Abstract
Peptidoglycan recognition proteins (PGRPs) are conserved from insects to mammals and function in antibacterial immunity. We have revealed a novel mechanism of bacterial killing by innate immune system, in which mammalian PGRPs bind to bacterial cell wall or outer membrane and exploit bacterial stress defense response to kill bacteria. PGRPs enter Gram-positive cell wall at the site of daughter cell separation during cell division. In Bacillus subtilis PGRPs activate the CssR-CssS two-component system that detects and disposes of misfolded proteins exported out of bacterial cells. This activation results in membrane depolarization, production of hydroxyl radicals, and cessation of intracellular peptidoglycan, protein, RNA, and DNA synthesis, which are responsible for bacterial death. PGRPs also bind to the outer membrane in Escherichia coli and activate functionally homologous CpxA-CpxR two-component system, which also results in bacterial death. We excluded other potential bactericidal mechanisms, such as inhibition of extracellular peptidoglycan synthesis, hydrolysis of peptidoglycan, and membrane permeabilization. In vivo, mammalian PGRPs are expressed in polymorphonuclear leukocytes, skin, salivary glands, oral cavity, intestinal tract, eyes, and liver. They control acquisition and maintenance of beneficial normal gut microflora, which protects the host from enhanced inflammation, tissue damage, and colitis.
Discovery and Functions of PGRPs
PGRPs function in antibacterial defenses and innate immunity. Insects have many PGRPs with diverse functions. Insect PGRPs sense bacteria and trigger host defense pathways, which generate antibacterial products, defend against infections, and regulate insects' microbiomes. Some insect PGRPs hydrolyze or nonenzymatically neutralize proinflammatory bacterial peptidoglycan and thus limit inflammation.9,33,34
Mammals have four PGRP genes, Pglyrp1, Pglyrp2, Pglyrp3, and Pglyrp4, which were initially named PGRP-S, PGRP-L, PGRP-Iα, and PGRP-Iβ (for “short,” “long,” or “intermediate” transcripts, respectively), by analogy to insect PGRPs. 29 PGLYRP1, PGLYRP3, and PGLYRP4 are directly bactericidal for both Gram-positive and Gram-negative bacteria,11,28,30,37,39 and PGLYRP2 is a peptidoglycan-lytic amidase14,40,45 and also has bactericidal activity. PGLYRP1 is mainly present in PMN's granules, PGLYRP2 is made in the liver and secreted into blood and is also induced in epithelial cells, and PGLYRP3 and PGLYRP4 are produced on the skin and mucous membranes, and in sweat, sebum, and saliva.11,26,28–30,35–40,45
This review will first focus on the question how PGRPs kill bacteria, and then will briefly discuss the in vivo consequences of antibacterial activity of PGRPs—their effect on microbiome and inflammation.
How do PGRPs Kill Bacteria?—The Initial Hypotheses
Our first hypothesis was that PGRPs kill bacteria by inhibiting the transglycosylation or transpeptidation steps in peptidoglycan synthesis10,30,39 because (a) PGRPs bind to the MurNAc-pentapeptide fragments present both in mature peptidoglycan and in peptidoglycan precursors used in these biosynthetic steps, and (b) crystallographic analysis showed that this binding locks peptidoglycan in a conformation that should prevent transpeptidation. 2 Our alternative hypotheses were that PGRPs kill bacteria by either hydrolyzing peptidoglycan and causing osmotic cell lysis, or by directly permeabilizing bacterial cytoplasmic membranes.
PGRPs Inhibit an Intracellular Step in Peptidoglycan Synthesis
Indeed, PGRPs completely inhibit total peptidoglycan biosynthesis in both Staphylococcus aureus and Bacillus subtilis, but they do not inhibit transglycosylation or transpeptidation, the two extracellular steps of peptidoglycan synthesis. 20 These results indicate that PGRPs inhibit an earlier intracellular step in peptidoglycan synthesis.
