Abstract
An algorithm was developed to statistically predict ischemic tissue fate on a pixel-by-pixel basis. Quantitative high-resolution (200 × 200μm) cerebral blood flow (CBF) and apparent diffusion coefficient (ADC) were measured on acute stroke rats subjected to permanent middle cerebral artery occlusion and an automated clustering (ISODATA) technique was used to classify ischemic tissue types. Probability and probability density profiles were derived from a training data set (
Introduction
Brain tissues with perfusion deficits below a critical threshold (Hossmann, 1994), as visualized on perfusion-weighted images (PWI), will experience metabolic energy failure that subsequently leads to a reduction in the apparent diffusion coefficient (ADC) of water in the brain, and thus hyperintensity on the diffusion-weighted images (DWI) (Moseley et al, 1990). During the first few minutes after ischemia in experimental stroke models or up to a few hours after ischemia in human strokes (i.e., the acute phase), the anatomic area defined by abnormal DWI is typically smaller than the area of perfusion deficit. However, most of the normal DWI-defined ischemic region expands and eventually coincides with the abnormal area defined by PWI. The difference in the abnormal region defined by the abnormal PWI and normal DWI, commonly referred to as the ‘perfusion—diffusion’ mismatch, is thought to approximate the ischemic penumbra and thus could potentially be salvageable (Warach et al, 1996).
The ability to readily and reliably
Magnetic resonance imaging data obtained early after stroke onset also offers the unique opportunity to statistically predict ischemic tissue fate. Welch et al (1995) used a threshold-based analysis and showed that the combination of T2 and ADC data provided improved prediction of infarction relative to either parameter alone in subacute stroke in humans. Wu et al (2001) reported the eloquent use of a generalized linear model to predict stroke outcomes based on DWI, PWI, and T2 data in humans. Lesions were defined using a threshold-based method to generate the training set. However, this approach appears less intuitive as the contribution of various parameters is difficult to assess.
The goal of this study was to develop and validate a simple and intuitive statistical algorithm for predicting ischemic tissue fate after acute stroke in a well-characterized rat stroke (permanent occlusion) model. We hypothesized that ischemic tissue fate can be statistically predicted with a high degree of accuracy and the prediction performance can be quantitatively evaluated. Quantitative high-resolution perfusion and diffusion imaging was obtained. A modified ISODATA cluster analysis (as opposed to a threshold-based analysis) was used to classify tissue types. Prediction of tissue infarct was made for
Methods
Animal Preparations
Twelve male Sprague—Dawley rats (300 to 350 g, Taconic Farms, NY, USA) were initially anesthetized with chloral hydrate (400 mg/kg, intraperitoneally: Sigma, St Louis, USA). Animals were subjected to permanent intraluminal middle cerebral artery occlusion (MCAO) as previously described (Shen et al, 20032004a). The left femoral artery was catheterized for blood-gas sampling, continuous blood pressure and heart rate monitoring. Blood pressure, heart rate, respiration rate, rectal temperature, blood gases were maintained within normal physiologic ranges (Shen et al, 2003, 2004a). Once the animal was in the magnet, the anesthetic was switched to ~1% isoflurane. All animals breathed spontaneously throughout without mechanical ventilation. At ~24 h after ischemia, the animals were killed and stained with 2,3,5-triphenyltetrazolium chloride (TTC) to determine infarct volume with edema correction (Shen et al, 2003, 2004a).
MR Experiments
Magnetic resonance imaging was performed on a Bruker 4.7-T/40-cm (Billerica, MA, USA) and a 20-G/cm gradient insert (ID = 12 cm, 120-μs rise time). The animal was placed into a stereotaxic headset and onto an animal holder, which consisted of an actively decoupled surface coil (2.3-cm ID) for brain imaging and a butterfly neck coil for arterial spin labeling (Shen et al, 2003, 2004a; Sicard et al, 2003). Coil-to-coil interaction was actively decoupled. Imaging was performed at 30, 60, 90, 120, and 180 mins.
High-resolution
Quantitative high-resolution CBF was measured using the continuous arterial spin-labeling technique (Silva et al, 1999; Duong et al, 2000) with gradient-echo EPI and parameters similar to the ADC measurement except
Data Analysis
Data analysis used codes written in Matlab (MathWorks, Natick, MA, USA) and displayed in STIMULATE (Strupp, 1996) software. Only six anterior slices were analyzed to avoid susceptibility distortion around the ear canals. All data were reported as mean±s.d.
