Abstract
Molecular diagnostic analysis and life science studies are dependent on the ability to effectively prepare samples for analysis. We report the development of a system that enables robust sample preparation of nucleic acids. To enable completely automated sample preparation, a consumable cartridge and consumable module system were developed to emulate every step of the sample preparation process. This included enzyme and reagent addition, temperature-controlled incubations, noncontact mixing of enzymes and reagents, buffer exchanges, and sample elution. Using this system, completely automated methods were developed for the purification of viral RNA and DNA from plasma and whole blood and of bacterial genomic DNA from water and whole blood. Extracted nucleic acids were detected and quantified using real-time PCR. The data indicate that automated viral DNA extraction was more efficient than sample extractions performed using a manual process, whereas automated total RNA extraction from the same samples was equivalent to controls. Additionally, we found that the process for bacterial genomic DNA extraction from either water or whole blood was equivalent to the manual extraction processes. We conclude the instrument, consumable cartridge, and reagent system enables easy, cost-effective, and robust sample preparation regardless of the experience of the operator.
Introduction
Sample preparation represents a significant bottleneck to a large proportion of molecular diagnostic and life sciences applications and is cited as the most important quality factor in gene expression studies. 1 Although most of the development in the field of molecular diagnostics is devoted to improving methods for the detection and identification of disease-related target analytes, less focus is devoted to the preparation of purified samples for these systems. Our goal in the following studies was to develop a systems wide approach to enable highly flexible, completely automated methods for the preparation of viral and bacterial target analytes from a wide variety of matrixes, ranging from water to whole blood. Here, we report the development an automated instrument (Fig. 1) to address many problems associated with complex methods including sample collection, lysis, separation of the biological macromolecules from a complex contaminant-rich environment, purification of these macromolecules from the remaining contaminants and, finally, elution of the sample in solutions that are compatible with downstream analysis processes. 2
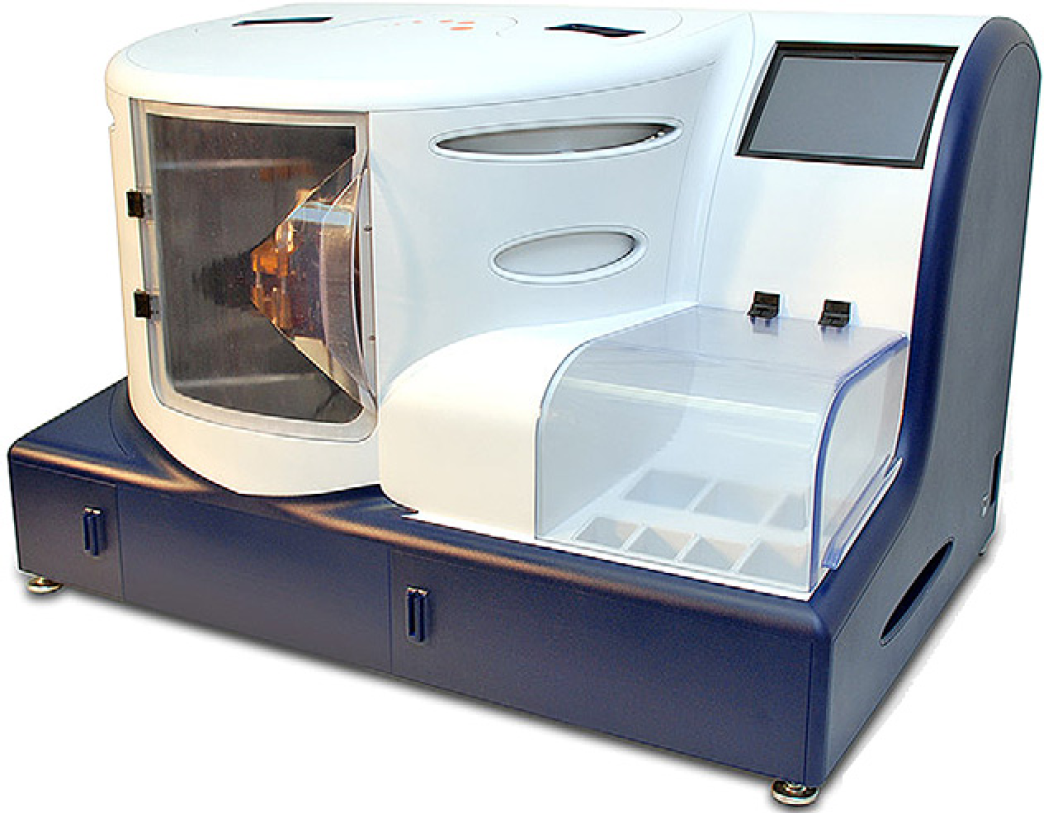

Xisyl instrument: The automated sample preparation instrument, in this embodiment, is a benchtop system that occupies approximately 12 ft3 (3″× 2″ × 2″). The system allows for front access for sample cartridge loading and removal and is controlled through the use of a touch screen located on the upper right side of the instrument. All solutions are housed in the front case, with the exception of the removable waste container, which is located at the front base of the instrument beneath the fluidic modules.