PGRPs Kill Gram-Positive Bacteria by Localizing to Cell Separation Sites
To selectively and directly inhibit an intracellular step in peptidoglycan synthesis, PGRPs would have to enter the cytoplasm. However, PGRPs do not enter the cytoplasm. Rather, PGRPs exclusively localize at the separation sites of the newly formed daughter cells in B. subtilis, S. aureus, and Listeria monocytogenes. 20 PGRPs do not co-localize with vancomycin; the latter localizes at the sites of new peptidoglycan synthesis, which is primarily at the synthesis of new septa. 20
Cell separation after cell division is carried out by dedicated peptidoglycan-lytic endopeptidases, which in B. subtilis are LytE, LytF, and CwlS13,43 and whose expression is limited to the cell separation sites. PGRPs co-localize with LytE and LytF in the cell separation sites. 20 This localization is necessary for bacterial killing, because mutants that lack LytE and LytF and do not separate after cell division are less efficiently killed by PGRPs than the wild-type (WT) strain. 20 These mutants also do not show specific binding of PGRPs, 20 suggesting that the cell-separating LytE and LytF enzymes are required for efficient PGRP binding to bacteria and bacterial killing. This effect is selective for LytE and LytF, because deficiencies in peptidoglycan-lytic amidase (LytC) and glucosaminidase (LytD), which function as autolytic but not cell-separating enzymes, have no effect on bacterial sensitivity to PGRP-induced killing. 20 Thus, in Gram-positive bacteria, PGRPs trigger their lethal effect from this extracellular site without entering the cytoplasm.
PGRPs Inhibit Protein, RNA, and DNA Synthesis
PGRPs also rapidly and completely inhibit protein, RNA, and DNA synthesis in S. aureus and B. subtilis. 20 Similar inhibition is caused by magainin (due to the loss of membrane integrity) and lysostaphin (due to the loss of peptidoglycan and immediate osmotic lysis). By contrast, antibiotics selectively inhibit either peptidoglycan, or protein, or RNA, or DNA synthesis. 20
PGRPs do not Kill by Osmotic Lysis or by Hydrolyzing Peptidoglycan
To determine whether PGRPs kill by hydrolyzing peptidoglycan, we used 0.5 M sucrose to prevent osmotic lysis. About 0.5 M sucrose prevents lysostaphin-induced, but not PGRP-induced, inhibition of protein, RNA, and DNA synthesis, indicating that PGRPs do not kill bacteria by hydrolyzing peptidoglycan and inducing osmotic lysis. 20 Consistent with these results, bactericidal PGRPs do not hydrolyze insoluble S. aureus or B. subtilis peptidoglycan, uncross-linked soluble polymeric peptidoglycan, synthetic peptidoglycan fragments, or heat-killed S. aureus and B. subtilis bacteria. 20 Thus, bactericidal PGRPs do not have amidase, carboxypeptidase, or any other peptidoglycan-hydrolytic activity. PGRP-induced killing is also not due to the activation of autolytic enzymes. 20
PGRPs do not Directly Permeabilize Cell Membranes
Direct permeabilization of bacterial cell membranes by PGRPs would explain their rapid and simultaneous inhibition of all biosynthetic reactions that is not prevented by hyperosmotic medium (and thus resemble the effect of membrane-permeabilizing peptides, such as magainin). However, PGRPs do not permeabilize bacterial cell membranes over a period of 6 hr, despite rapid killing that exceeds 99% in 2–4 hr and is not prevented by 0.5 M sucrose. 20
Thus, the mechanism of bactericidal activity of PGRPs is distinct from the bactericidal activity of antibiotics that inhibit peptidoglycan, protein, RNA, or DNA synthesis and is also distinct from membrane-permeabilizing peptides and from enzymes that hydrolyze the bacterial cell wall.