Receiver operating characteristic curves were also generated for each algorithm by plotting TPR (sensitivity) against the false-positive ratio (FPR, which is defined as 1–specificity). The area under the ROC curve was calculated as an indicator of the algorithm performance (Hanley and McNeil, 1982). Optimal operating point on the ROC curve (Wu et al, 2001; Obuchowski, 2003) was obtained by determining the combined sensitivity and specificity that maximizes the function,

where

Assuming equal probability of infarcted (0.5) and noninfarcted (0.5) voxels and equal costs of false-positives and false-negatives,
Results
Figure 1A shows the
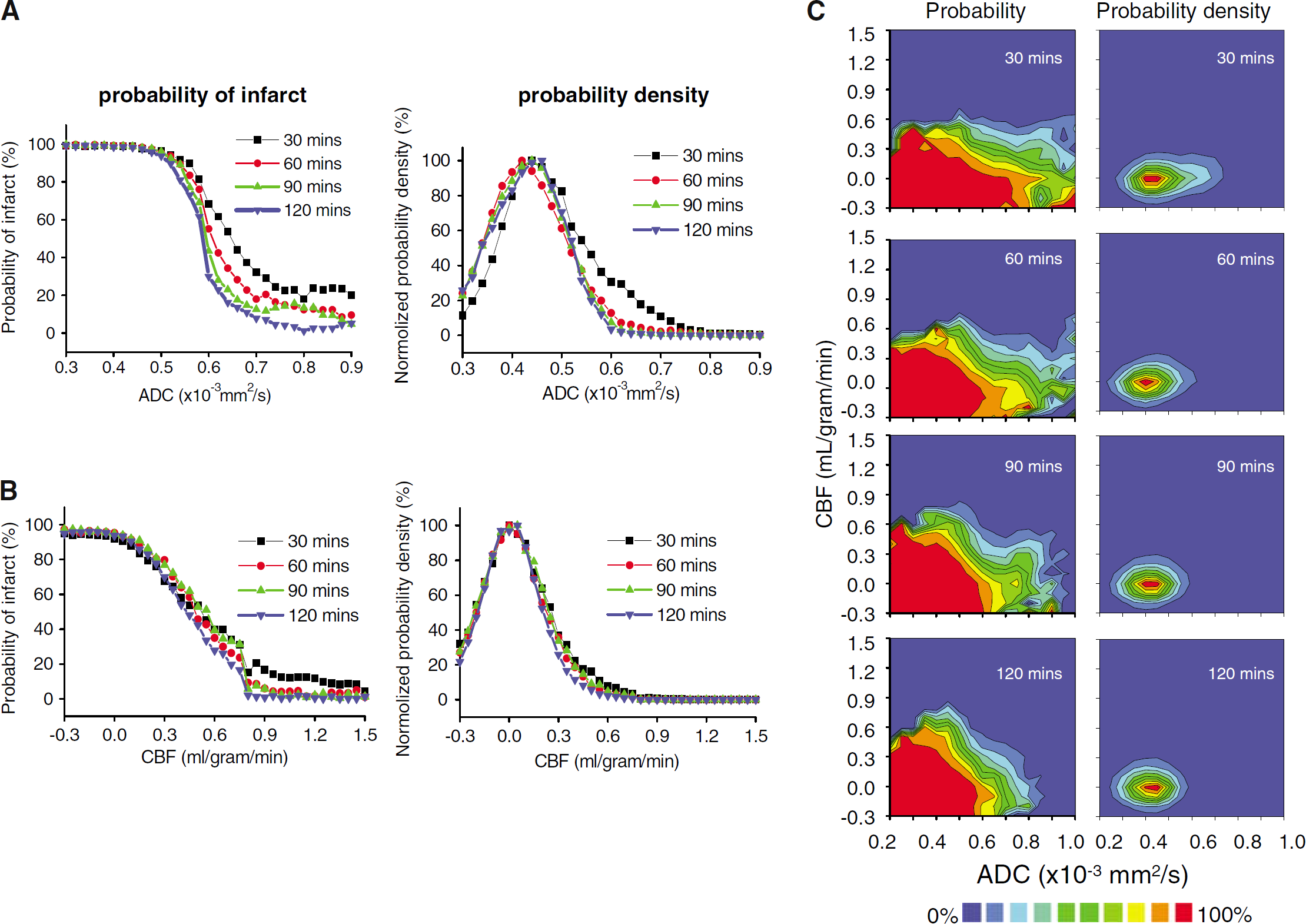
Profiles of probability of infarct and probability density of infarct as a function of (
Two-dimensional
Probability maps of risk of subsequent infarction were computed on a separate group of animals (Group B,
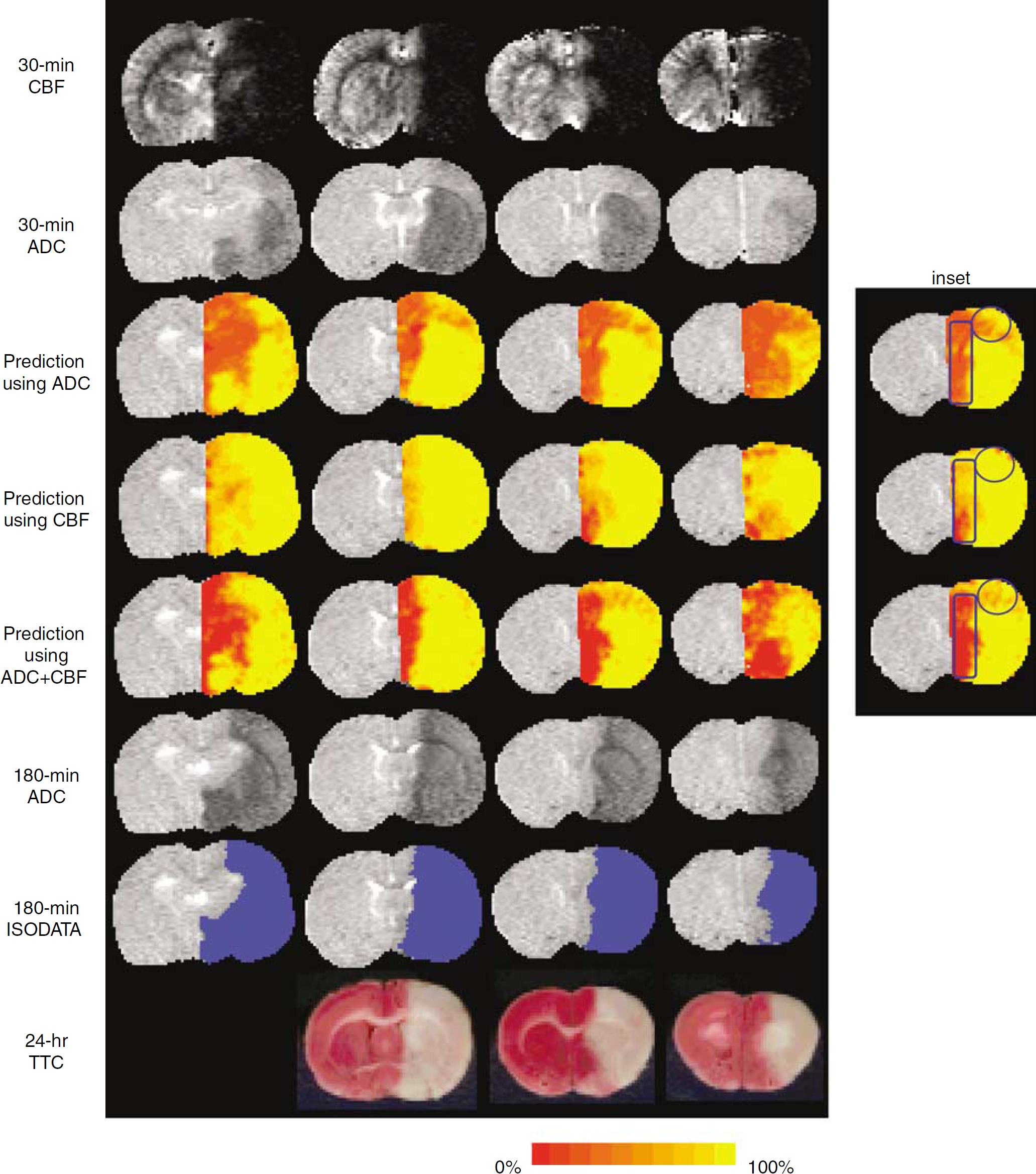
Probability maps of risk of future infarction determined based on the 30-min apparent diffusion coefficient (ADC) data alone, cerebral blood flow (CBF) data alone, and ADC + CBF data from a representative animal (experimental Group B). Multislice images are displayed from left to right as posterior to anterior slices. Also shown are the 30-min ADC and CBF maps, 180-min ADC maps, ISODATA analysis at 180 mins, and 24-h TTC staining. Hypointensities in the ADC and CBF maps indicate regions of reduced ADC and CBF values, respectively. Red-yellow color bar indicates the probability of infarct ranging from 0% to 100% in steps of 10%. The inset shows prediction using ADC alone underestimated infarct volume whereas CBF alone overestimated infarct volume (circular ROI). Prediction using ADC + CBF showed ‘normal’ tissues having a high degree of certainty of not going into infarct (low probability, rectangular ROI) relative ADC or CBF alone.
