Abstract
Ctenocephalides felis were killed and collected from 92 cats in Alabama, Maryland, and Texas. The fleas and blood from the corresponding cat were digested and assessed in polymerase chain reaction assays that amplify DNA of Ehrlichia species, Anaplasma phagocytophilum, Neorickettsia risticii, Mycoplasma haemofelis, ‘Candidatus M haemominutum’ and Bartonella species. DNA consistent with B henselae, B clarridgeiae, M haemofelis, or ‘Candidatus M haemominutum’ was commonly amplified from cats (60.9%) and their fleas (65.2%). Results of this study support the recommendation to maintain flea control on cats in endemic areas.
Ctenocephalides felis commonly infests cats and is known to harbor a number of infectious agents including Bartonella quintana, B koehlerae, B henselae, B clarridgeiae, Rickettsia felis, Wolbachia pipientis, and ‘Candidatus Mycoplasma haemominutum’ (Bergmans et al 1997, Comer et al 2001, Ishida et al 2001, La Scola et al 2002, Rolain et al 2003, Shaw et al 2004). Fleas can transmit B henselae infection between cats and may play a role in the transmission of Bartonella species between cats and people (Chomel et al 1996, Foil et al 1998). ‘Candidatus Mycoplasma haemominutum’ and M haemofelis have been shown to be ingested by C felis when allowed to feed on experimentally infected cats and DNA of both haemoplasmas has been detected in flea faeces or eggs (Woods et al 2005). Recently, clinically ill cats have been shown to be infected with an Ehrlichia canis like organism as well as Anaplasma phagocytophilum (Bjoersdorff et al 1999, Beaufils et al 2002, Breitschwerdt et al 2002, Lappin et al 2004). In addition, cats become clinically ill when inoculated experimentally with Neorickettsia risticii (Dawson et al 1988). To our knowledge, whether C felis is involved in the transmission of these organisms has not been assessed.
Numbers of studies of pathogen carriage by fleas collected from client-owned cats are small and have been performed in countries other than the United States (Bergmans et al 1997, Ishida et al 2001, La Scola et al 2002, Parola et al 2003, Rolain et al 2003, Kelly et al 2004, Shaw et al 2004). To our knowledge, only one study has reported the results of Bartonella species testing of cats and their fleas (La Scola et al 2002), no study has attempted to directly correlate results of haemoplasma species testing between cats and their fleas, and no study has assessed C felis for the presence of Ehrlichia species, N risticii, or A phagocytophilum DNA. Thus, the purpose of this study was to attempt to amplify Bartonella species DNA, haemoplasma species DNA, Ehrlichia species DNA, N risticii DNA, and A phagocytophilum DNA from the blood of cats and their fleas from three states in the United States.
Materials and methods
Cat blood and fleas
Veterinarians in Alabama (BG and AR), Maryland (Brunt), and Texas (Burney) were requested to kill fleas seen on cats with pyrethrin spray, collect the fleas, and store them at −20°C. All fleas were C felis based on morphology (Menier and Beaucornu 1998). At the same client visit, 1 ml of blood was drawn by jugular venepuncture from the cat with fleas, placed in a 1.5 ml draw EDTA tube, and stored at −20°C. The blood of the cats and their fleas were batched until shipped on cold packs by overnight express to Colorado State University. On arrival, the samples were stored at −70°C until assayed.
Polymerase chain reaction (PCR) assays
After being thawed at room temperature, blood and fleas from each cat (varying from 1 to 14 fleas per cat) were prepared for PCR assays as previously described (Woods et al 2005). A multiplex PCR assay that amplifies the DNA of known, sequenced Ehrlichia species, A phagocytophilum, N risticii, M haemofelis, and ‘Candidatus M haemominutum’ (Jensen et al 2001, Lappin et al 2004, submitted for publication) and a PCR assay that amplifies the DNA of seven Bartonella species were used (Jensen et al 2000). Both haemoplasma species and each Bartonella species give a different band size and these sizes are used to determine which species is present in the sample. Appropriate positive and negative controls were included in all assays.
