Abstract
The cerebral vasculature ensures proper cerebral function by transporting oxygen, nutrients, and other substances to the brain. Distribution of oxygenated blood throughout the neuroaxis takes place at the level of the circle of Willis (CW). While morphologic and functional alterations in CW arteries and its main branches have been reported in cerebrovascular and neurodegenerative diseases, accompanying changes in protein expression profiles remain largely uncharacterized. In this study, we performed proteomics to compile a novel list of proteins present in mouse CW arteries and its ramifications. Circle of Willis arteries were surgically removed from 6-month-old wild-type mice, proteins extracted and analyzed by two proteomics approaches, gel-free nanoLC-mass spectrometry (MS)/MS and gel-based GelLC-MS/MS, using nanoAcquity UPLC coupled with ESI-LTQ Orbitrap XL. The two approaches helped maximize arterial proteome coverage. Six biologic and two technical replicates were performed. In all, 2,188 proteins with at least 2 unique high-scoring peptides were identified (6,630 proteins total). Proteins were classified according to vasoactivity, blood–brain barrier specificity, tight junction and adhesion molecules, membrane transporters/channels, and extracellular matrix/basal lamina proteins. Furthermore, we compared the identified CW arterial proteome with the published brain microvascular proteome. Our database provides a vital resource for the study of CW cerebral arterial protein expression profiles in health and disease.
INTRODUCTION
A healthy cerebral circulation is essential for maintaining brain perfusion and function. 1 In human, oxygenated blood to the brain is delivered by the internal carotid and vertebrobasilar arteries, and is distributed throughout the neuroaxis at the level of the circle of Willis (CW), a ringlike arterial structure located in the subarachnoid space at the base of the brain. The CW is formed by the confluence of the anastomotic branches of the two internal carotid arteries, rostral portion of the vertebrobasilar artery, and the anterior and posterior communicating arteries. 2 The arterial wreath allows for communications between the anterior and posterior circulations providing blood to the forebrain and the hindbrain, respectively, and insures redundancies in the cerebral circulation. 3 The three principal brain arteries arising from the CW are the left and right anterior, middle, and posterior cerebral arteries, cortical branches of which penetrate the brain parenchyma to irrigate the cerebral cortex and deep structures of the brain. 2 In mice, the CW is similarly located along the ventral aspect of the brain extending from the pons-midbrain junction to the anterior cerebrum and involves the same major arteries. 4
Several studies have reported morphologic and protein expression changes in the CW and its surface branches in healthy aging 5 as well as pathologic conditions, such as cerebrovascular and neurodegenerative diseases (e.g., Alzheimer's disease).5,6 Arterial wall thickening, loss of elasticity, and alterations in perivascular innervation are common changes observed in aged cerebral arteries. 5 Pathology-associated alterations observed in brain arteries/arterioles resemble an admix of accelerated vascular aging and disease-specific hallmarks, such as amyloid deposits in Alzheimer's disease. To date, in comparison with the plethora of structural and functional changes documented in CW arteries of human and animal models, only a limited number of differentially expressed proteins have been identified. The main aim of our study was to generate, and compile a database of the mouse CW arterial proteome that will provide a valuable resource to study differential protein expression patterns in healthy and diseased brain arteries. In this paper, we have developed methodologies for efficient, mass spectrometry (MS)-compatible arterial protein extraction and peptide detection, and compared the CW arterial proteome with a previously published 7 brain microvascular proteome.
MATERIALS AND METHODS
Mice
Eighteen 6-month-old C57BL/6J wild-type (WT) mice from two different breedings (B1 and B2) of nine mice each were used in this study. Furthermore, two 6-month-old WT mice from a third breeding were used for protein validation. To eliminate potential gender-related differences in brain vasculature only males were used. Mice were housed under a 12-hour light-dark cycle, in a temperature (23°C) and humidity (50%) controlled room, with food and tap water available
Surgical Extraction of Circle of Willis Cerebral Arteries
Mice were anesthetized (65 mg/kg sodium pentobarbital, intraperitoneal) and transcardially perfused (0.9% saline, 4°C) for 5 minutes to clear the brain vasculature of blood contaminants. After each perfusion, brain was extracted, immersed in a precooled (4°C) plexiglass-dissection bath containing 0.9% saline, and secured (ventral side up first) with two pins to the transparent silicone cushion at the bottom of the bath. The CW, cerebral arteries (anterior, middle, and posterior), and their main ramifications were surgically removed from the pia mater under a dissecting microscope. Thereafter, isolated arteries underwent a second round of attached pia mater removal. Similarly prepared ‘pure’ cerebral arteries from three mice were combined (constituted one biologic replicate) and stored at –80°C. A total of six (B1a, B1b, B1c, B2a, B2b, and B2c) biologic replicates were prepared for this study.
Protein Extraction from Circle of Willis Arteries
Proteins were extracted from surgically isolated arteries using Rapigest SF (Waters, Milford, MA, USA), a MS-compatible acid-labile surfactant that enhances solubilization and tryptic digestion efficiencies of proteins in complex biologic mixtures.
