Abstract
Background
The worldwide incidence of rheumatic heart disease is at least 15.6 million cases and is responsible for around 233,000 deaths per year. The pathogenesis of rheumatic heart disease is complex and involves genetic factors that predispose a person to the development of autoimmune reactions. Although Arg753Gln polymorphism of the TLR2 gene was considered to be related to acute rheumatic fever susceptibility in child, two groups have identified that there were no relations between this Arg753Gln polymorphism and rheumatic heart disease susceptibility.
Methods
In this study, we scanned the full length 826 bp of 3′ untranslated region of TLR2 in a Chinese-Han population and found that the rare allele G of rs35514500 was highly related to rheumatic heart disease.
Results
Results of dual-luciferase assay indicated that this T->G variation weakens the interaction between miR-340 and TLR2 3′ untranslated region and up-regulated firefly luciferase expression. Further results about the patients’ serum cytokines concentration detection constructed a relationship between G allele of rs35514550 and up-regulated serum IL-6 and TNFα.
Conclusions
These findings may give insight into understanding of rheumatic heart disease development and create an opportunity to approach the diagnosis and treatment of rheumatic heart disease.
Introduction
Rheumatic fever (RF) is caused by the Gram-positive bacteria Streptococcus pyogenes and follows a non-treated throat infection in susceptible children and teenagers (3–19 years old). The incidence of acute RF (ARF) in some developing countries exceeds 50 per 100,000 children. 1 The worldwide incidence of RHD is at least 15.6 million cases and is responsible for around 233,000 deaths/year. 2 Rheumatic heart disease (RHD), the most serious complication, occurs in 30–45% of RF patients and leads to chronic valvular lesions. The pathogenesis of RF/RHD is complex and involves genetic factors that predispose a person to the development of autoimmune reactions.
In recent years, much attention has been paid to Toll-like receptors (TLR) and their role in disease. TLRs are a family of proteins that play an important role in the innate immune response. Unlike the specific nature of adaptive immune response, TLRs are preformed receptors that recognize motifs common to microbial pathogens including lipids, nucleic acids, and proteins known as pathogen associated molecular patterns (PAMPs). Currently, 11 mammalian TLRs have been identified, each of which responds to a specific class of PAMP. 3 Genetic association with streptococcal infections has been shown with the toll-like receptor 2 (TLR2) gene, which encodes the principal receptor recognizing PAMPs found in Gram-positive bacteria.4,5 TLR2 can be activated by a variety of microbial components including lipoproteins, peptidoglycan, and lipoteichoic acid, especially antigens of Gram-positive bacteria including streptococci. 4
MicroRNA (miRNA) is a kind of short non-coding RNAs that suppress the expression of protein coding genes by partial complementary binding, especially to the 3′ untranslated regions (UTRs) of mRNAs. According to the predicted results of bioinformatics, there may be more than 60% of protein coding genes the expression of which is regulated by miRNA.
Although there is no relation between Arg753Gln polymorphism of the TLR2 gene (which confirmed related to ARF susceptibility in child) and RHD, the functional single nucleotide polymorphisms (SNPs) in TLR2 3′UTR have not been well studied.6,7
Materials and methods
Patients
The study group consisted of 110 patients with RHD and 126 healthy controls. Patients with RHD were recruited from the Department of Cardiology, PLA General Hospital, Beijing, China. RHD is defined by a positive medical history along with ecocardiographic documentation of valvular disease and/or surgical replacement of the heart valve(s). An ethnically matched group of healthy blood donors, who did not have a history of cardiac disease, was used as control. The study was approved by the Ethics Committee of the PLA General Hospital, and a written informed consent was obtained from all subjects before inclusion into the study.
Samples
A total of 10 mL heparinized peripheral venous blood was withdrawn after informed consent of the subjects. The processing of these blood samples was started within 30 min after collection. Portion of blood sample was processed for serum separation which was stored at −20℃ until used.
DNA collection and genotyping
DNA from blood samples was extracted by using TIANamp Blood DNA Kit (TIANGEN, Beijing, China). DNA specimens were amplified by using standard PCR protocols. The PCR products were sequenced in forward direction with the ABI 3730xl sequencing platform. The sequencing results were analyzed by using DNAMAN and Chromas Lite software.
