Abstract
Background
Recently, several studies have shown that microRNAs are present in high-density lipoprotein, and high-density lipoprotein-microRNA may be a promising disease biomarker. We investigated the stability of high-density lipoprotein-microRNAs in different storage conditions as this is an important issue for its application to the field of clinical research.
Methods
microRNAs were extracted from the high-density lipoprotein fraction that was purified from the serum. miR-135 a and miR-223, which are known to be present in high-density lipoprotein, were quantified by quantitative real-time PCR. The influence of preanalytical parameters on the analysis of high-density lipoprotein-miRNAs was examined by the effect of RNase, storage conditions, and freezing and thawing.
Results
The concentrations of microRNA in high-density lipoprotein were not altered by RNase A treatment (0–100 U/mL). No significant change in these microRNAs was observed after storing serum at room temperature or 4℃ for 0–24 h, and there was a similar result in the cryopreservation for up to two weeks. Also, high-density lipoprotein-microRNAs were stable for, at least, up to five freeze–thaw cycles.
Conclusions
These results demonstrated that high-density lipoprotein-microRNAs are relatively resistant to various storage conditions. This study provides new and important information on the stability of high-density lipoprotein-microRNAs.
Keywords
Introduction
MicroRNAs (miRNAs) are small non-coding RNAs that play a pivotal role in the regulation of gene expression at the post-transcriptional level. 1 In humans, it is estimated that at least 30% of genes are regulated by miRNAs, and miRNA-mediated gene silencing pathways play important roles in cell functions such as differentiation. 2
Recent reports demonstrated that serum/plasma miRNAs, which are present in exosomes, microvesicles (MVs), high-density lipoprotein (HDL), etc., circulate and that such circulating miRNAs are stable in the bloodstream.3,4 Dysregulation of serum/plasma miRNA expression may be involved in various human disorders.5–7 It has been already demonstrated that the alteration of the serum/plasma miRNA expression profile is associated with some cancers in humans.8–10
According to some recent reports, not only serum/plasma miRNAs but also miRNAs in exosomes and MVs (parts of serum/plasma) could be unique biomarkers.11–14 For example, some MVs-miRNAs, which are abundant in gliomas, were easily detected from patients, suggesting that miRNAs in MVs may provide information about disease state. Exosomal miRNAs also reflect the progression of ovary and lung cancer, indicating their utility for diagnosis. More recently, it was shown that miRNA profiles in HDL are different from those in exosomes, and the human HDL-miRNA profile of normal subjects is significantly different from that of familial hypercholesterolaemia. 4 In addition, Wagner et al. reported that the concentrations of several miRNAs in HDL isolated from patients with an acute coronary syndrome are significantly different compared with healthy subjects. 15 Thus, in some cases, it is important for diagnosis to analyse exosomal, MVs and HDL-miRNA separately from serum/plasma miRNAs. Also, the amount of HDL-miRNA is quite low compared with the amount of serum/plasma miRNA. Thus, the significant alteration of HDL-miRNA expression profile may be masked using a serum/plasma miRNA sample. For the adequate quantification of HDL-miRNA, it may be necessary to analyse purified HDL-miRNA from serum/plasma.
The stability of circulating miRNA.
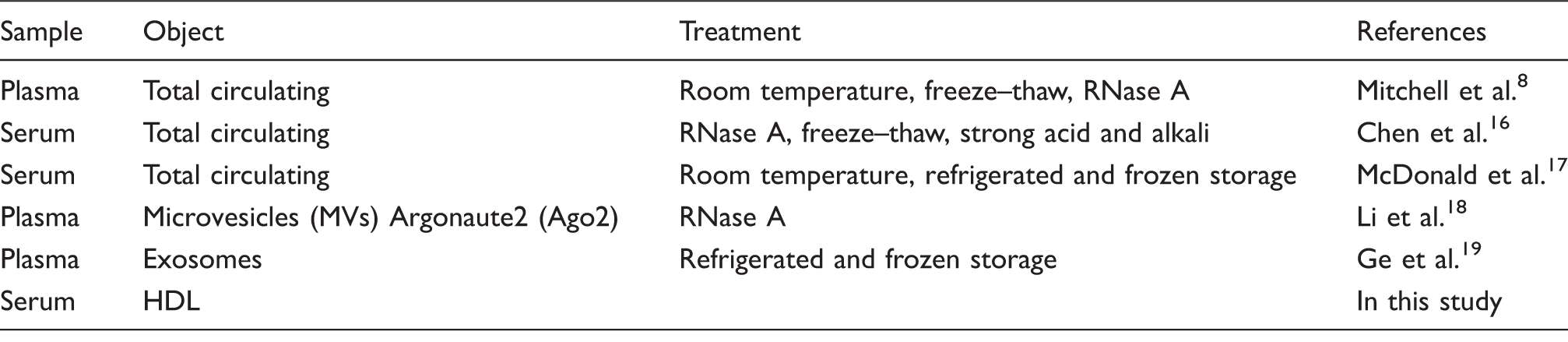

HDL: high-density lipoprotein.