Although a plethora of sample preparation platforms exist, 3 –6 few have truly integrated all of these functions together into one user-friendly and flexible platform. Most of the sample preparation systems are based on magnetic bead technology 7 or spin columns, 8 which require multiple benchtop steps to perform the individual functions of the process and are prone to user error depending on the experience of the laboratory technician. With the increasing dependence on automation of molecular diagnostic testing in the clinical environment, more standardized, reproducible methods and systems for sample preparation are required. Several high-throughput systems have been recently developed to address this challenge. 9 –12 However, these systems require some form of preprocessing for various sample types, such as for the extraction of gram-positive bacterial nucleic acids from whole blood, which requires a combination of enzymatic digestions before the analyte capture, purification, and elution. There is a need for the development of truly integrated technologies to perform these functions and provide consistent samples for molecular diagnostic assays. Additionally, our goal in these studies was to develop a system that could serve diagnostic laboratories with a wide variety of applications and sample types. We designed the system to address a range of user requirements addressing sample turnaround time and system flexibility.
Our previous studies focused on novel methods for the disruption of cellular organisms including bacterial spores using ultrahigh temperatures 13 and improved methods for the selective isolation of nucleic acid species, such as mRNA 14,15 and sequence specific target capture. 16 These previous developments heavily influenced the system designed in the studies presented here. To construct a highly flexible platform that could perform a variety of applications, we have designed a novel device that is capable of integrating every step required for the lysis of viral, bacterial, and eukaryotic cells and the extraction and purification of the target analytes from a variety of sample types, including large volumes of whole blood, plasma, and water. The resultant consumable device enables the user to simply load the sample into a cartridge and place that cartridge on the instrument to enable automated sample preparation. The heart of the system is a consumable module designed to enable the addition of enzymes, buffers, washing, and elution reagents, and perform multiple temperature-controlled incubation and mixing steps necessary for efficient target analyte purification. In the current embodiment, the instrument is designed to house 12 consumable modules, which operate independently from one another, enabling random access method execution. This allows the user to process several different sample types (e.g., water, plasma, blood) with a variety of methods (e.g., RNA, DNA) during a single run.