8
To each surgically isolated sample, we added 100 μL of 0.2% (w/v) Rapigest SF in 50 mmol/L ammonium bicarbonate. The mixture was then exposed to three 20-second pulses of sonication (550 Sonic Dismembrator; Fisher Scientific, Ottawa, ON, Canada) to further facilitate protein extraction. The above steps were all performed on ice. After sonication, each preparation was incubated at 95°C for 10 minutes, followed by a 20-minute centrifugation (Eppendorf Centrifuge 5415D; Brinkmann Instruments, Westbury, NY, USA) at 10,000
Proteomics Methods for Analyses of Cerebrovascular Proteins
We used two different approaches, gel-based and gel-free, to maximize CW arterial proteome coverage. Both approaches reduced sample complexity by fractionating either (1) at the level of the protein or (2) at the peptide level, respectively. It should be noted that while all six biologic replicates underwent gel-free proteomics, only three samples (B1a, B1b, and B1c) were additionally subjected to the gel-based method.
NanoLC-MS/MS Analysis
For each of the six biologic replicates, each SCX and/or in-gel-digest fraction was analyzed twice (technical replicates) by nanoLC-MS/MS. This culminated to a total of 96 nanoLC-MS/MS runs. In detail, each fraction was analyzed using ESI-LTQ-Orbitrap-XL mass spectrometer (Thermo) coupled to a NanoAcquity UPLC system (Waters). Samples were loaded on a NanoAcquity symmetry C18 trap (Waters) and separated on a 10 cm × 100 μm I.D. C18 column (Waters, 1.7 μm BEH130C18) at 250 nL/min using a 68-minutes gradient: 0% to 45% solution B (100% ACN/0.1% formic acid) over 66 minutes and 45% to 95% solution B over 2 minutes. Ten-minute washes in 40% solution B were followed by 30 minutes blank gradients between each sample to minimize carryover effects. The mass spectrometer was set to automated MS/MS analysis with an MS scan at 60 k resolution in the Orbitrap analyzer and data-dependent turbo MS/MS scans on the top three ions in the trap with dynamic exclusion (180 seconds).
Protein Identification and Bioinformatics