Receiver operating characteristic curves were used to evaluate the accuracy of the predictions made by using ADC alone, CBF alone, and ADC + CBF (Figure 3, Group B). The optimal operating point, sensitivity, specificity, area under the ROC curve, and PAI are summarized in Table 1. Predictions made using ADC + CBF data generally showed higher sensitivity, specificity, area under ROC curve, and PAI than those using ADC alone or CBF alone. At the optimal operating points, combined ADC + CBF predicted tissue infarction with 86%±4% sensitivity and 89%±6% specificity.
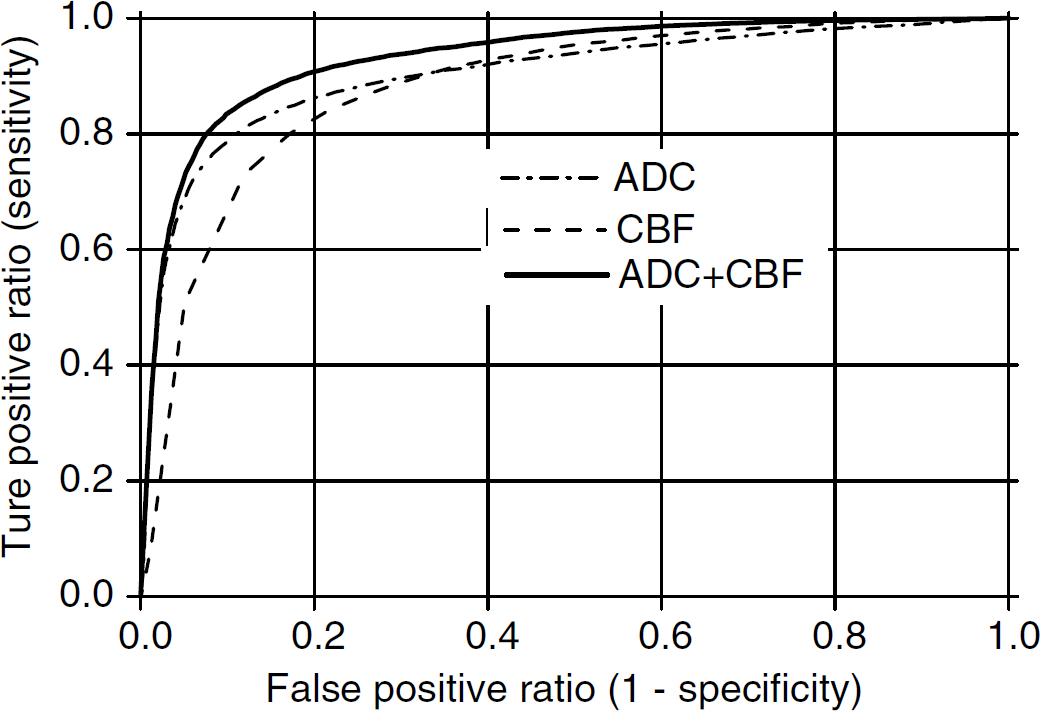
Receiver operating-characteristic (ROC) curves of sensitivity versus (1–specificity) of the overall prediction of tissue fate. Data were derived from incorporating the apparent diffusion coefficient (ADC) data, cerebral blood flow (CBF) data, and ADC + CBF data (Group B,
Group-average performance measures of the prediction algorithm of overall tissue fates (
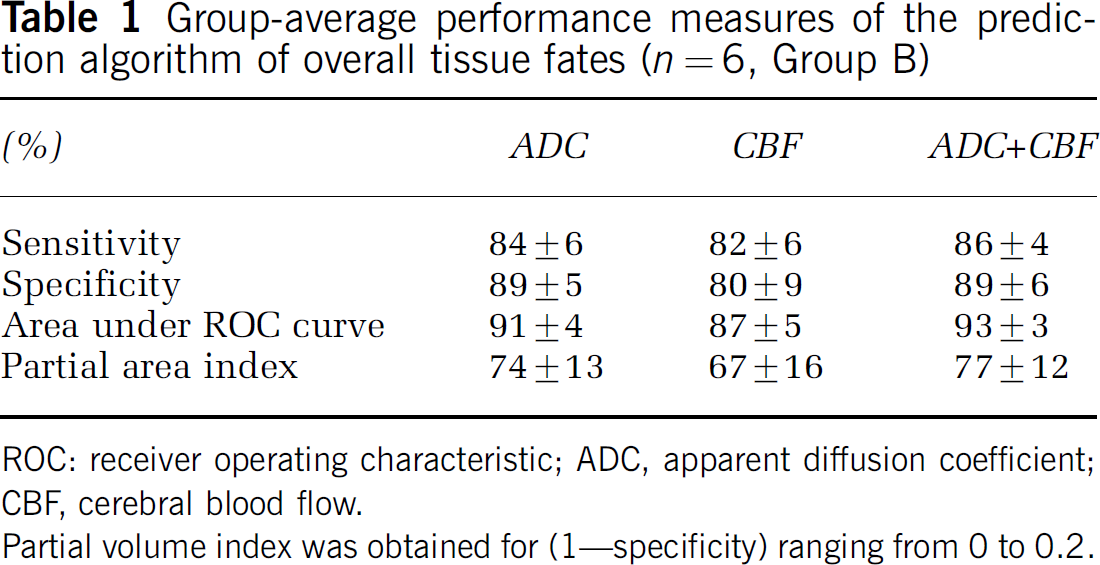
ROC: receiver operating characteristic; ADC, apparent diffusion coefficient; CBF, cerebral blood flow.
Partial volume index was obtained for (1–specificity) ranging from 0 to 0.2.
To further evaluate the performance of the prediction algorithm, predicted infarct volumes were derived at the optimal operating point and correlated with TTC infarct volumes. The optimal operating point is effectively an objective threshold (similar to viability threshold) and may be used in clinical decision making. The predicted infarct volume derived from ADC + CBF (189±44 mm3) provided better correlation with TTC infarct volumes (190±51 mm3), whereas ADC alone (187±64 mm3) slightly underestimated and CBF alone (206±43 mm3) slightly overestimated infarct volumes. The large uncertainties largely reflected cross-subject variations. The correlation coefficients were 0.88, 0.92, and 0.96 for CBF alone, ADC alone, and ADC + CBF, respectively.
