Abstract
The events that occurred following the mailing of Bacillus anthracis-laced envelopes through the postal system has highlighted the need to perform biological screening on large numbers of environmental samples. High-throughput screening that relies on integrated robotic systems to speed analysis has been undertaken to handle the surge in samples requiring testing in events involving weapons of mass destruction. These automated screening systems require DNA extraction methods capable of handling environmental samples that contain inhibitors and have target organisms at low concentrations. This study describes the development of a method for the detection of the biological warfare agent simulants Erwinia herbicola and Bacillus subtilis var. niger spores using paramagnetic bead-based resin with an automated liquid handler and environmental samples.
Keywords
Introduction
Events in the United States during 2001 have renewed interest in routine biological monitoring of the environment for biological warfare agents. As previously predicted, public health laboratories were ill-prepared to deal with a large influx of samples during a national incident, and a high-throughput sample screening solution was needed. 1 As a model, diagnostic laboratories are faced with high numbers of urgent samples each day in which the results must carry a high level of confidence. Diagnostic samples are presented, however, in preordained sample matrices such as blood, urine, nasal swabs, or feces. Although these matrices can often prove a challenge for sample cleanup and nucleic acid isolation, the limited number of matrices allows for standardized approaches in methodology to prepare nucleic acids and remove inhibitors. When samples are taken from the environment that come from aerosol samplers or surface wipes of unknown samples, there is a wide spectrum of potential inhibitors that must be considered. 2
Based on our previous work of extracting DNA from environmental samples, bead-based methods were preferable to avoid organic extractions, centrifugation, or filtration steps that can result in clogging. Magnetic bead resins were advantageous because they allow the option of aggressive washes and fluid-like dynamics that can aid in the removal of particulate matter and potential PCR inhibitors as part of a fully automated system. Magnetic bead-based DNA extraction has been used to successfully isolate nucleic acids from a variety of sample types that contain PCR inhibitors.3–6 MagneSilTM (Promega, Madison, WI) was selected for study as a paramagnetic bead-based nucleic acid extraction system that offers a variety of porosities and size options for use with food, blood, and forensic samples. 7
The automated liquid handler system selected was required to be versatile and programmable for DNA extraction as well as PCR assay setup. The full integration of the system was required since the final marriage of magnetic bead and automated liquid handler was only one piece of a larger robotic system for screening large numbers of samples for biological weapons. The Automated Biological Agent Testing System (ABATS) (patent pending), initiated by the Department of Defense in response to homeland defense initiatives, is designed to combine PCR and electrochemo-luminescent (ECL) immunoassay to analyze each environmental sample, rather than relying on just one technology or the other (Fig. 1).
The Automated Biological Agent Testing System (ABATS). The ABATS utilizes the Biomek® FX liquid handling robotic system to set up immunoassay and PCR reactions and also performs DNA extraction.
The Biomek® FX liquid-handling robotic system (Beck-man-Coulter, Fullerton, CA) allows the ABATS to use a CORE system to link the Applied Biosystems ABI Prism® 7900HT thermocycler and the ORIGE N® M8 Analyzer (IGEN, Gaithersburg, MD) to form an integrated screening laboratory. The overall goal of the ABATS program is a three-fold increase in sample throughput with a two-fold reduction in cost per sample when compared with current manual screening operations.
To realize this goal, manual procedures requiring time savings were sample chain of custody paperwork, nucleic acid extraction, reaction setup, and result interpretation. In this study, we investigate time-saving measures pertaining to nucleic acid extraction of two accepted biological agent simulants in an effort to reduce labor and speed analysis of unknown environmental samples by addressing bottlenecks in the analysis process.
