Abstract
To make progress in understanding peptidoglycan metabolism, we will reconstitute in vitro the assembly process and the molecular machineries that carry out this formidable task. We review here the reports of isolation of complexes comprising penicillin-binding proteins (PBPs), the enzymes that synthesize the peptidoglycan from its lipid-linked precursor.
It is self-evident that a complicated dynamic structure such as the murein sacculus cannot be created by enzymes constitutively churning out peptidoglycan. Assembly and hydrolysis of the peptidoglycan have to be tuned and turned on and off at the appropriate times and places to produce cells of the right size, shape, and strength. Interactions with other proteins within functional complexes certainly provide a large share of the required regulation. In this respect, it may be telling that reconstitution of peptidoglycan assembly in vitro has rarely been reported with recombinant enzymes. Polymerization of glycan chains has been shown a number of times with recombinant class A PBPs from diverse organisms (e.g., references in Sauvage et al. 25 ). The crosslinking of glycan chains by transpeptidation was observed only with recombinant class A PBPs from Escherichia coli.3,5 This fundamental reaction has never been reproduced in vitro with pure proteins from a Gram-positive organism, or with class B PBPs from any bacteria. Beside the trivial explanation that the right conditions have not been found (pH, cations, detergent, substrate, etc.), it is likely that PBPs need interacting partners to function properly. As PBPs and their likely partners are membrane proteins, the isolation of functional complexes is expected to be a difficult task.
Numerous interactions have been inferred from localization experiments, where the localization of one protein is shown to depend on the presence of another one (e.g., in Errington et al. 9 ). Two- or triple-hybrid method has also been used to reveal a lot of interactions (e.g., Karimova et al. 12 ). State-of-the-art fluorescence resonance energy transfer (FRET) experiments in E. coli confirmed the close contacts between PBP3 and other proteins. 2 On the genetic front, comprehensive synthetic lethality and chemical genomic screens have recently pinpointed novel PBP regulators.23,27 However, to the biochemist at heart, the way forward is the preparation of purified complexes to reconstitute and study peptidoglycan metabolism in vitro. Here, we wish to review reports of the physical isolation of protein complexes that include PBPs. Many approaches were taken, yielding various insights.
Ideally, the most biologically relevant method would be the purification of complexes from a native microorganism expressing its endogenous set of unmodified genes. Surprisingly, this most simple approach has almost never been reported, perhaps indicating that PBPs are not involved in very stable complexes that could resist detergent solubilization of the membranes and subsequent purification steps, or that complexes are in low abundance. The exception is the immunoprecipitation of the DivIVA homologue Wag31 in Mycobacterium smegmatis, which co-purified the septal class B PBP3 that was identified by mass spectrometry and subsequent western blotting. 19 DivIVA (Wag31) is a highly conserved cytoplasmic protein of Gram-positive organisms and mycobacteria. 22 It is localized at the septum and poles, and plays a role in the determination of the division site and in chromosome partitioning. DivIVA (Wag31) appears to recognize the spherical membrane curvature of the septum and poles.
Purification of complexes from native organisms has been achieved on several occasions after chemical crosslinking of the proteins. In a very original work, PBPs from Haemophilus influenza were first labeled with fluorescent penicillin; proteins were then crosslinked by exposition of cell suspensions to cyanogen gas. 1 Cyanogen crosslinks salt bridges' residues. Two complexes of about 600 and 450 kDa were then isolated by reverse-phase high performance liquid chromatography (HPLC) of the detergent-solubilized proteins, and monitoring the fluorescence of bound penicillin. Individual PBPs could be attributed to the two complexes by comparing HPLC chromatograms of tryptic digests of the complexes with that of the individual PBPs labeled with fluorescent penicillin. However, the identity of the individual PBPs was established by size-exclusion chromatography in the presence of SDS rather than by SDS-polyacrylamide gel electrophoresis (PAGE). 1 On this basis, one complex was proposed to comprise PBP1a, PBP3, PBP4, and PBP6, and the other to contain PBP1b, PBP2, PBP4, and PBP5. Given the ambiguous identification of the PBPs, the proposed composition of the complexes must be regarded with caution, as they are at odds with more recent data.