In addition to predicting
(A) Group-average probability of infarct (
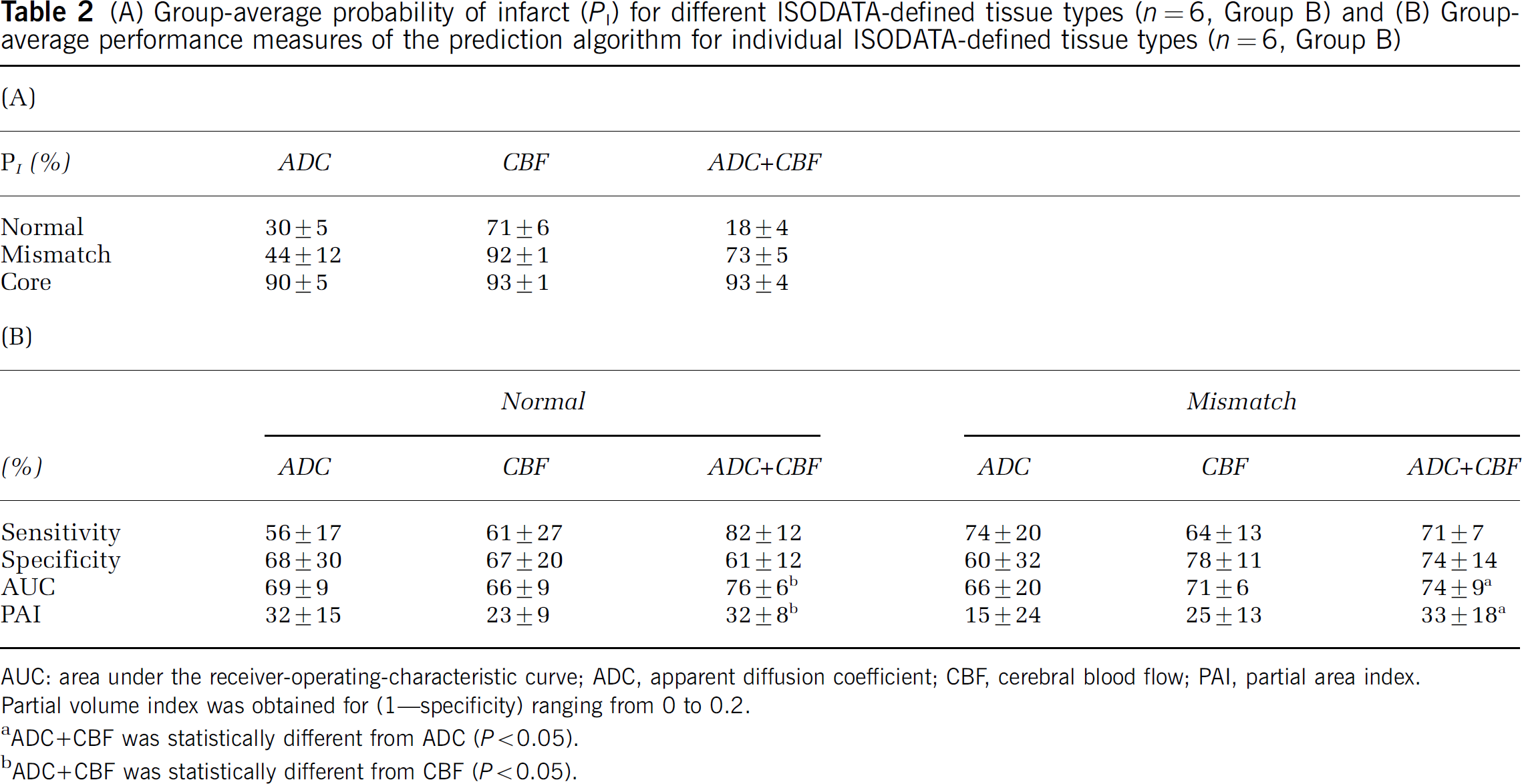
AUC: area under the receiver-operating-characteristic curve; ADC, apparent diffusion coefficient; CBF, cerebral blood flow; PAI, partial area index. Partial volume index was obtained for (1–specificity) ranging from 0 to 0.2.
ADC + CBF was statistically different from ADC (
ADC + CBF was statistically different from CBF (
Receiver operating characteristic analyses of the predictions made for individual ISODATA cluster are shown in Figure 4 (Group B). The group-average optimal operating point, sensitivity, specificity, area under the ROC curve and PAI are summarized in Table 2B. For the normal cluster, performances of the prediction made with ADC + CBF showed clear improvement over those made with ADC or CBF alone. For the mismatch cluster, performances of the prediction also showed clear improvement over those made with ADC or CBF alone. ROC analysis of the core cluster was not performed because the false-positive ratio is essentially zero (i.e., essentially all core pixels stayed core). Area under the ROC curve and PAI of the prediction made on ADC + CBF data showed the best performance for all tissue types.
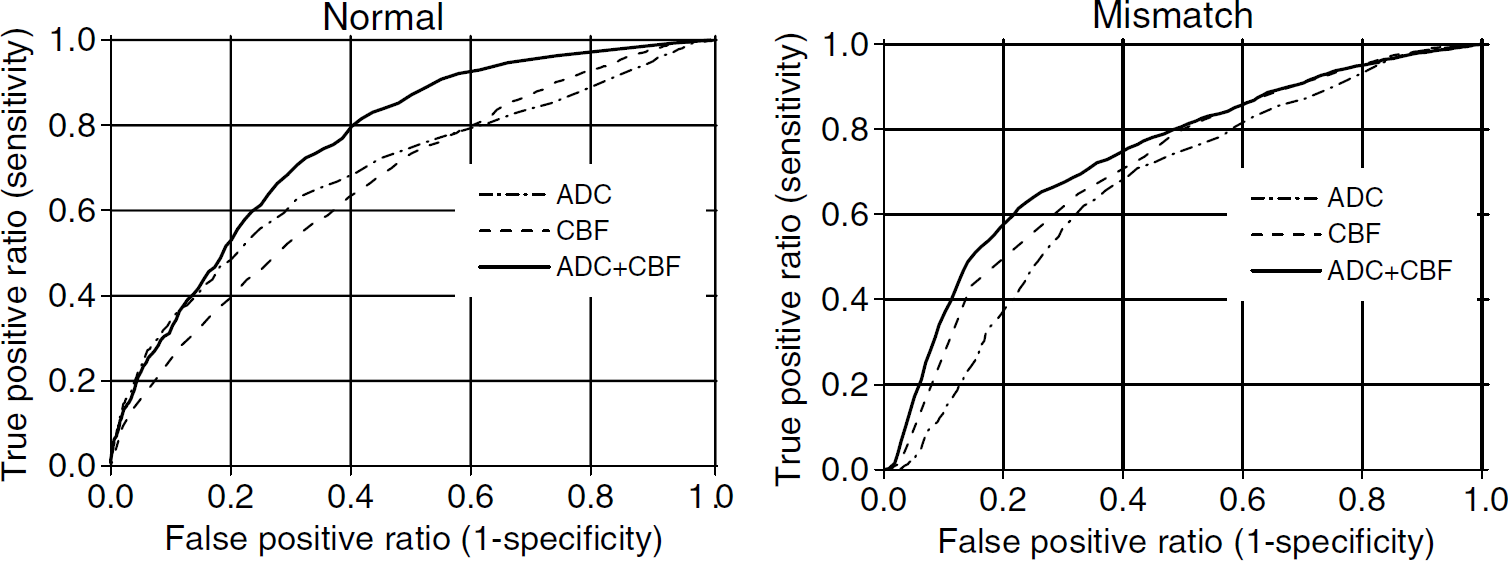
Receiver operating-characteristic (ROC) curves of sensitivity versus (1–specificity) of each ISODATA-derived (normal, mismatch and core) pixel clusters derived from incorporating the apparent diffusion coefficient (ADC) data, cerebral blood flow (CBF) data and ADC±CBF data (Group B,