Dual luciferase assay
Full length of 826 bp TLR2 3′UTR was cloned into downstream of firefly luciferase coding region in pmirGLO vector (Promega, Madison, WI, USA) to generate luciferase reporter vector. For luciferase reporter assays, HEK293T cells were seeded in 48-well plates. MiR-340 mimic and luciferase reporter vector were co-transfected by using lipofectamine 2000 (Invitrogen, Carlsbad, CA, USA). Two days after transfection, cells were harvested and assayed with the Dual-Luciferase Assay (Promega, Madison, WI, USA). Each treatment was performed in triplicate in three independent experiments. The results were expressed as relative luciferase activity (Firefly LUC/Renilla LUC).
Western blotting
Protein extracts were boiled in SDS/β-mercaptoethanol sample buffer, and 20 μg samples were loaded into each lane of 8% polyacrylamide gels. The proteins were separated by electrophoresis, and the proteins in the gels were blotted onto PVDF membranes (Amersham Pharmacia Biotech, St. Albans, Herts, UK) by electrophoretic transfer. The membrane was incubated with rabbit anti-TLR2 polyclonal antibody (Abcam, Cambridge, MA, USA) and mouse anti-β-actin monoclonal antibody (Santa Cruz Biotechnology Inc., Santa Cruz, CA, USA) for 2 h at 37℃. The specific protein–antibody complex was detected by using horseradish peroxidase conjugated rabbit anti-mouse or goat anti-rabbit IgG. Detection by the chemiluminescence reaction was carried using the ECL kit (Pierce, Appleton, WI, USA). The β-actin signal was used as a loading control.
Measurement of serum concentration of miR-340
Quantitative RT-PCR analysis was used to determine the relative expression level of miR-340. Total RNA was extracted from serum samples, using Trizol LS reagent (Invitrogen, Carlsbad, CA, USA) according to the manufacturer’s instructions. The expression level of miR-340 was detected by TaqMan miRNA RT-Real Time PCR. Single-stranded cDNA was synthesized by using TaqMan MicroRNA Reverse Transcription Kit (Applied Biosystems, Foster City, CA, USA) and then amplified by using TaqMan Universal PCR Master Mix (Applied Biosystems, Foster City, CA, USA) together with miRNA-specific TaqMan MGB probes (Applied Biosystems, Foster City, CA, USA). The miR-16 was used for normalization. Each sample in each group was measured in triplicate and the experiment was repeated at least three times for the detection of miRNAs.
Measurement of serum concentration of IL-6, IL-10, and TNF-α
Commercial enzyme-linked immunosorbent assay (ELISA) kits for IL-6, IL-10, and TNF-α (R & D Systems, Inc., USA) were purchased, and testing was performed according to the manufacturers’ instructions.
For serum IL-6 detection, we used the solid phase sandwich ELISA kit with 96-well strip plate. Sample volume for each well is 100 μL. Assay range: 3.12–0.3 μg/L. Sensitivity: 0.0007 μg/L. Intra-assay precision: 2.04%. Inter-assay precision: 3.77%.
For serum IL-10 detection, the solid phase sandwich ELISA kit with 96-well strip plate was used. Sample volume for each well is 200 μL. Assay range: 7.8–0.5 μg/L. Sensitivity: 0.0039 μg/L. Intra-assay precision: 1.70%. Inter-assay precision: 5.88%.
For serum TNF-α detection, the solid phase sandwich ELISA kit with 96-well strip plate was used. Sample volume for each well is 200 μL. Assay range: 0.015–6–1 μg/L. Sensitivity: 0.0055 μg/L. Intra-assay precision: 4.60%. Inter-assay precision: 5.38%.
Statistical analysis
Allele frequencies of the patients and healthy controls were compared with a Pearson’s chi-square test, and ORs with 95% confidence intervals were calculated. The data of luciferase assay, serum cytokines, and serum miR-340 expression were analyzed by using SPSS Statistical Package version 16. Un-paired Student’s t-test was employed. P < 0.05 was considered statistically significant.