Experimental Details
Consumable Design and Operation
To enable completely automated sample preparation, a consumable was designed to allow all sample preparation steps to be performed in a single cartridge (Fig. 2). In the fields of molecular diagnostics and environmental sampling, typically large sample sizes are required to attain the necessary assay sensitivity. As a result, the consumable module was designed to process samples as large as 1.0 mL of raw sample input using conventional chemistries, and will be capable of processing considerably larger samples using hybridization capture methodologies. Most of the conventional methods, typically using chaotropic salts and alcohol-based extraction methods, require a vessel that has a volume of approximately 10× of the original sample size, to accommodate the addition of buffers and solvent and leave enough head space to manipulate the sample. A starting volume of 1.0 mL requires that the consumable cartridge be designed to contain a volume of approximately 10.0 mL of fluid volume. To enable lysis, preparation, and purification of robust organisms, several design attributes are necessary, the most important of which is the ability to effectively mix the sample with the variety of enzymes, buffers, and solvents added to the sample. As seen in the exploded view (Fig. 2A), the multiple component consumable is designed to achieve this functionality. To enable sample containment and mixing, we made use of a magnetic disk that serves two functions. The first function the disk serves is to seal the cartridge after sample loading and during transport of the consumable to the instrument. This sample containment is critical when processing samples that contain potentially hazardous pathogenic agents. To accomplish this, the disk was designed to be press fit inserted in to the cartridge cap, to prevent liquid flow out of the top of the consumable during transport to the instrument (Fig. 2A, B). Once loaded on to instrument, this disk is ejected from the sample cap (Fig. 2C) to enable its second and primary function. Having been ejected from the sample cap, the disk is now located within the sample chamber, and can be actuated by an external magnetic field. Depending on the extraction method, a series of buffers are injected into the consumable cartridge. During these filing steps, the magnet can be rotated to enable noncontact mixing of the enzymes, buffer, and alcohols within the consumable cartridge, significantly reducing or eliminating the possibility of sample-to-sample carryover and cross contamination.
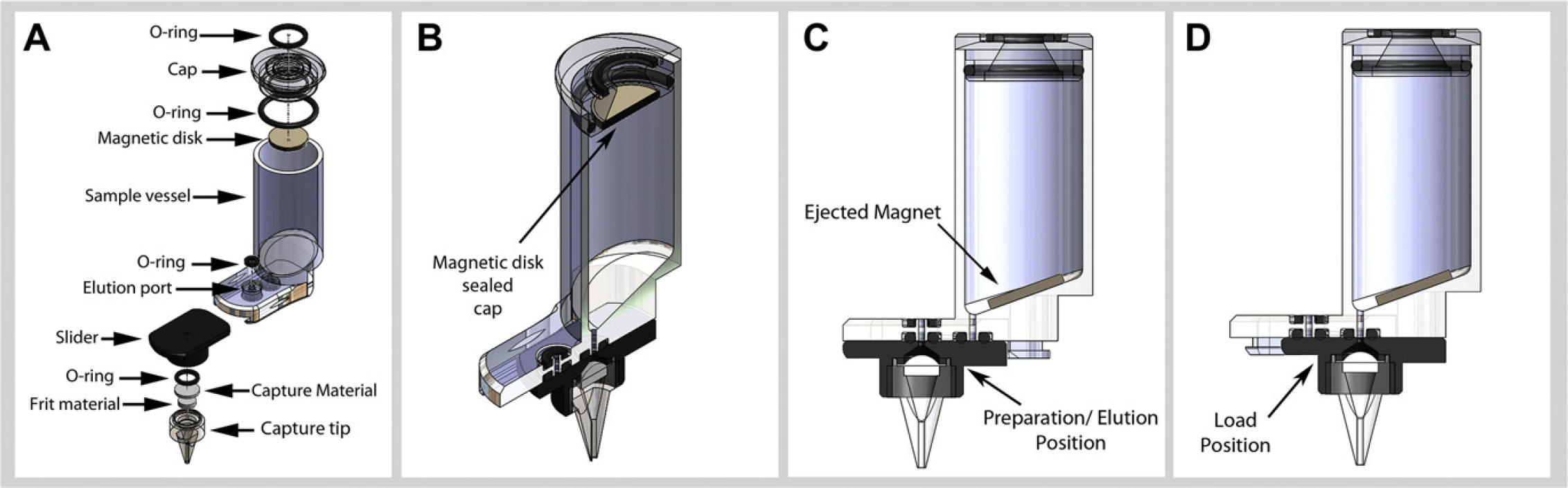
Consumable cartridge: Samples are processed in a self-contained multicomponent sample cartridge. The single use consumable contains a tubular reaction chamber, which is connected through a slide to a capture material that facilitates sample extraction. This slide allows the capture material to be positioned to incubate/mix, capture, or elute the purified sample. (A) Shows the exploded view of the consumable. In (B), the cartridge is positioned to receive a sample, with the disk secured in the consumable cap to allow for full-sample containment inside the consumable. In (C), the magnet is ejected from the consumable cap to enable mixing of sample and reagents. In (D), the slider is located in the purge position, with the capture material positioned to be in fluid communication with the reaction chamber.