The raw data generated from LTQ-Orbitrap were converted to mzXML format using ReAdW program (http://tools.proteomecenter.org). MGF files were then generated from the mzXML file using MzXML2Search from the Trans Proteomics Pipeline project (http://tools.proteomecenter.org) and the resulting MS/MS spectra were searched against
Characterization of Arterial Protein List
Two protein lists (List 1 and List 2) of varying stringency were generated. All proteins with
Western Blotting
Western blots were performed to validate List 1 proteins discoidin domain receptor tyrosine kinase 2 (
Frozen CW arteries from each mouse (
Comparison with Microvascular Proteome
List 2 arterial proteins were compared with the previously released cerebral microvessel protein data set.
7
Inclusion criteria used for microvascular proteins were the following: (1) the presence of ≥2 unique tryptic peptides, (2) the presence of protein in all technical replicates, and in at least one biologic replicate, and (3) an average spectral count of ≥5. Common and unique proteins to each data set were identified. The PANTHER Classification System was used to classify proteins by pathways. Proteins were listed as (1)
RESULTS
Extraction and Solubilization of Cerebral Arteries
The CW and its ramifications (Figure 1A) were extracted in a reproducible manner using the surgical procedure described (see Materials and methods). Figure 1B outlines the protein solubilization and digest workflow developed in-house for efficient protein extraction from CW cerebral arteries. On average, 87 μg (ranged from 70 to 110 μg) proteins were extracted per biologic replicate. Figure 1C shows the reproducibility of two technical and biologic replicates, where the median relative standard deviation between technical replicates is 6% and between biologic replicates is 18%, values similar to that published for gel-free analysis. 16
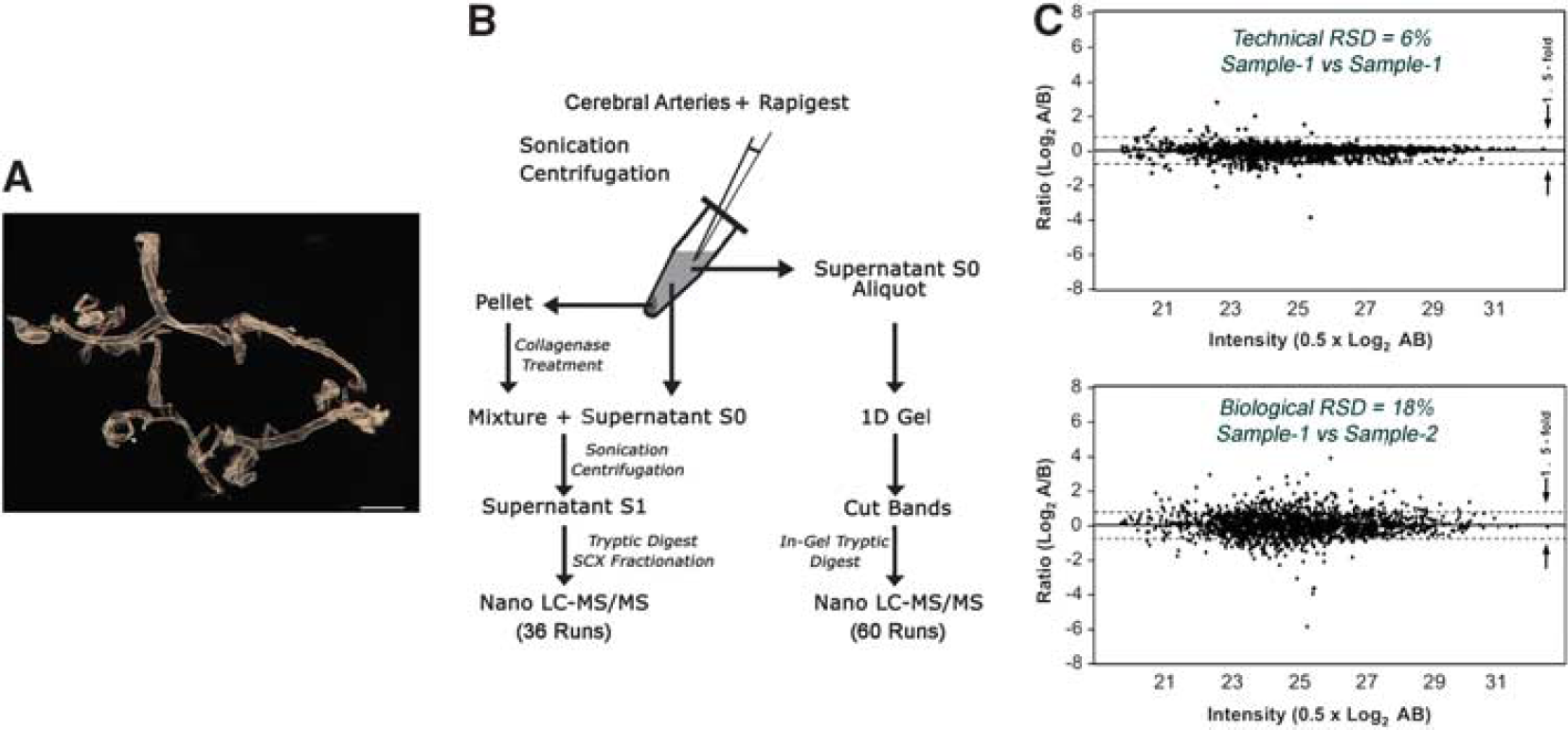
Generation of the circle of Willis (CW) cerebral arterial database.
Proteomic Profile
A total of 6,630 CW cerebral arterial proteins (List 1 proteins, Supplementary Table 1) were identified at
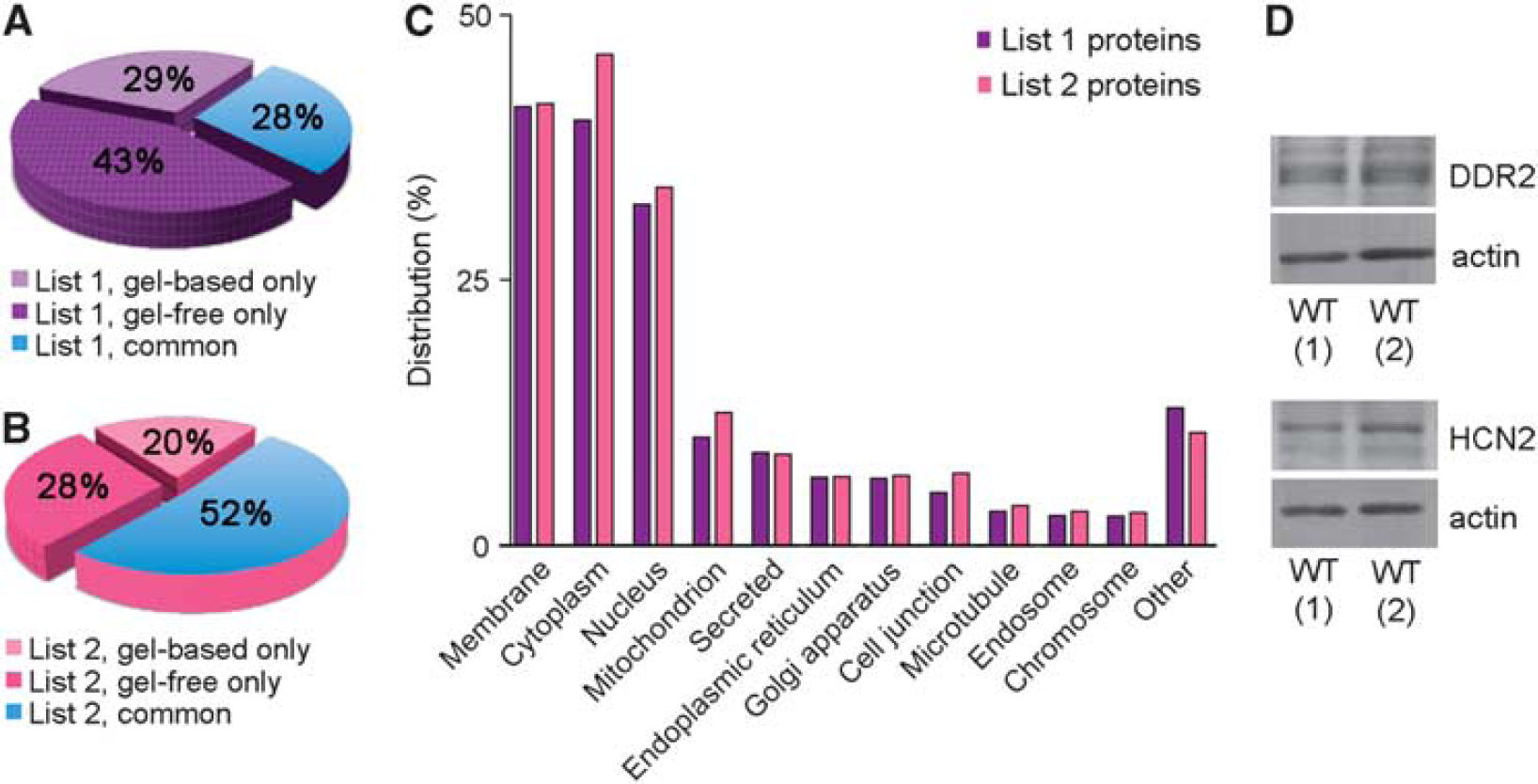
Circle of Willis (CW) cerebral arterial protein database.