Results
Genotypes and risk of RHD
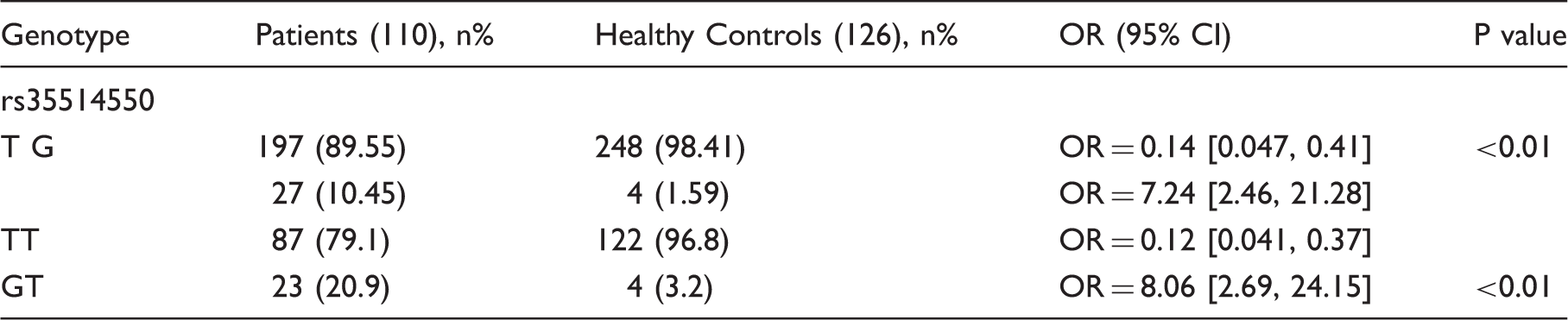
To investigate the relations between TLR2 and RHD susceptibility, we scanned the full length of 826 bp of TLR2 3′UTR in 110 RHD patients and 126 healthy controls. The mean ages of 110 RHD patients is 42 (62 male, 48 female) and the mean age of 126 healthy controls is 40 (74 male, 52 female). All of the study group were successfully genotyped for the TLR2 gene 3′UTR. Only one SNP was found in this region and the genotype frequencies of rs35514550 in controls and patients conformed to the Hardy–Weinberg equilibrium (P = 0.22 for 110 patients without inflammatory cardiac disease and P = 0.86 for 126 healthy controls). After statistical analysis, we found the rare allele G of rs35514550 was associated with RHD occurrence significantly (P < 0.01) (Table 1).
Rs35514550 is included in the miR-340 binding region
To further unveil the biological function of rs35514550, we analyzed the possible miRNAs binding sites in 3′UTR of TLR2 by using miRanda—an online bioinformatics tool (http://www.microrna.org). We found that this polymorphism site may exist in the miR-340 binding region and the U->G variation may interfere with the direct interaction between miR-340 and TLR2 mRNA (Figure 1(a)). Since there are no reports about the fact that TLR2 expression was repressed by miR-340 and the binding site of miR-340 in TLR2 mRNA was not confirmed, our subsequent study focused on investigating the relation between miR-340 and TLR2.