Chemical crosslinking applied prior to immunoprecipitation has been used in several instances to demonstrate particular interactions in native bacteria, revealing ligands by western blotting. These experiments do not permit the discovery of unknown interactions, but the confirmation of suspected interacting partners. For example, immunoprecipitates of PBP3, after crosslinking with DTSSP applied on whole E. coli cells, showed the presence of PBP1b. 4 Similarly, immunoprecipitates of E. coli FtsN revealed the presence of PBP3 and PBP1b, and conversely immunoprecipitates of PBP1b contained both PBP3 and FtsN. 20 FtsN is a bitopic membrane protein of the division of proteobacteria that binds peptidoglycan and stabilizes the divisome assembly. 18
Also in E. coli, immunoprecipitation of PBP1a and PBP1b after crosslinking was used to demonstrate interactions with LpoA and LpoB, respectively. 27 LpoA and LpoB are outer-membrane proteins thought to be activators of peptidoglycan assembly.23,27
In this review, we shall qualify experiments as “pull-down” when a solid phase is used to capture a tagged protein, and the resulting precipitate is analyzed, usually by western blotting to reveal expected interactors. Due to the presence of the tag, the protein and the organism studied are not strictly native as in the true immunoprecipitations reviewed above. However, when the tagged version complements the deletion of the endogenous protein, the system is still very close to a natural state. For example, formaldehyde crosslinked complexes from Bacillus subtilis were isolated by Ni-affinity chromatography of his-tagged MreB. Several PBPs labeled with fluorescent penicillins were found to be associated with MreB, in particular, PBP1, and possibly PBP2a, PBP2c, and PBP4. 13 MreB is an actin homologue that forms rings and helical cytoskeletal filaments on the cytoplasmic side of the membrane of rod-shaped bacteria. 28 MreB is thought to organize the elongation of bacilli. 16
When anti-tag antibodies and a resin that captures them are used to isolate the complexes, a “pull-down” is in practice an immunoprecipitation. The interaction of the septal PBP3 from Mycobacterium tuberculosis with FtsW and FtsZ was demonstrated in this way. 6 Tagged proteins (myc-, his-, and S-tags) were co-expressed in E. coli and immunoprecipitated with anti-tag antibodies. Analysis was performed by western blotting. Alternatively, tagged proteins were overexpressed independently, and purified membranes were mixed prior to solubilization. Interestingly, when his-tagged FtsW from M. tuberculosis was expressed in M. smegmatis, the endogenous PBP3 was pulled down by immunoprecipitation, indicating that the interaction was preserved with 66% of sequence identity (75% similarity) between the M. tuberculosis and M. smegmatis proteins. 6 The integral protein FtsW is a flippase that allows translocation across the plasma membrane of the precursor of peptidoglycan synthesis. 17 FtsW is dedicated to the septal peptidoglycan assembly, whereas its paralogues RodA and SpoVE are involved in elongation and sporulation, respectively.
In a very similar experiment, immunoprecipitation showed the interaction of E. coli PBP3 and FtsW, GFP- and HA-tagged, respectively, overexpressed in separate E. coli strains. Detergent-solubilized membranes were mixed before immunoprecipitation with anti-tag antibodies and analysis by western blot. 11 The paralogous interaction between the class B PBP SpoVD and SpoVE during the sporulation of B. subtilis was shown by immunoprecipitation of FLAG-tagged SpoVD and GFP-tagged SpoVE. 10 In this case, the system was very close to native as both tagged proteins were complementing.
