Abstract
Apramycin and florfenicol are two antimicrobial agents exclusively used in veterinary medicine. Resistance determinants to these antimicrobial agents have been described in several staphylococci, yet no inhibition zone-based epidemiological cutoff (ECOFF) values are available to detect populations harboring resistance mechanisms. In this study, we propose disk diffusion inhibition zone ECOFF values of Staphylococcus aureus for apramycin and florfenicol. The susceptibility to apramycin and florfenicol was evaluated by disk diffusion of five S. aureus collections, comprising 352 isolates of animal (n = 265) and human (n = 87) origin. The aggregated distributions of inhibition zone diameters were analyzed by the normalized resistance interpretation method to obtain normalized wild-type (WT) population distributions and corresponding ECOFF values. The putative WT populations of S. aureus were characterized by an inhibition zone ≥15 mm (ECOFF = 15 mm) for apramycin and ≥21 mm for florfenicol (ECOFF = 21 mm). Five nonwild-type (NWT) isolates were detected for apramycin, all without inhibition zone and harboring the apmA gene, whereas five NWT isolates were identified for florfenicol, all carrying the fexA gene. The proposed ECOFF values for apramycin and florfenicol may be a valuable tool in future antimicrobial resistance monitoring and surveillance studies to identify S. aureus NWT populations toward these antimicrobial agents.
Introduction
S
Apramycin is an aminocyclitol antimicrobial agent produced by Streptomyces spp. approved only for use in veterinary medicine for clinical purposes in Member States of the European Union. 5 It is administered in oral formulations in feed or drinking water for the treatment of enteric infections caused by, for example, Salmonella enterica or Escherichia coli in pigs, calves, lambs, poultry, and rabbits.5,6 Florfenicol is a fluorinated thiamphenicol derivative. Similarly to apramycin, its use is restricted to veterinary medicine and it is administered parenterally for the treatment of respiratory bacterial infections in pigs, cattle, and ovine as well as in oral formulations in feed and drinking water for the treatment of bacterial infections in pigs, poultry, and fish.6,7 Florfenicol is also available, in combination with an antifungal and a steroid, for the topical treatment of otitis externa in dogs caused by Staphylococcus pseudintermedius and occasionally S. aureus. 6 Importantly, both antimicrobial agents belong to classes that are considered as veterinary critically important antimicrobials, 8 as well as critically or highly important antimicrobials in human medicine. 9
Although apramycin is not used for the treatment of infections caused by staphylococci in animals, the presence of a resistance determinant to this antibiotic, apmA, was identified in MRSA isolates from cattle and pigs belonging to the clonal complex CC398. 10 Since then, apmA has been described in MRSA isolates collected from a broiler, 11 from a nasal swab of a human working on a broiler farm and the environment of his residence 11 and from the environmental dust of a pig farm. 12 This gene was also identified in a Staphylococcus lentus (recently reclassified as Mammaliicoccus lentus) 13 isolate from a veal calf 14 and in a Staphylococcus sciuri (recently reclassified as Mammaliicoccus sciuri) 13 isolate from an environmental sample of a pig farm. 15 The apmA gene, which is usually located on small plasmids or large multiresistance plasmids, 16 encodes an acetyltransferase and is associated with elevated minimum inhibitory concentrations (MICs) of apramycin (32 to ≥128 mg/L) and gentamicin (8 mg/L).10,12,16
The florfenicol resistance gene fexA was first identified in a bovine S. lentus isolate. 17 To date, this determinant has been described in various S. aureus and other staphylococcal isolates (including S. pseudintermedius and Staphylococcus simulans) collected from pigs, 18 cattle,14,19 poultry, 20 cats, 21 a dog, 22 a horse, 23 and a marmot 24 as well as from soil surrounding pig farms. 25 The fexA gene has also been reported in MRSA isolates from nasal swabs of pig farmers. 26 This gene is located in the Tn558 transposon or its variants, which can be found on small or large plasmids,3,27 or even integrated in the staphylococcal chromosomal DNA. 23 It encodes the efflux pump FexA, a 14 transmembrane segment transporter of the Major Facilitator Superfamily and is associated with florfenicol MIC values ranging from 16 to 64 mg/L.17,23,28 Other determinants can convey resistance to florfenicol, namely, the cfr, optrA, and the poxtA genes. The cfr gene encodes an rRNA methylase and was first identified in a bovine S. sciuri isolate, 29 but its presence is now reported in several staphylococci, including S. aureus, from various animal origins. 