The G allele of rs35514550 weakens the interaction between miR-340 and TRL2 mRNA 3′UTR. (a) Rs35514550 exists in the putative miR-340 binding region. By using online bioinformatics tool miRanda, we found rs35514550 existed in the miR-340 binding region and the T->G variation may disrupt the G-U base pair. (b) Dual luciferase assay was used to confirm miR-340 repress TLR2 by directly binding to the TLR2 3′UTR. HEK293T cells were co-transfected with miR-340 mimic or inhibitor and reporter vector which contains TLR2 3′UTR sequence. Forty-eight hours after transfection, cells were lysed and luciferase activity was detected. The results were shown as relative luciferase activity (firefly luciferase activity/renilla luciferase activity). The results were analyzed using Student’s t-test and P < 0.05 was considered statistically significant. *P < 0.05, **P < 0.01. (c) miR-340 represses endogenous TLR2 expression. A549 cells were transfected with miR-340 mimic or inhibitor. Fort-eight hours after transfection, western blot was employed to detect the TLR2 expression. (d) The T->G variation weakens the interaction between miR-340 and TLR2 mRNA. HEK293T cells were co-transfected with miR-340 and pmirGLO-TLR2-TT or pmirGLO-TLR2-GG. Forty-eight hours after transfection, luciferase activity was detected. *P < 0.05.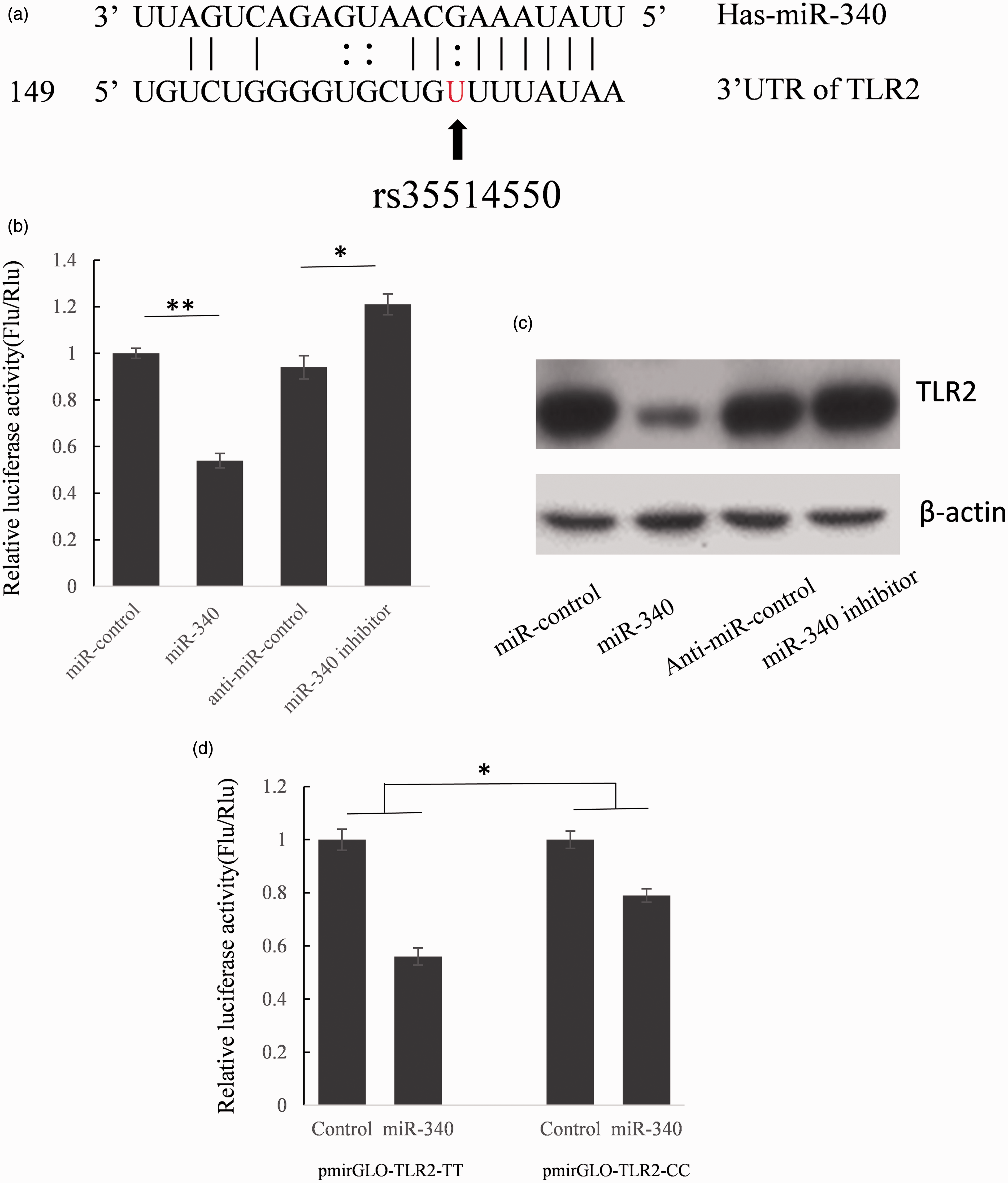
We cloned the full length of 826 bp TLR2 3′UTR into the pmir-GLO plasmid following the firefly luciferase gene coding region to construct TLR2 3′UTR reporter vector. Mir-340 mimic or inhibitor was co-transfected into HEK293T cells with TLR2 3′UTR reporter vector and 48 h after transfection, the luciferase activities were detected. As shown in Figure 1(b), the firefly luciferase activity was down-regulated to 54.1% by miR-340 mimic and meanwhile, up-regulated to 122.4% when co-transfected with miR-340 inhibitor.
