Abstract
The hemodynamic response function (HRF) describes the local response of brain vasculature to functional activation. Accurate HRF modeling enables the investigation of cerebral blood flow regulation and improves our ability to interpret fMRI results. Block designs have been used extensively as fMRI paradigms because detection power is maximized; however, block designs are not optimal for HRF parameter estimation. Here we assessed the utility of block design fMRI data for HRF modeling. The trueness (relative deviation), precision (relative uncertainty), and identifiability (goodness-of-fit) of different HRF models were examined and test–retest reproducibility of HRF parameter estimates was assessed using computer simulations and fMRI data from 82 healthy young adult twins acquired on two occasions 3 to 4 months apart. The effects of systematically varying attributes of the block design paradigm were also examined. In our comparison of five HRF models, the model comprising the sum of two gamma functions with six free parameters had greatest parameter accuracy and identifiability. Hemodynamic response function height and time to peak were highly reproducible between studies and width was moderately reproducible but the reproducibility of onset time was low. This study established the feasibility and test–retest reliability of estimating HRF parameters using data from block design fMRI studies.
INTRODUCTION
The hemodynamic response function (HRF) reflects the regulation of regional cerebral blood flow in response to neuronal activation. Accurate modeling of the HRF is of interest in a number of areas of research.1, 2, 3, 4, 5 The HRF has a key role in the analysis of functional magnetic resonance imaging (fMRI) data 3 and variation in the HRF between individual subjects and between brain regions has prompted the estimation of subject-specific HRFs as a means of enhancing the accuracy and power of fMRI studies. 1 Characteristics of the shape of the evoked hemodynamic response such as height, delay, and duration may also be used to infer information about intensity, onset latency, and duration of the underlying neuronal activity. 4 Accurate estimation of the HRF is essential if fMRI is to be used to detect fine differences in the timing of neuronal activation as a means of understanding the temporal sequences of brain processes. Hemodynamic response function modeling also enables noninvasive investigation of neurovascular coupling, which changes with brain aging and may have a pathophysiological role in dementia and cerebrovascular disease.6, 7