Proteins involved in vasomotricity
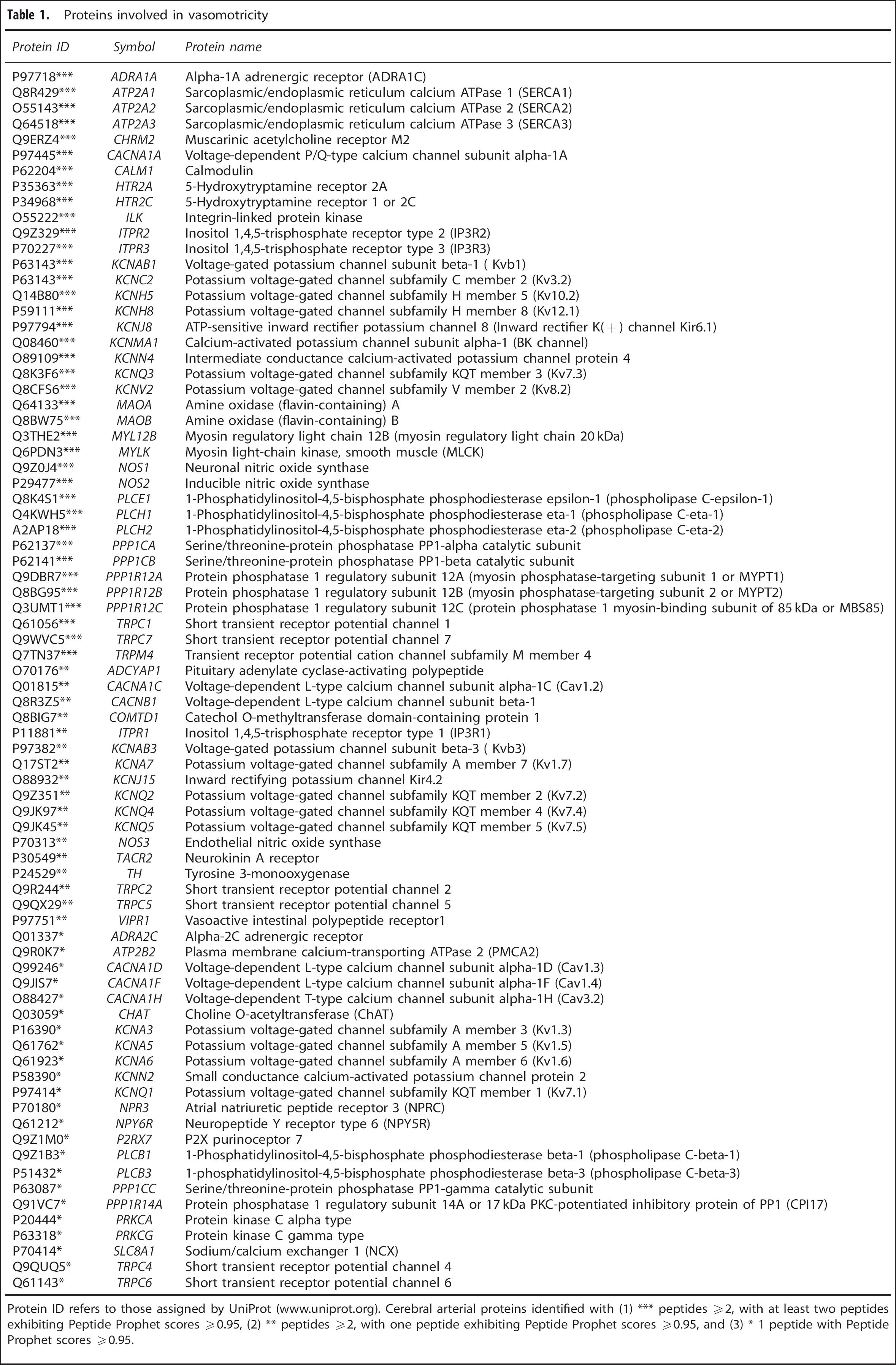
Protein ID refers to those assigned by UniProt (www.uniprot.org). Cerebral arterial proteins identified with (1) *** peptides ≥2, with at least two peptides exhibiting Peptide Prophet scores ≥0.95, (2) ** peptides ≥2, with one peptide exhibiting Peptide Prophet scores ≥0.95, and (3) * 1 peptide with Peptide Prophet scores ≥0.95.
A total of 2,188 List 1 proteins (33%) were identified with ≥2 peptides each and were cataloged as List 2 proteins (Supplementary Table 1). Approximately half (52%) of these were detected using both gel-based and gel-free approaches (Figure 2B). The top five subcellular locations of List 2 proteins were cytoplasm (46.3%), membrane (41.7%), nucleus (33.7%), mitochondrion (12.6%), and secreted (8.6%) (Figure 2C). A total of 466 proteins (21% of List 2 proteins) belonged to curated pathways in the PANTHER database. Ten pathways with >20 proteins were observed. In descending number of proteins they were as follows:
Vasoactivity
We detected 78 proteins shown to regulate the cerebral circulation or exert vasomotor effects on cerebral arteries (Table 1; Supplementary Table 2 for references and additional information). A few of these, specifically, neuronal mediators of vasoactivity and their vascular receptors detected in our arterial data set have been highlighted in the subsections below, and are illustrated in Figure 3.