Materials and Methods
Biological Agent Simulants
Cell suspensions of Bacillus subtilis var. niger spores (BG spores) and Erwinia herbicola vegetative cells were prepared for use as biological simulants. BG spores were prepared by inoculating 1-g spores of B. subtilis var. niger (Merck & Co., Whitehouse Station, NJ) in 100-mL irrigation water (McGAW, Irvine, CA). The suspension was mixed with a vortex mixer (Barnstead Thermolyne Type 37600; Bebu-que, IA) for 10 min. The mixture was placed in an orbital shaker water bath at 65 °C and 50 rpm for 10 min. The suspension was plated on 15 plates of AK Agar #2 Sporulating Agar (Cat. No. 210912; Becton Dickinson, Sparks, MD) using a swab to cover the whole plate. The plates were incubated at 30 °C for 7 days. The cells were examined periodically via phase contrast microscopy to examine spore development and formation. The spores were scraped from the plates using sterile swabs and suspended in 400-mL sterile phosphate buffered saline (PBS) (Catalog No. 1000-3; Sigma, St. Louis, MO) and were washed three times with PBS. A titer was established by plating serial dilutions of the washed spore suspension on Tryptose Agar media (Cat. No. 264100; BD, Franklin Lakes, NJ). The final solution was adjusted to contain 1 × 106 CFU/mL in PBS. Aliquots of 1 mL were frozen at —20 °C until needed. E. herbicola suspensions were prepared by inoculating 1 L of sterile Trypticase Soy Broth (Cat. No. 211771; Becton Dickinson, Sparks, MD) with E. herbicola (ATCC No. 33243). The culture was incubated at 25 °C in a Model G25 orbital shaker (New Brunswick Scientific, Edison, NJ) at 200 rpm for 48 h. The cells were harvested by low-speed centrifugation and washed three times in sterile PBS at 4 °C. Serial dilutions of the cells on Trypticase Soy Agar media were plated to determine the titer of the cells. The harvested cells were adjusted to 1 × 106 CFU/mL with PBS, and 1-mL aliquots were flash-frozen at —20 °C until needed.
DNA Isolation
All DNA extraction procedures were carried out on a Biomek® FX (Beckman Coulter, Fullerton, CA). DNA purification kits for food (Cat. No. FF3750) and blood (Cat. No. MD1360) were kindly provided by Promega (Madison, WI). For each extraction sample, either the BG spore suspension or E. herbicola cell suspension was aliquoted into one well of a 96-well, 2-mL deep-well plate (Costar 3960). After filling the wells, the deep-well plate was placed onto the deck of the Biomek® FX, where all subsequent pipetting was performed robotically. Also placed onto the deck of the Biomek® FX in deep-well plates were all reagents, including isopropanol, 70% ethanol, water, and MagneSilTM Yellow, and either the blood kit or food kit lysis buffers. Following DNA extraction using the Food Kit Protocol, Blood Kit Protocol, or Hybrid Protocol, a shallow 96-well plate containing eluted DNA was frozen at —20 °C or used immediately for PCR analysis.
Food Kit Protocol for Biomek® FX
Extraction using the food kit was performed as follows: 250 μL of a sample was added to 500-μL Lysis Buffer A and shaken at 1400 rpm for 1 min; 250 mL of Lysis Buffer B was added and shaken at 1300 rpm for 1 min, followed by incubation at room temperature for 10 min. MagneSil™ Yellow (60 μL) was added and shaken at 1200 rpm for 1 min, followed by the addition of 248-μL isopropanol and shaken at 1100 rpm for 5 min. The MagneSilTM particles were removed from the solution in two half-volume steps by 1 min incubations on a deep-well MagnaBot® (Promega V3031, V8581). The MagneSil™ particles were washed with an additional 250 μL of Lysis Buffer B, shaken at 1500 rpm for 1 min, and recollected with the MagnaBot® (1 min). The MagneSilTM particles were washed with 70% ethanol (200 μL 70% ethanol, shaken at 1500 rpm for 1 min, recollect with MagnaBot® for 1 min) three times. Particles were dried for 13 min at room temperature. The MagneSil™ particles were combined with 100-μL water, incubated at room temperature for 1 min, shaken at 1500 rpm for 1 min, and the MagneSilTM was removed using the MagnaBot® (1 min). The eluted DNA in water was transferred into a shallow 96-well plate.