The design of fMRI studies affects both estimation efficiency, a measure of the ability to estimate the shape of the HRF, and detection power, a measure of the ability to detect activation. Previous studies have demonstrated that there is a fundamental tradeoff between these characteristics such that designs maximizing detection power necessarily have minimum estimation efficiency and designs that achieve maximum estimation efficiency cannot attain maximum detection power.8, 9 For example, event-related designs offer high estimation efficiency but poor detection power, whereas block designs offer good detection power at the cost of low estimation efficiency. 9
A considerable body of fMRI data has already been acquired using block designs to maximize signal-to-noise ratio (SNR) and to increase the likelihood of detecting a response. In some of these studies, it would be of interest to examine the HRF explicitly even though study designs were not optimized with this in mind. In particular, studies involving large number of subjects,10, 11 multiple sites,10, 12, 13 special subject groups,11, 14 and longitudinal designs
14
would be difficult and expensive to replicate. Our specific interest in estimating HRF parameters from a block design study was stimulated by a large twin study. Here we were interested in how robustly HRF parameters could be estimated as a prelude to examining the heritability of the HRF. We approached this question first by comparing the performance of different HRF models. We then examined the test–retest reliability of HRF parameter estimation. This allowed us to identify which HRF traits remain stable over time, providing some insight into the extent to which individual parameters serve as variables reflecting a biologic trait or an experimental state. Test–retest reliability was examined in 82 healthy young adult twins tested with the same fMRI paradigm on two occasions. We examined the following parameters of the HRF: height (
MATERIALS AND METHODS
Hemodynamic Response Function Model and Features
The blood oxygenation level dependent (BOLD) fMRI signal was modeled as the convolution of the stimulus function and the HRF. Before assessing test–retest reliability in human data, we assessed the identifiability of five HRF models, summarized in Table 1, in simulated fMRI time series. Model I was the canonical HRF used in SPM8 (The Wellcome Trust Centre for Neuroimaging, London, UK), in which only the height parameter
Summary of HRFs a
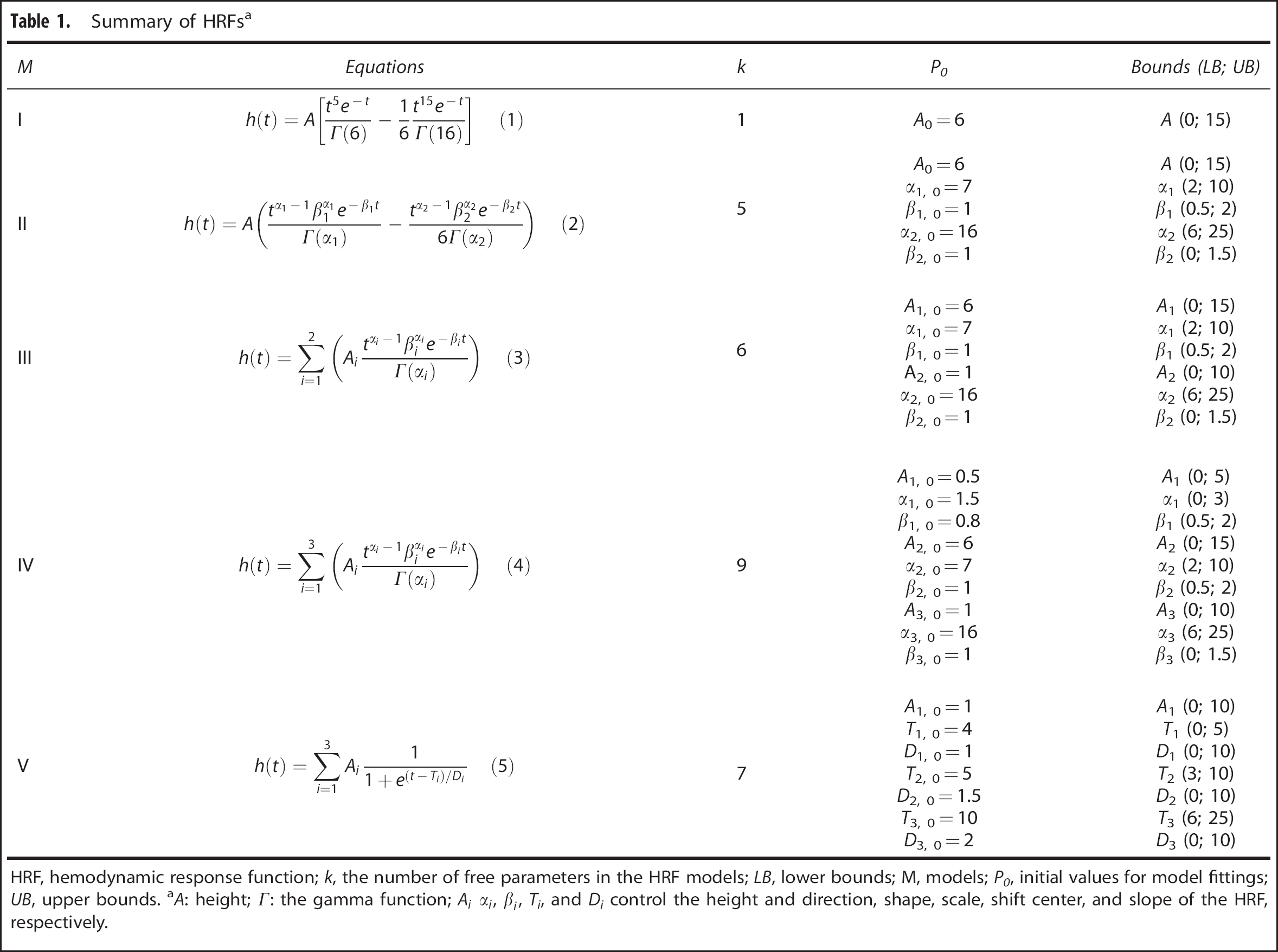

HRF, hemodynamic response function;
We used the following summary HRF features: height (H), defined as the maximum signal change during the peristimulus time window (30 seconds was used for this study); time to peak (
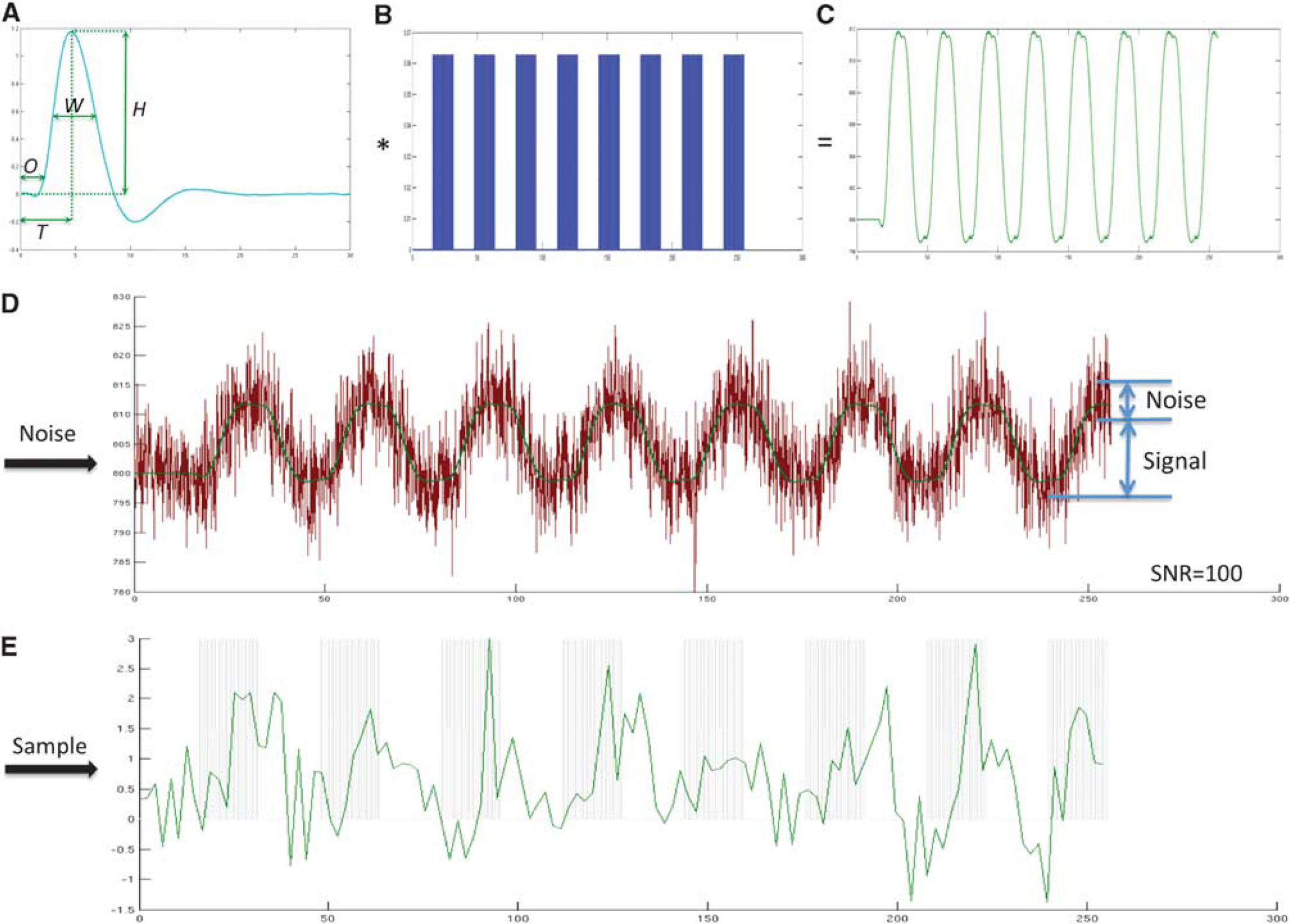
The procedure used to simulate the functional magnetic resonance imaging (fMRI) signal time course. (
Simulation of Functional Magnetic Resonance Imaging Signal Time Course
