (0.871). HCV Ag concentrations and RNA-to-Ag ratio of the groups for HCV RNA concentrations ≤105 and >105 IU/mL were both significantly different from each other (P < 0.05).
Conclusion
The Architect HCV core Ag assay may be an alternative method for hepatitis C diagnosis, performed on the same analytical platform and sample as the anti-HCV assay, shortening the diagnostic window period, demonstrating good correlation with HCV RNA assay with high specificity and positive predictive value.
Introduction
Hepatitis C virus (HCV), a single-strand, positive-sense RNA virus, causes slowly progressive diseases: chronic hepatitis, liver cirrhosis and hepatocellular carcinoma.1,2 The World Health Organization (WHO) estimates that approximately 170 million people are infected with HCV worldwide, with about 75–80% developing chronic disease.3–5 Virological diagnosis and monitoring of HCV infection are based on the use of serologic assays detecting specific anti-HCV antibodies. 6 However, HCV RNA detection, viral load qualification and HCV-genotyping are also widely used in clinical practice.7,8
The anti-HCV assay is simple, inexpensive and rapid as a screening test, but lacks detection sensitivity in the early window period of 45 to 68 days after infection.9–11 Furthermore, anti-HCV assay is unable to distinguish between active infection and past infection. 9 HCV RNA detection techniques are used to overcome this shortcoming. 12 The HCV RNA assay is a reliable method, which has high sensitivity, specificity and low detection limits and can give quantitative results. 13 However, it involves considerable technical skill, high costs and longer turn-around time and may lead to false positive results due to contamination.4,14 Therefore, the availability of an accurate, simpler and cheaper test would be appealing. Tests for HCV core antigen detection were first reported in the 1990s 15 and the detection effectiveness has improved over the subsequent decades. The majority of the previous assays used enzyme-linked immunosorbent assays (ELISAs) or enzyme immunoassay (EIAs) to detect HCV core Ag. 16 Recently, a fully automated chemiluminescent immunoassay (CLIA) was introduced, which allows for a quantitative determination of HCV core antigen in human serum and plasma. 17
In this study, we aimed to evaluate the CLIA and compare it with a quantitative RNA assay and anti-HCV assay to verify its utility as an alternative test for hepatitis C diagnosis.
Methods
Patients
A total of 229 serum samples were studied, which were derived from the Laboratory diagnosis center of Beijing Tiantan Hospital, Capital Medical University. All samples had been previously tested for anti-HCV concentrations by the Architect anti-HCV assay and were stored at −70℃ prior to analysis. Thereafter, all samples were selected for HCV RNA testing of which 125 specimens were tested for HCV Ag. Of the 229 specimens tested, 208 were anti-human immunodeficiency (HIV) negative (21 samples not tested).
Measurements
The Architect anti-HCV assay
The Architect anti-HCV assay is a two-step immunoassay, using chemiluminescent microparticle immunoassay (CMIA) technology, for the qualitative detection of anti-HCV in human serum and plasma by an Abbott Architect i2000SR analyzer (Abbott Laboratories, Abbott Park, IL, USA) with an HCV Ab assay kit (Abbott Laboratories, Abbott Park, IL, USA). In the first step, sample, recombinant HCV antigen-coated paramagnetic microparticles and assay diluent are combined. Anti-HCV present in the sample binds to the HCV-coated microparticles. After washing, anti-human acridinium-labelled conjugate is added in the second step. Following another wash cycle, Pre-trigger and trigger solutions are added to the reaction mixture. The resulting chemiluminescent reaction is measured as relative light units (RLUs). AS/CO < 1 means ‘not reactive’, and >1 ‘reactive’, where S is the signal of the sample and CO is the cut-off control. The quality control material used was Bio-Rad (California, USA) QC level 1: GBW (E) 090044, level 2: GBW (E) 090045.