3 It is usually located on plasmids but can also be found in the staphylococcal chromosome. 3 The optrA gene codes for an ABC-F protein that has a ribosomal protection effect and has been described in S. sciuri isolates from cats, dogs, and pigs. 3 Recently, it has also been reported in MRSA isolates collected from pigs and poultry. 27 The poxtA gene codes also for an ABC-F protein, which, however, is only distantly related to OptrA. 30 Besides resistance to florfenicol, all three genes also mediate resistance to oxazolidinones. The gene cfr confers additional resistance to lincosamides, pleuromutilins, and streptogramin A. 3 The poxtA gene also confers elevated MICs to tetracycline. 30
Despite the occurrence of S. aureus strains carrying apramycin and florfenicol resistance genes, there are no inhibition zone-based epidemiological cutoff (ECOFF) values established to identify such strains in a S. aureus population during antimicrobial resistance surveillance studies. Hence, this study focused on proposing ECOFFs of S. aureus for apramycin and florfenicol based on inhibition zones that may allow a rapid identification of isolates with resistance mechanisms toward these two antimicrobial agents.
Materials and Methods
Bacterial isolates
The collection studied included 352 S. aureus isolates, 87 of human origin (from 2006 to 2007 and 2014), and 265 of animal origin (from 2001 to 2020). The isolates of human origin were collected from several infection sites of hospitalized patients (n = 53)31,32 and ambulatory patients (n = 34) in the Lisbon area. The S. aureus of animal origin were collected from cattle (n = 83), dogs (n = 46), cats (n = 44), pigs (n = 41), horses (n = 22), rabbits (n = 16), poultry (n = 6), dolphins (n = 4), and a bird (n = 1). The animal host species was not known for two isolates.
These isolates were analyzed at five independent laboratories; two located in Portugal (cities of Lisbon [Lab1] and Oeiras [Lab2]) and three located in Germany (two in Berlin [Lab3, Lab4], another in Wunstorf [Lab5]). In total, the collection comprised 178 MRSA isolates and 174 methicillin-susceptible S. aureus isolates.
This study involved only bacterial strains that were already isolated and thus, no ethics approval was necessary.
Antimicrobial susceptibility testing
Inhibition zone diameters were determined for apramycin (15 μg) and florfenicol (30 μg) disks by the Kirby–Bauer method according to EUCAST (Lab1, for human isolates) or CLSI standards.33,34 Antibiotic disks were acquired from MAST Group Ltd. (Liverpool, United Kingdom) or Thermo Scientific™ Oxoid™ (Basingstoke, United Kingdom or Wesel, Germany). In brief, fresh overnight cultures were obtained for each isolate on Tryptic Soy Agar (Thermo Scientific Oxoid) or blood agar plates (Thermo Scientific Oxoid), from which isolated colonies were transferred to 0.85% (p/v) NaCl to obtain a cellular suspension with turbidity equivalent to 0.5 McFarland. The cellular suspension was swabbed onto Mueller–Hinton agar (Thermo Scientific Oxoid) plates and the antibiotic disks were placed on the inoculated media within 15 minutes. After 5 minutes, plates were inverted and placed in an incubator at 35°C ± 1°C. After incubation for 18–20 hours, inhibition zone diameters were measured in millimeters. In agreement with the CLSI and EUCAST recommendations, S. aureus ATCC®25923™ and S. aureus ATCC®29213™ were used as quality control strains in this study.33,34
Determination of ECOFF values
The inhibition zone-based ECOFF values were estimated using the normalized resistance interpretation (NRI) method.35,36 This method uses the distributions of inhibition zone diameters to make a least-square regression analysis to determine the putative wild-type (WT) population, the mean inhibition zone diameter and the associated standard deviation (SD) for each species-antimicrobial agent combination. The ECOFF corresponds to the smallest inhibition zone diameter presented by the putative WT population and is calculated at 2.5 × the SD above the mean value and rounded up to the lowest absolute value.35,36 Thus, the ECOFF allows the distinction between putative WT populations (devoid of phenotypically detectable acquired resistance mechanisms) and nonwild-type (NWT) populations (with phenotypically detectable acquired resistance mechanisms). 37 The ECOFF estimated by the NRI method will include 99.4% of the WT population. 36 The NRI method was used with permission from the patent holder, Bioscand AB, TÄBY, Sweden (European Patent No. 1383913, U.S. Patent No. 7,465,559). The automatic and manual excel programs were made available through courtesy by P. Smith, W. Finnegan, and G. Kronvall at www.bioscand.se/nri/.