PGRPs induce Membrane Depolarization and •OH Production
We next considered whether the loss of membrane potential is responsible for inhibition of intracellular biosynthetic reactions and killing of bacteria by PGRPs, because all these reactions require energy from ATP, whose production is largely dependent on the ATP synthase driven by the proton gradient maintained by the membrane potential.8,16
Indeed, PGRPs at bactericidal concentrations induce rapid and sustained membrane depolarization in B. subtilis. Membrane depolarization was accompanied by intracellular production of toxic hydroxyl radicals (•OH), which are part of the bacterial stress response. 20 Similar effect is also seen in S. aureus. 20
To determine whether •OH production is responsible for the PGRP-induced bacterial killing and how •OH is generated, we tested the effect of dipyridyl and thiourea on PGRP-induced •OH production and bacterial killing. Dipyridyl is a membrane-permeable selective Fe2+ chelator that inhibits Fenton reaction–mediated •OH production, whereas thiourea is an •OH scavenger.22–24 Both dipyridyl and thiourea significantly inhibit PGRP-induced bacterial killing. These results indicate that •OH is generated through the Fenton reaction from FeS clusters and is required for PGRP-induced bacterial killing. 20
Membrane depolarization is sufficient to kill bacteria, because chemical membrane potential decoupler carbonyl cyanide 3-chlorophenylhydrazone (CCCP) induces membrane depolarization and kills bacteria, and the killing is not inhibited by dipyridyl. However, membrane depolarization and •OH have synergistic bactericidal effect. 20
PGRPs Kill B. Subtilis by Activating the CssR-CssS System
We then tested whether the CssR-CssS two-component system in B. subtilis is involved in PGRP-induced membrane depolarization, •OH production, and bacterial killing, because a functionally homologous CpxA-CpxR two-component system in Escherichia coli detects misfolded proteins in antibiotic-treated bacteria and is responsible for antibiotic-induced membrane depolarization, •OH production, and killing.23,24
PGRP-induced membrane depolarization and •OH production is significantly reduced in both ΔcssS and ΔcssR mutants compared with isogenic WT B. subtilis. 20 Membrane depolarization by CCCP is similar in WT and ΔcssS and ΔcssR mutants, indicating that these mutants do not have an inherent defect in maintaining membrane potential and that CssS and CssR selectively mediate the effect of PGRPs. These results indicate that both membrane depolarization and •OH production induced by PGRPs are mediated through the CssR-CssS two-component system.
PGRPs also cause rapid high-level induction of htrA mRNA in WT B. subtilis, but not in ΔcssS and ΔcssR mutants (HtrA is membrane-bound protease directly regulated by CssR). These results indicate that PGRPs induce CssS- and CssR-dependent activation of the CssR-CssS two-component system and activate the regulator (the transcription factor CssR) through the sensor (CssS). 20
To test whether the CssR-CssS two-component system is responsible for the bactericidal activity of PGRPs, we compared the sensitivity of ΔcssS and ΔcssR mutants and isogenic WT B. subtilis to PGRPs. Whereas WT B. subtilis is readily killed by all PGRPs, ΔcssS and ΔcssR mutants are ∼100 times more resistant to killing by all four PGRP proteins. 20 We confirmed these results by complementing ΔcssS and ΔcssR mutants with cssS- or cssR-expressing plasmids. Thus, the CssRS two-component system in B. subtilis is required for bacterial killing by all four PGRP proteins. 20
PGRPs Kill E. Coli by Activating the CpxA-CpxR System
To determine whether PGRPs also kill Gram-negative bacteria through protein-sensing two-component systems, we compared the sensitivity to PGRPs of WT E. coli and its isogenic ΔcpxA and ΔcpxR mutants. E. coli CpxA-CpxR is a functional homologue of B. subtilis CssR-CssS4,15,17,18,24; both systems sense extracytoplasmic misfolded proteins and induce membrane depolarization, •OH production, and stress response. 20 WT E. coli bacteria are readily killed by PGRPs, whereas ΔcpxA and ΔcpxR mutants are 100–1,000 times more resistant to PGRP-mediated killing. 20 We confirmed these results by complementing ΔcpxA and ΔcpxR mutants with cpxA- and cpxR-expressing plasmids. 20
PGRPs activate the CpxA-CpxR two-component system, as shown by PGRP-induced triggering of rapid high-level induction of cpxP mRNA (a stress-response gene directly regulated by CpxR7,24) in WT E. coli, but not in ΔcpxA and ΔcpxR mutants. 20 These results indicate that PGRPs activate the regulator (the transcription factor CpxR) through the sensor (CpxA).