Functional magnetic resonance imaging signal time courses were simulated using known HRFs to test the performance of different HRF models. Three ground truth HRFs were created (Supplementary Figure S1). In Simulation 1, the HRF was generated using the balloon model.17, 18 The HRF was generated using a single input at 1.5 seconds with 0.1-second duration and using typical parameters obtained empirically in a previous study: neuronal efficacy 0.54, signal decay 1.54, autoregulation 2.46, transit time 0.98, stiffness parameter 0.33, resting oxygen extraction 0.34, and resting blood volume fraction 0.06. 18 The HRF in Simulation 2 was generated using the sum of four inverse logit functions to create four segments of HRF time course:
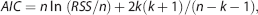
in which
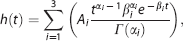
in which
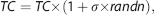
where
Hemodynamic Response Function Model Selection
Hemodynamic response function modeling was performed using an in-house MATLAB (MATLAB R2010a, The MathWorks, Natick, MA, USA) toolkit, sHRF, which is publicly available at our website (http://www.cai.uq.edu.au/shrf-toolkit). The fMRI time courses were high-pass filtered with a 128 seconds discrete cosine basis set and then modeled as the convolution of the HRF (Table 1) and the stimulus function: 22
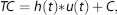
where
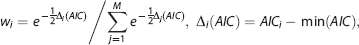
in which
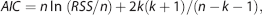
where
Parameter Estimation from Block Design Functional Magnetic Resonance Imaging Data
The noise in measured fMRI time series and sparse sampling in acquired data may lead to HRF parameter mis-specification. To examine this, we used the balloon model simulation17, 18 as the ground truth. The HRF parameters were varied in three ways: (1)
Variations in the Block Design Paradigm
To investigate the influence of variations in the block design paradigm on the trueness and precision of HRF modeling, the experimental conditions of the block design were systematically varied: (1) the number of blocks was varied from 1 to 8 with one block increments; (2) block length was varied from 4 to 16 seconds with 2-second increments; (3) stimulus duration was varied from 0.2 to 6.2 seconds with 1-second increments and a stimulus duration of 16 seconds in individual blocks (long block design) was also evaluated; (4) the gap between individual stimuli in each block was varied from 0.8 to 6.8 seconds with 1-seconds increments. Simulation of the fMRI time course was carried out as described above using the balloon model to generate ground truth, a SNR of 100, and Model III to estimate the HRF. Each simulation was executed 100 times. Trueness and precision for each paradigm were calculated as described above.