The ECOFF values generated in this study are based on five data sets from five independent laboratories located in two countries, which provided data for 142 (Lab 1), 33 (Lab 2), 108 (Lab 3), 32 (Lab 4), and 37 (Lab 5) S. aureus isolates.
Results
The five data sets generated in this study were analyzed individually before aggregation (Supplementary Table S1). Each distribution was validated, abiding the minimum number required of WT isolates (at least n = 15) 37 and an SD below the acceptable SD upper limit of 3.38 mm, recommended by the NRI method.
The aggregated distributions of inhibition zone diameters of apramycin and florfenicol for the 352 S. aureus isolates included in this study are shown in Fig. 1. Both aggregated distributions were bimodal, with inhibition zone diameters ranging from 6 ( = growth until the disk) to 30 mm for apramycin and from 9 to 38 mm for florfenicol. The aggregated distributions of the putative WT populations and associated ECOFF values of both antimicrobial agents were calculated using the NRI method (Table 1). Both normalized distributions of the WT populations were validated as they included >100 observations in the putative WT distribution 37 and the SDs were below the acceptable SD upper limit of 3.38 mm (Table 1).
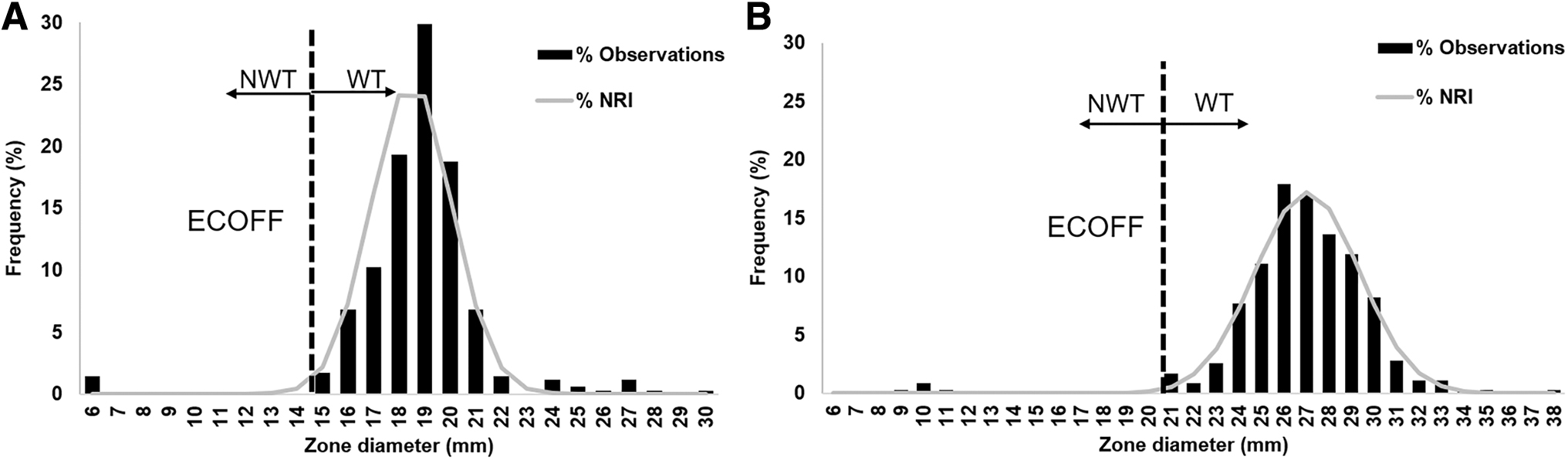
Aggregated distributions of inhibition zone diameters of apramycin
Epidemiological Cutoff Values of Staphylococcus aureus for Apramycin and Florfenicol Estimated Using the Normalized Resistance Interpretation Method
The ECOFF values were determined based on the aggregated distributions of inhibition zone diameters for 352 S. aureus isolates.