Materials and methods
Sample collection
Blood from a healthy non-fasting male volunteer was taken, in the morning, into a serum separator tube (Terumo Co., Ltd, Tokyo, Japan). Serum was separated by centrifugation at 1700 g for 10 min at 4℃. The pooled serum was prepared from the serum samples. Written informed consent was obtained from the volunteer. The study protocol was approved by Ethics Committee of Fujita Health University (no. 11-101).
HDL purification
The HDL fraction was highly purified from human pooled serum using a four-step protocol consisting of an ultracentrifugation method (368,000 g, 3 h, 4℃) using Optima MAX-X (Beckman Coulter, Inc., Brea, CA, USA) to remove the chylomicron and VLDL fractions, a polyethylene glycol precipitation method to remove the LDL fraction, 20 an exosome precipitation method using ExoQuick (System Biosciences, Mountain View, CA, USA), and a fast protein liquid chromatography system (FPLC; GE Healthcare, Amersham, UK) was used to remove other proteins. Superose 6 10/300GL column was used with EDTA-free FPLC buffer (10 mM Tris-HCl, 150 mM NaCl and 0.02% sodium azide, pH 8.0). FPLC analysis was performed at room temperature using two P-500 pumps, a ValveV-7 multi-injector with a 500 μL loop, an optical unit monitor UV-1, a FRAC-200 fraction collector and a GP-250 Plus controller. The flow rate was 0.5 mL/min, and 500 μL of the sample was injected. All operations were monitored by absorbance at 280 nm. Each obtained HDL fraction was measured for cholesterol concentration using the Cholesterol E-Test Wako Kit (Wako Pure Chemical, Osaka, Japan). The protein concentration in the HDL fractions was also measured in duplicate using the Protein Assay Rapid Kit Wako (Wako Pure Chemical, Osaka, Japan) to confirm the purification efficiency of HDL. These items were measured in accordance with the manufacturer's instructions.
Western blot assay
For sodium dodecyl sulphate-polyacrylamide gel electrophoresis (SDS-PAGE), the aliquot of the HDL fraction was solubilized in 2× Laemmli buffer and electrophoresed in a 10% polyacrylamide gel using AE-7350/W CompactPAGE (Atto Corporation, Tokyo, Japan). To detect apolipoprotein A-I (apoA-I) and CD63, markers of HDL and exosome, these proteins were transferred to a PVDF membrane using AE-7500 CompactBlot (Atto Corporation, Tokyo, Japan). This membrane was divided into two membranes using scissors, and then these were blocked with Can Get Signal (Toyobo Co., Ltd., Osaka, Japan). All antibodies were purchased from Abcam (Cambridge, UK). Western blotting was performed using an apoA-I antibody (ad17278) or a CD63 antibody (ab118307) diluted at 1 μg/mL overnight at 4℃, followed by 1 h incubation with a horseradish peroxidase-conjugated secondary antibody (ab99603 or ab97085) diluted at 1:2000, respectively. Protein bands were detected using SuperSignal West Pico Chemiluminescent Substrate (Thermo Scientific, Rockford, IL, USA) and visualized using an ImageQuant LAS 4000mini imager (GE Healthcare, Amersham, UK).
MiRNA extraction
We isolated HDL-miRNAs using TRIzol reagent (Life Technologies, Carlsbad, CA, USA), in accordance with the manufacturer's instructions. Protein was denatured in 900 μL of TRIzol reagent to 300 μL of HDL solution, vortexed immediately for 15 s, then incubated at room temperature for 5 min. Aqueous and organic phases were separated by adding 240 μL of molecular grade chloroform, vortexing at the maximum setting for 5 s, followed by centrifugation at 12,000 g for 15 min at 4℃. The aqueous phase was rapidly transferred to a new tube. To precipitate RNA, 20 μg of RNase-free glycogen (Roche, San Francisco, CA, USA) and 600 μL of 100% isopropanol were added to the aqueous phase, and the mixture was incubated at room temperature for 10 min, then centrifuged at 12,000 g for 10 min at 4℃. Next, the supernatant was removed from the tube, leaving only the RNA pellet and the RNA wash procedure continued. Finally, the total RNA was dissolved in 10 μL of RNase-free water and stored at −80℃ until further use.
Target miRNAs
The miR-135a and miR-223 that were reported to exist in HDL were measured by the quantitative real-time PCR (qRT-PCR) method. MiR-153a is most present in the HDL of healthy subjects. miR-223 is the fourth largest in that, and is increased dramatically in the HDL of familial hypercholesterolaemia patients. These primers were purchased from Qiagen (Valencia, CA, USA).