Experimental Functional Magnetic Resonance Imaging Time Course Data
The data we analyzed were acquired as part of a prior fMRI study, 11 the Queensland Twin Imaging Study. 27 Retrospective utilization of the data was approved by the Research Ethics Committee of the Queensland Institute of Medical Research and The University of Queensland in compliance with the Australian National Statement on Ethical Conduct in Human Research. Functional MRI data from 30 male and 52 female healthy young adult twins of mean age 22.5±2.5 (s.d.) years were included. For all participants, the fMRI scans were repeated 2 to 6 months later (mean 117±56 days).
Participants performed the 0- and 2-back versions of the
The three-dimensional T1-weighted MR image and echo planar imaging data were acquired on a 4T Bruker Medspec whole body scanner (Bruker, Hanau, Germany). Three-dimensionalT1-weighted images were acquired using an MP-RAGE pulse sequence (TR=2500 milliseconds, TE=3.83 milliseconds, T1=1500 milliseconds, flip angle=15 degrees, 0.89 × 0.89 × 0.89 mm). For each participant, 127 sets of echo planar imaging data (TR=2.1 seconds, TE=30 milliseconds, flip angle=90, 3.6 × 3.6 × 3.0 mm) were acquired continuously during the tasks.
The fMRI data were analyzed using Statistical Parametric Mapping (SPM8, the Wellcome Trust Centre for Neuroimaging, London, UK). The first five echo planar imaging volumes were discarded to ensure that tissue magnetization had reached steady state. The spatial preprocessing included 2-pass motion correction
29
and spatial normalization to the average brain T1 template
30
implemented in SPM8. Normalized volumes were smoothed with an 8 × 8 × 8 mm full width half maximum Gaussian kernel. In a first-level analysis of individual subject data, we determined locations of activation using the general linear model with a finite impulse response basis function. The 2-back minus 0-back contrast images were then entered into a group level, random-effects one-sample
Four regions identified at the group level were selected for HRF modeling: left and right middle frontal gyrus and left and right angular gyrus (Supplementary Figure S3). Volumes of interest were defined as the overlap of an existing probabilistic atlas of each structure in stereotaxic coordinate space
31
with group activation regions. For each participant, the fMRI time course was extracted by averaging the signal intensity at each time point in the voxels with the top 12.5% of SPM
Analysis of Test–Retest Reliability
For each brain region and each HRF parameter, the intraclass correlations of the two experimental sessions were calculated using SPSS20 (IBM) using a two-factor mixed effects model and tests of significant difference in ICC from zero performed. 32 Because the participants were biologically related monozygotic and dizygotic twins, the intraclass correlations of the HRF heights from the first-born participant in each twin pair were also evaluated to assess whether relatedness between participants affected the reliability results. The reliability of the performance measure used in the experiment has been reported previously. 11
RESULTS
Simulation Study
Figure 2 summarizes the HRF parameter estimates from the simulations. Parameter estimation was considered accurate if estimated values did not differ significantly (i.e.