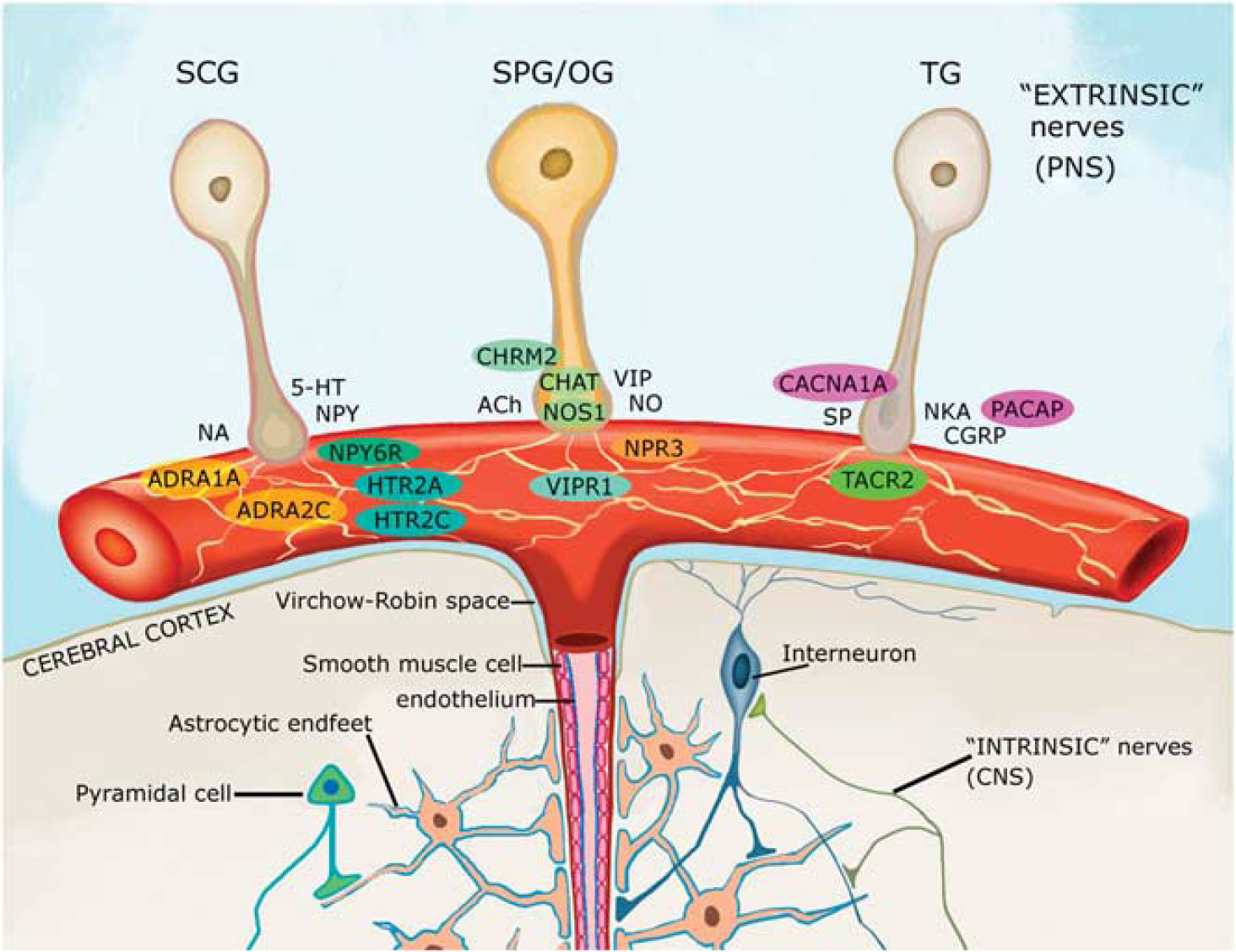
Neuronal mediators of vasoactivity. Perivascular nerve fibers originating from various peripheral ganglia release vasoactive mediators that act on specific receptors in the vascular wall, many of which were detected in our arterial data set and have been highlighted with colored circles. Protein IDs in figure refer to those assigned by UniProt (www.uniprot.org). In alphabetical order, detected proteins were as follows: ADCYAP1 or PACAP, pituitary adenylate cyclase-activating polypeptide; ADRA1A and 2C, the α-1A and –2C adrenergic receptors; CACNA1A, alpha1A subunit of the voltage-dependent P/Q-type calcium channel; CHAT (or ChAT), choline o-acetyltransferase; CHRM2 (or mAChR2), muscarinic acetylcholine receptor M2; HTR2A and 2C, serotonin receptors 2A and 2C; NOS1, neuronal nitric oxide synthase; NPR3, natriuretic peptide receptor C (also known to bind VIP); NPY6R, neuropeptide Y receptor 6; VIPR1, vasoactive intestinal polypeptide receptor 1; TACR2, neurokinin A receptor. 5-HT, 5-hydroxytryptamine; ACh, acetylcholine; CGRP, calcitonin gene-related peptide; CNS, central nervous system; NA, noradrenaline; NKA, neurokinin A; NO, nitric oxide; NPY, neuropeptide Y; PNS, peripheral nervous system; SCG, superior cervical ganglion; SP, substance P; SPG/OG, sphenopalatine and otic ganglia; TG, trigeminal ganglion; and VIP, vasoactive intestinal polypeptide. Adapted from Hamel 1 with permission from author (EH).