Blood Kit Protocol for Biomek® FX
Extraction using the blood kit was performed as follows: 200 μL of sample was combined with 50 μL of MagneSil™ and 310 μL of Blood Kit Lysis Buffer and shaken at 900 rpm for 1 min, followed by a room temperature incubation for 6 min. MagneSilTM particles were collected using two successive MagnaBot incubations (1 min and 30 s) with effluent discarded between each incubation. Additional Blood Kit Lysis Buffer (360 μL) was added to the MagneSilTM particles, shaken at 1250 rpm for 1 min, and the particles were again collected in two successive MagnaBot® incubations (1 min and 30 s). The MagneSil™ particles were washed using a salt wash solution (add 180-μL salt wash, shake at 1250 rpm for 1 min, incubate at room temperature for 1 min, add 180-μL salt wash, shake at 1250 rpm for 1 min, incubate at room temperature for 1 min, incubate on MagnaBot® (1 min), remove effluent, incubate on MagnaBot® (30 s), remove all remaining effluent, and repeat twice). MagneSil× particles were then washed using an alcohol wash solution (add 180-μL 2X alcohol wash, shake at 1250 rpm for 1 min, incubate at room temperature for 1 min, add 180-μL 2X alcohol wash, shake at 1250 rpm for 1 min, incubate room temperature for 1 min, incubate on MagnaBot® (1 min), remove effluent, incubate on MagnaBot® (30 s), remove all remaining effluent) and repeat three times. MagneSilTM was then dried for 15 min at room temperature, and purified DNA was eluted by the addition of 210-μL elution buffer, shaken at 1300 rpm for 1 min, incubating at room temperature for 5 min, and removing MagneSil× particles with a 1-min incubation on the MagnaBot®. Two hundred milliliters of effluent was drawn off and transferred to a shallow 96-well plate.
Hybrid Protocol for Biomek® FX
The Hybrid Protocol was performed as follows: 625 μL of a sample was added to 625-μL Blood Kit Lysis Buffer and shaken at 900 rpm for 1 min, followed by an incubation at room temperature for 10 min. MagneSil™ Yellow (60 μL) was added and shaken at 900 rpm for 1 min, followed by the addition of 250-μL isopropanol and mixing by pipetting, on the Biomek, for 5 min. The MagneSil™ particles were removed from the solution in two half-volume steps by 1 min incubations using the MagnaBot® . The MagneSil™ particles were washed with an additional 250 μL of Blood Kit Lysis Buffer, shaken at 1500 rpm for 1 min, and recovered with the MagnaBot® for 1 min. The MagneSilTM particles were washed with 70% ethanol (200-μL 70% ethanol, shaken at 1500 rpm for 1 min, recovered with MagnaBot® for 1 min) three times. Particles were dried for 13 min at room temperature. Purified DNA was eluted into water (100-μL water, incubated at room temperature for 1 min, shaken at 1500 rpm for 1 min, heated at 65 °C for 3 min, shaken at 1500 rpm for 1 min, and MagnaBot® for 1 min) and transferred into a shallow 96-well plate. A diagram of the Biomek® FX deck layout is pictured in Figure 2.
A diagram of the Biomek® FX deck layout for the hybrid method.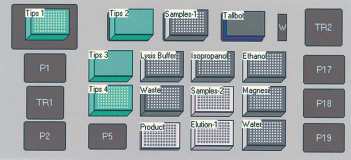
Physical Disruption of BG Spores
A dilution series of BG spores and E. herbicola were combined in a 2-mL screw cap microfuge tube with 1-mL cell solution, 0.16 g of 0.1-mm glass beads, and 0.03 g of 2-mm zirconium beads (Cat. No. 11079110, 11079124zx; BioSpec Products, Bartlesville, OK). Samples were processed in a Mini-BeadBeater-8 (BioSpec Products) for 30 min in PBS + T (0.3% Tween20, Cat. No. P-9416; Sigma, St. Louis, MO) at 3200 rpm. Processed samples were removed and pelleted in a microcentrifuge at 12,000g for 2 min. The supernatant was removed, and 625 μL was processed in the hybrid method as described.
Manual DNA Extraction
Manual DNA extraction followed the same protocol as the optimized automated extraction, with the exception that all pipetting steps were performed manually with a single-channel and an eight-channel pipette.