PGRPs Bind to the Entire E. Coli Outer Membrane
How do PGRPs access the CpxA-CpxR two-component system in E. coli? The entire E. coli outer membrane uniformly binds PGRPs at all stages of growth, 20 which contrasts selective localization of PGRPs to the cell separation sites in Gram-positive bacteria. Similarly to Gram-positive bacteria, PGRPs do not enter the cytoplasm in E. coli. 20 Thus, in Gram-negative bacteria, PGRPs bind to the outer membrane and are sensed by the CpxA-CpxR two-component system, consistent with the ability of E. coli CpxA-CpxR to sense bacterial proteins in the outer membrane. 4
The Model: PGRPs Kill Bacteria by Activating Two-Component Systems
PGRPs kill bacteria by activating bacterial two-component stress response defense systems 20 (Fig. 1), which detect extracytoplasmic misfolded and aggregated proteins. Such proteins are generated in bacteria under stress, exported from the cell, and degraded by proteases.4,5,17,23,24,42
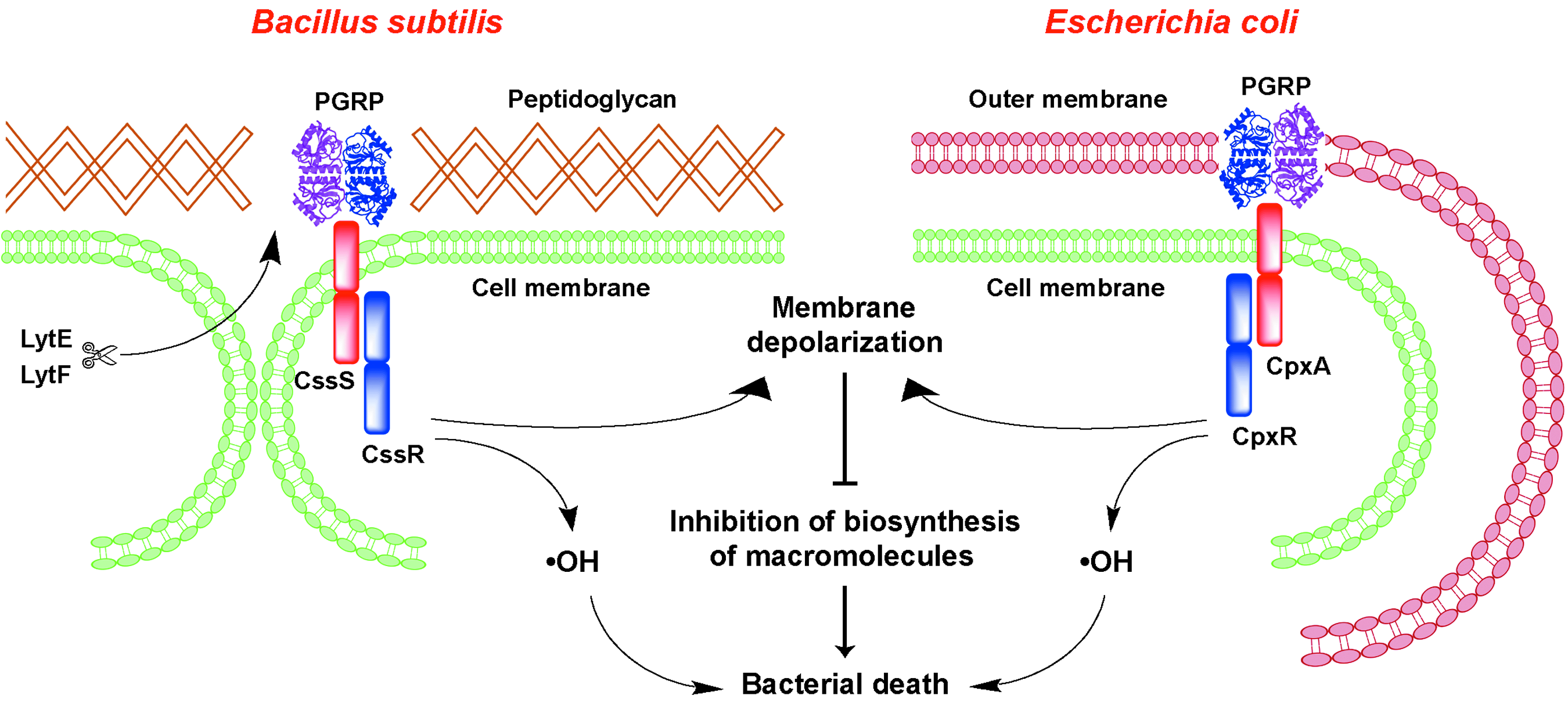
The model of PGRP-induced killing of bacteria by activation of two-component systems, induction of membrane depolarization, production of •OH, and inhibition of biosynthesis of macromolecules. PGRP, peptidoglycan recognition protein.