Discussion
An intuitive statistical algorithm was developed to predict ischemic tissue fate in acute experimental stroke based on quantitative high-resolution perfusion and diffusion imaging. This algorithm was tested in a well-established experimental rat stroke model during the acute phase and at different time points of the ischemic progression. Probability, probability density profiles, and prediction maps of risk of future infarction provided intuitive information of ischemic tissue fate and showed excellent correlation with end point imaging and histology. Furthermore, by combining with an automated ISODATA tissue segmentation analysis, probabilities of infarct for different tissue types (i.e., normal, mismatch, and core pixels) were also determined on a pixel-by-pixel basis. Performance measures demonstrated that the prediction based on ADC + CBF data generally outperformed those based on ADC or CBF alone.
Multi-Parametric Cluster Analysis
Accurate classification of tissue types is important to properly generate a training set for probability prediction. At the minimum, ‘core’ pixels must be defined. The criteria used to classify such ‘core’ pixels could affect the probability distribution and, thus, the outcome of prediction. Threshold-based criteria for ADC, CBF, and T2 changes as well as follow-up imaging to define the ‘core’ pixels had been used. Viability thresholds were typically defined as being one or two standard deviations below the normal values or are based on correlation with a histological ‘gold standard,‘ which may be difficult to validate precisely. ISODATA cluster analysis, instead of threshold-based