MiRNA qRT-PCR analysis
MiRNAs analysis was carried out using a miScript PCR System (Qiagen, Valencia, CA, USA) that included specific primers for miRNAs, and following the manufacturer's instructions. 21 The reverse transcription (RT) reaction system contained 1 μL of miScript Reverse Transcriptase Mix, 2 μL of 5× miScript HiFlex Buffer and 1 μL of 10× Nucleics Mix; samples were allowed to react for 60 min at 37℃, and 5 min at 95℃ in the 2720 Thermal Cycler (Applied Biosystem, Foster City, CA, USA), and then diluted with 10 μL of RNase free Tris-EDTA buffer (pH 7.4). The reaction system of qRT-PCR contained 5 μL of SYBR Green PCR Master Mix (Qiagen, Valencia, CA, USA), 1 μL of miScript universal primer, 1 μL of specific primer, 2 μL of cDNA and 1 μL of RNase-free water; PCR cycles were completed at 95℃ for 15 min, 40 cycles at 94℃ for 15 s, 55℃ for 30 s and 70℃ for 30 s. qRT-PCR was performed with an ABI PRISM 7900HT Sequence Detection System (Applied Biosystems, Foster City, CA, USA).
RNase A treatment of HDL fraction
The HDL fraction was purified by the above mentioned method, and this fraction was treated with 10, 50 and 100 U/mL RNase A (Macherey-Nagel GmbH & Co., KG, Dueren, Germany) for 15 min at 37℃. Furthermore, it was treated to degrade by severe conditions in the following ways: 0.5% Triton X-100 for 2 min, 20 μg/mL Proteinase K (Takara Bio Inc., Shiga, Japan) for 15 min, then the addition of 10 M NaOH and the incubation for 1 h at 100℃. miRNAs extracted from these samples were measured by the qRT-PCR method.
Stability of HDL-miRNAs
We purified the HDL fraction from serum immediately after centrifugation of the blood; this aliquot served as the 0 h control. The remainders of the serum were maintained at room temperature or refrigeration (4℃) for 3, 6, 12 or 24 h. Those were also stored at −20℃ or −80℃ for 1 week or 2 weeks. Then, the HDL fractions were purified from them. To study the effect of freeze–thaw cycles, serum samples frozen at −20℃ were subjected to a total of five freeze–thaw cycles, and then the HDL fractions were purified from the serum of 1, 3 and 5 freeze–thaw cycles. These independent experiments were repeated three times. The extracted miRNAs were measured by a qRT-PCR analysis that was performed in duplicate.
Statistical analysis
The resulting miRNA concentrations are reported as raw cycle threshold (Ct) values and presented as mean ± standard error (SE) for three independent experiments. Statistical calculations for this study were carried out using a software package ‘Prism4’ purchased from GraphPad Software (San Diego, CA, USA). We calculated the miRNAs stability using a repeated-measures analysis of variance test to compare the miRNA concentrations at various time points (room temperature, 4℃, −20℃, −80℃ and freeze–thaw cycle). A P value <0.05 was considered statistically significant.
Results
Purification of HDL fraction
The FPLC profile of the purified HDL is shown in Figure 1(a). A large absorbance peak at 280 nm has been observed in this profile, and the HDL fractions were further estimated from the distribution of total cholesterol in the FPLC fractions. Consequently, we collected from 33 to 42 fractions and pooled them together. Next, apoA-I and CD63 in the pooled HDL fraction were detected using Western blotting. ApoA-I, that is a marker protein of HDL, was detected in this fraction, but exosomal protein marker CD63 was negative (Figure 1(b)). Therefore, this sample was regarded as the purified HDL fraction. The Ct values of miR-135a and miR-223 were 33.1 ± 0.63 and 33.7 ± 0.60 (mean ± SE), respectively (Figure 1(c)). From these results, it was confirmed that the miRNAs exist in the purified HDL fraction.