The interaction between the same two proteins was also shown by a pull-down experiment where GST-tagged SpoVD and GFP-tagged SpoVE from B. subtilis were co-overexpressed in E. coli. A glutathione affinity resin was used to capture GST-tagged SpoVD, and a western blot confirmed that GFP-tagged SpoVE was also retained on the resin. 10
The interaction between PBP3 and Wag31 from M. tuberculosis was investigated in a slightly different manner. Both proteins were overexpressed in separate E. coli strains, but solubilized his-tagged PBP3 was first captured on a metal affinity matrix, which then retained S-tagged Wag31 that was revealed by western blotting. 19
With the same tags, but using antibodies against both, GST-tagged PBP3 and GFP-tagged FtsQ from E. coli could be co-immunoprecipitated when co-expressed in the same organism. 8 More surprising, the same result was obtained with GFP-tagged homologue DivIB from Streptococcus pneumoniae, which was co-immunoprecipitated with E. coli GST-tagged PBP3, in the same system, suggesting that the interaction was conserved despite the near absence of sequence identity between homologous DivIB and FtsQ (about 10%; see Masson et al. 15 ). The same system was also used to show an interaction between S. pneumoniae PBP2x and FtsL. 14 FtsQ/DivIB and FtsL are bitopic membrane proteins involved in cell division that are required for the recruitment of the septal class B PBP. As the precise function of FtsQ/DivIB and FtsL remains unknown, they have been proposed to be regulators of the PBP activity at the septum.21,31
Affinity chromatography, whereby a cell extract is passed upon a solid phase functionalized with a purified recombinant protein of interest, has revealed a number of complexes with PBPs. In early experiments, the E. coli lytic transglycosylase Slt70 was found to retain PBP3, PBP2, PBP1b, and PBP1c from solubilized membranes and PBP7, a low-molecular-weight PBP, from a periplasmic extract.24,30 The other E. coli lytic transglycosylase MltB retained PBP3, PBP1b, and PBP1c. 30 A third E. coli lytic transglycosylase MltA also retained PBP3, PBP2, PBP1b, and PBP1c. 29 Conversely, PBP1b retained MltA. 20 Another E. coli protein eluted from the MltA affinity chromatography, MipA, was found to retain PBP1b when used itself as a solid-phase chromatographic medium. 29 This ensemble of results established the notion that peptidoglycan synthetic and degradative enzymes work together within complex machineries.
Another protein used to capture PBPs by affinity chromatography was E. coli FtsN, which trapped PBP3, PBP1b, and PBP1a. 20
MreC is a bitopic membrane protein that functions in bacilli with MreB to orchestrate the elongation process. 16 Together with MreD, another membrane protein, MreC, is present in most ovococci, despite the absence of MreB. The precise role of MreCD is not known. When immobilized MreC from Caulobacter crescentus was used for affinity chromatography of solubilized membranes, several PBPs were retained (PBP2, PBP1a, and PBP1b). 7
PBPs from E. coli have themselves been immobilized on resin to capture partners in chromatography experiments. PBP3 retained PBP1b. 4 Conversely, PBP1b captured itself, PBP3, FtsN, MltA, and AmiC.20,26
In most of the affinity chromatography experiments mentioned previously, eluates were analyzed by SDS-PAGE after fluorescent labeling of the PBPs or by western blot. In both cases, only proteins that are being looked after are observed. Affinity chromatography, however, offers the potential to discover new partners if eluates are analyzed with appropriate proteomic tools such as N-terminal sequencing or mass spectrometry. The discovery of MipA is a case in point. 29 LpoA and LpoB were also identified following affinity chromatography with immobilized E. coli PBP1a and PBP1b, respectively. 27
With the various approaches presented previously, the only conclusion that can be drawn is the following: proteins X and Y are part of a complex. Although the trivial conclusion is that proteins X and Y are interacting, the presence of one or several bridging partners cannot, usually, be excluded formally. Actual co-purification is required to demonstrate physically the direct interaction of two or more proteins. Results of the purification should not only show the presence of the desired proteins, as obtained by western blotting or specific labeling of the PBPs, but also show more importantly the absence or low abundance of contaminant, as revealed by protein staining after SDS-PAGE. Moreover, obtaining complexes in large amount is a prerequisite for the ultimate goal of reconstituting various steps of the peptidoglycan metabolism in vitro, and the determination of their structure.
The direct interaction of FtsN and PBP1b was demonstrated with independently purified recombinant proteins. Purified recombinant FtsN was captured on a matrix functionalized with murein fragments, which then was shown to pull down purified PBP1b. 20 However, 10 times less PBP1b than FtsN was recovered from the matrix, as determined by a semiquantitative dot blot. Thus, although direct binding of PBP1b onto FtsN was demonstrated, the complex was not prepared quantitatively in this case.
