Abstract
DNA-encoded libraries (DELs) have demonstrated to be one of the most powerful technologies within the ligand identification toolbox, widely used either in academia or biotech and pharma companies. DEL methodology utilizes affinity selection (AS) as the approach to interrogate the protein of interest for the identification of binders. Here we present a high-throughput, fully automated AS platform developed to fulfill industrial standards and compatible with different assay formats to improve the reproducibility of the AS process for DEL binders identification. This platform is flexible enough to virtually set aside all kinds of DELs and AS methods and conditions using immobilized proteins. It bears the two main immobilization methods to support of the proteins of interest: magnetic beads or resin tip columns. A combination of a broad variety of protocol options with a wide range of different experimental conditions can be set up with a throughput of 96 samples at the same time. In addition, small modifications of the protocols provide the platform with the versatility to run not only the routine DEL screens, but also test covalent libraries, the successful immobilization of the proteins of interest, and many other experiments that may be required. This versatile AS platform for DEL can be a powerful instrument for direct application of the technology in academic and industry settings.
INTRODUCTION
DNA-encoded libraries (DEL) are positioned as a very useful technology for ligand identification within the pharmaceutical industry, 1 –3 having reduced costs per screening campaign compared with more classical screening technologies. 4 –6
There is a current need of affinity selection (AS) platforms with higher throughput and versatility for the different experimental conditions required for relevant outputs. Technology development examples of DEL screenings focus on the relevance of testing different experimental conditions for one target, 7,8 screen multiple targets in the same experiment, 9 –11 or highlight the importance of replicate experiments to assess the statistical relevancy of the results. 12,13 Also, increasing complexity of DEL inputs, with billion compounds and pools of multiple libraries makes the reproducibility of AS data challenging. Liquid handling accuracy and throughput are critical to obtain high-quality data in this context.
We previously described the development of technologies covering critical steps of DEL workflow, including a database-centric platform to track every step in the experimental process involved in the development and production of DELs, a method to normalize AS results, and an algorithm for the deconvolution of sequencing results. 14 –16
Herein, we aim to develop a platform for automated AS of DELs, highly parametrizable, with higher throughput than previously reported platforms 17 and yielding consistently reproducible results through runs.
MATERIALS AND METHODS
Platform Configuration
This high-performance platform has been configured to be versatile. Based on a TECAN Freedom EVO 200 liquid handler, it has the capacity for processing 96 samples at the same time. The worktable enables the use of multiple plates to fit the requirements of the different steps of the protocols. Different reagents can be placed in each plate position and even in the different wells of the different plates, thus giving the option of using multiple selection conditions within the same experiment. The deck is fully configurable; therefore, it can be adapted for each protocol to match the setup needed to assay the screening conditions (Figure 1).
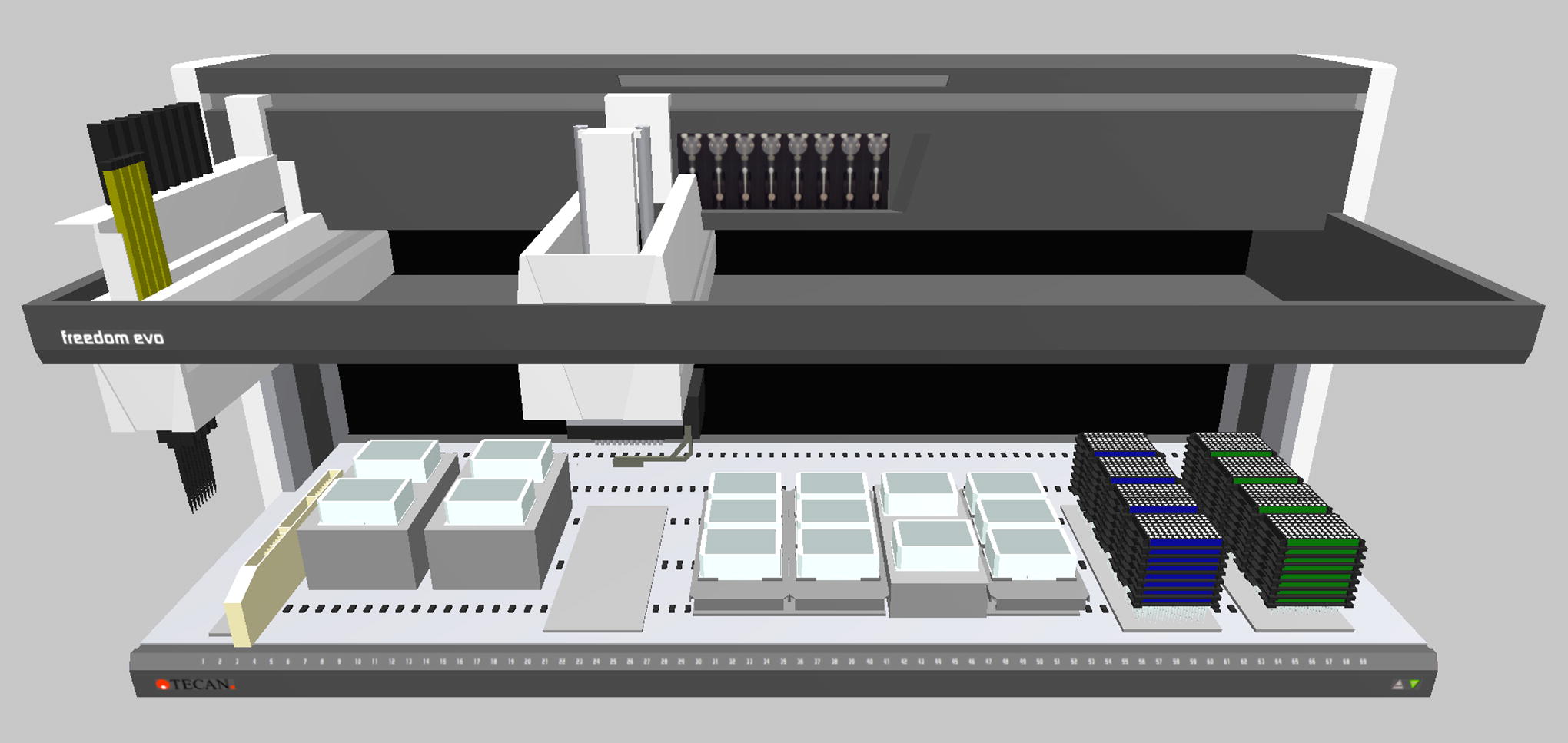
From left to right, the TECAN instrument setup includes an 8-tip liquid handling arm, a multichannel pipetting arm that is able to work with 96 tips simultaneously and with an integrated gripper to move plates between the different table positions. On the deck there are, DiTi waste station-6 trough positions carrier, 4 Inheco incubators with temperature control, a DiTi waste station, 12 plate positions and 8 positions for nested tip boxes.