HDL-miRNA analysis. (a) The FPLC distribution of cholesterol and protein concentration in HDL fractions (left axis; mg/dL, Right axis; mAU, milli-absorbance unit). (b) The detection of apoA-I and CD63 by Western blot. (c) The concentrations of miR-135a and miR-223 in the purified HDL fraction. The resulting miRNA concentrations are reported as raw Ct values. Vertical axis represents Ct value.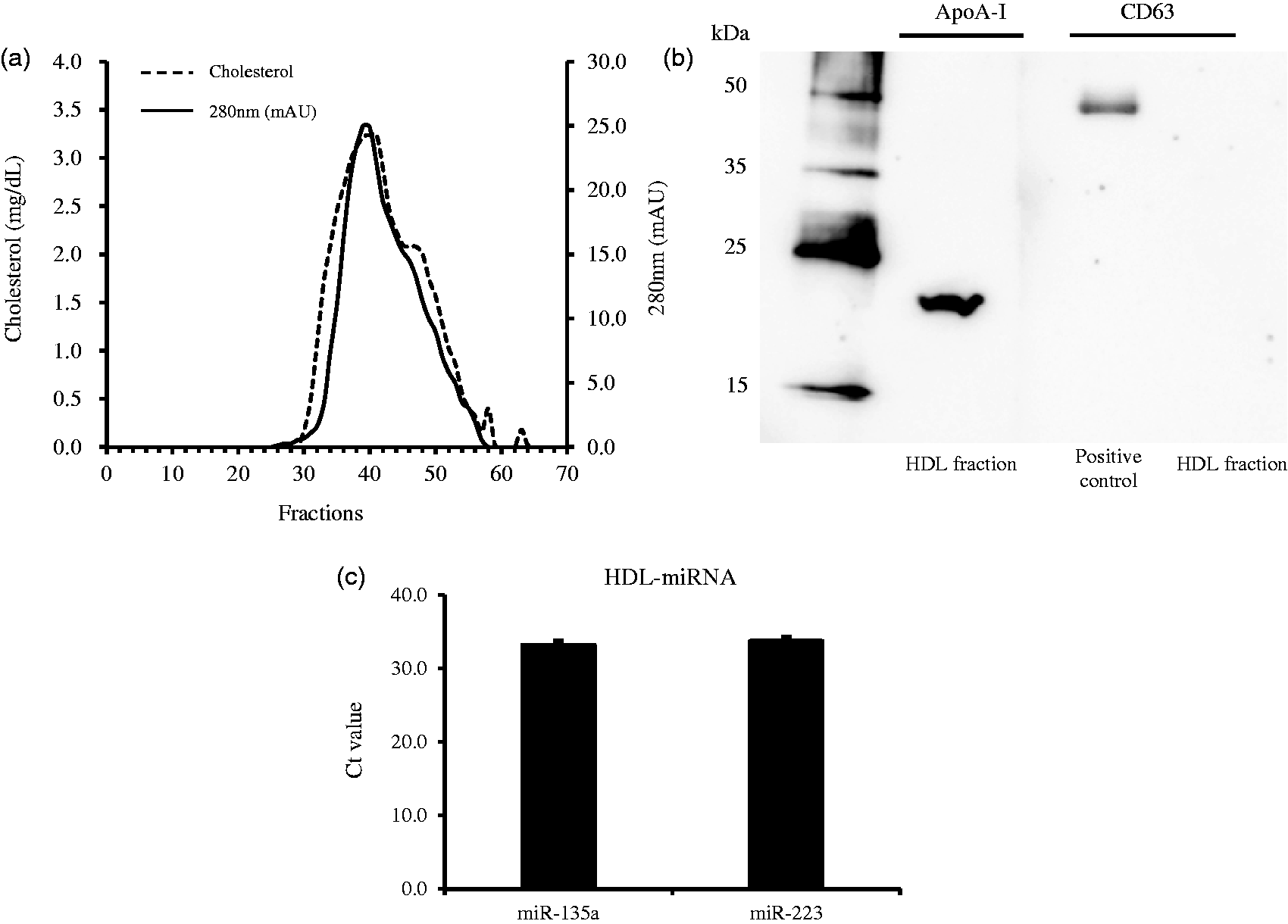
Effect of RNase A on HDL-miRNAs
To characterize the stability of miRNAs in HDL, we investigated the stability of miRNAs in the purified HDL fraction against RNase A and other severe conditions, such as extreme temperature and pH. We found that when the purified HDL fraction was treated with 0–100 U/mL RNase A for 15 min, miR-135a and miR-223 in HDL proved to be quite stable (Figure 2(a) and (b)). Next, we investigated whether the HDL-miRNAs are stable from the degradation of HDL; when we destroyed the HDL in the above severe conditions, miR-135a and miR-223 were not detected in this sample (Figure 2(c) and (d)).