Genetic sequencing
It was recently reported that Bartonella species PCR assays that use the 16S–23S rRNA intergenic region can also amplify Mesorhizobium species DNA resulting in a 420-bp amplicon (Maggi and Breitschwerdt 2005). While none of the Bartonella species amplicons in this study were of that size, we chose to perform genetic sequence analysis on Bartonella species amplicons from 10 cats and 10 flea groups using a commercially available genetic sequencing service (Macromolecular Resources, Colorado State University, Fort Collins, CO 80523).
Results
Blood and at least one flea were available from 92 cats; 54 were from a humane shelter in Alabama, 5 were from a veterinary clinic in Alabama, 25 were from a veterinary clinic in Maryland, and 8 were from a specialty hospital in Texas. Because of small sample sizes, we did not attempt to compare between states or correlate to the presence or absence of clinical disease.
DNA consistent with B henselae, B clarridgeiae, M haemofelis, or ‘Candidatus M haemominutum’ was commonly amplified (Table 1) from cats (60.9%) and their fleas (65.2%). All of the Bartonella species amplicons assessed by genetic sequencing were the appropriate Bartonella species as predicted by the amplicon size (Jensen et al 2000). DNA of Ehrlichia species, A phagocytophilum, or N risticii was not amplified from the blood of any cat or their fleas.
Distribution of Bartonella species and haemoplasma species PCR assay results from cat blood and flea digests in the United States
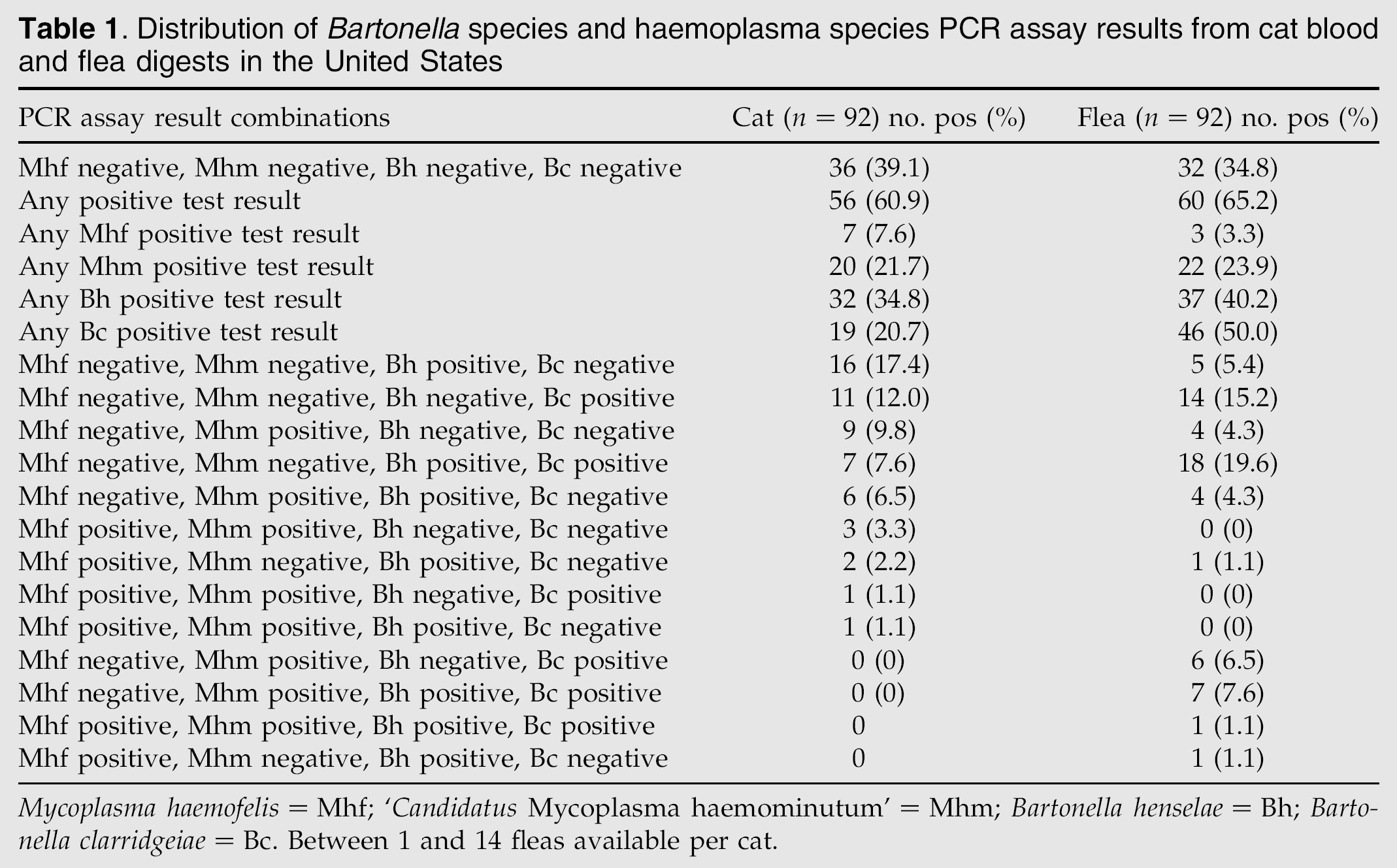
Mycoplasma haemofelis=Mhf; ‘Candidatus Mycoplasma haemominutum’=Mhm; Bartonella henselae=Bh; Bartonella clarridgeiae=Bc. Between 1 and 14 fleas available per cat.