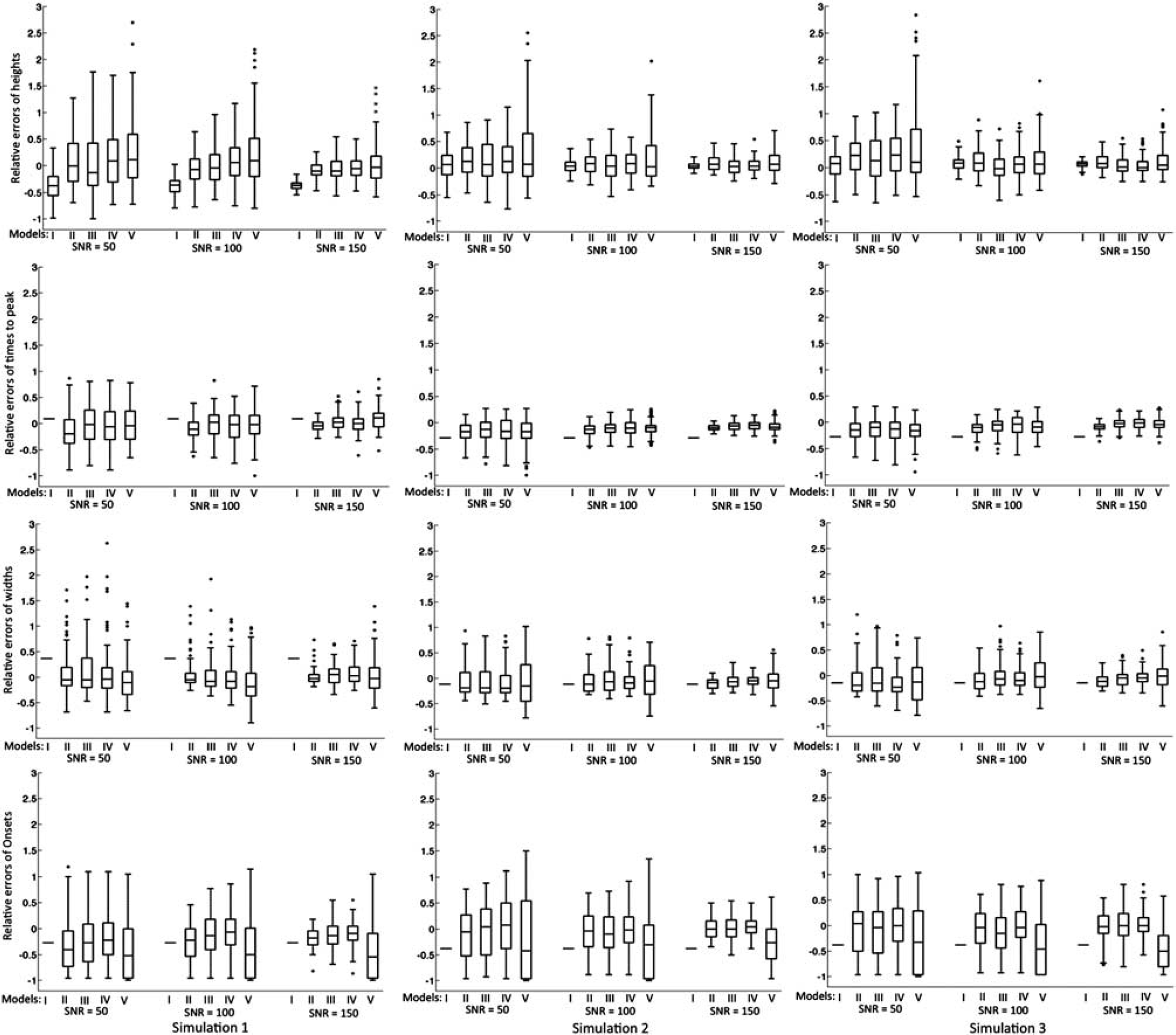
Box-and-whisker plot of the relative differences of estimated hemodynamic response function (HRF) parameters from the true values in the simulation study. Outliers were defined as exceeding three times the s.d.; all data including outliers were included in the analysis. Rows from top to bottom are the results for height (
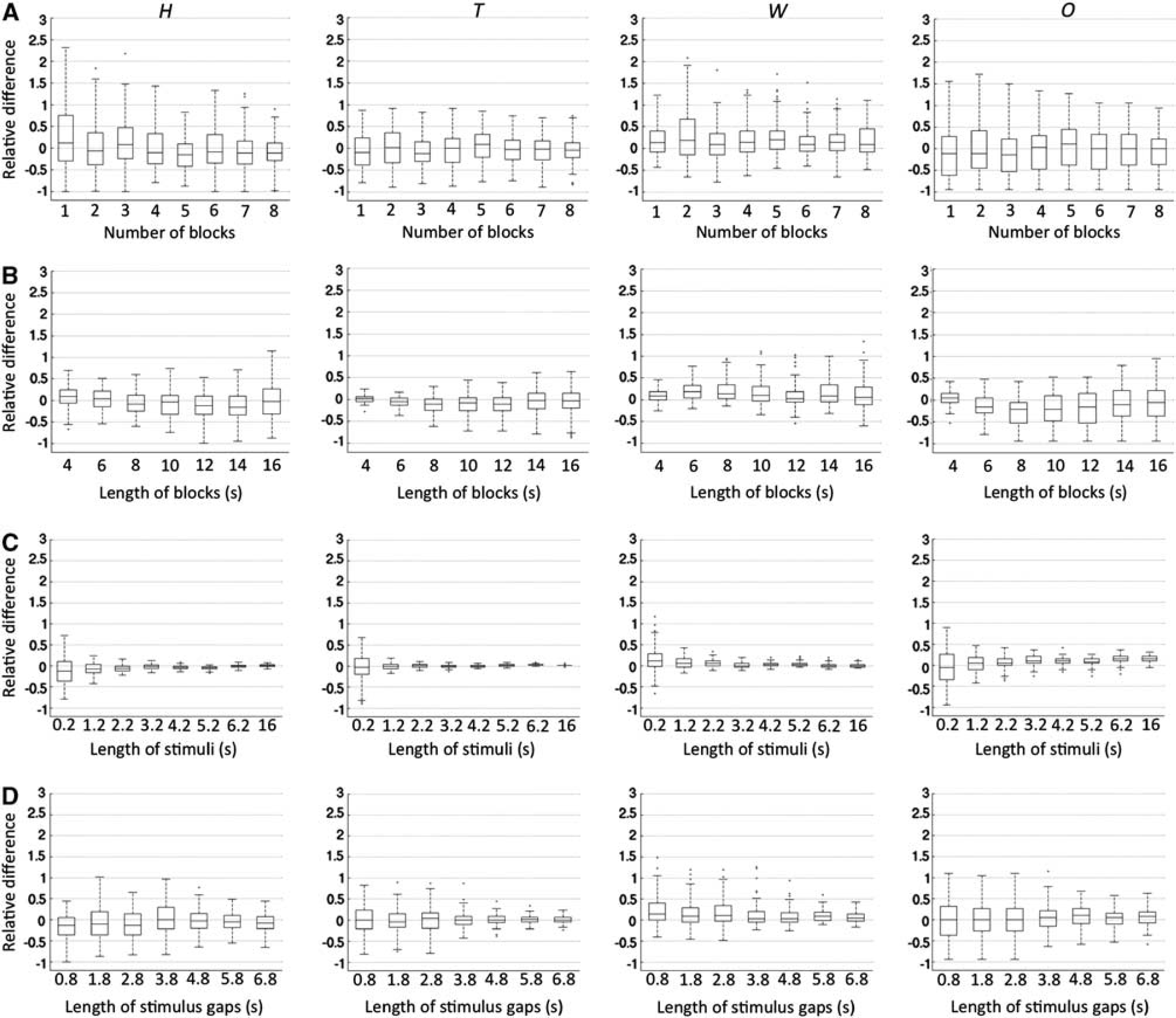
Boxplots of estimated hemodynamic response function (HRF) features with systematic variation in attributes of the block design paradigm. The