The Architect HCV Ag assay
The Architect HCV Ag assay is a two-step immunoassay, using CMIA technology, with flexible assay protocols referred to as Chemiflex, for the quantitative determination of core antigen of HCV by an Abbott Architect i2000SR analyzer (Abbott Laboratories, Abbott Park, IL, USA) with an HCV Ab assay kit (Abbott Laboratories, Abbott Park, IL, USA). In the pre-treatment step, sample, pre-treatment reagent 1 and pre-treatment reagent 2 are combined. An aliquot of the pre-treated sample is aspirated and dispensed into a new reaction vessel. The pre-treated sample, assay-specific diluent and anti-HCV-coated microparticles are combined. HCV Ag present in the pre-treated sample binds to the anti-HCV-coated microparticles in the first step. After washing, acridinium-labelled anti-HCV conjugate is added in the second step. Following another wash cycle, pre-trigger and trigger solutions are added to the reaction mixture. The resulting chemiluminescent reaction is measured as RLUs. The cut-off value is 3.0 fmol/L; HCV Ag concentrations below 3.0 fmol/L were therefore considered nonreactive.
HCV RNA detection and quantification
Quantitative detection of HCV RNA by real-time polymerase chain reaction (RT-PCR) was performed using the HCV real time RT-PCR kit (Shanghai ZJ Bio-Tec