Resistance of HDL-miRNA to degradation by RNase A and the degradation of HDL-miRNA under severe conditions. Results are reported as raw CT values. (a) miR-135a, (b) miR-223, (c)miR-135a and (d) miR-223. n.d.: not detected.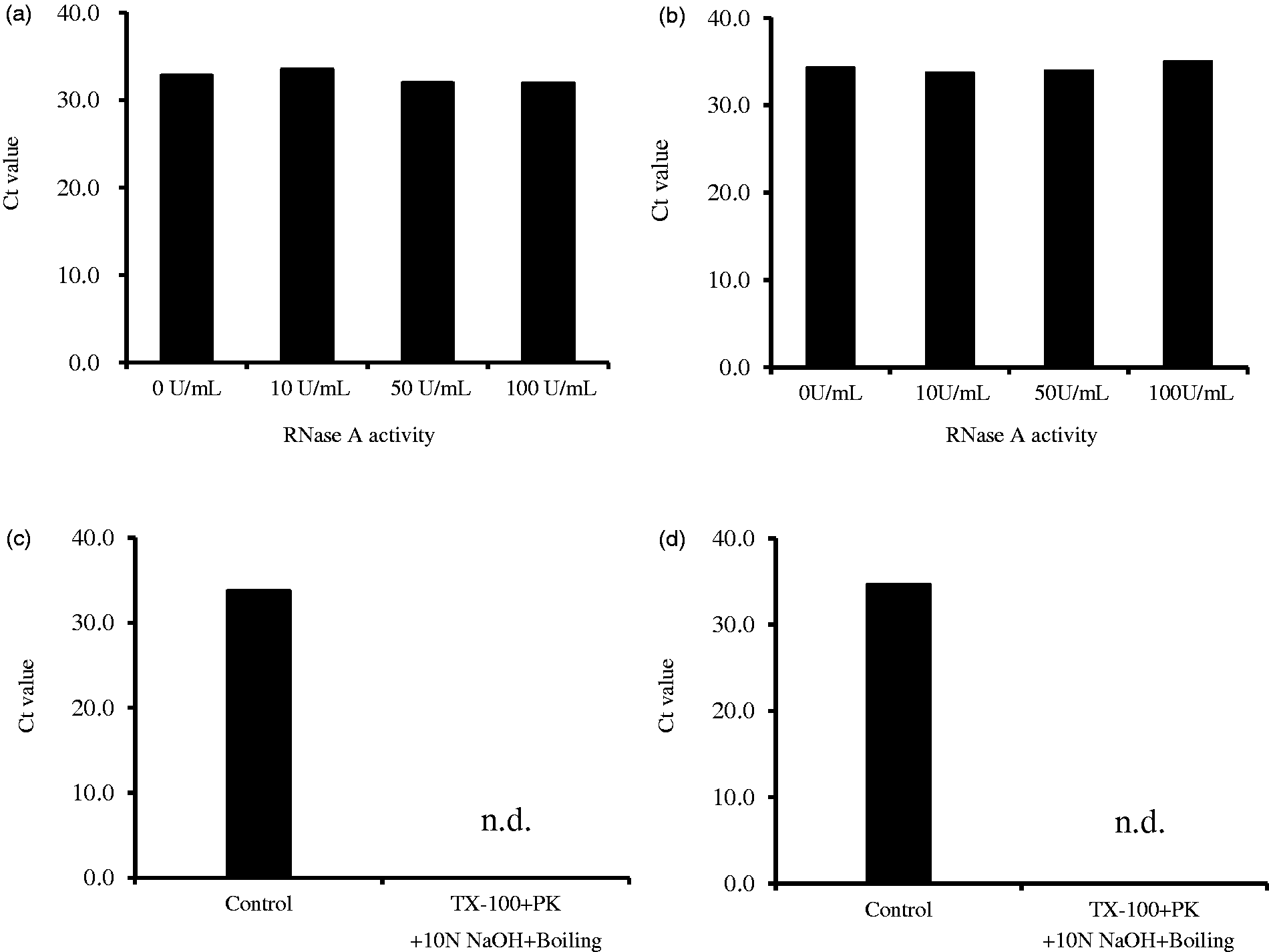
Stability of HDL-miRNAs in various storage conditions
To evaluate the influence of differential storage temperature and time on HDL-miRNAs stability, the freshly isolated serum from a healthy volunteer was stored at room temperature or refrigeration (4℃), and the HDL fraction was separated from the stored serum after 0, 3, 6, 12 or 24 h. After HDL purification, the protein concentration in the HDL fraction was measured. The protein concentrations of room temperature storage and cold storage were 0.93 ± 0.04 mg/mL and 0.98 ± 0.02 mg/mL, respectively. Furthermore, the protein concentrations of frozen storage at −20℃ and −80℃ were 0.90 ± 0.05 mg/mL and 0.92 ± 0.07 mg/mL, respectively. The results of the protein concentration revealed that the protein was largely not lost in the purification process of the HDL fraction.