All the references of the components of the liquid handler, as well as a list of reagents and plastic labware used to execute the AS protocols, are listed in the Supplementary Data S1. The plate adaptors, plates, and tips are changed based on the protocol needs and are described for each protocol in the corresponding script section.
This platform has been built using a TECAN liquid handler as the base instrument, but any other liquid handler device with similar capabilities can be used to build an AS platform, including but not limited to Hamilton, Beckman Coulter, Echo, and others. The included protocols can be directly imported into the TECAN software, and they could be easily translated to any other liquid handler software.
Protocol Outlines and Configuration
We describe three different protocols in this article (Figure 2): one protocol that uses chromatography resin tip columns (PhyTips) and two protocols using magnetic beads as protein immobilization support. Each of these protocols can be configured to achieve specific experimental needs and is included as an importable file of scripts in the Supplementary Data S2.
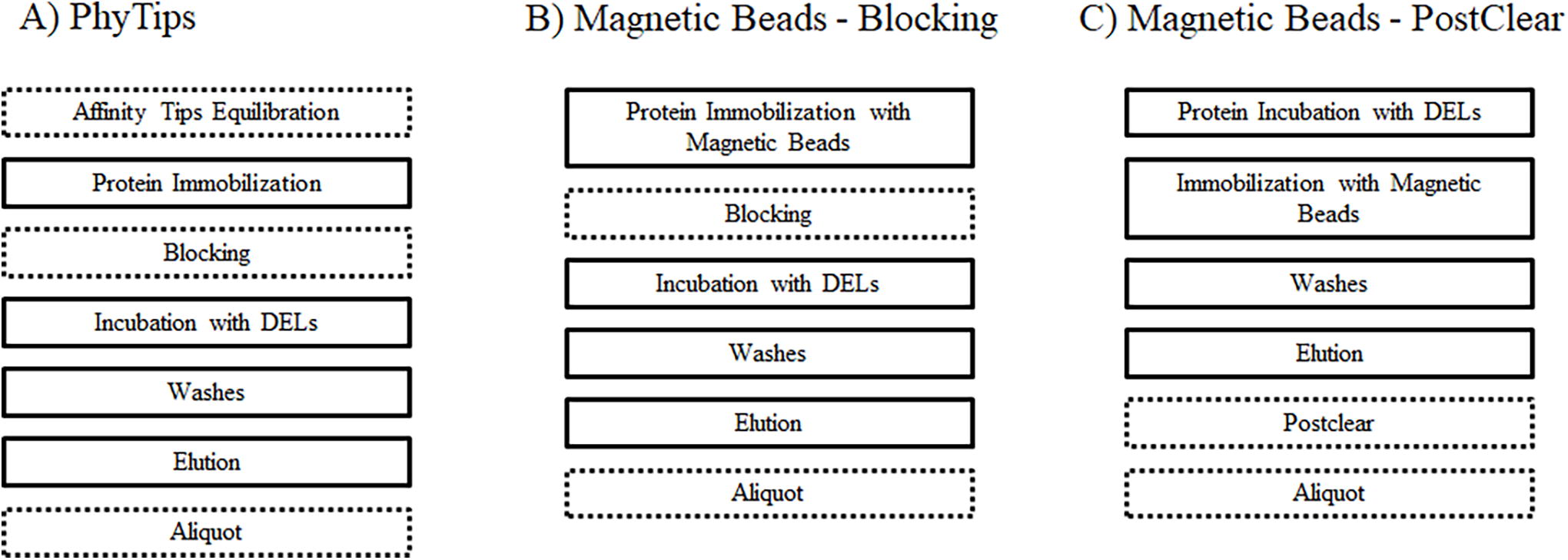
Protocol outlines for the Scripts for PhyTips
Figure 2 reflects the general outline of the three protocols, including the most relevant steps in the AS process.