The results of luciferase assay indicated that miR-340 can repress luciferase activity by binding to TLR2 3′UTR, but whether the expression of endogenous TLR2 is repressed by miR-340 is still not understood. A549 cells were transfected with miR-340 mimics or inhibitor to see whether the dysfunction of miR-340 can affect endogenous TRL2 expression. Compared with corresponding control, the concentration of TLR2 protein was significantly suppressed by miR-340 mimics and up-regulated by miR-340 inhibitor indicated that TLR2 is the direct target of miR-340 (Figure 1(c)).
T->G variation weaken the interaction between miR-340 and TLR2 3′UTR
To understand the biological roles of rs35514550 in the regulation of TLR2 expression, we constructed the mutant reporter plasmid which contains the TLR2 3′UTR with the rare allele of rs35514550. As shown in Figure 1(d), miR-340 repressed the luciferase activity to 56.2% when co-transfected with pmir-GLO-TLR2-TT. But when the reporter vector contained GG genotype TLR2 3′UTR, the luciferase activity was only reduced to 80.1% by miR-340 relative to scramble non-sense RNA control. The results suggest that the T->G variation up-regulated luciferase expression by weakening the interaction between miR-340 and TLR2 3′UTR.
Rs35514550 T->G variation enhance the patients’ serum IL-6 and TNF-α concentration
Because autoimmune response is recognized to play the key role in the RHD pathogenesis and the activation of TLR2 by S. pyogenes can elevate the serum cytokines concentration. To understand the function of T->G variation in vivo, we detected three serum cytokines concentration in different genotypes patients’ serum samples (15 GT genotype patients, 15 age and sex paired TT genotype patients). As shown in Figure 2(B), the serum IL-6 and TNF-α concentration in GT patients was higher than TT genotype patients significantly (P < 0.05). Meanwhile, there was no significant difference between these two groups in the serum miR-340 concentration. These results indicated that the rare allele G of rs35514550 is related to RHD by over-activating host immune system.