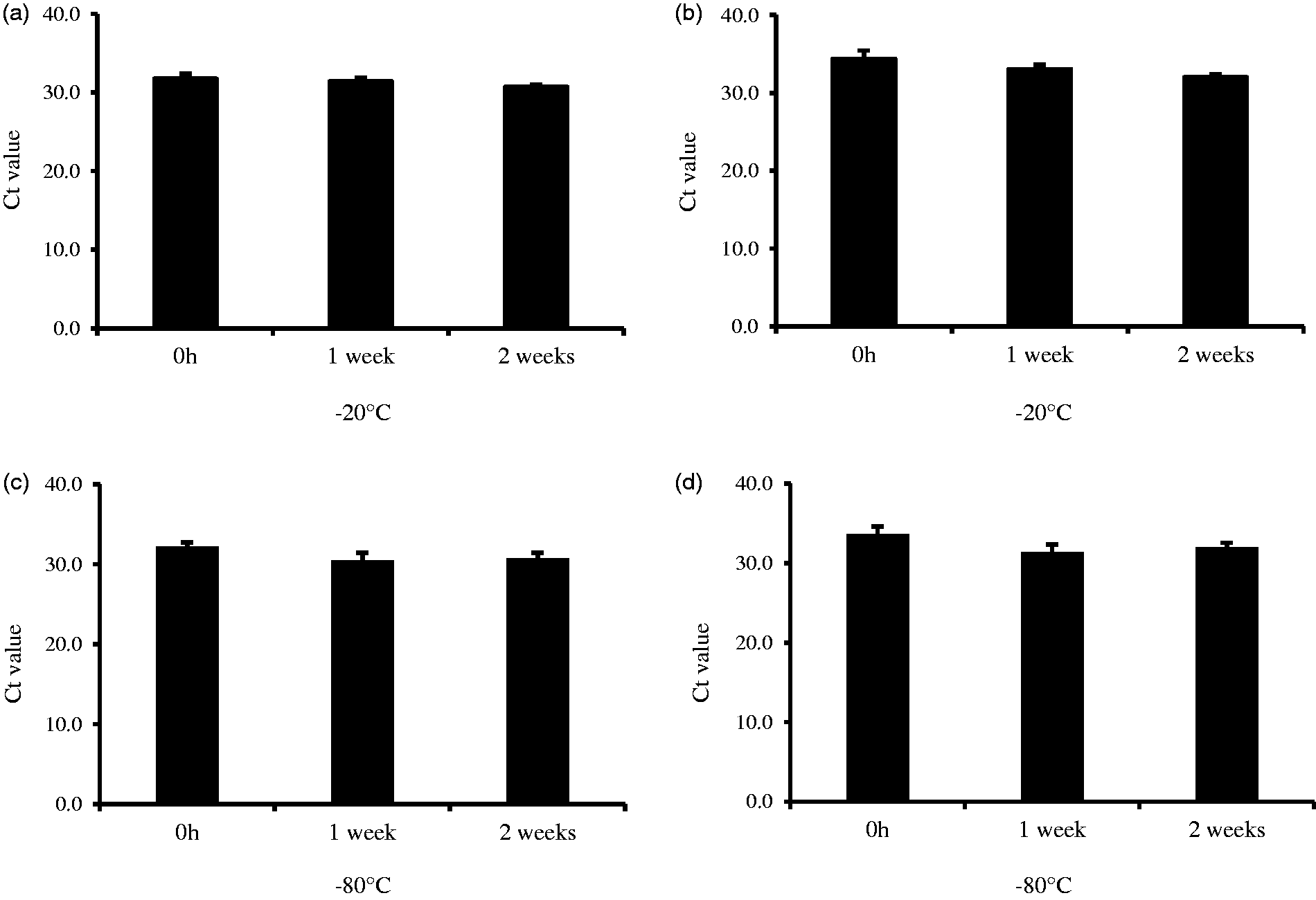
The Ct values of HDL-miR-135a and miR-223 at the storage of room temperature and refrigeration contain some variability for up to 24 h. However, the storage times were found to have no significant differences between them (Figure 3). Similarly, even if serum was stored frozen for up to two weeks at −20℃ or −80℃, there was no significant change in the Ct values of the HDL-miRNA (Figure 4).
Stability for HDL-miR-135a and HDL-miR-223 with storage at room temperature and under refrigeration. Stability for HDL-miR-135a and HDL-miR-223 during frozen storage (−20℃ and −80℃).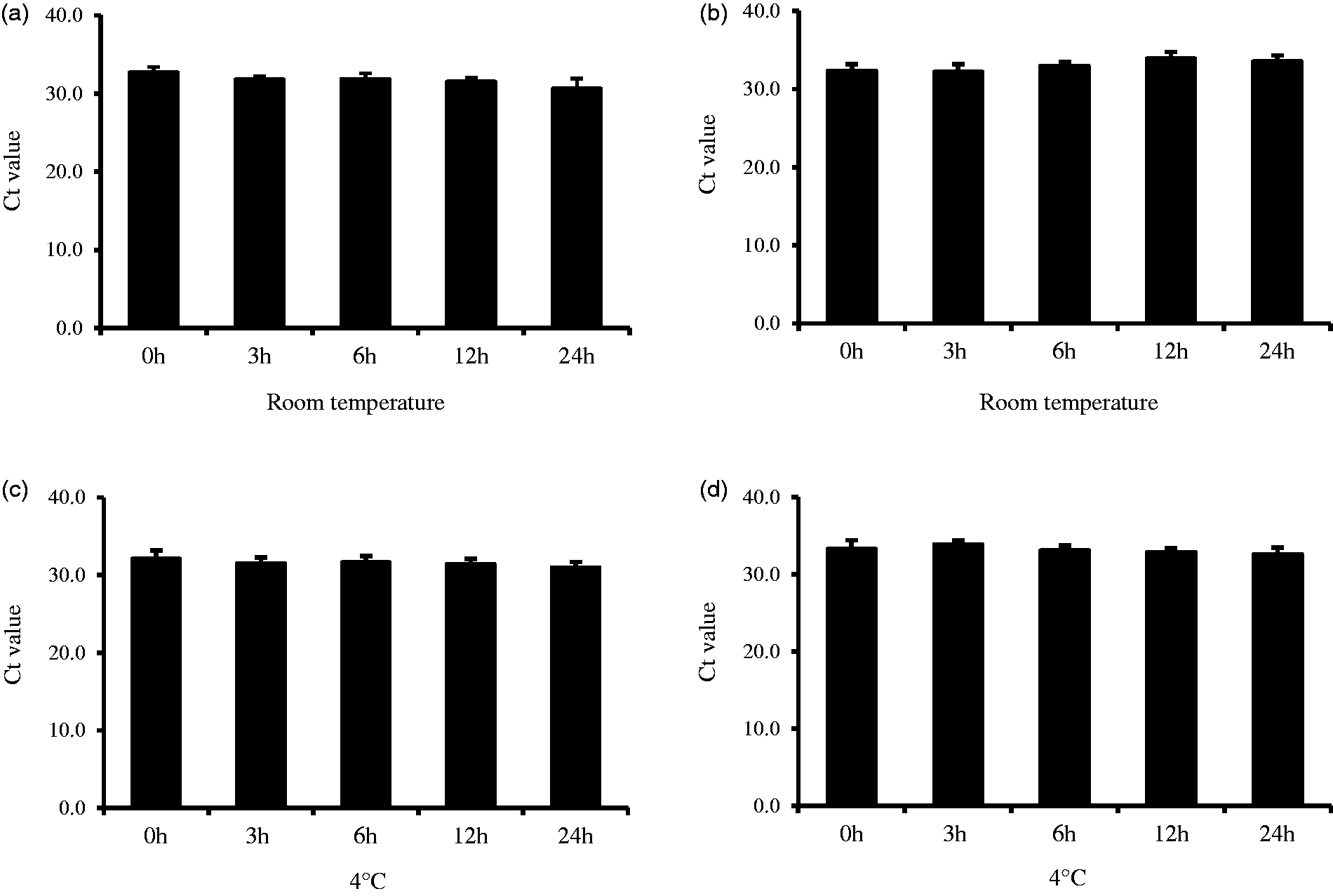
Effect of repeated freeze–thaw cycles
Serum samples were subjected to assess the influence of repeated freeze–thaw cycles on HDL-miRNAs stability. After the purification