Co., Ltd) according to the manufacturer’s instructions on Roche light cycler real-time PCR system (Roche Diagnostics GmbH, Mannheim, Germany). The results were expressed in HCV RNA IU/mL. The lower detection limit is 1 × 103 IU/mL. HCV RNA results were accepted as the gold standard when performing the comparisons. The quality control material purchased from Shanghai ZJ Bio-Tec Co., Ltd (Shanghai, China).
Statistical analysis
Statistical analysis was performed using SPSS 20.0 for Windows (SPSS Inc., Chicago, IL, USA). The correlation coefficients between HCVAg and HCVRNA were calculated by Spearman’s rank test and a comparison between the groups was carried out using the Mann-Whitney U test. A P value of less than 0.05 was considered statistically significant.
Results
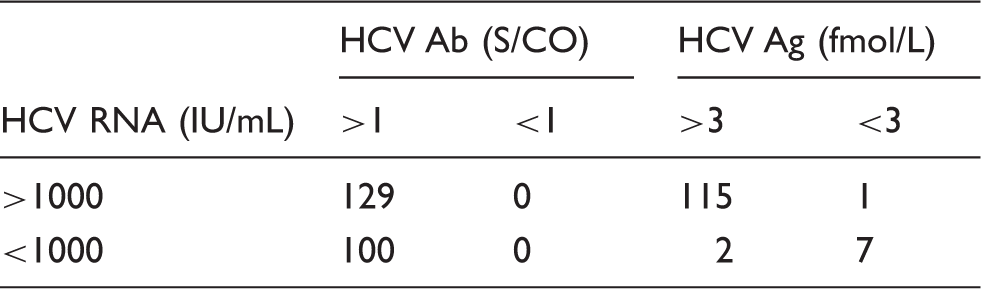
HCV Ab, HCV Ag and HCV RNA test results.
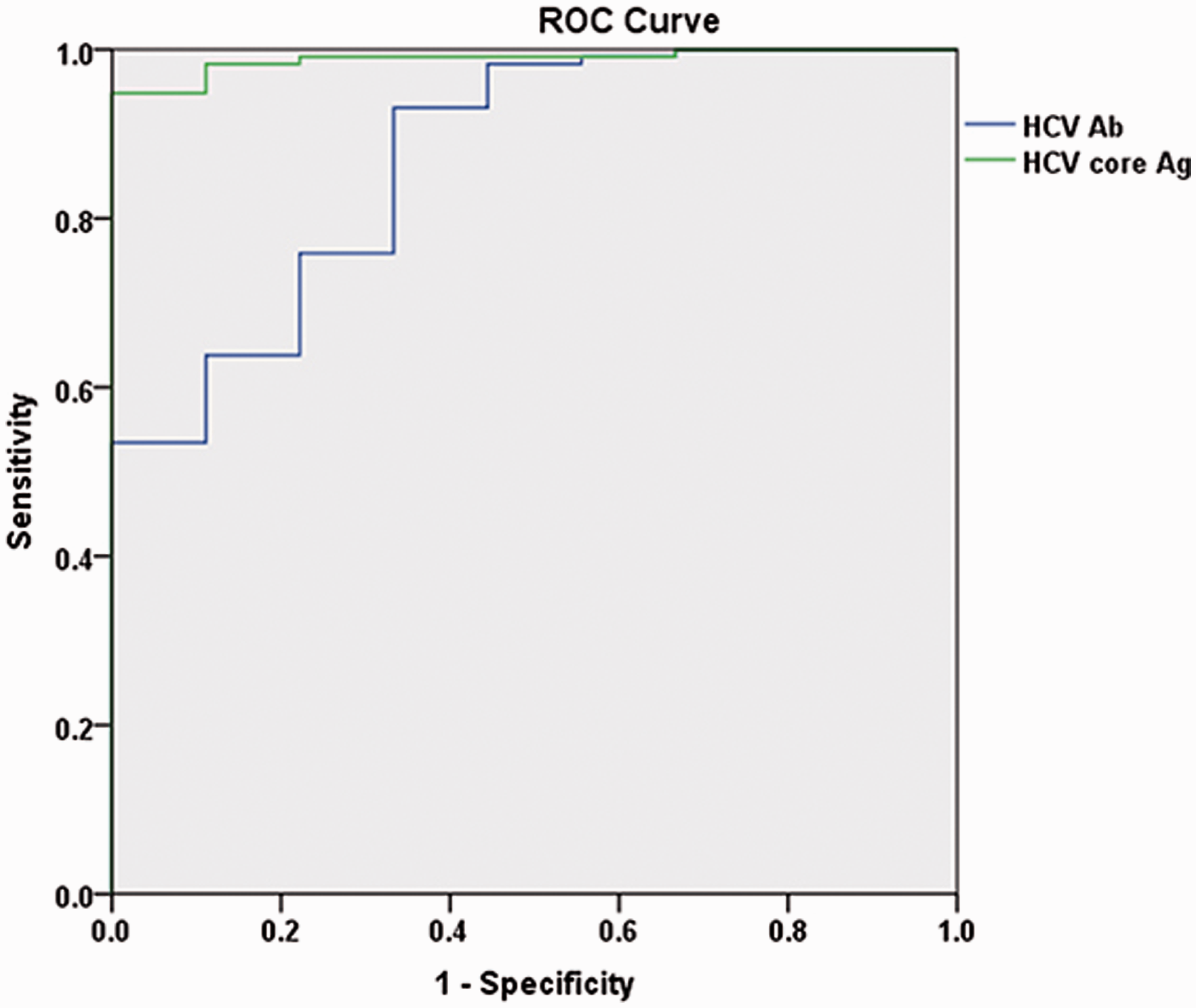
ROC analysis illustrating that the detection efficacy of HCV core Ag was better than HCV Ab for hepatitis C (AUC = 0.989 > 0.871, P < 0.05).
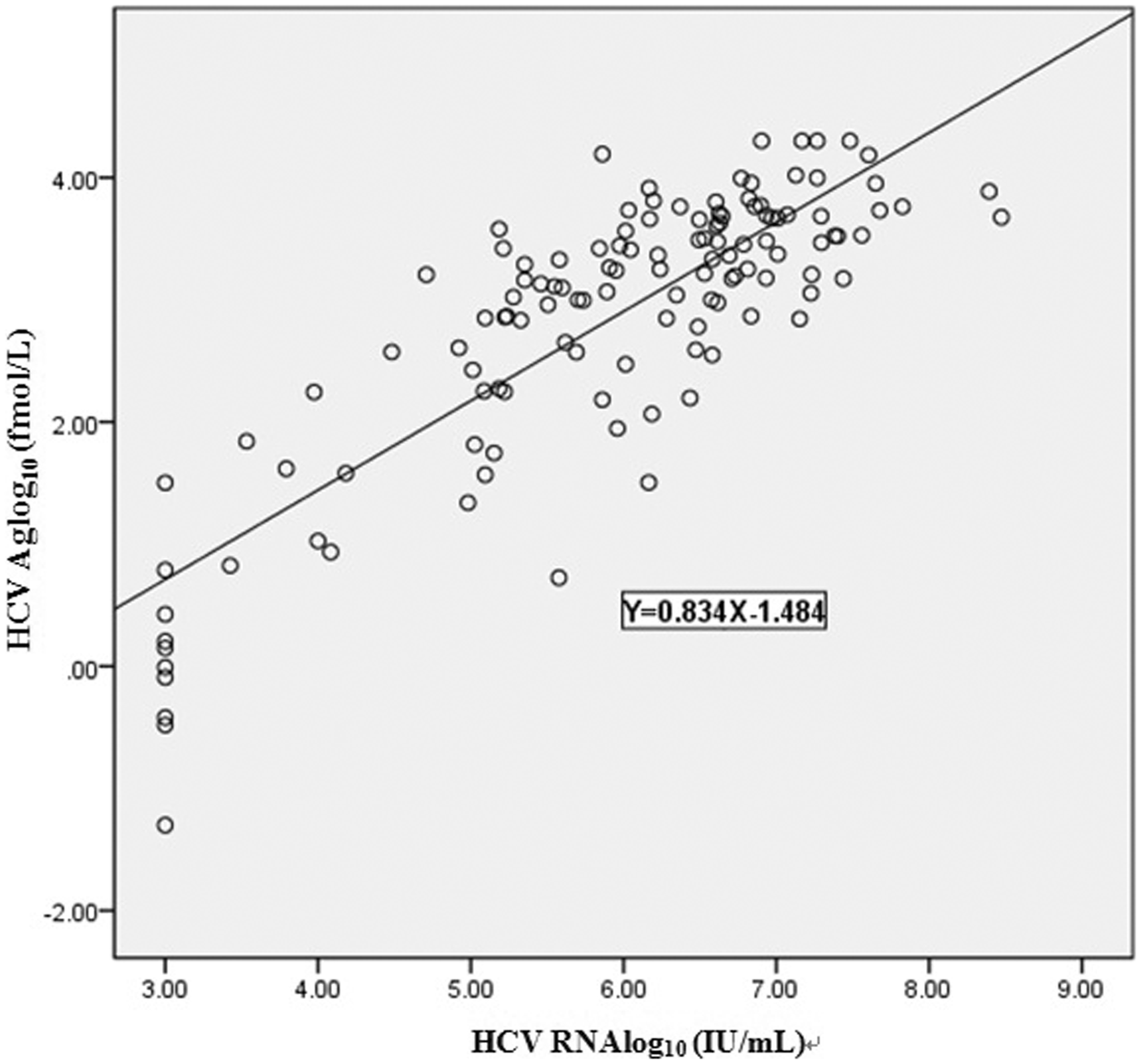
Correlation between HCV Ag and HCV RNA concentrations. The concentrations of HCV Ag were highly correlated with those of HCVRNA by Spearman’s rank correlation test (r = 0.834, P < 0.05).
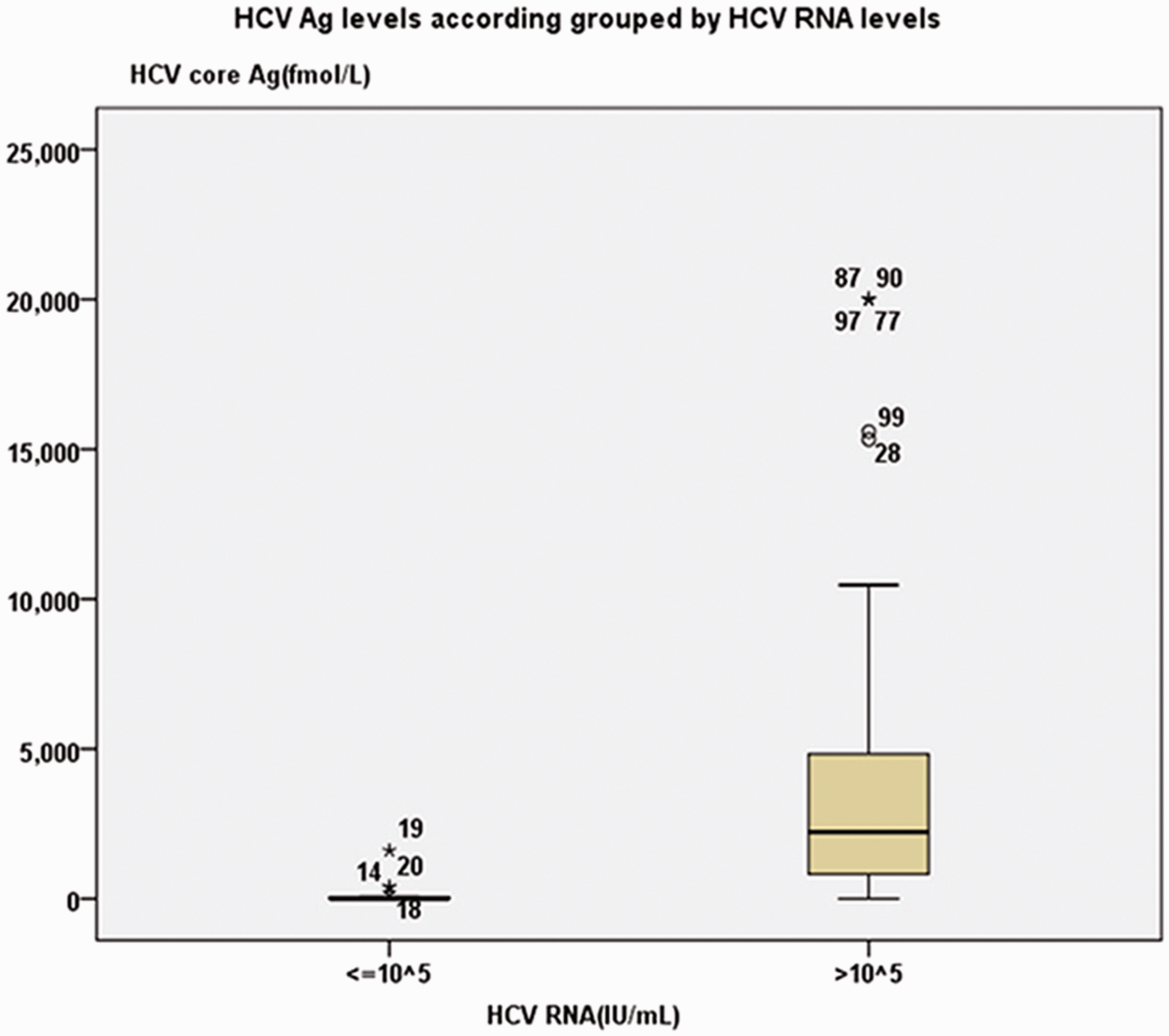