Extraction and purification of the target analytes is performed using a capture material located at the base of the consumable cartridge. This capture material is contained within a movable sliding component; we term the “slider” (Fig. 2A). The slider is connected to the consumable base on a rail system that enables the capture material to be spatially located at one of two positions in the current embodiment of the consumable cartridge. In the first position, the capture material is positioned directly under the elution port preventing sample flow out of the main chamber (Fig. 2C). The slider is located in this position while all preparation steps before sample capture are performed. At the conclusion of the temperature-controlled enzymatic and buffer incubation steps, this slider is actuated to move into a second position directly below the main sample chamber where target analyte extraction on the capture material is conducted (Fig. 2D). During this extraction step, the consumable module is positioned directly beneath an air input line that, when contacted to the consumable, enables pressurization of the cartridge by the instrument. This pressurization forces the prepared sample through the capture material enabling analyte extraction. Once extracted, several washes of the internal vessel of the consumable cartridge are conducted by alternating the dispensing of solutions with the pressurization of the consumable. When these washes are completed the consumable is dried with air while being heated. This step is critical for the removal of ethanol, which is especially inhibitory to various enzymes used in downstream molecular diagnostic assays such as PCR and sequencing. When this capture step is complete, the slider is again moved into the first position directly underneath the elution port on the consumable base (Fig. 2C) to elute the purified sample from the consumable device. The ability to isolate flow paths during capture and elution are critical to processing samples free of contaminants.
Sample preparation in the cartridge is performed in coordination with the consumable module (Fig. 3). A critical feature of any sample preparation system or molecular diagnostic analysis platform is the ability to limit or eliminate sample cross contamination. Essential to this effort is the ability to perform noncontact filling and mixing, which was accomplished by designing a consumable module (Fig. 3) that is able to externally direct the internal workings of the cartridge. As an example, the loading of buffer and solvents is performed using a nozzle that is positioned to allow the solutions to be added to the consumable cartridge without contacting the sample itself. To enable rotation of the magnetic disk, a rare earth magnet is located at the base of the cartridge approximately 2 cm from the base of the consumable cartridge. This magnet can be rotated on axis at variable speeds for a duration that is scripted to the individual consumable module. Additionally, the consumable module has a geared DC motor capable of actuating the slider of the consumable. Thermal control of the consumable and its contents are achieved with an aluminum sleeve that makes contact with the exterior of the consumable. Thermal heating is accomplished using a resistive board mounted heater, thermistor monitoring of temperature, and a miniature fan for forced air convective cooling.

Consumable module: Consumables are used in conjunction with a module that performs capture/elution tip positioning, noncontact mixing, and thermocontrol of the cartridge. This module was designed (A) to perform stand-alone sample preparation by having onboard proportional integral derivative control of the module functions and fluid connectivity to outgoing waste stream. The module is spring loaded to enable reproducible compression of the consumable to the system fluid manifold. (B) Shows a working prototype module with consumable.
The individual modules are operated independently using a Controller Area Network communication system (Fig. 4). Controller Area Network is a message-based protocol, designed specifically for automotive applications but now also used in other areas such as industrial automation and medical equipment. 17 This system enables each consumable module to independently operate an individual method script during a single-sample preparation run. Each consumable module contains discrete electronics hardware and software packages that are slaved to the central controller. This electronic package enables the central controller or motherboard to execute high-level commands communicated to the slaved consumable module that can operate proportional integral derivative control of the temperature, mixing, and slider motion control. In this way, the motherboard contains all of the high-level commands for the various methods that can be operated in tandem or in parallel.