In Gram-positive bacteria, PGRPs bind to peptidoglycan in bacterial cell walls at the sites of hydrolysis by the enzymes that separate daughter cells after cell division (LytE and LytF in B. subtilis). In Gram-negative bacteria, PGRPs bind to the entire outer membrane, which is composed of lipopolysaccharide (LPS). It should be noted that the name PGRPs is somewhat of a misnomer, because PGRPs, in addition to binding peptidoglycan, also bind LPS and other molecules1,21,27–29,37 using binding sites outside of the peptidoglycan-binding groove. Binding of PGRPs to peptidoglycan or LPS probably induces their oligomerization into ribbon-like structures, 27 which are then detected by B. subtilis CssR-CssS or E. coli CpxA-CpxR two-component systems. 20 Inhibition of binding of PGRPs to bacteria by addition of isolated peptidoglycan or LPS inhibits PGRP-mediated killing of Gram-positive and Gram-negative bacteria, respectively.30,37,39
Activation of the protein-sensing two-component systems by PGRPs induces a sustained high-level stress response in bacteria, which induces membrane depolarization, the production of toxic •OH, and inhibition of all major intracellular biosynthetic reactions. 20 The latter are probably caused by depleting of energy stores, such as ATP, whose generation depends on the membrane potential.6,8,12,16
Some PGRPs also bind to and kill unicellular fungi, such as Cryptococcus neoformans. 37 Because fungi also have envelope stress response systems (but animals do not), activation of these systems by PGRPs could explain fungicidal activity of PGRPs.