The automated protocols included are designed to screen up to 96 samples using different experimental conditions in every individual well, such as different proteins or protein constructs, several protein tag immobilizations, diverse buffer conditions, empty beads as negative controls, competition assays, or any other combination of conditions. In addition, technical replicates can be included to assess the reproducibility of the binder identification for each specific condition.
The first protocol is annotated as “A) PhyTips.” This protocol uses purification tips with a variety of affinity resins to immobilize the proteins of interest. Here, we immobilize the protein in the tip and incubate the DELs by slowly pipetting up and down the libraries, allowing the interaction of the compounds with the immobilized protein.
The second protocol is annotated as “B) Magnetic Beads-Blocking.” This protocol uses magnetic beads for the protein immobilization and then incubates the DELs together with the immobilized protein.
The third protocol is annotated as “C) Magnetic Beads-PostClear.” This protocol uses magnetic beads for the protein immobilization after the incubation of the DELs with the protein.
All the protocols have “PCR Plate Adaptors” (black) situated in positions 1–5 and four nested 200 µL tips (dark blue) in positions 17–20, but the other positions will vary between the different table configurations. The other variable positions include 50 µL tips (green), nested 50 µL tips (dark green), PhyTips (cyan), Deep Well Plates of 0.5 mL (white), 1 mL (green) or 2 mL (light blue). Also, for the Magnetic Beads protocols, a Magnum™ EX (Alpaqua) is needed in the “Magnet” position (black).
Before running the protocols, the worktable must be filled with the corresponding plate or tip box. Positions marked as “Empty” in the figure will only have the PCR Plate Adaptor, as those positions will be used to move PCR Plates from different positions during runtime.
General Considerations and Experimental Outline
Typically, a capture test is done prior to the AS to assess the amount of protein immobilized on the solid support. It involves the use of a tagged protein, the solid support, and a similar protocol used for the AS screen except for the incubation of the DELs. In this assay, the different eluents from the immobilization and washing steps can be recovered and monitored using protein gels. Afterwards, the immobilized protein can be removed from the solid support through washings, for instance, with a high concentration of Imidazole (HIS-Tagged proteins), to assess the amount of protein immobilized.
For the regular AS assay (already including the DELs incubation), if your assay design does not include all the wells of the 96 well plates, only those that are involved in the plan will be included in the protocol, keeping the remaining wells empty to save protein, buffer, magnetic beads, and resin tips columns. The only exception is the resin tips included in the box in the protocol “1-DEL-PhyTips” (Figure 3-A, position 24), because the 96 arm needs to utilize the full box to ensure the proper attachment of the tips through homogeneous pressure along the box surface. Indeed, a shortcut to avoid wasting resin tips, the empty positions of the box could be filled with empty tips of similar size, at least in the corners and central positions.

Table Layout for
For every protocol, the AS buffer to be placed in the “Reservoir” position refers to the specific selection buffer required for each protein, which is usually determined by a biophysical technique (i.e., thermal shift assay) screening different buffers and additives for each protein. Once the selection buffer for each protein is identified, between 0.1 and 1 mg/mL bovine serum albumin, 0.1 to 1 mg/mL external DNA, and 0.01 to 0.1% Tween-20 are added to create the selection buffer for the AS experiments. These buffer conditions can be tested using an orthogonal binding or biochemical assay if already available. To confirm the correct protein conformation and immobilization, an assay with an on-DNA tool compound marked with a fluorophore can be run. The same protocols and conditions used for the regular AS with DELs can be employed in the automated platform for confirmation using the fluorescent tool compound instead of the DELs. In addition, multiple assay conditions (buffers), with and without the protein of interest to test the specific binding of the compound to the immobilized protein could be run in parallel, using a fluorescence reader to determine the optimal conditions.
The selected screening buffer is used to dilute the protein of interest to the desired assay concentration, for instance, 1 µM in the magnetic beads protocol or 4 µM in the columns protocol. The proteins are pipetted manually into a PCR plate, and the plate is placed in the “Protein” position on the TECAN platform, which previously has been warmed or cooled to the required temperature (set up in the protocol).
Between 1 and 10 µL of the pool of DELs at 1 mM concentration (1 to 10 nanomoles) or the desired amount for each experiment is pipetted into a PCR plate and placed in the “Incubation” position at the temperature used for the incubation of the protein with the libraries, for example, at 22°C.
For the magnetic beads protocol, the volume of the specific beads needed for the immobilization of the protein, depending on the protein tag used, is added to a PCR plate. In our case, we have successfully worked with several protein tag immobilizations: Streptavidin beads for the biotinylated proteins, Ni/Co Beads for the HIS-tagged proteins, or Protein A for the Fc-tagged proteins. Each position in the plate can have different immobilization beads depending on the protein tag of the corresponding position on the protein plate. The plate must be placed at the desired temperature in the “Beads” position for the protocols in which the magnetic beads are used.
For the column-based protocol, the corresponding tips, depending on the protein tag used, need to be added: Streptavidin tips for the biotinylated proteins, IMAC tips for the HIS-tagged proteins, or Protein A tips for the Fc-tagged proteins. Each position in the box can have tips with different immobilization resin depending on the protein tag for the corresponding position on the protein plate. The tips need to be placed in the box in the “PhyTips” position. If the experiment is not going to use the whole box filled with tips, it is important to add empty tips with the same size of the resin-loaded tips to the corners and the center of the box to avoid any problem in the tip charge with the MCA96 arm.