Cerebral artery proteome classified into BBB-specific proteins, tight junction and adhesion molecules, membrane transporters and channels, and ECM and basal lamina proteins
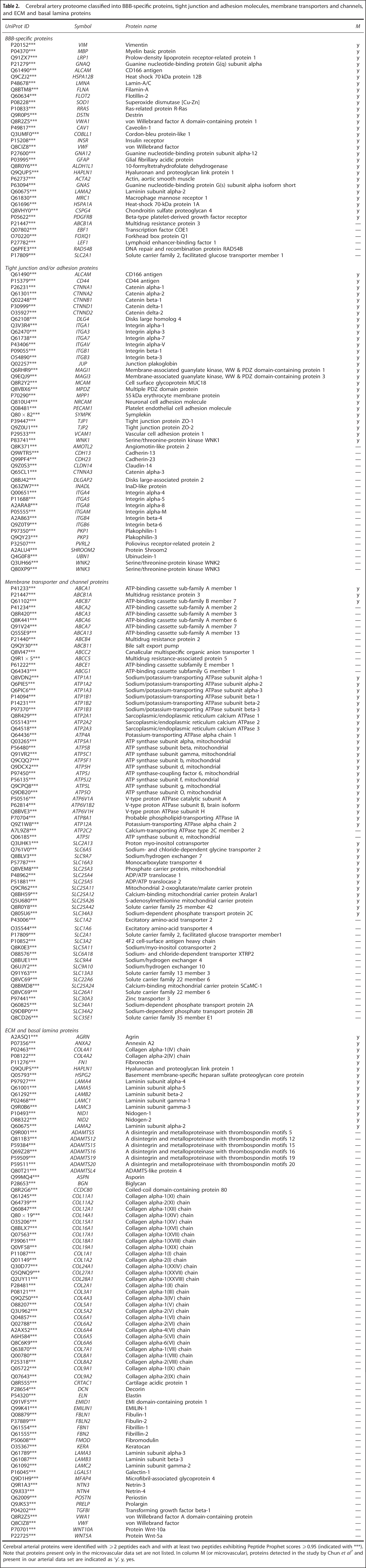
Cerebral arterial proteins were identified with ≥2 peptides each and with at least two peptides exhibiting Peptide Prophet scores ≥0.95 (indicated with ***). Note that proteins present only in the microvascular data set are not listed. In column M (or microvascular), proteins detected in the study by Chun
In addition, we detected various smooth muscle and endothelial mediators of vasoactivity (Table 1; Supplementary Table 2 for references and additional information), including endothelial NO synthase (
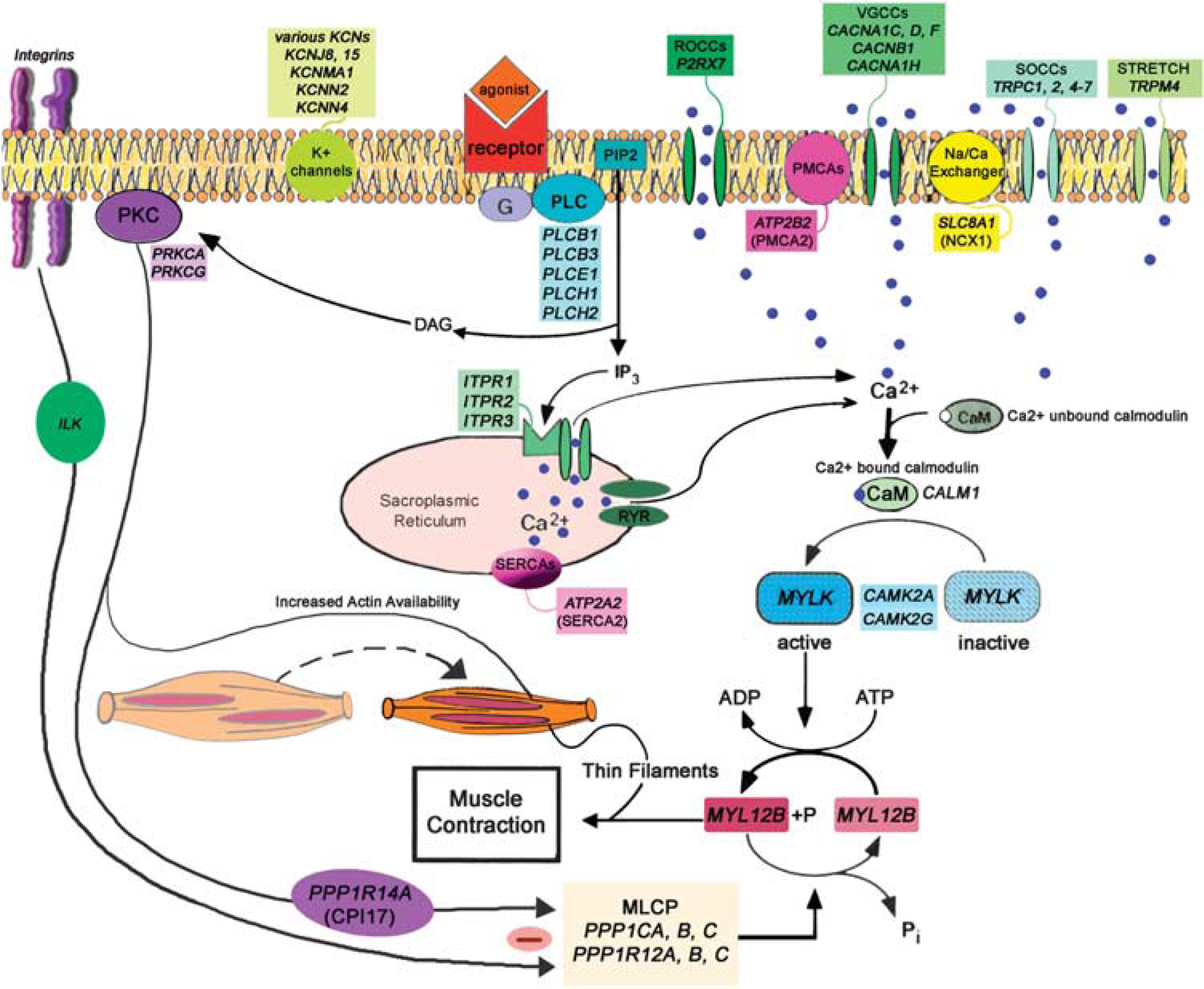