Polymerase Chain Reaction
Amplification, data acquisition, and data analysis were carried out on an Applied Biosystems model 7900HT sequence detection system (Applied Biosystems, Foster City, CA). Primer and probe sequences for BG detection were developed at the Edgewood Chemical and Biological Center (Edgewood, MD). Primer and probe sequences for E. herbicola detection were developed by the Naval Medical Research Center (Silver Spring, MD). PCR reactions were performed in 20-μL volumes in 384-well microtiter plates. Each reaction was set up using Taqman® Universal Master Mix (4304437; Applied Biosystems, Foster City, CA), 500 -nM forward and reverse primer, 200-nM FAM/TAMRA-labeled fluorogenic probe, and 8.5 μL of extracted DNA product per reaction. The thermocycler conditions were as follows: 50 °C for 2 min, 95 °C for 10 min, and 50 cycles of 95 °C for 15 sec, and 60 °C for 1 min. Analysis was performed using Sequence Detection Software v.2.0.
Environmental Sample Analysis
Surface swipes were collected using a Biological Sampling Kit (BiSkit) (Edgewood Chemical Biological Center, Edge-wood, MD), which dispensed the sample in approximately 4-mL phosphate-buffered saline, pH 7.4, plus 0.3% Tween-20 (PBS + T). Air samples were collected in metropolitan areas using a Dry Filter Unit (DFU) 1000 (Joint Program Executive Office for Chemical and Biological Defense, Department of Defense) for 12 h at a flow rate of 300-600 L/min. Filters were removed and placed into 10 mL of sterile PBST and vortexed for 1 min to generate filter washes. Six soil samples provided by the Soil Society of America's North American Proficiency Testing Program (Alabama, Big Horn, Brookings, Keenesburg, Teller, and Whitewater) and two uncharacterized soil samples (Baghdad and Kahmasia) were mixed with PBST at the ratio of 1 mL of PBST per 0.5-g soil. Samples were mixed by inversion for 10 min at room temperature. Soil particles were removed by centrifugation, and the supernatant was designated as soil washes. The inherent presence of PCR inhibitors was determined by using soil washes, air filter washes, and surface swipe samples as described above and adding them directly into a PCR reaction for E. herbicola at 1/10 of the final volume. PCR was run as described, and the change in CT value was compared with the CT value of the same reaction without environmental matrix added. Serial 10-fold dilutions were created and tested likewise until a non-inhibitory dilution (ΔCT = 3 × StDev) was found for each sample matrix. From the filter washes, surface samples, and soil washes, 620 μL were removed and added to 5 μL of 1.25 × 108 CFU/mL of E. herbicola suspensions to generate a final concentration of 1 × 106 CFU/mL. The 625 μL of spiked washes were used for DNA extraction using the hybrid protocol. For BiSKit samples and soil washes that showed inhibition following DNA extraction, dilutions were generated at the ratios 1:5 and 1:25 for BiSKit samples, and 0.5 g/5 mL and 0.5 g/25 mL for soil washes, and retested.
Results
Biomek® FX DNA Extraction
Promega MagneSil™ DNA purification resin with a mac-roporous composition is commercially available for use with food and blood, and these kits were used as a starting point to develop a purification method for environmental samples to extract DNA from E. herbicola vegetative cells and from B. subtilis var. niger (BG) spores. Real-time PCR was used to evaluate the effectiveness of the nucleic acid extraction for the two organisms, as the total quantities of DNA isolated were low, and the overall objective was to more efficiently integrate the extraction process with PCR analysis and increase the level of detection using PCR. 8 TaqMan® polymerase chain reaction is a probe hydrolysis method that measures the progress of PCR amplification by measuring light in a technique called fluorescence resonance energy transfer (FRET). 9 10 Researchers from the U.S. Army and U.S. Navy provided the TaqMan® PCR primer and probe sequences that specifically amplify genes within E. herbicola and BG. The Applied Biosystems model 7900HT thermocycler is a TaqMan® -based instrument in which the interpretation of a positive response is measured when a reaction breaches a threshold (CT) when referenced against a non-responsive baseline. Thresholds that occur at earlier cycles for the same PCR assay can be correlated with a higher starting concentration of template DNA in the sample. Thus, a sample that breaches the threshold at cycle 30 can be inferred to have contained a higher amount of starting DNA than a sample that crosses the threshold at cycle 42. Using the unmodified food kit, average CT scores of 29 and 40 were obtained for E. herbicola (1 × 105 CFU/mL) and BG (1 × 106 CFU/mL), respectively (Table 1). Using the unmodified blood kit, CT scores of 27 and 44 were observed. An optimized hybrid protocol of these two kits that used the method from the food kit and the chaotropic lysis buffer from the blood kit proved to be the better combination, resulting in improved CT values of 26 and 36, respectively. The blood lysis buffer has a more concentrated formulation of guanidinium salts than is found in the food kit (personal communication, Promega Corp.), and that component may facilitate more efficient lysis or recovery of DNA from environmental samples.