The selection buffer is placed in the buffer “Reservoir” filled in as many wells as protein wells are used. In the Magnetic Beads Protocols “B) Magnetic Beads-Blocking” and “C) Magnetic Beads-PostClear,” the reservoir is a Deep Well plate of 1 mL with 800 µL of buffer in “position 9” and 400 µL of Elution Buffer (or water) in the Deep Well Plate of 0.5 mL in “position 8” (Figure 3).
In the Columns Tips Protocol “A) PhyTips,” the reservoir is a Deep Well Plate of 2 mL with 1.8 mL in “position 8.” Also, a Deep Well Plate of 0.5 mL in “position 10” with 400 mL of buffer and a Deep Well Plate of 0.5 mL in “position 9” with 400 µL of Elution Buffer (or water) are needed. An optional step is to add a blocking reagent (such as biotin or imidazole) to the plate in “position 11” to quench the affinity resin after the protein immobilization (Figure 3).
RESULTS
The automated protocols described above for the AS screening platform were designed to screen from 1 to 96 samples using different assay conditions or to assess the reproducibility through the introduction of technical replicates. In addition, to ensure the proper performance of the platform, a control protein with known binders can be included in all the AS campaigns to confirm the consistency of the results over a long period of time. To analyze possible cross-contamination due to the proximity between the wells within the plate, all the adjacent wells were tested to detect similar enriched patterns between unrelated protein targets or no protein controls. In any case, cross-contamination between wells in any direction was detected (Supplementary Data S1).
Here, we include representative examples of AS results to assess the reproducibility of the process using this automated platform. We include a set of results for a bromodomain target using similar conditions but with 12 months of time difference. These results will demonstrate the reproducibility and consistency of the results using the same automated protocol over a long period of time. We also include a set of results using a protein-protein interactions (PPI) target using different immobilization methods (magnetic beads and resin tips) and therefore different automated protocols. These results will demonstrate the reproducibility of the different immobilization methods and the different protocols of this automated platform.
The fully codified positive and negative controls can also be used to set up the conditions of the screening. For troubleshooting of the AS, these controls can be included in the experiments to confirm the correct enrichment of the positive control and the absence of enrichment of the negative control. For the PPI target, we included fully codified positive and negative controls. The positive control was included at the same amount as each compound of the DEL pool, while the negative control was included at a concentration 100 times higher. The positive control showed clear enrichments (between 1,300 and 4,000 reads depending on the experiment), and the negative control showed only between 2 and 5 reads, clearly within the noise level of the experiment. The successful enrichment of the positive control compound confirmed that the AS was performed correctly.