process of the HDL fraction, the protein concentration was 0.96 ± 0.08 mg/mL. The HDL fraction could be stably purified in freeze–thaw for up to five times. Since the Ct values of HDL-miRNA have no significant change among the number of freeze–thaw cycles, the HDL-miRNAs were stable for at least up to five freeze–thaw cycles (Figure 5).
Stability for HDL-miR-135a and HDL-miR-223 for up to five freeze–thaw cycles.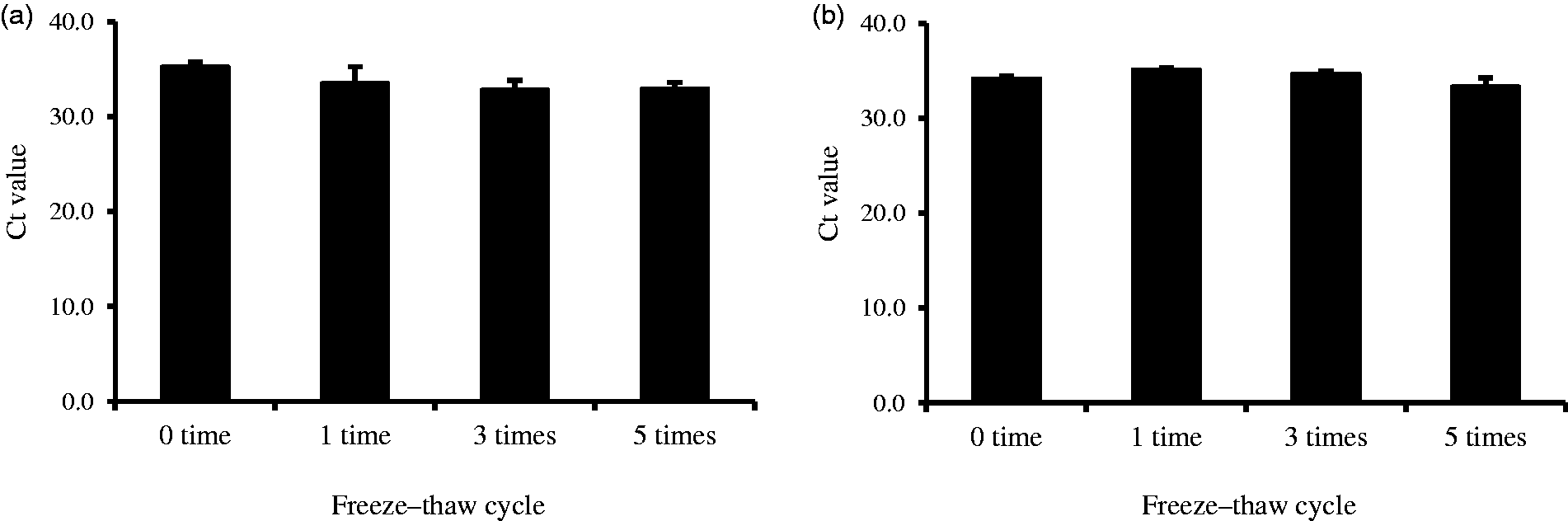
Discussion
Here, we demonstrated that the miRNAs in HDL are stable under different sample storage conditions. Specifically, the results of this study have clarified that HDL-miRNAs are stable in the RNase A treatment and the various storage conditions of the serum sample, such as room temperature, refrigeration, frozen storage and repeated freeze–thaw cycles.