HCV Ag concentrations grouped by HCV RNA concentrations. Median HCV core Ag concentrations were 133.5 and 3694.2 fmol/L for the groups of RNA concentrations ≤105 and >105 IU/mL respectively and there were significant differences between them (P < 0.05).
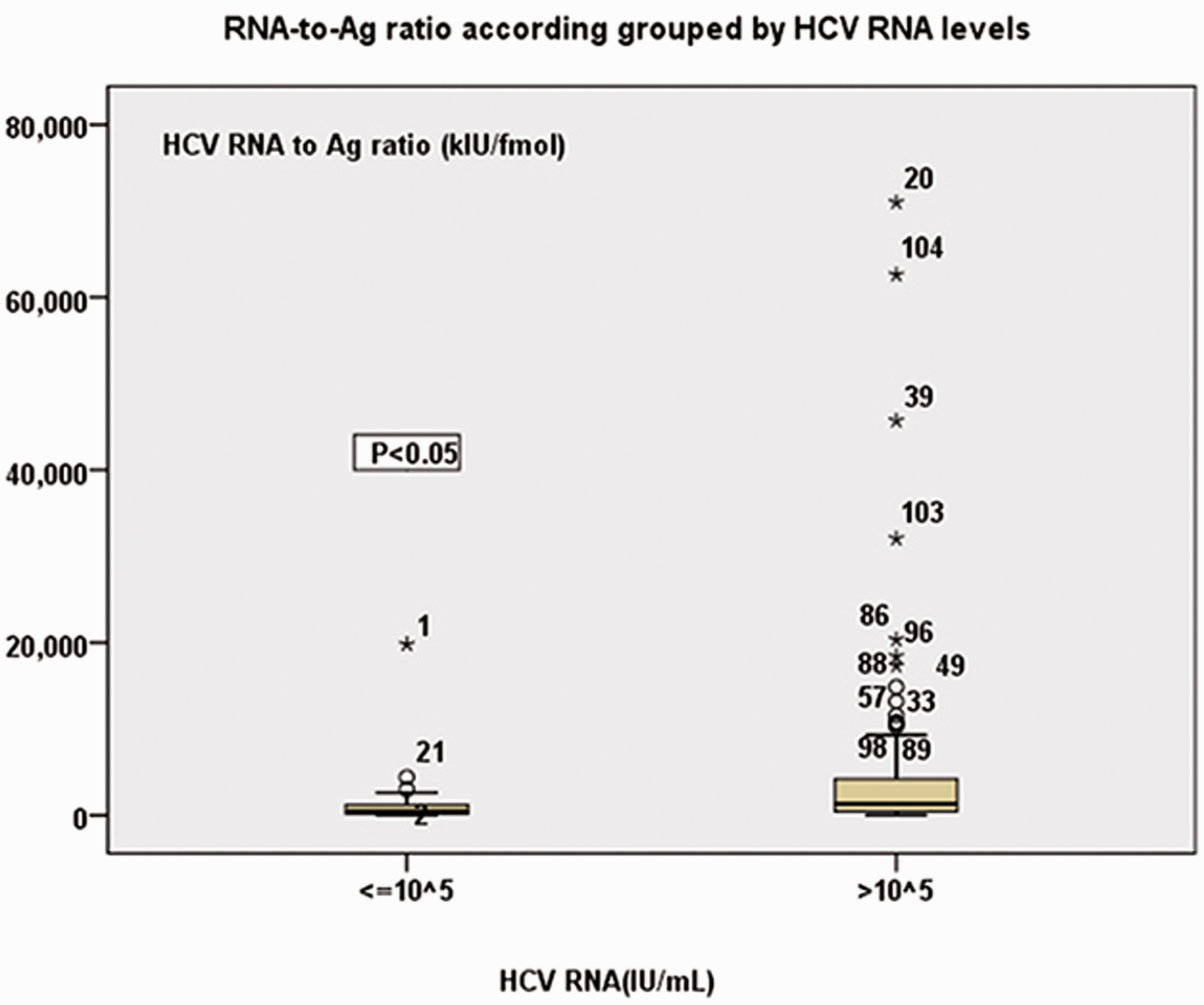
RNA-to-Ag ratio accordingly grouped by HCV RNA concentrations. The RNA-to-Ag ratios of the groups of RNA concentrations ≤105 and >105 IU/mL were significantly different from each other (P < 0.05).
The diagnostic efficacy evaluation of HCV Ag and HCV RNA assay for hepatitis C.
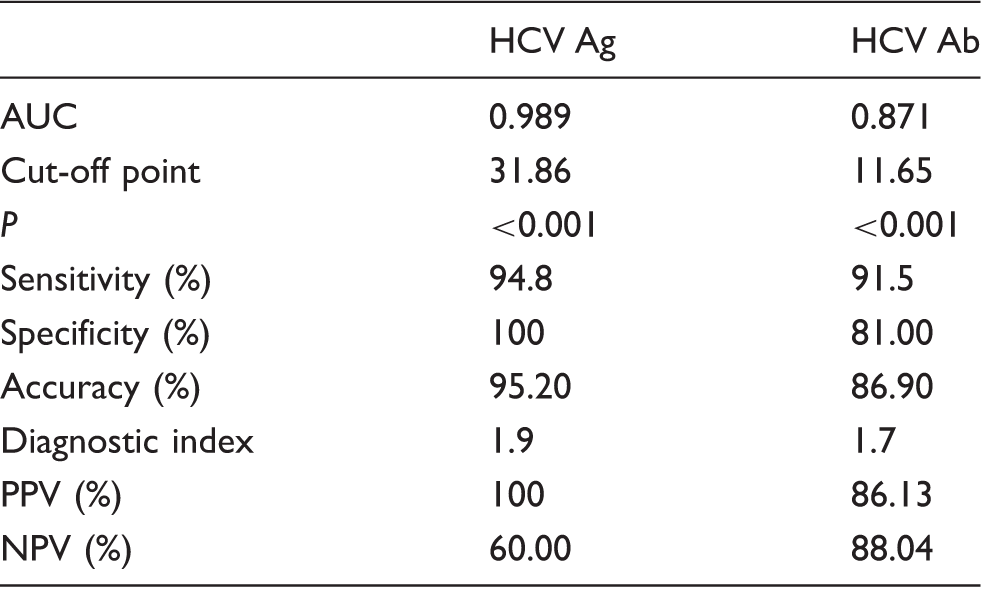
Note: The HCV RNA assay was used as the gold standard.