The samples after AS were sequenced using next-generation sequencing (NGS). The process involves determining the total number of cycles needed for preparative PCR based on quantitative PCR (qPCR) results, organizing samples into subgroups to ensure optimal cycling, and performing barcoding of each sample within the preparative PCR and PCR cleanup steps before sequencing. Samples are prepared into an equimolar DNA mix, diluted for sequencing, and PhiX is added to introduce diversity for sequencing. Data from the sequencer are processed using tagFinder, 16 with the ratio of “Raw” to “Deduped” counts indicating sequencing depth. Higher ratios suggest better sample representation, while lower ratios may indicate issues with DNA recovery or PCR duplicates. Complete procedures for the sequencing process can be found in the supplementary material from a recent publication by our group. 18 The number of sequencing reads for the samples in these examples can vary from 0.5M reads to 15M reads depending on the number of samples including in each sequencing run.
Reproducibility of Data within 12 Months
Here we show a visual representation of consistent results obtained for the same target run regularly over a one-year time frame, under similar screening conditions: buffers, number of DELs in the screening, and number of copies per compound (Figure 4).
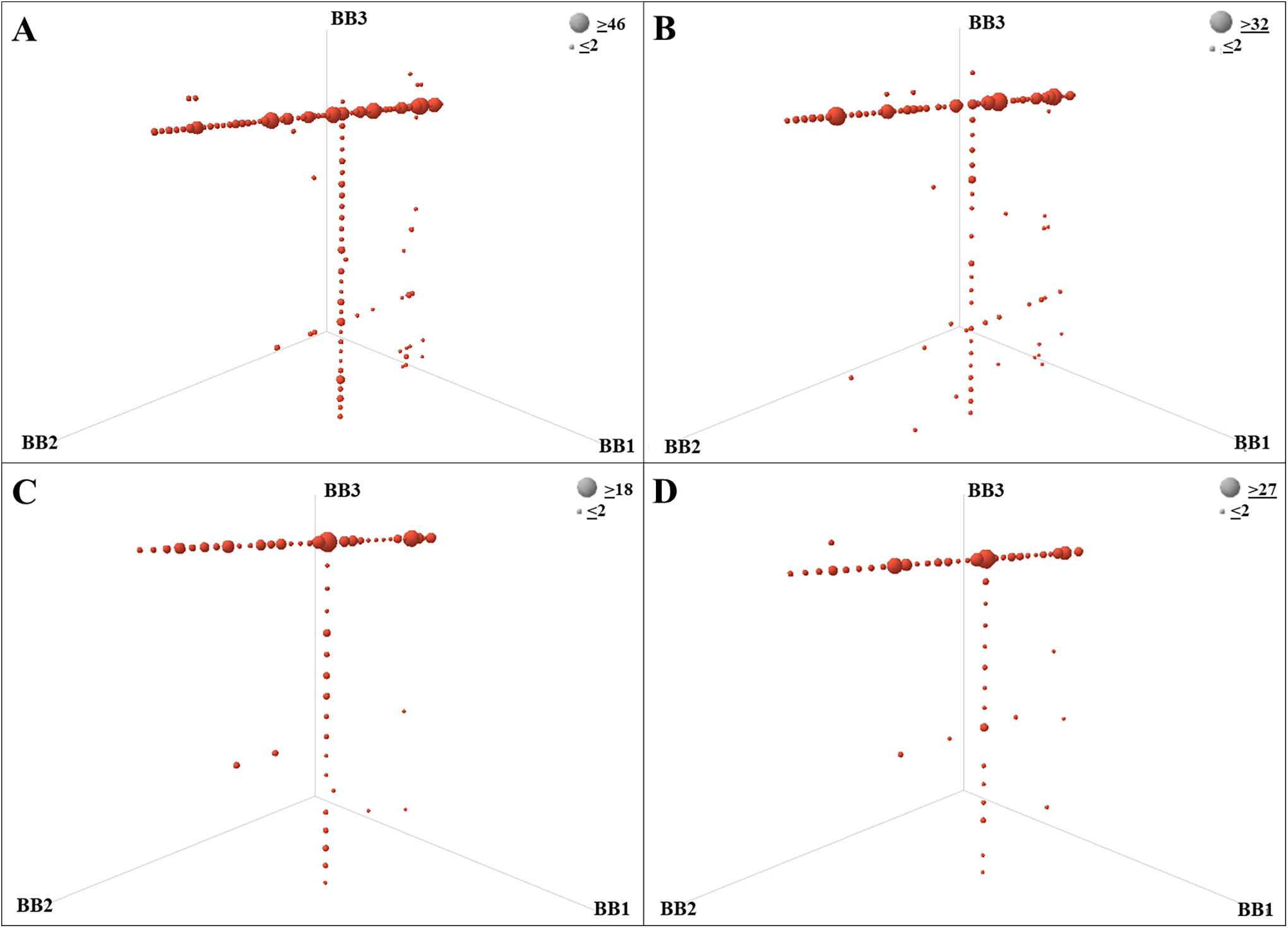
Example of affinity selection results with the enriched compounds from a library of 3.5 million compounds (DEL-18) versus a bromodomain target within a pool of multiple libraries. Immobilization of HIS tagged protein using Cobalt coated magnetic beds.
The AS in this example was performed against a bromodomain target using the protocol options and conditions described in the Supplementary Data S1. In summary, a volume of 60 µL of 1 µM protein (approximately 5 µg of total protein) was used for each experimental condition and mixed with approximately 10 µL of 1 mM of a pool of DELs (or 10 nmol of the DEL pool). The protein and libraries were incubated for 60 min with continuous mixing (pipetting three times every 5 min). Subsequently, 20 µL of the His-Tag Isolation magnetic beads were added for the immobilization of the protein and mixed for 30 min. After the protein immobilization, three washing steps with 100 µL of selection buffer were performed to remove the nonbinders. The bound molecules were eluted in 50 µL of water by heat denaturation at 80°C by mixing for 5 min. An aliquot of the eluate was used for quantitative PCR by mixing 4 µL of water and 2 µL of the eluate. The remaining volume of eluate was used for a subsequent round of AS, or to submit for high-throughput sequencing after the third round of selection.
Figure 4 shows the experiment replicates run with 12 months of time difference using similar experimental conditions: the same Structure-Activity Relationship (SAR) trends with similar enrichments can be observed consistently. Each dot stands for a single compound built by combining a building block (BB) of cycle 1, cycle 2, and cycle 3, represented on the axes of the figure. The size of the dots on the cube representation corresponds to the number of unique sequencing counts for each enriched compound after removal of PCR duplicates using a Unique Molecular Identifier (UMI). Analysis of the experimental replicates
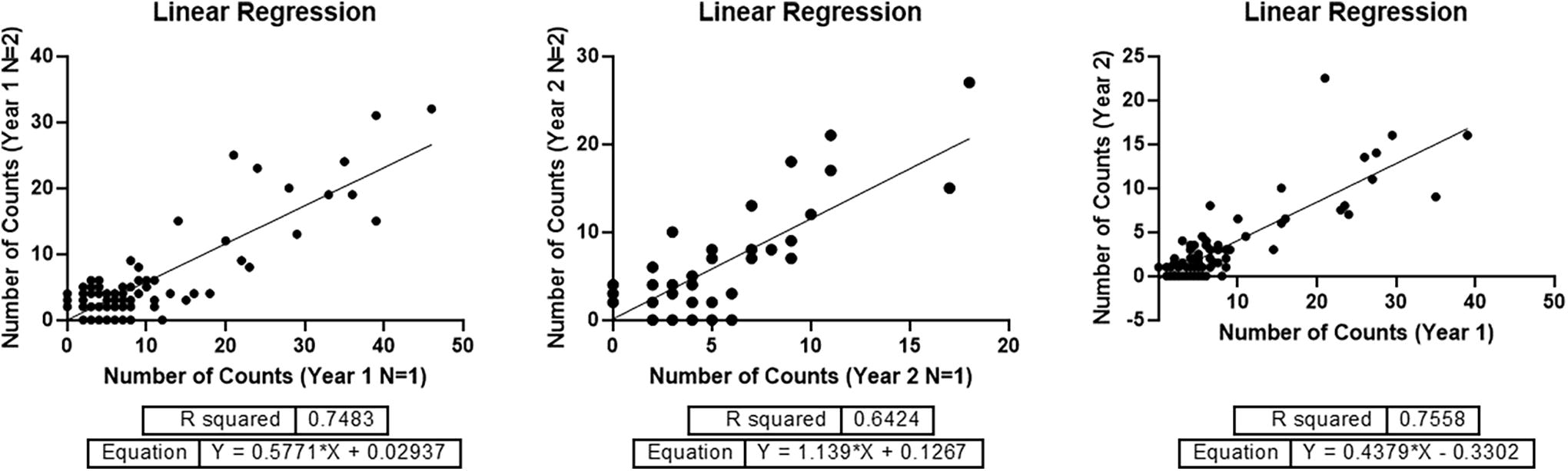
Linear regression for datasets of Figure 4. Each dot represents a single compound, and each axis represents the number of sequencing counts for each sample or experiment. Year 1 (n = 1) corresponds to the dataset of Figure 4-A, Year 1 (n = 2) to Figure 4-B, Year 2 (n = 1) to Figure 4-C, and Year 2 (n = 2) to Figure 4-D. The reproducibility of the overall experiments after one year was analyzed using the average counts of each replicate of Year 1 versus the average counts of Year 2.