Threshold of detection (CT) for the purification of Bacillus subtilis var. niger and Erwinia herbicola DNA using Promega Blood Method, Promega Food Method, and Hybrid Method
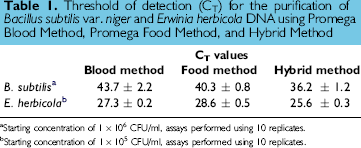

Starting concentration of 1 × 106 CFU/ml, assays performed using 10 replicates.
Starting concentration of 1 × 105 CFU/ml, assays performed using 10 replicates.
Detection Limits
Using the optimized hybrid protocol, it was determined that the detection limit for E. herbicola vegetative cells was between 1 and 10 CFU/mL (Table 2). Detection was not observed with a sample of 0.1 CFU/mL (data not shown). BG spores were detected at 100,000 CFU/mL in 91.7% of samples (11/12), and at 10,000 CFU/mL in 8.3% of samples (1/12) using the hybrid protocol. For both BG and E. herbicola, the CT values increased by approximately 3.3 for each 10-fold dilution, which is indicative of optimal thermocycling efficiency under the conditions tested (Applied Biosystems, User Bulletin #2: ABI Prism 7700 Sequence Detection System, December 11, 1997).
Comparison of Bacillus subtilis var. niger and Erwinia herbicola extraction using a Biomek® FX, manual pipetting steps, and physical disruption prior to Biomek® FX extraction
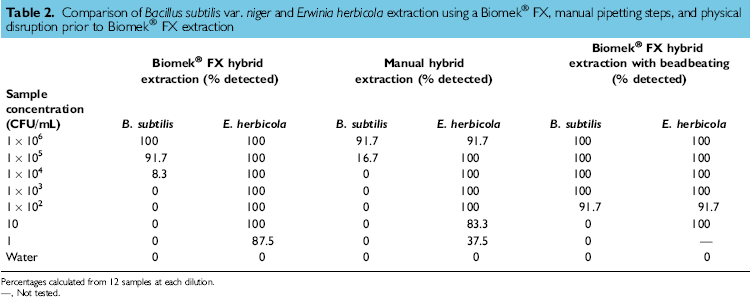

Percentages calculated from 12 samples at each dilution.
—, Not tested.
In an effort to drive detection thresholds even lower for spore-containing samples, additional pretreatment lysis methods were tested to augment the hybrid extraction protocol. Boiling the cell suspensions for 30 min prior to extraction was tested in an effort to disrupt the tough exosporium that surrounds Bacillus species of bacteria. This process did not significantly contribute to enhanced PCR detection (data not shown). Bead-beating BG spore suspensions prior to performing the hybrid method was successful in increasing the detection limit of BG down to 1 × 102 CFU/ mL 91% of the time and had no deleterious affect on the purification of E. herbicola DNA. These results suggest that the hybrid protocol with no bead-beating pretreatment is likely purifying residual DNA that remains trapped on the exosporium of the BG spore, whereas bead-beating probably disrupts the BG spores.
Manual Extraction
In order to compare the sensitivity and reliability of the automated procedure, the hybrid process was performed manually on the same samples using an eight-channel multipipettor and testing by PCR. The manual extraction results were very similar to the results obtained with the Biomek® FX with respect to CT scores, although the manual extraction was less consistent in detecting both bacteria (Table 2). Manual extraction was less effective than automated extraction with BG spores, with a reduced detection limit of 1 × 105 CFU/mL for 17% of the samples. Furthermore, performing the hybrid extraction protocol manually took approximately 150 min, whereas the automated Biomek® FX protocol took only 70 min.