G allele of rs35514550 is related to high serum IL-6 and TNF-α concentration in RHD patients. (A) Serum miR-340 concentration was detected by using stem-loop qRT-PCR. Total RNA was extracted from serum samples that derived from 15 GT genotype patients and 15 TT genotype and age, sex paired patients. The expression level of miR-340 was detected by TaqMan miRNA RT-Real Time PCR. The results were analyzed using Student’s t-test and P < 0.05 was considered statistically significant. (B) Serum IL-6, IL-10, and TNF-α concentrations were detected using ELISA kit. The results were analyzed using Student’s t-test and P < 0.05 was considered statistically significant.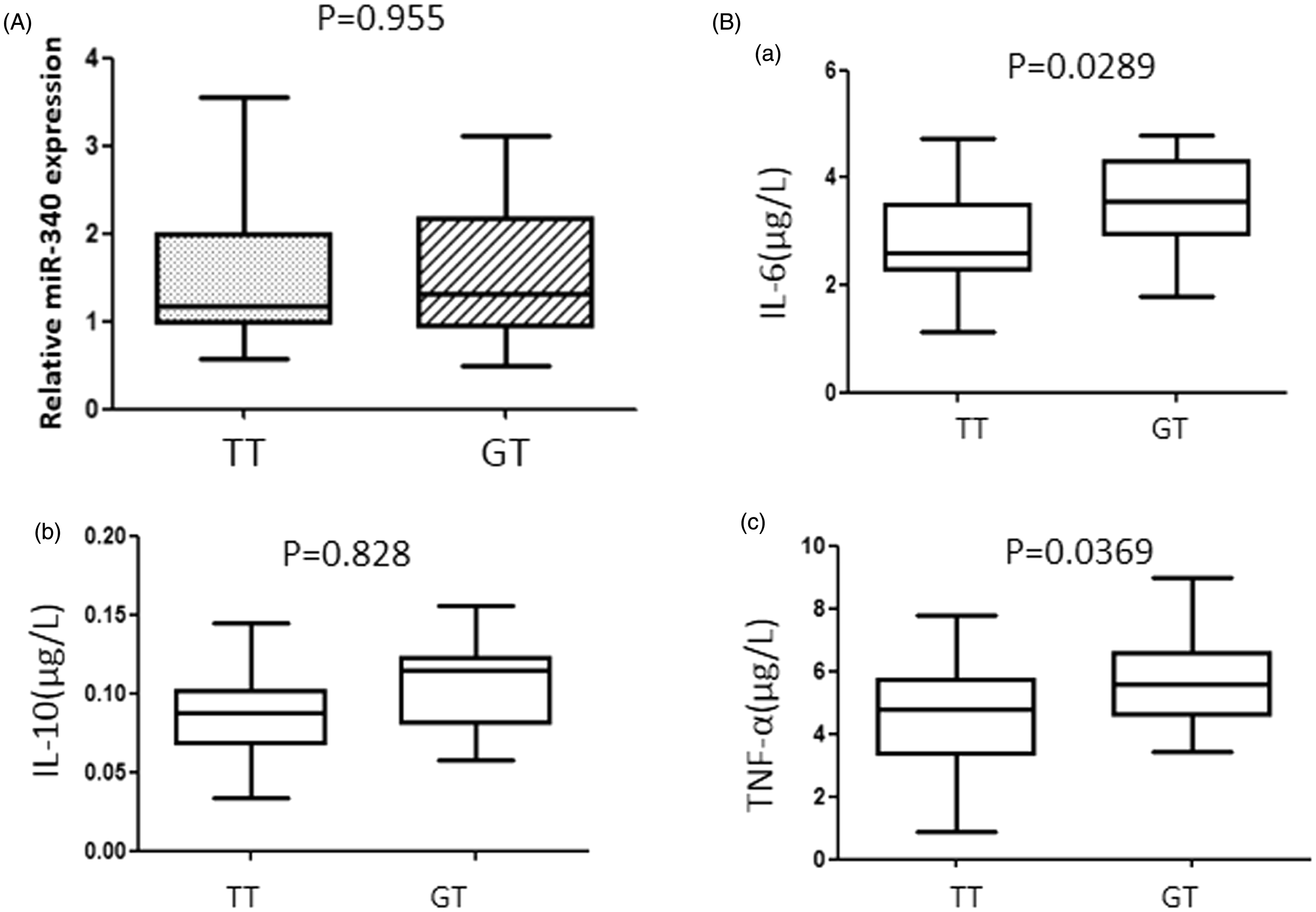
Discussion
TLR2 was considered to be activated by streptococcus which is the major pathogen for RF. Although Arg753Gln polymorphism of the TLR2 gene was confirmed to be related to ARF susceptibility in child, two groups have identified that there were no relations between this Arg753Gln polymorphism and RHD pathogenesis. In this study, we scanned the full length of 826 bp TLR2 3′UTR sequence in 110 RHD patients and found that the rare allele G of rs35514550 is highly related to RHD susceptibility. By using online bioinformatics tools, we found that this site exists in the putative miR-340 binding region. So we assume that this one nucleotide variation may weaken the interaction between miR-340 and TLR2 mRNA and related to over-activated immune response. After confirming TLR2 is a direct target gene of miR-340 using luciferase assay and western blot, we detected the serum cytokines concentration. Serum IL-6 and TNF-α concentration was significantly higher in the rs35514550 GT genotype patients compared with rs35514550 TT genotype patients. These results gave an indirect proof that the rare allele G of rs35514550 is related to RHD by up-regulating host immune response through raising TLR2 expression. But further direct evidence has to be collected in the patients’ heart tissue samples.
Genotype and allele frequencies of rs35514550 in 3′UTR of TLR2 gene in patients with rheumatic heart disease (RHD) and healthy controls (HC).
Footnotes
Declaration of conflicting interests
None declared.
Funding
This research received no specific grant from any funding agency in the public, commercial, or not-for-profit sectors.
Ethical approval
The study was approved by the Ethics Committee of the PLA General Hospital, and a written informed consent was obtained from all subjects before inclusion into the study.
Guarantor
YC.
Contributorship
YL and WX wrote the manuscript, acquired, and analyzed the data; XG and QX performed data analysis, SZ and YC were involved with study design and data analysis. All authors have revised the manuscript and contributed to its final version.