PGRPs are Antibacterial In Vivo
Do PGRPs have antibacterial activity in vivo? In zebrafish, PGRPs protect the embryos from infection. 25 In mice, PGRPs protect mice from infection. Sequential intranasal application of PGRPs, followed by a challenge with S. aureus or E. coli, protects mice from lung infection; it results in reduced numbers of bacteria recovered from the lungs 30 (Fig. 2). This effect is synergistic with antibacterial peptides, such as phospholipase A2, which is consistent with synergistic bactericidal activity of PGRPs and antimicrobial peptides. 39
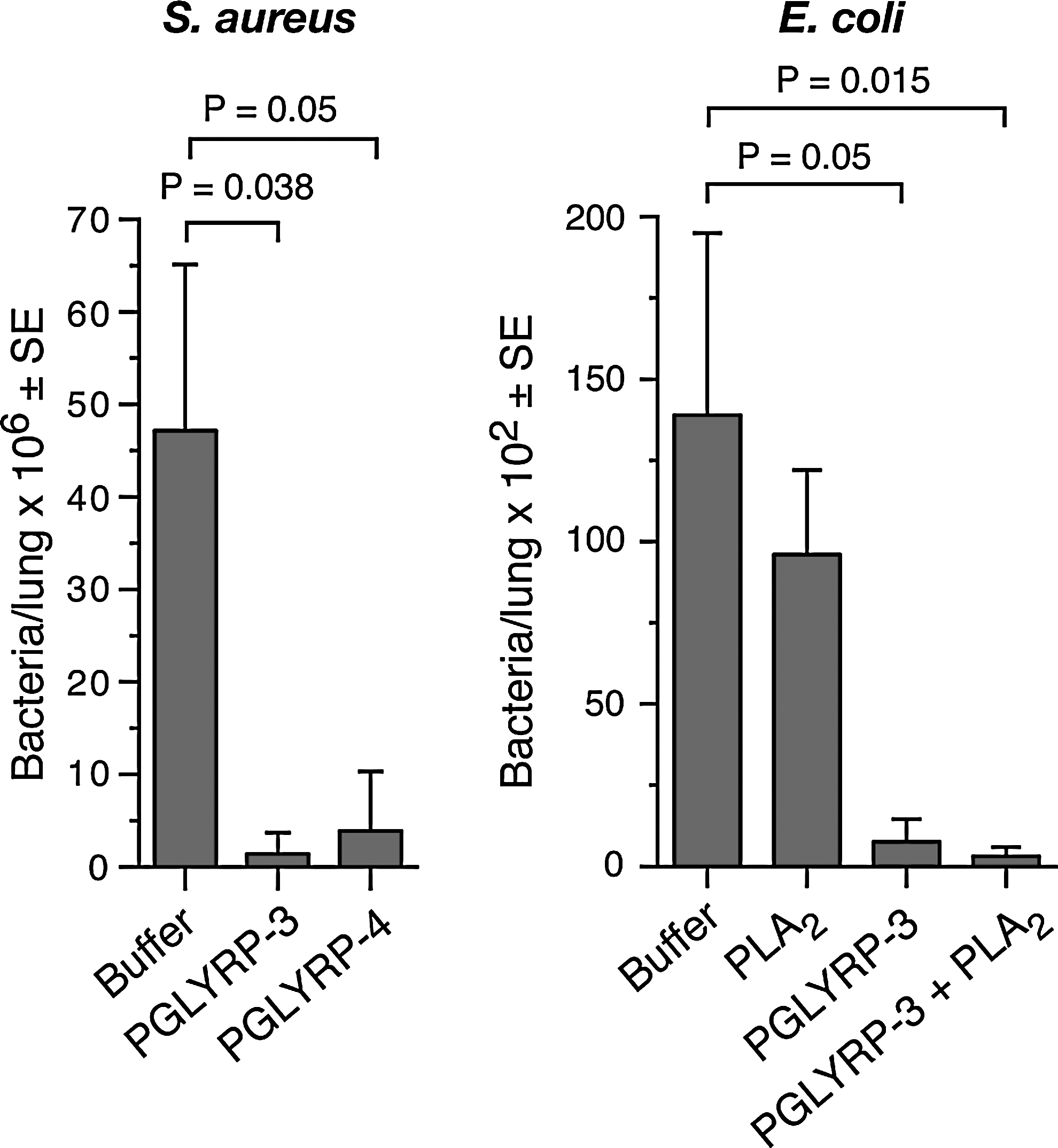
PGRPs protect mice from Staphylococcus aureus and Escherichia coli lung infection. Intranasal application of PGLYRP3 (4.5 μg) or PGLYRP4 (4.5 μg) followed after 15 min by 3×108 S. aureus (left), or of PGLYRP3 (6 μg) or PGLYRP3 (6 μg)+phospholipase A2 (PLA2, 4 μg) followed after 30 min by 4×104 invasive E. coli (strain Bort, right) significantly reduces the numbers of bacteria recovered from the lungs at 4 hr postinfection (S. aureus, n=6; E. coli, n=7).