AUC: area under the curve; PPV: positive predictive value; NPV: negative predictive value
Discussion
Hepatitis C remains a major health care burden globally with a high prevalence in developing countries. 14 Intravenous drug use, blood transfusion and haemodialysis are major risk factors for the spread of HCV.18–20 Thomas et al. 21 have shown that an increasing number of people living with human immunodeficiency virus (HIV) are co-infected with the HCV. HIV co-infection complicates the course of hepatitis C with subjects more likely to develop cirrhosis over a shorter period of time. 22 Furthermore, co-infection with HCV may lead to a more rapid progression to acquired immune-deficiency syndrome (AIDS). 23 There is therefore a need for early HCV diagnosis in HIV patients so that optimum treatment may be offered. Anti-HCV testing is a commonly used screening test for HCV infection, but which is not sensitive and does not discriminate active from past infection. Although HCV RNA testing has been recommended to confirm active infection, its use in clinical practice has been limited by the expensive and time-consuming nature of the assay and the high degree of technical expertise required. For these reasons, HCV core Ag testing has been extensively used, as it was considered to be quick, simple, less expensive and cost-effective. Aghemo et al. 24 demonstrated that measurement of HCV core Ag can be used to monitor the efficacy of direct-acting antiviral (DAA) therapy for HCV infection. Moreover, Florea et al. 9 showed that HCV core Ag assay may diagnose active HCV infection.
The present study found a good correlation between HCV RNA and HCV core Ag concentrations (correlation coefficient = 0.834) which was in keeping with other studies comparing the Architect HCV core antigen test with a range of HCV RNA measurement methods and which found correlation coefficients in the range 0.713 to 0.915.10,25–27 A previous study also found a strong correlation (r = 0.864) between the HCV Ag and HCV RNA as measured by the Qiagen HCV RNA tes (Quiagen, Hilden). 10 Using HCV RNA assay as the gold standard, the receiver operator characteristic (ROC) curve analysis showed the area under the curve of HCV core Ag (0.989) is greater than HCV Ab (0.871) indicating that the detection efficacy of HCV core Ag was better than HCV Ab. The specificity and positive predictive values were both 100% with a sensitivity was 94.82%, in keeping with previous studies.1,10 Our finding of a negative predictive value of 60% (lower than that reported in previous studies) means that negative results should be confirmed by HCV RNA testing.
Our results showed large fluctuations in HCVRNA/core Ag ratio ranging from 31.48 to 70,998.12 kIU/fmol, but increased as the concentration of RNA increased. In the present study, 1 pg/mL of total HCV core Ag is equivalent to approximately 6607 IU/mL RNA. In previous reports, the 1 pg/mL HCV core Ag equivalent has varied between 428 and 8000 IU/mL RNA.6,11,27,28 In one of the two studies it was investigated that HCV genotype was found to have an effect on the HCV RNA/core Ag equivalence value.27,28 Other possible reasons for differences in the equivalence values between different studies may include differences between assays and differences between subjects; this requires further investigation. The present study has several limitations. Firstly, the study was conducted using anti-HCV antibody-positive serum only and we were uncertain whether early HCV infection in antibody-negative subjects could be detected by the HCV core Ag assay. Secondly, the limit of detection of the HCV RNA assay was 1000 IU/mL and lower than that used by other investigators, which may have resulted in differences in sensitivity, specificity, PPV and NPV of the HCV Ag assay.
In summary, the Architect HCV core Ag assay showed a good correlation between the logarithmic values of HCV RNA and HCV Ag (r = 0.834), with 100% specificity and PPV, and 94.82% sensitivity. Moreover, HCV core Ag testing can use the same platform and sample with anti-HCV assay, which is simpler, faster and more convenient than the HCV RNA tests.
Footnotes
Acknowledgement
Lijuan Wang thanks Hong Lv and Guojun Zhang who have supported the author with sincere gratitude.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Ethical approval
Not required.
Guarantor
GZ.
Contributorship
All three authors contributed to the design of the study. LW researched the literature, wrote the first draft of the paper and edited the final manuscript. GZ was responsible for the overall design of experiments and performed all statistical analyses. HL was responsible for the initial data acquisition and also contributed to the first draft of the paper. All authors agreed with the final draft of the manuscript.