columns from left to right relate to estimated height, time to peak, width, and onset, respectively. The rows from top to bottom relate to variation in the number of blocks (
Table 2 summarizes the AIC of different models for data with different noise levels. Model III had the highest AIC weights and Model I the lowest across different simulations and noise levels.
Akaike weights (
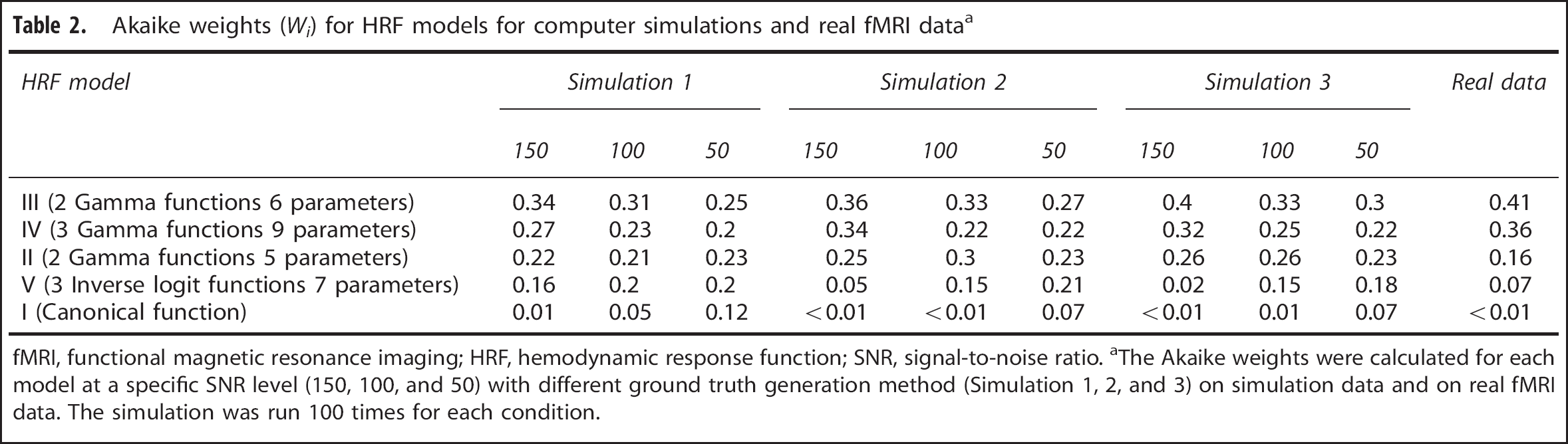
fMRI, functional magnetic resonance imaging; HRF, hemodynamic response function; SNR, signal-to-noise ratio.