The main function of HDL is to remove excess cholesterol from peripheral tissues and return it to the liver for storage and excretion. HDL mainly participates in the regulation of the cholesterol level in the body. In addition, HDL also has a significant role in anti-inflammatory action, an antioxidative effect and improves endothelial function.22,23 Tabet et al. 24 reported that the anti-inflammatory action of HDL is mediated by the transfer of miR-223 to endothelial cells. 24 Given these reports, further research will clarify the association between the level of HDL-miRNAs and the various diseases. Along with this, the analysis of HDL-miRNAs profile would become more important for estimating the onset of various diseases. Taking into consideration the high stability of HDL-miRNA, its availability may depend on the turnover of HDL in vivo. Additionally, we showed that HDL-miRNA is resistant to frozen storage and repeated freeze–thaw cycles. Thus, regarding the analysis of HDL-miRNAs, we can store serum samples until use. Cuhadar et al. reported that HDL-cholesterol shows adequate stability after 3 months of storage in serum samples at −20℃ or up to 10 times of freeze–thaw cycle. 25 Therefore, we suggest that miRNAs encapsulated in HDL are stable as with cholesterol in HDL.
For the clinical application of HDL-miRNA, it is necessary to quantify the miRNAs after purifying HDL because blood cells contain some miRNAs which are enriched in HDL. Otherwise, the change of HDL-miRNA may be masked. Indeed, miR-223, which exists widely in HDL, is also expressed in macrophages. Another key point for clinical application is to develop a purification method for HDL-miRNA. Currently, this method is not simple enough to apply to laboratory examinations. In this study, we used this conventional method, which is widely adopted for HDL purification but is time-consuming and cumbersome. Therefore, it is necessary to establish a simpler HDL purification method. Also, the evaluation method of HDL-miRNAs has not been determined. The evaluation issues of HDL-miRNAs will become a major obstacle in preparing the profile of HDL-miRNAs for various diseases. For example, miRNA concentrations in the tissue are often quantified by U6B, a small nuclear RNA, which is constantly expressed in the cell as an internal standard.26–28 Since HDL varies by various pathological conditions, it is necessary to consider the internal standard substance in HDL. To establish a quantitative analysis of HDL-miRNAs using an external and internal standard method is a future research challenge.
Although many miRNAs exist in HDL, 4 we have not confirmed the stability about all those miRNA in this study. However, it has been reported that miRNAs encapsulated in exosomes or MVs are stable to a certain degree.29–31 Similarly, we infer that the miRNAs existed in HDL are also stable.
In conclusion, this study provides new and important information on the stability of HDL-miRNAs. Given the fact that the HDL contains various miRNAs, some miRNA might be sensitive to storage. Although we analysed the stability of representative HDL-miRNAs (miR-223 and miR-135), it is considered that most HDL-miRNAs appear to be stable, taking into consideration the fact that no degradation of miR-223 and miR-135 was observed under various sample storage conditions and addition of RNase. Therefore, we conclude that HDL-miRNAs are extremely stable and can be used in various experiments.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This research was funded by Tsukuba Medical Laboratory of Education and Research (TMER-25.1.05).
Ethical approval
The ethics Committee of Fujita Health University approved the protocol of this study (no. 11-101).
Guarantor
HI.
Contributorship
HI researched literature and wrote the first draft of the manuscript. HI, HY and RT gained ethical approval and conceived this study. NT and KK purified HDL from serum and examined the stability of HDL-miRNA. AN and MY extracted the miRNA and measured it. KO and EM performed the degradation experiment of HDL-miRNA. YA prepared the pooled serum. KS was involved in the data analysis. All authors reviewed and edited the manuscript and approved the final version of the manuscript.