Very convincing isolations of pure complexes that contain a PBP are those of PBP1a or PBP1b with His-tagged LpoA or LpoB from E. coli, respectively. 23 When equivalent amounts of pure His-tagged LpoA and PBP1a were loaded onto a Ni-affinity matrix, both proteins were quantitatively eluted, under nondenaturing conditions. The same result was obtained with LpoB and PBP1b. Amounts of proteins were significant as samples were analyzed by Coomassie-stained SDS-PAGE. Structural characterization of these complexes is likely underway.
We have ourselves pursued the isolation of recombinant complexes that include PBPs from S. pneumoniae. Proteins were co-overexpressed in E. coli and purified from detergent-solubilized membranes. Preliminary results show that it is possible to partially purify complexes that include DivIB, FtsL, DivIC, as well as PBP2x and FtsW. Localization and genetic data have long suggested that these division proteins interact together. 31 DivIB was His-tagged and PBP2x was strep-tagged; Fig. 1A shows that two successive streptactin and Ni-affinity chromatography could co-purify co-expressed PBP2x, DivIB, DivIC, and FtsL. Using the same method, a complex formed by pneumococcal strep-tagged PBP2x and His-tagged FtsW could be purified, as shown in Fig. 1B.
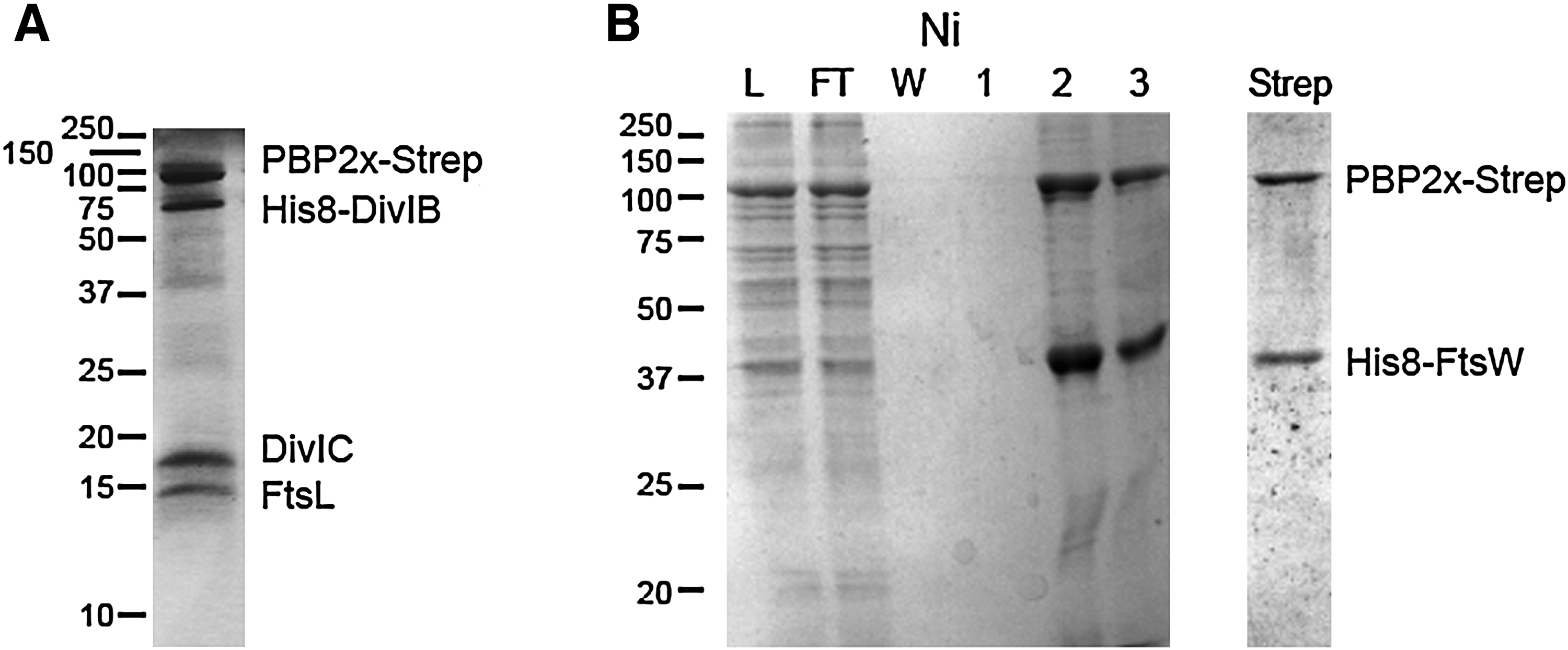
Coomassie-stained SDS-polyacrylamide gel electrophoresis (PAGE) analysis of purification of recombinant membrane protein complexes comprising PBP2x from pneumococcus. Proteins were co-expressed in Escherichia coli. Detergent-solubilized membranes were submitted to successive Ni- and Streptactin-affinity chromatography.