In addition to PGRPs, mammals have many bactericidal defense mechanisms, including oxidative killing by phagocytic cells, the complement system, antimicrobial peptides, and antibacterial lectins present in phagocytic cells, on body surfaces, in secretions, and in tissue fluids. The loss of one antibacterial system is often compensated by other systems. Thus, in general, Pglyrp-deficient mice do not suffer from serious infections, although Pglyrp1−/− mice are more sensitive than WT mice to infections with some Gram-positive bacteria. 11
PGRPs Maintain Healthy Gut Microbiome and Protect the Host from Colitis
PGRPs are expressed in the salivary glands and on mucous membranes in the mouth, throat, esophagus, and intestines and they play a role in maintaining healthy gut microbiome. 34 Pglyrp1−/−, Pglyrp2−/−, Pglyrp3−/−, and Pglyrp4−/− mice all have significant and distinct changes in the abundance of various bacterial groups in the stools, and especially they have decreased numbers of Lactobacillus/Lactococcus group and Clostridium perfringens group, 35 which are generally considered beneficial for the host. Pglyrp1−/−, Pglyrp2−/−, Pglyrp3−/−, and Pglyrp4−/− mice are also more sensitive than WT mice to ulcerative colitis induced by epithelial-cell-damaging agent, dextran sulfate sodium. 35 This increased sensitivity is due to the loss of protective microflora in the gut and the dominance of predisposing microflora in Pglyrp-deficient mice, because germ-free WT mice gavaged with stools from WT mice show resistance to colitis, whereas germ-free WT mice gavaged with stools from Pglyrp-deficient mice are sensitive to colitis. 35 Thus, in WT mice, PGRPs promote maintenance of the beneficial gut microbiota that protects the colon from developing a severe colitis following intestinal injury (Fig. 3).
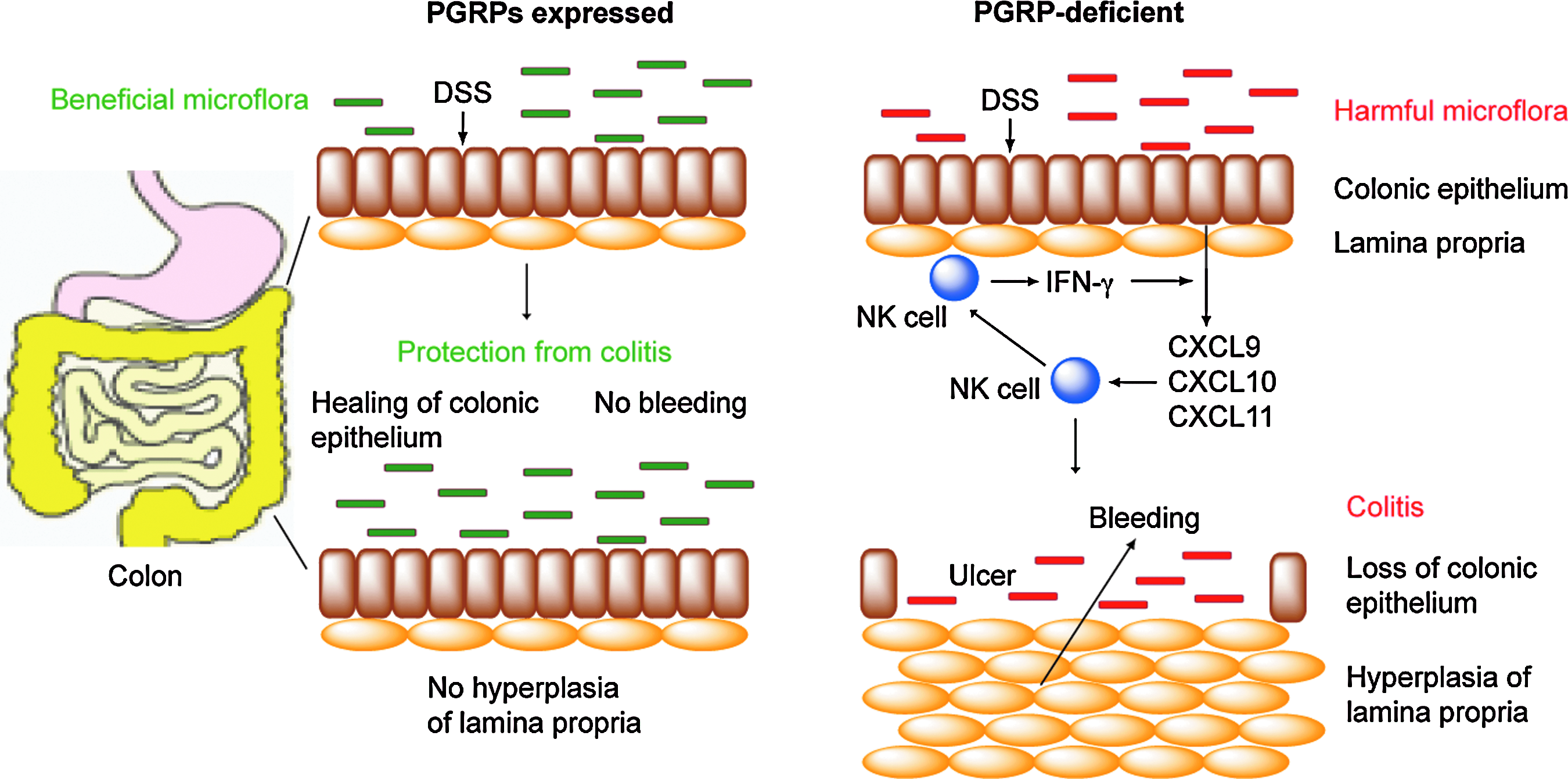