Reproducibility of Data Using Different Solid Supports
In a different assay, we compared the protocols (PhyTips and Magnetic Beads) using different solid support immobilization methods for the same protein and the same pool of libraries. The experimental conditions for the magnetic beads are identical to the conditions described in the previous section, but changing the buffer and the type of magnetic beads as the protein is a PPI target with an Fc protein tag. The protocol options and conditions are described in the Supplementary Data S1.
For the protocol using the resin tips or PhyTips, the tips are equilibrated by mixing in 70 µL of selection buffer for 30 min. Then, the protein is immobilized by mixing 100 µL of 2 µM of the protein solution (approximately 8 µg of total protein) with the resin tips for 15 min. Then the DELs are incubated with the immobilize protein by mixing 70 µL of the DEL pool (10 µL of the DEL pool was previously mixed with 90 µL of selection buffer) for 30 min. After the compounds had the possibility to bind to the target protein, the washing steps were performed by mixing one time with 70 µL of fresh selection buffer in each of the plates labeled as “Buffer 3” to “Buffer 7” to remove the nonbinders. The bound molecules were eluted in 100 µL of water by heat denaturation at 80°C by mixing 70 µL for 10 min. An aliquot of the eluate was used for quantitative PCR by mixing 4 µL of water and 2 µL of the eluate. The remaining volume of eluate was used for a subsequent round of AS, or to submit for high-throughput sequencing after the third round of selection.
Similar enrichment trends were observed in all cases as exemplified in Figure 6.
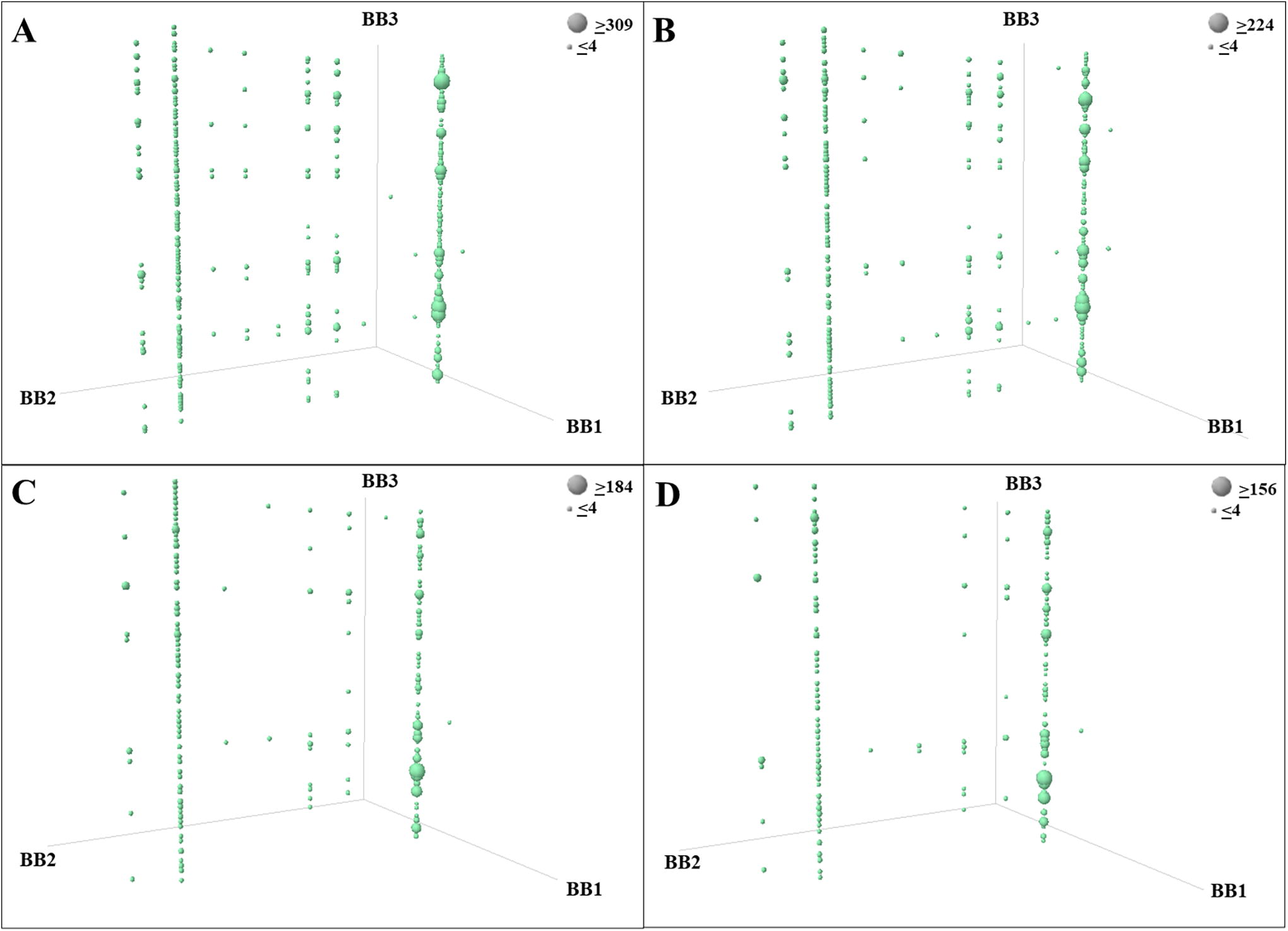
Example of affinity selection results for a library of 1.3 million compounds (DEL-3) for a Fc-tag immobilized protein within a pool of multiple libraries (530 million compounds and 9 million copies of each compound).