Beside the isolation of complexes, the physical interaction between proteins can be demonstrated by numerous biophysical techniques. Few have been applied to PBPs so far, certainly due to the fact that membrane proteins need to be solubilized by detergents, which often hamper measurement.
Size-exclusion chromatography in the presence of detergent was applied to the complex of mycobacterial PBP3 and Wag31, showing that the complex resisted the dilution inherent to the method. 19 Wag31 was also found to reduce quenching of the tryptophan fluorescence of PBP3 by acrylamide. 19
Surface plasmon resonance was used to characterize diverse interactions of PBPs, using Biacore™ instruments. One protein is immobilized on a surface, while a soluble partner is injected in a mobile liquid phase. This method demonstrated the binding of M. tuberculosis PBP3 onto immobilized Wag31. 19 However, the best-characterized interactions by surface plasmon resonance are those of E. coli PBP1b. Immobilized PBP1b was found to bind itself and MipA,3,29 whereas solubilized PBP1b was found to bind to immobilized PBP3, 4 MipA, 29 or FtsN. 20
Interestingly, affinity chromatography had revealed that the MltA lytic transglycosylase was part of a complex with PBP1b. Detailed surface plasmon resonance experiments have shown that there was no direct binding of MltA to PBP1b, but that this interaction was mediated by MipA, which binds to both proteins. 29
Having listed the methods that have been used to physically demonstrate interactions between PBPs and other proteins, the emerging picture is one of great complexity. Considering only PBP1b from E. coli, by many measures the best characterized PBP, it was found to take part in complexes with 12 different proteins, including 5 PBPs (counting PBP1b itself), and 3 lytic transglycosylases (Table 1). Isolation after crosslinking or affinity chromatography against an immobilized protein can capture whole molecular complexes, and does not single out direct interactions. Moreover, a PBP can be part of different complexes during the cell cycle, or at different cellular localization. Using co-purification, monitored by Coomassie-stained SDS-PAGE, or surface plasmon resonance, PBP1b was demonstrated to interact directly with four proteins, itself, PBP3, MipA, and LpoB.
Bold characters indicate that the detected interaction is unambiguously direct.
AC, affinity chromatography; IPXL, immunoprecipitation after crosslinking; IPOE, immunoprecipitation after overexpression; PBPs, penicillin-binding proteins; PP, purified proteins; SPR, surface plasmon resonance; ?, unknown or putative.
Although PBPs undoubtedly interact with various partners, these interactions rarely result in stable complexes that can be isolated for further biochemical characterization. Indeed, chemical crosslinking was used in most isolation experiments from nonrecombinant bacteria. So far the exceptions are the complexes of the E. coli PBP1a and PBP1b with their respective outer-membrane regulators LpoA and LpoB, and the complexes of the class B PBPs PBP2x and PBP2b from S. pneumoniae with the flippases FtsW and RodA, as well as of PBP2x with the trimer DivIB, FtsL, and DivIC. These complexes are now available for biochemical and structural studies that will likely enhance our understanding of the complicated process, that is, peptidoglycan metabolism.
Footnotes
Acknowledgment
This work was supported by grant PneumoPG ANR-08-BLAN-020.
Disclosure Statement
No competing financial interests exist.