Protein effectors of arterial smooth muscle contraction and dilation. Simplified representation of proteins and protein pathways involved in vascular smooth muscle contraction. Protein effectors present in our arterial database are indicated in
Cerebral Arterial Versus Microvascular Proteomes
A total of 3,411 unique proteins were present in the arterial (List 2, 2,188 proteins total) and microvascular (1,824 proteins total) data sets combined (Figure 5A). Of these, 1,587 (47%) and 1,223 (36%) proteins were unique to the arterial and microvascular data sets, respectively, whereas 601 (18%) were common to both (Figure 5A). These common proteins comprised 28% of the List 2 CW cerebral arterial data set and 33% of the microvascular data set. Analysis of common proteins yielded a total of 62 PANTHER pathways, the top 5 being
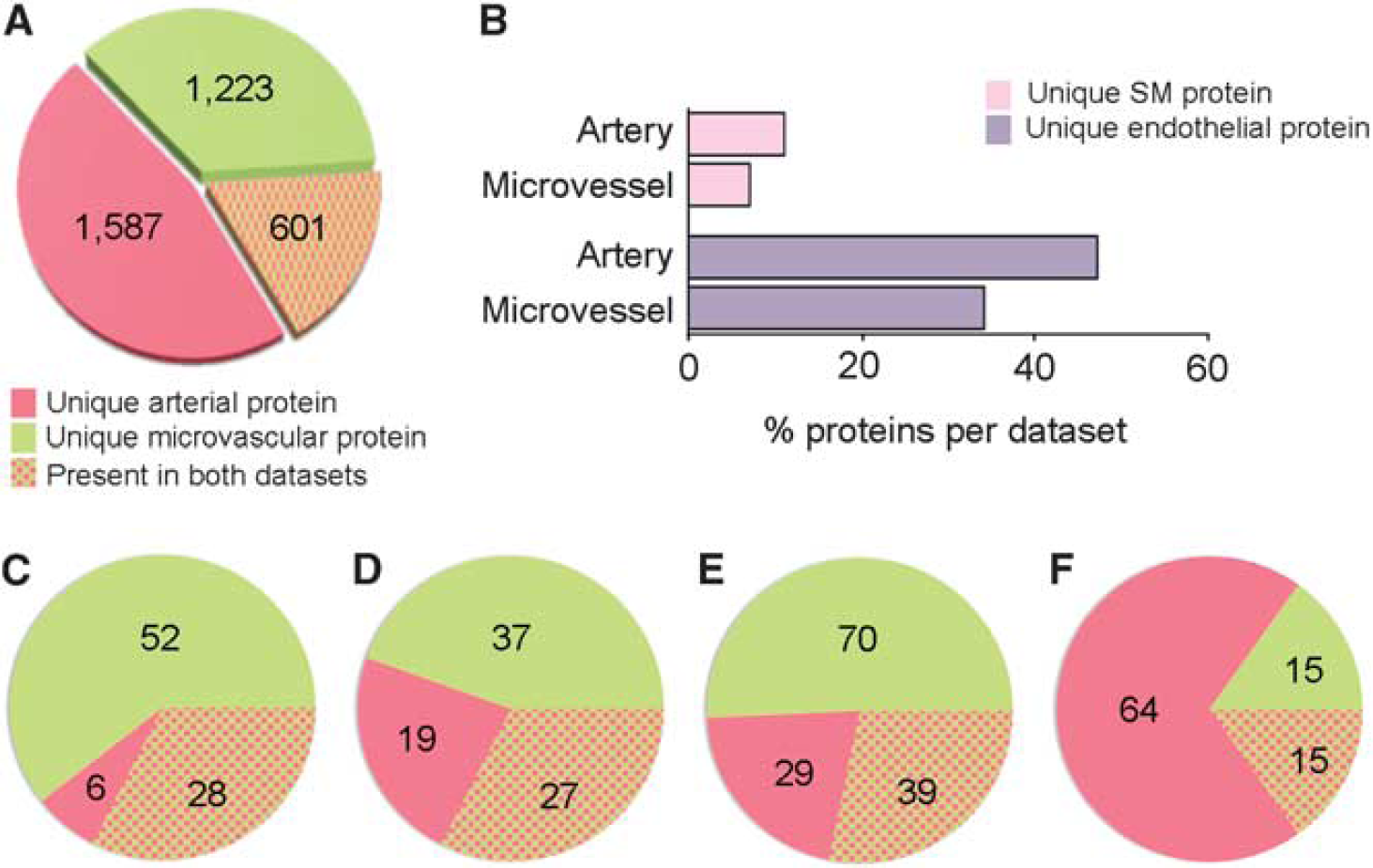
Cerebral arterial versus microvascular proteomes.