analysis, was used in this study to define ‘core’ pixels. ISODATA cluster analysis is an automated, unsupervised multiparametric clustering method, which requires minimal user intervention, and the infarct volume derived with this method showed excellent correlation with the histological infarct volume as shown previously in the same animal stroke model (Shen et al, 2004b). Only ADC and CBF data were used because they were informative during the acute phase; changes in T1 and T2 relaxation times did not occur until a few hours after stroke onset coinciding with the advent of the vasogenic edema.
Prediction Made with ADC or CBF Alone, and ADC + CBF
Apparent diffusion coefficient and CBF values at which there is a 50% probability of infarct,
Performance measures allowed further comparisons to be made. Prediction based on ADC data alone showed a relatively high false-negative prediction that resulted in a relatively low prediction sensitivity and thus underestimated the infarct volumes. This is because of the presence of the ‘mismatch’ pixels at the early time points, which became infarcted in a time-dependent manner. The sensitivity of prediction based on ADC data alone was indeed poorer at the earlier time points. On the contrary, prediction based on CBF data alone had relatively high false-positive prediction, resulting in relatively low specificity and thus overestimated the infarct volumes. This is because not all pixels with mild CBF reduction become infarcted (Shen et al, 2004a, b ) (i.e., the persistent ‘mismatch’). Predictions made using combined ADC + CBF data yielded a balance between sensitivity and specificity, resulting in an overall improvement in the algorithm performance.