Of the 32 cats from which B henselae DNA was the only Bartonella species DNA amplified from blood (overall prevalence rate of 34.8%), 21 cats (65.6%) had fleas from which B henselae DNA was amplified. Of the 19 cats from which B clarridgeiae DNA was the only Bartonella species DNA amplified from blood (overall prevalence rate of 20.7%), 18 cats (94.7%) had fleas from which B clarridgeiae DNA was amplified. Of the seven cats from which both B henselae and B clarridgeiae DNA were amplified from blood, six (85.7%) had fleas from which both B henselae and B clarridgeiae DNA were amplified. Both B henselae and B clarridgeiae were amplified from the blood of some cats and their fleas in all three states.
Of the seven cats from which M haemofelis DNA was the only haemoplasma species DNA amplified from blood (overall prevalence rate of 7.6%), none had fleas from which M haemofelis DNA was amplified. Of the 20 cats (overall prevalence rate of 21.7%) from which ‘Candidatus M haemominutum’ DNA was the only haemoplasma species DNA amplified from blood, 3 (15%) had fleas from which ‘Candidatus M haemominutum’ DNA was amplified. Of the five cats from which both M haemofelis and ‘Candidatus M haemominutum’ DNA were amplified from blood, one (20%) had fleas from which both M haemofelis and ‘Candidatus M haemominutum’ DNA were amplified. While ‘Candidatus M haemominutum’ DNA was amplified from the blood of cats and their fleas from all three states, M haemofelis DNA was only amplified from the blood of cats and their fleas that resided in Alabama.
Discussion
Results of this study show that DNA of M haemofelis, ‘Candidatus M haemominutum’, B henselae, and B clarridgeiae can be amplified from the blood of cats and from whole body digests of C felis that were killed while found on cats in three states in the United States. The results document that infection with one organism does not induce cross protection against the others and that co-infections are common. We failed to amplify DNA of Ehrlichia species, A phagocytophilum, or N risticii from cats or fleas in this study.
Bartonella henselae is associated with feline red blood cells and it has been previously shown that C felis is a mechanical vector for this organism (Chomel et al 1996). Multiple studies from throughout the world show an increased risk for B henselae infection in cats with fleas (Foley et al 1998, Al-Majali 2004, Guptill et al 2004). Viable B henselae is passed in faeces of the flea (Higgins et al 1996, Finkelstein et al 2002) and infected flea faeces can result in infection of cats via inoculation (Foil et al 1998). Bartonella henselae DNA has been amplified from flea digests in a number of other countries including the United Kingdom (Shaw et al 2004), Thailand (Parola et al 2003), Japan (Ishida et al 2001), New Zealand (Kelly et al 2004) and France (La Scola et al 2002, Rolain et al 2003). Bartonella clarridgeiae has been grown from the blood of cats or amplified from flea digests in Italy (Fabbi et al 2004), France (La Scola et al 2002, Rolain et al 2003), Thailand (Parola et al 2003), New Zealand (Kelly et al 2004), and the Netherlands (Bergmans et al 1997) but not the United Kingdom (Shaw et al 2004). We believe this is the first report documenting B clarridgeiae DNA in C felis in the United States and infection was common. To our knowledge, prevalence rates for Bartonella species infections in cats and the fleas that parasitize them have only been reported once. In that study, flea digests from 10 cats were assessed; some Bartonella species infected cats were parasitized by Bartonella species negative fleas and some Bartonella species negative cats were parasitized by Bartonella species positive fleas (La Scola et al 2002). In our study, the majority of Bartonella species infected cats were parasitized by Bartonella species infected fleas; 94.7% of the B clarridgeiae infected cats carried B clarridgeiae infected fleas. These results suggest, but do not prove, that C felis is a vector for B clarridgeiae.