DISCUSSION
We have generated the first extensive database of 6,630 proteins expressed in the wall of fresh cerebral arteries. An advantage of using freshly isolated over cultured tissue is that it allows for maximal preservation of the tissue microenvironment experience
Arterial Proteome Coverage
To generate a comprehensive arterial protein database, we developed highly reproducible methodologies that allowed for (1) rapid surgical isolation of fresh cerebral arteries, (2) efficient protein extraction using the MS-compatible surfactant Rapigest, with yields ~3 × higher compared with SDS-based extraction (unpublished data), and (3) optimal detection and quantification of low-abundant peptides. Concomitant use of gel-based and gel-free approaches contributed to increased detection of peptides (and proteins) as showcased by (1) the presence of EC layer proteins, which are harder to detect since unlike the SMC and adventitia layers, the EC layer is a single layer of cells and (2) good agreement between the breakdown of detected arterial proteins by subcellular localization and the relative sizes of cellular compartments (e.g., membrane > mitochondria).
Cerebral Arteries and Vasoactivity
Large arteries as well as smaller arterioles at the brain surface constitute resistance vessels that control blood flow and influence global perfusion.
5
Vascular resistance is essential for pushing blood through the cerebral circulation, and is increased through contraction of the vessel muscular wall, and decreased
We also detected nine neuronal mediators of vasoactivity (Table 1; Supplementary Table 2 for references; and Figure 3). Of these, the pituitary adenylate cyclase activating peptide (
Increases in iCa2+ are a major determinant of arterial smooth muscle contractility.
19
In SMCs, rise in iCa2+ is facilitated by (1) Ca2+ release from intracellular stores in the smooth sarcoplasmic reticulum and (2) Ca2+ entry from the extracellular space
Several pathways of Ca2+ entry from the extracellular space have been suggested, and include passive leak Ca2+ channels, as well as channels activated by membrane depolarization (voltage gated), agonist binding (receptor operated), depletion of Ca2+ from intracellular stores (store operated), or stretch.
28
Voltage-gated Ca2+ channels enriched in SMCs of major cerebral arteries and arterioles are of L, and to a lesser extent, T type.
29
We detected four α-1 subunits belonging either to the high-voltage-activated L-type (
In contrast to smooth muscle contraction, its relaxation is accompanied by a decrease in iCa2+ levels to resting values. Decrease in iCa2+ is facilitated by (1) reuptake into intracellular stores by Ca2+ ATPase pumps (smooth endoplasmic reticulum calcium ATPases, SERCAs) on the sarcoplasmic reticulum membrane and (2) extrusion from the cell by the plasma membrane Ca2+ ATPases (PMCAs) and the Na+/Ca2+ exchanger (Figure 4). We detected three SERCAs (
Insights into the Proteomes of Arteries and Microvessels
The recent release of the cerebral microvessel proteomics study 7 provided an opportunity to compare and contrast protein-based findings in pial/extracerebral CW arteries (this study) and intraparenchymal brain vessels. Vessel diameters in our sampling of mouse cerebral arteries were ~100 μm and lower, with the majority being large diameter vessels. The wide range of arterial diameters in our sampling is representative of the varied vessel sizes present at the brain surface, starting with the CW and its major arteries ranging from 75 to 100 μm in diameter. 36 Furthermore, it has been shown that peripheral branches of the major arteries range between 50 and 70 μm in diameter, while their ramifications coursing over the brain surface, reach 35 μm or lower in diameter, just before perforating as penetrating intracerebral arterioles.36,37 In comparison with our arterial sample, vessel diameters in the microvascular sampling 7 ranged between >20 μm and <100 μm. Thus, it can be concluded that while the majority of vessels in the microvascular sample tended toward small diameters, some overlap in vessel diameter size exists between the two studies.
Comparatively, the most striking difference between the arterial and microvascular data sets was the ~9 × increase in the number of
In summary, we have compiled a novel resource database for proteins present in the arterial wall of mouse CW and its surface ramifications. To ensure detection and identification of an extensive and comprehensive list of CW cerebral arterial proteins, we used fresh unfixed tissue and employed two parallel, custom proteomic approaches. Our database provides an excellent resource for the study of protein expression profile in healthy cerebral arteries and its perturbations in cerebrovascular diseases. Furthermore, we shed light on the difference and the similarities between brain arteries and microvessels at the level of the proteome.
Footnotes
The authors declare no conflict of interest.
ACKNOWLEDGMENTS
The authors thank Drs A Pshezhetsky and E Kanshin (CHU Sainte-Justine, Research Center, Montréal, QC, Canada) for initial protein extraction and detection trials, Ms CE Delaney and Mr L Tessier (IBS-National Research Council of Canada, Proteomics and Mass Spectrometry group, Ottawa, ON, Canada) for their technical assistance with protein isolation and mass spectrometry, Dr Xinkang Tong (Montreal Neurological Institute, Montréal, QC, Canada) for actin western blots, and Mr I Markovic for help with graphic illustrations.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