The same trends (lines) are clearly enriched similarly in the different replicates and experimental conditions, while PhyTips replicates
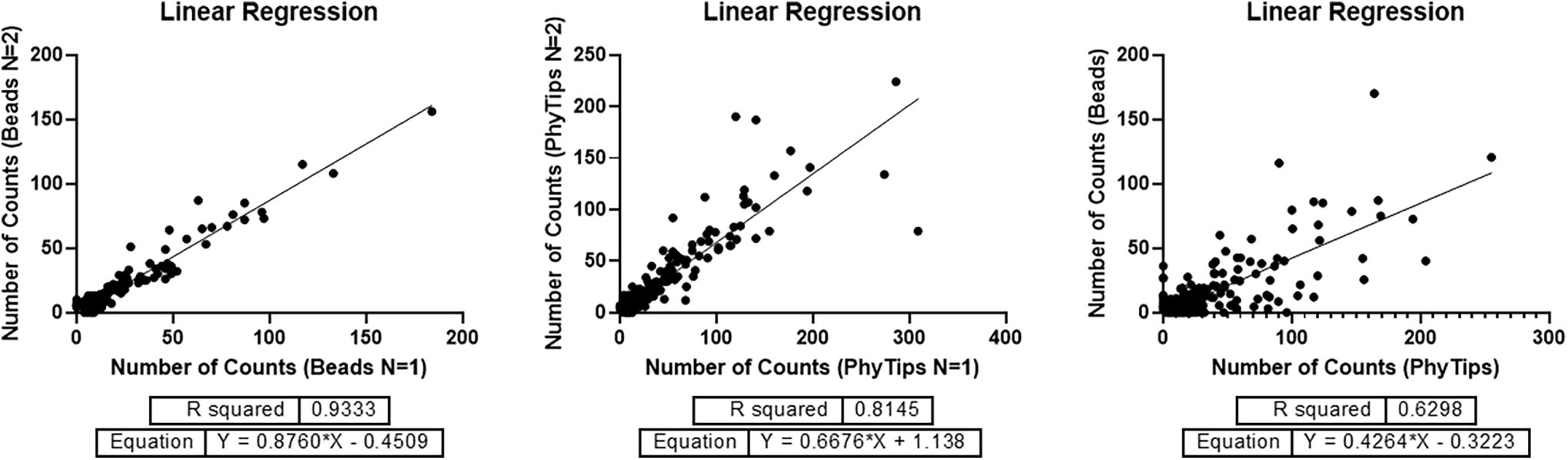
Linear regression for datasets of Figure 5. Each dot represents a single compound, and each axis represents the number of sequencing counts for each sample or experiment. PhyTips (n = 1) corresponds to Figure 5-A, PhyTips (n = 2) to Figure 5-B, Beads (n = 1) to Figure 5-C, and Beads (n = 2) to Figure 5-D. The reproducibility between the different immobilization methods was analyzed using the average counts of each replicate of the PhyTips experiments versus the average counts of the Beads experiments.
The trends shown in Figures 4 and 6 come from the selection with multiple libraries, but the examples of confirmed trends are from the libraries DEL-3 and DEL-18 (Figure 8).