To our knowledge, this is the first report of M haemofelis DNA detected in flea digests from a naturally exposed cat; DNA of the organism was not detected in fleas from cats in a similar study in the United Kingdom (Shaw et al 2004). However, this may only reflect overall haemoplasma prevalence rates. As in previous studies, the prevalence of ‘Candidatus M haemominutum’ was greater than that of M haemofelis in cats (Jensen et al 2000, Tasker et al, 2003, 2004, Luria et al 2004) and fleas (Shaw et al 2004). In the study described here, M haemofelis DNA was only detected in one cat and its fleas (both were coinfected by ‘Candidatus M haemominutum’) and ‘Candidatus M haemominutum’ DNA was amplified from only 15% of fleas collected from positive cats. We have shown that both haemoplasmas are ingested by C felis while feeding and that DNA from both haemoplasmas can be detected in flea faeces (Woods et al, submitted for publication); however, it is unknown whether the organisms are alive. While we were able to show transmission of M haemofelis to one cat by flea haemophagous activity, our attempts to transmit ‘Candidatus M haemominutum’ to cats by flea haemophagous activity or either haemoplasma by feeding fleas and flea faeces containing haemoplasma DNA to cats have failed to date (Woods et al, submitted for publication). Results of epidemiological studies assessing the association between haemoplasmosis and fleas have also varied (Nash and Bobade 1986, Grindem et al 1990, Hackett et al, submitted for publication). Thus, the role C felis plays in the transmission of haemoplasmas is still unclear.
While all potential vectors for Ehrlichia species, A phagocytophilum, and N risticii infections in cats are unknown, Ehrlichia species and A phagocytophilum are generally associated with ticks and N risticii is associated with aquatic insects (Mott et al 2002). In this study, DNA of the organisms was not detected in the blood of cats or their fleas. We also failed to amplify DNA from the organisms from feral cats in Florida (Luria et al 2004). As Florida, Alabama, and Texas are known to be endemic for E canis infections in dogs, we believe these results suggest that cats are more resistant to infection than dogs, that cats are less commonly exposed to appropriate vectors than dogs, or that cats remove the vector before transmission can occur. Alternately, the negative results may only reflect the sample size of this study. Anaplasma phagocytophilum infections are generally restricted geographically by the presence of Ixodes species ticks which are not common in Florida, Texas, or Alabama. While Ixodes ticks are present in Maryland, our failure to detect A phagocytophilum infections of cats may only reflect the small sample size (25 cats).
The patterns of Bartonella species or haemoplasma species DNA detected in individual cats and their fleas frequently varied in this study. However, it is impossible to know how long each flea parasitized an individual cat and so discordant results may indicate that fleas killed on an individual cat may have just recently parasitized the cat. In addition, some variability in the results of this study may relate to overall sensitivity of the PCR assays because of the differences in numbers of fleas available from each cat for assessment. However, in experimentally infected cat studies, DNA of either haemoplasma species is consistently detected in digests of five fleas (Woods et al, submitted for publication). Lastly, it is possible that a cat blood sample could be contaminated by flea feces during venepuncture. Regardless, haemoplasma species and Bartonella species infections of cats and fleas were common in the states studied. Haemoplasma species and Bartonella species are associated with clinical disease in some cats and Bartonella species are associated with clinical disease in some people. While it has not been shown that use of flea products lessens risk of infection by either genera to date, we believe that our results support the recommendation to maintain flea control on cats (Brown et al 2003).