ECOFF, epidemiological cutoff; NWT, nonwild-type; SD, standard deviation; WT, wild-type.
For apramycin, the NRI analysis characterized a WT population ≥15 mm with an associated SD of 1.55 mm (Table 1). Applying the estimated ECOFF = 15 mm, an NWT population was identified comprising five isolates (1.4%) collected from cattle (n = 2) and pigs (n = 3). All these isolates were MRSA, showed no inhibition zone, and carried the apmA gene for apramycin resistance.
For florfenicol, a WT population with inhibition zone diameters ≥21 mm was estimated with an associated SD of 2.30 mm (Table 1). The application of the calculated ECOFF = 21 mm identified an NWT population of 1.4%, corresponding to five MRSA isolates collected from cattle (n = 2), pigs (n = 2), and a dog (n = 1). These isolates presented inhibition zone diameters ranging between 9 and 11 mm and all carried the fexA gene mediating resistance to florfenicol in staphylococci. 11
Discussion
The aggregated distribution analyzed in this study revealed an NWT population (1.4%) toward apramycin. This low frequency of NWT isolates is not surprising since dissemination of apramycin resistance determinants is still rare in S. aureus and is mainly associated with food-producing animals and humans with professional contact to them.3,11 In fact, all the five NWT isolates for apramycin in this study were collected from food-producing animals, either cattle or pigs. Apramycin is not affected by most aminoglycoside-modifying enzymes and in staphylococci, a single acetyltransferase encoded by the apmA gene has been identified as mediating resistance to this antimicrobial agent. 3 In this study, the five isolates carrying apmA showed no inhibition zone for apramycin. All isolates of human origin were categorized as WT for apramycin, in agreement with Truelson et al., who analyzed the distribution of apramycin MICs for a collection of >100 S. aureus (mainly of human origin), proposing an MIC-based cutoff value of 32 mg/L and also finding no NWT population among human S. aureus. 38
For florfenicol, the application of the ECOFF value proposed in this study detected the presence of an NWT population (1.4%), comprising five isolates collected from cattle, pigs, and a dog, all harboring fexA. The NWT MRSA isolate of canine origin belongs to the clonal lineage ST398 and presents a multidrug resistance phenotype, showing additional resistance to fluoroquinolones and tetracyclines. 39 The ECOFF estimated in this study may complement the MIC-based ECOFF established by EUCAST for the combination florfenicol—S. aureus, ECOFFEUCAST = 8 mg/L.
The finding of NWT populations for these antimicrobial agents confirms the presence of apramycin and florfenicol resistance determinants in S. aureus isolates from food-producing and companion animals.
In this study, we propose ECOFF values for two antimicrobial agents used exclusively in veterinary medicine, apramycin and florfenicol, based on the inhibition zone diameter distributions obtained for a collection of 352 S. aureus independent isolates of animal and human origin from different geographic regions. The application of the proposed ECOFF values to other collections by other laboratories will be valuable in antimicrobial resistance monitoring and surveillance studies to identify S. aureus NWT populations toward these antimicrobial agents, in a One Health context.
Footnotes
Disclosure Statement
No competing financial interests exist.
Funding Information
This study was supported by Project BIOSAFE funded by FEDER through the Programa Operacional Factores de Competitividade–COMPETE and by Fundação para a Ciência e a Tecnologia (FCT, Portugal)—Grant LISBOA-01-0145-FEDER-030713, PTDC/CAL-EST/30713/2017 and by FCT through funds to GHTM (UID/04413/2020), CIISA Project (UID/CVT/00276/2020), and Project PTDC/CVT-CVT/28469/2017. The contributions of Andrea T. Feßler and Stefan Schwarz were financially supported by the German Federal Ministry of Education and Research (BMBF) under project numbers 01KI1727D and 01KI2009D as part of the Research Network Zoonotic Infectious Diseases. Part of this research was supported by Cost Action CA18217: European Network for Optimization of Veterinary Antimicrobial Treatment (ENOVAT).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