The Akaike weights were calculated for each model at a specific SNR level (150, 100, and 50) with different ground truth generation method (Simulation 1, 2, and 3) on simulation data and on real fMRI data. The simulation was run 100 times for each condition.
Results for HRF parameter estimation are summarized in Table 3. The selected HRF model (Model III) was able to identify parameter differences exceeding 10% with SNR of 100. There were no instances in which significant changes in one parameter resulted from variation in another parameter.
Estimation of HRF parameter variation a
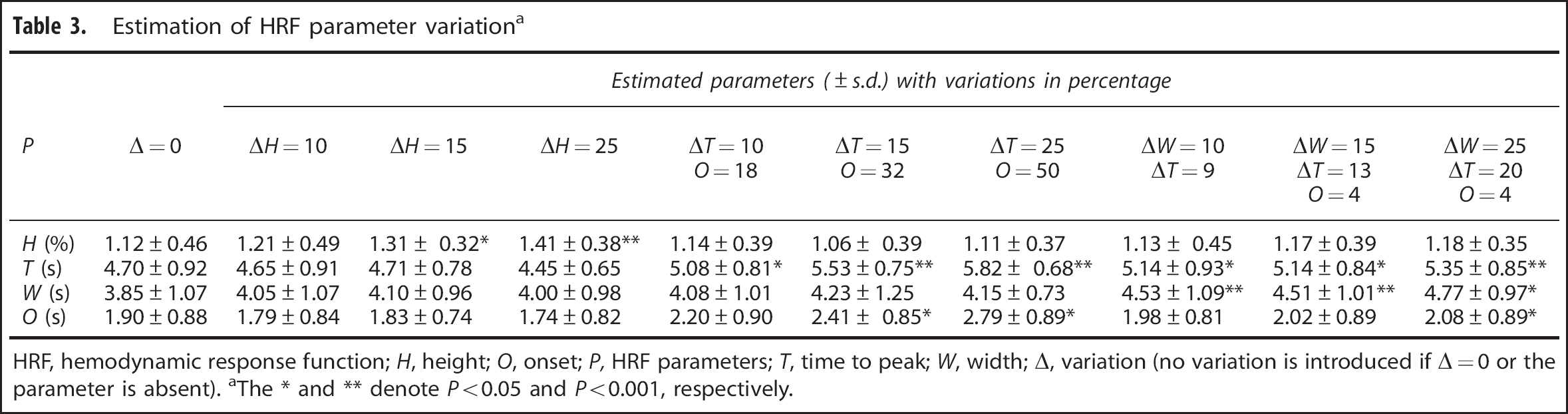
HRF, hemodynamic response function;
The ∗ and ∗∗ denote
Results of parameter estimation with different block design paradigms are summarized in Figure 3 using boxplots constructed in the same manner as Figure 2. For less than six blocks, mean estimated
Real Functional Magnetic Resonance Imaging Data
Akaike Information Criterion weights for the models using real fMRI data were consistent with results from simulations (Table 2). Model III had the highest AIC weights. Activation maps between the two experimental sessions were highly reproducible (Supplementary Figure S3). Quantitative assessment of the repeatability of the activation map between two experimental sessions has been reported previously.