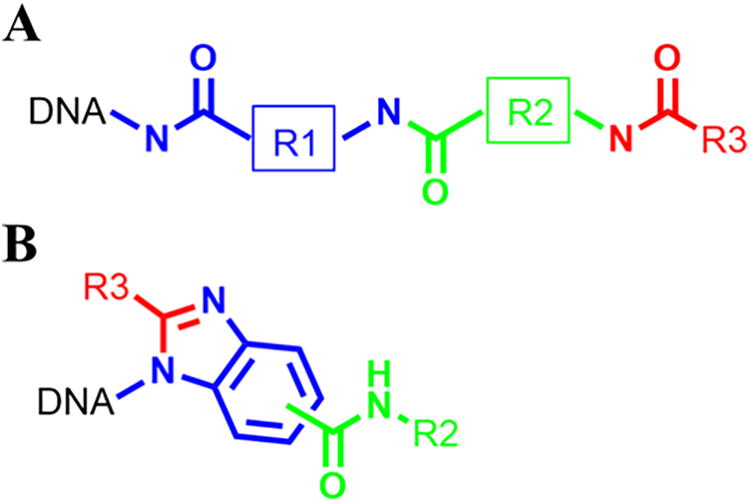
Schematic representation of DEL-3
Correlation Analysis
The different chemotypes enriched by the ASs are defined by the combination of the BBs that are enriched. Therefore, in the results, we enriched several lines (combination of 2 BBs), in which each line represents a different chemotype. Even if the same chemotypes are clearly enriched in the different datasets used as examples for Figures 4 and 6, an analysis based on each individual compound presence was done to prove the overall reproducibility of the results using the automated platform described in the publication.
The axes of every graphical representation correspond to the number of unique sequencing counts of each single compound after removal of the PCR duplicates using a UMI. In every experiment, two replicates for each AS condition were analyzed using linear regression analysis (GraphPad Prism V9.3) to calculate the R-squared. After the analysis of each dataset with the experimental replicate, the average of the counts of the replicates was calculated and used to analyze the reproducibility of each experimental condition using linear regression analysis again.
Overall, all the datasets showed good linear correlations between the experimental replicates and between the different variables related to experimental conditions, including time and immobilization supports used.
DISCUSSION
The AS protocols and conditions for DEL screening are still in continuous evolution, and every academic or pharma group uses its own protocols, platforms, or experimental conditions depending on the automation availability and the specific requirements for each protein of interest. As presented in the Results section, we show a robust, reproducible, and very flexible screening platform that can be used for a wide range of experimental conditions.
Another advantage of this platform is its adaptability for using multiple protein immobilization systems (resin tip columns and magnetic beads), the option to screen multiple experimental conditions for each target of interest, or the possibility to run multiple targets in a single run, reducing the time required to make large screening campaigns using DELs. We envision that this platform can be modified to accommodate very different selection conditions, providing a versatile screening tool for almost every DEL practitioner and can be modified to cover a wide range of experimental conditions without requiring automation skills.
The capacity to screen up to 96 different conditions at the same time is also an important feature of the platform, allowing to perform replicates of the same condition to assess the reproducibility of the experimental data. The automated high throughput is critical to make improved statistical analysis of the results for the deconvolution, facilitating the identification of the real binders out of the noise level based on the reproducibility of the enrichments of the different families of compounds detected. Indeed, testing the reproducibility of data by running automated protocols and calculating R-squared coefficients obtained in linear regression analysis demonstrates not only that the experiment replicates in terms of similar enrichment trends in all cases when run with a 12-month time difference, but also by comparing PhyTips and Magnetic Beads solid support immobilization methods. This feature allows the study of additional proteins with confirmed binders from different libraries to verify the technical performance, reproducibility, and robustness of the automation process.
The platform’s versatility is demonstrated as it can be used for the DEL screening of Single-Pharmacophore, Dual-Pharmacophore, Covalent DEL Libraries, or any other type of DELs with minimum modifications of conditions and reagents. For example, for screening of a covalent DEL library, the protocols included can be configured to add more washing steps and can include a denaturing agent in the corresponding buffer to ensure the recovery of only the covalent binders.
In summary, we describe an automated platform amenable to run any kind of DEL screening for immobilized proteins by using basic liquid handling capabilities.
CONCLUSIONS
This high-throughput automated AS platform is versatile, highly configurable to accommodate very different experiment designs, compatible with different assay formats, and demonstrates reproducibility in the AS results for DEL binder identification.
This is an advance in comparison with other DEL screening platforms or methodologies. Handmade protocols rely on the experience, pipetting reliability of the scientist, and are prone to human mistakes. Other screening platforms are only available for the use with magnetic beads and the degree of configuration is limited.
This platform can run screening protocols lasting several hours without any human intervention, therefore saving time and effort. Also, the addition of automation of the screening process allows DEL practitioners to test different experimental conditions with almost no extra effort in contrast to handmade protocols.
In summary, we believe this platform can increase the throughput of the screening, can save time and effort, all without compromising the reliability of the results obtained.
Footnotes
ACKNOWLEDGMENTS
The authors thanks all the Lilly DEL group for their continuous support and help.
AUTHORS’ CONTRIBUTIONS
R.R.-G. Conceptualization, Methodology, Software, Validation, Writing-Original Draft, Writing-Review & Editing. E.D. Writing-Original Draft, Writing-Review & Editing. M.I.L. Supervision, Writing-Review & Editing. M.J.L.: Supervision, Writing-Review & Editing. J.D.B. Writing-Original Draft, Writing-Review & Editing. M.A.T. Conceptualization, Writing-Review & Editing. R.H. Conceptualization, Writing-Original Draft, Writing-Review & Editing.
CONFLICT OF INTEREST
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article. The authors received no specific financial support for the research, authorship, and/or publication of this article.
DISCLOSURE STATEMENT
No competing financial interests exist.
FUNDING INFORMATION
No specific funding was received for this article.
SUPPLEMENTARY MATERIAL
Supplementary Data S1
Supplementary Data S2