Environmental Sampling
Washes from air filters, soil, and surface wipes were generated to test the effectiveness of the MagneSil® Hybrid protocol and its ability to remove potential PCR inhibitors commonly associated with environmental samples. The initial inhibition present within each sample was gauged by changes in the threshold (CT) when the samples were tested by E. herbicola PCR and compared to the threshold observed with buffer alone. Each sample was diluted systematically until it no longer inhibited PCR. Dilutions ranged from 1:10,000 to 1:10,000,000, with a higher dilution factor indicating a higher concentration of inhibitors within the starting sample (Table 3). To examine the hybrid protocol's ability to clean the environmental samples of these inhibitors, the environmental washes were spiked with 1 × 106 CFU/mL of E. herbicola cell suspension, and the DNA extraction protocol was performed in the same manner as the previous experiment. No significant PCR inhibition was observed in undiluted air filter washes, three surface swipe samples, and one soil type (Table 3). Most soil washes and two swipe samples had to be diluted (resuspended in a larger volume of buffer) to show minimal inhibition, although for this study, the samples containing the most inhibitors (Bighorn soil and mailbox surface swipe) showed at least a 2000-fold removal of PCR inhibitors using the hybrid method. It is apparent that the hybrid protocol was effective in removing the vast majority of PCR inhibitors from all samples. We plan to test a dilution of each sample or use an internal positive control to test for inhibition when evaluating environmental samples.
Detection of Erwinia herbicola in environment samples using the hybrid purification method and real-time PCR analysis
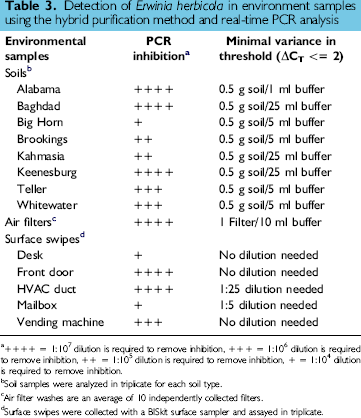
++++ = 1:107 dilution is required to remove inhibition, +++ = 1:106 dilution is required to remove inhibition, ++ = 1:105 dilution is required to remove inhibition, + = 1:104 dilution is required to remove inhibition.
Soil samples were analyzed in triplicate for each soil type.
Air filter washes are an average of 10 independently collected filters.
Surface swipes were collected with a BISkit surface sampler and assayed in triplicate.
Discussion
Biological warfare agent detection in environmental samples requires rapid diagnostic techniques that can give results in hours, as opposed to the days required for traditional culturing methods. Sample analysis by real-time PCR requires effective nucleic acid purification and removal of inhibitors. Through the combination of two magnetic bead-based DNA purification kits, an effective automated protocol was created for the purification of nucleic acids from spore-forming and vegetative bacteria, by using the biological warfare agent simulants B. subtilis var. niger and E. herbicola. The military often employs biological simulants to represent highly pathogenic biological weapons and to perform research and testing functions in a safe and cost effective manner. E. herbicola is a biological simulant for gram– negative vegetative bacteria such as Yersinia pestis, and B. subtilis var. niger spores, also called BG spores, are a gram-positive spore-forming bacterial simulant meant to mimic Bacillus anthracis. 11
Although detection of microorganisms by real-time PCR can be accomplished without a separate DNA purification step for some types of samples,12–13 the removal of PCR inhibitors from dirty environmental samples necessitates DNA purification techniques. Paramagnetic bead-based DNA purification has proven effective in generating “clean” PCR templates and has been adapted for automated platforms developed to screen samples by the U.S. military. Bead-based methods allow fluid-like dynamics, and the beads can be fully resuspended during washing steps to facilitate efficient removal of potential inhibitors.6, 7, 14
The hybrid method described here utilizes a macroporous silica resin that surrounds a ferrous oxide core. This resin was developed for food matrices, but the hybrid method improved its use for environmental samples by the incorporation of the lysis buffer from a blood kit. The blood kit lysis buffer contains a higher concentration of guanidinium chaotrope when compared to the two-part lysis buffer from the food purification kit (personal communication, Promega Corp.). This higher concentration of chaotropic salt may result in improved recovery of nucleic acid material and the reduction in the thermocycling detection threshold (CT score) when processing biological simulants. An additional advantage is using only one lysis buffer, resulting in a faster processing time and fewer consumables. It is likely that the chaotrope produces more efficient lysing of vegetative cells and more efficient recovery of trapped DNA that is within the spore sample.