11
The ICC(3,1) of HRF parameters estimated in the two experimental sessions using Model III are summarized in Table 4. Height and
The ICC(3,1) of HRF parameters between the two experimental sessions a
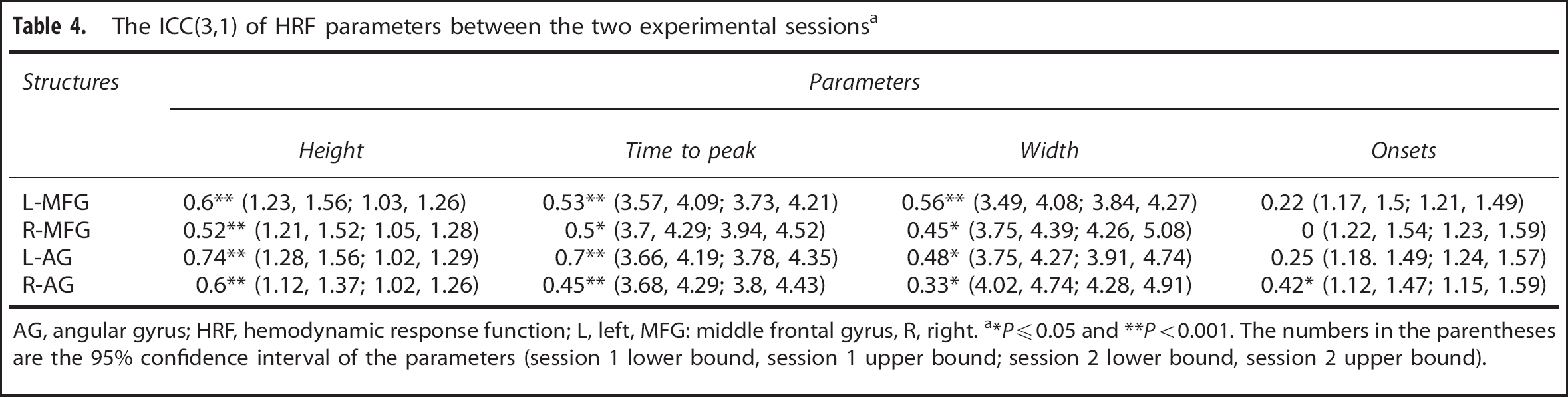
AG, angular gyrus; HRF, hemodynamic response function; L, left, MFG: middle frontal gyrus, R, right.
∗
DISCUSSION
Modeling of the HRF using fMRI is a noninvasive method to investigate the brain's hemodynamic response to neuronal activation. In this study, we found that with block design fMRI paradigms, most HRF models were able to estimate HRF parameters except for Model V that had difficulty estimating onsets. The sum of two gamma functions with six free parameters had the greatest identifiability of the five models tested but the accuracy of parameter estimation varied with block design attributes.
We used computer simulations with a known ground truth to assess the performance of the HRF models. In the simulations, Gaussian noise was added at each SNR level without consideration of autocorrelation. However, fMRI noise typically exhibits temporal dependence (autocorrelation); therefore, the SNR level is slightly lower than in practice. Parrish
A key consideration in HRF model selection is the number and nature of
In this study, we represented neural activity by the stimulus time course, an assumption underlying commonly used methods for statistical parametric mapping. To the extent that this assumption is violated, it is likely that the HRF model parameters estimated here contain a component of neural activity in addition to vascular reactivity. Several techniques have been recently proposed to dynamically filter fMRI time courses to estimate neural activity as well as the vascular response explicitly.39, 40 Application of these techniques in future studies may allow the reproducibility of the HRF to be estimated more specifically.
Many HRF models have been proposed in the literature. Our study was restricted to five models that are commonly used or have been shown in prior studies to be less susceptible to mis-specification errors (sum of inverse logit functions) or parameter bias (sum of two gamma functions). In this study, we examined vascular responses to neural activity evoked by a single cognitive function in four brain structures that demonstrated robust activation. Further studies using other activation tasks are needed to assess generalizability of our findings to other brain regions and brain functions.
Block designs are not optimal for HRF estimation and random event-related designs and
In conclusion, we investigated trueness, precision, model identifiability, and test–retest reproducibility of parameter estimates for HRF models with a block design paradigm using simulated and real fMRI data. The HRF features of height, time to peak, and width were reproducible between test sessions and may be useful as measures to characterize the coupled vascular response to neural activity in individual subjects.
Footnotes
The authors declare no conflict of interest.
ACKNOWLEDGMENTS
The authors thank the twins for participating in this study, research nurses Marlene Grace and Ann Eldridge at QIMR for twin recruitment, and radiographers Matthew Meredith, Peter Hobden, and Aiman Al Najjar from the Centre for Advanced Imaging, The University of Queensland, for data acquisition, and research assistants Kori Johnson and Lachlan Strike, Queensland Institute of Medical Research, for preparation and management of the imaging files.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