Comparison with Existing Prediction Algorithms
Reports on pixel-by-pixel and statistical prediction of ischemic tissue fate in form of probability maps are sparse. Zhao et al (1997) developed ‘infarct frequency distribution maps’ to show the probability of tissue infarction based on coregistered histological slides. Welch et al (1995) developed a signature model based on diffusion- and T2-weighted imaging during the subacute stroke phase and used it to predict reversible and irreversible ischemic brain injury. More recently, Wu et al (2001) propose a generalized linear model (GLM) analysis of multiparametric data to predict ischemic brain injury. Perfusion imaging using the Gd-DTPA bolus tracking technique was used to derive CBF, cerebral blood volume, mean transit time, and time to peak data. Diffusion and T2 data were also measured. Tissue classification in the training data set was based on ROIs of tissues that were clearly infarcted or normal. Probability maps of infarction were derived. Their prediction algorithm showed 66% sensitivity and 84% specificity. One major advantage of GLM application to multispectral analysis is computational speed because it involves solving a system of linear equations instead of performing iterative regression analysis on multiple parameters; the latter is computationally expensive as the number of parameters in the basis set grows. One drawback is that the predictions were relatively less intuitive, and contributions of various measured parameters with different signal-to-noise and contrast-to-noise ratios were difficult to evaluate. The highly variable outcomes in human stroke relative to animal stroke models contributed to relatively poor sensitivity and specificity, despite reasonable selection of stroke patients with similar stroke characteristic. The low sensitivity reported relative to specificity suggests that the prediction model is dominated by ADC and/or T2 parameters because ADC and T2 are the better indicators of final infarction and thus contributed more to specificity compared with other MR parameters. Only prediction of the
There are several advantages of the prediction algorithm and the animal stroke model used herein. First, the statistical prediction algorithm was based on quantitative high-resolution ADC and CBF data that offered a simple and intuitive means to predict ischemic tissue fate
Secondly, by combining the modified ISODATA clustering algorithm, probability of infarct could be evaluated for individual tissue types, namely, ‘normal’, ‘mismatch,’ and ‘core’ pixels. Although performance measures of overall tissue fate prediction made with ADC + CBF generally (but not always) showed improvement over those made with ADC or CBF alone, the improvement was surprisingly small. This is because the performance measures based on the prediction of
It was anticipated that prediction of the ‘normal’ tissue should improve with combined ADC + CBF and this was indeed observed. Contrary to expectation, prediction of the ‘mismatch’ tissue was not significantly improved with combined ADC + CBF over ADC or CBF alone. Although ADC + CBF has high sensitivity, it also showed increased false-positive prediction rate because not all mismatch pixels became infarcted. Furthermore, since the number of mismatch pixels was relatively small compared with other clusters, the false-positive prediction of the mismatch cluster had a relatively larger effect in the ROC analysis.
Finally, another advantage of the current approach is that the sample size of the training basis set can be improved by adding more subjects to the training set and prediction of infarct can be achieved by simply using the look-up table in a computationally inexpensive manner. To improve performance of the prediction algorithm, it is important to carefully select samples to be included in the training data set. Adding more parameters to the prediction algorithm does not necessarily improve prediction unless they are informative. The samples to be used in the training data also need to be properly categorized. For example, permanent and transient ischemia with or without therapeutic intervention need to be distinguished and/or different prediction algorithms need to be generated for different conditions. Poor performance of the prediction algorithm is likely to be caused by not properly selecting samples to be included in the training data set and not properly applying the appropriate prediction algorithm to specific type of ischemic injury. Establishing proper training basis sets and selecting the proper algorithm, however, may not be trivial in clinical practice. Combined tissue classification and probability prediction nonetheless offer the potential to accurately predict ischemic tissue fate in the clinical settings in a computationally inexpensive manner and is currently under investigation.
Conclusion
This study showed a simple and intuitive pixel-by-pixel probability-based algorithm to statistically predict ischemic tissue fate of different tissue types based on early quantitative perfusion and diffusion data. The accuracy of infarct prediction could be quantitatively evaluated. This algorithm was tested on an established experimental stroke model. Performance measures demonstrated that prediction made based on ADC + CBF data outperformed those based on ADC data alone or CBF data alone. Predicted tissue fate based on ADC + CBF data showed remarkable correlation with histological and ISODATA-defined lesion volumes on a pixel-by-pixel basis. This approach provides a novel means to analyze stroke data. The implication is that prediction of tissue infarct could be achieved by establishing catalogs of training data sets for different types of ischemic injury and by developing an algorithm to statistically determine which of the training data set should be applied to specific stroke patients in a computationally inexpensive manner.