The detection limit observed for E. herbicola is in close agreement with previously published results 15 and was observed with other vegetative rods 16 and viruses. 17 The detection limit observed with BG spores following processing with the hybrid method is in agreement with some studies12,18,15 but higher than the 25–250 spores/mL reported in other studies.19–20 It is important to note that these lower claims came from performing PCR directly from relatively inhibitor-free samples and without a procedure for the removal of extraneous non-nucleic acid material from environmental samples. The study by Kuske et al. 21 employed the use of bead mill homogenization to effectively lyse Bacillus spores. Manual steps such as bead-beating and sonication are undesirable to a completely automated approach and have additional problems like the creation of aerosols of biological materials. They also require disruption facilitators such as silica or zirconium beads that have to be removed from each sample downstream in the process. In spite of its limitations, bead-beating was shown to greatly increase the limit of detection of BG spores in our study and would likely be an essential part of biological agent detection. It is important to note that the detection limits in this study were determined using E. herbicola and BG spore preps where every effort was made to remove all extracellular debris through a series of aggressive washing steps. It is likely that real-life environmental samples would have some vegetative growth, thus improving the detection limit when clumped, and dirty spore preparations are extracted using this system. 11
The developed hybrid protocol was effective in removing PCR inhibitors from washes of air filters, surface swipes, and eight different soil types, as observed by the minimal shift in CT values observed in spiked samples. The level of detection that our method obtains meets our internal requirements for environmental samples, and the DNA extraction step is accomplished in only 70 min without requiring a spore germination step. 22 In combination with an integrated automation system, the entire protocol is performed without user intervention, greatly reducing our staffing requirements, and thus reducing our overall cost.
The Automated Biological Agent Testing System (ABATS) has been designed to reduce the turnaround time for analysis of unknown environmental samples as part of a routine bio-surveillance operation or to serve as a replacement for manual laboratories in the event of sample surge at high-impact testing sites. As part of the labor reduction effort, we have developed a hybrid protocol for extraction of nucleic acids from environmental samples that can be fully automated and represents a significant increase in sample throughput compared to manual DNA preparation methods. The ABATS system can realize a two-thirds reduction in the cost per sample, based upon our current operational capability in which 300 samples are processed per day for six bio-threat agents. Higher sample numbers that process 1000 samples a day are possible and will reduce cost per sample even more when using this system. The utility of the magnetic bead resins is in their capacity for full automation, their ability to handle real-world environmental samples, and their ability to isolate total nucleic acids. The use of this purification method on the Biomek® FX will be useful for screening sites in which smaller numbers of samples will be processed. This method is being used by the Department of Defense as part of a concept called Stations of Robotic Monitoring (STORM), which seeks to employ islands of automation to automate only the slowest steps in a testing process. Clearly, the linkage of automated processing systems with traditional laboratory methodologies has allowed for an increase in the efficiency of our operations and can provide a cost savings to national homeland defense efforts.
Footnotes
Acknowledgments
The authors wish to acknowledge the contribution of Agnes Akinyemi and Kevin Kearns for supplying cell cultures and bead-beating equipment and for helpful discussions. We also thank Sanjiv Shah at Edgewood Chemical Biological Center for providing primer and probe sequences for B. subtilis var. niger and Joan Gebhardt at the Naval Medical Research Center for primer and probe sequences for E. herbicola detection. For technical suggestions and discussions, we thank Evan Skowronski (LLNL) and Mike Ehret (MRI). The authors would also like to thank Marc Goldstein and Dawna Houston at Beckman-Coulter for all of their assistance. For document editing, we thank Jessica Dang. This work was supported in part by the Joint Program Executive Office for Chemical and Biological Defense.