Mammalian PGRPs promote beneficial intestinal microflora and protect from colitis. PGRPs promote beneficial gut microflora, which does not induce cytokines and chemokines and does not cause recruitment of inflammatory cells, and thus protect against colitis. Lack of PGRPs results in the overgrowth of harmful intestinal microflora, which upon intestinal injury with DSS results in increased production of natural killer (NK) cell-attracting and activating chemokines (CXCL9, CXCL10, and CXCL11), recruitment of NK cells to the colonic epithelium, production of interferon (IFN)-γ by NK cells, activation of IFN-inducible genes (including NK cell-attracting chemokines), and eventually colitis, manifested by hyperplasia of the lamina propria, loss of colonic epithelium, ulceration, and bleeding. DSS, dextran sulfate sodium.
PGRPs Protect the Host from Excessive Skin Inflammation
PGRPs are also expressed in the skin and play a role in protecting the skin from excessive inflammatory response to chemicals and allergens. However, various PGRPs have differential effects in various models of inflammation.31,32 Pglyrp2−/− mice (but not Pglyrp1−/−, Pglyrp3−/−, or Pglyrp4−/− mice) are more sensitive to 12-O-tetradecanoylphorbol 13-acetate (TPA) and develop more severe TPA-induced psoriasis-like inflammation than WT mice. 31 By contrast, Pglyrp3−/− and Pglyrp4−/− mice (but not Pglyrp1−/− or Pglyrp2−/− mice) develop more severe oxazolone-induced atopic dermatitis than WT mice. 32 In both models of inflammation this increased sensitivity to TPA or oxazolone is due to reduced recruitment of regulatory T cells and enhanced production and activation of Th17 cells in the affected skin. Thus, in WT mice PGRPs prevent an aberrant activation of Th17 cells by promoting accumulation of regulatory T cells at the site of inflammation, and this protects the skin from an exaggerated inflammatory response. It is not known, however, whether these effects of PGRPs are due to their effects on the microbiome (similar to colitis), or due to their direct effect on the inflammatory response in the skin.
Footnotes
Acknowledgment
This work was supported by USPHS Grants AI028797 and AI073290 from NIH.
Disclosure Statement
The authors have no conflicts of interest and no competing financial interests.