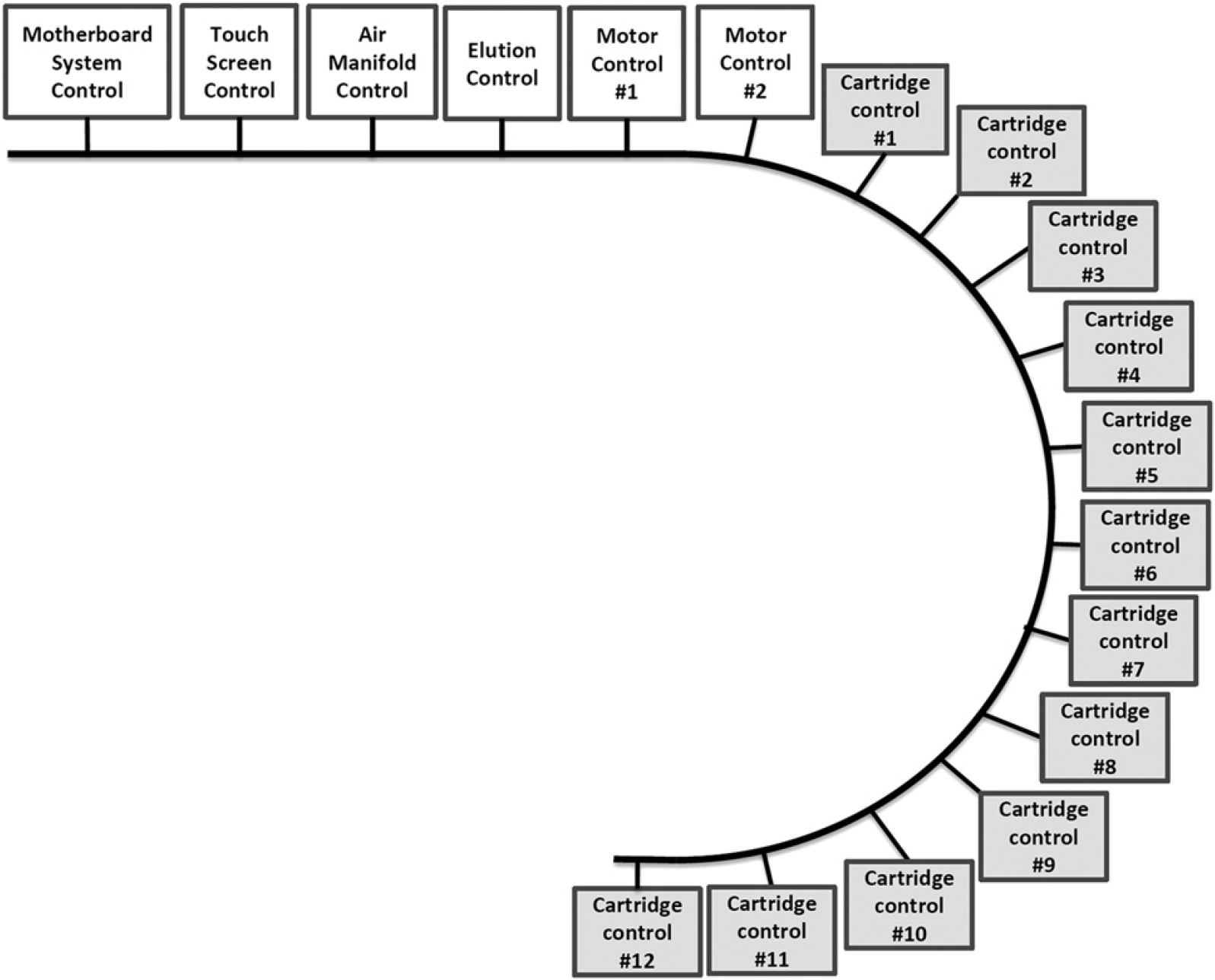
CAN system layout: A Controller Area Network (CAN) was used to perform system communication. This communication was selected for it's flexibility and it's ability to perform parallel control of the slaved consumable modules. This communication protocol is essential to independently control each module and provides the ability for random access control.
In addition, the modules are connected to a waste manifold at the base of the instrument (Fig. 5). This manifold evacuates waste from the capture tip under vacuum, without contacting the capture tip. We have found in previous prototype systems, this was critical system feature as contaminants such as chaotropic buffers and alcohol can accumulate on the external surface of the capture tip. If these contaminants are not removed, they can inhibit the downstream molecular diagnostic analysis. During the extraction of the samples, the capture tip interfaces with this manifold to remove the chaotropic buffers and alcohols by directing them to a waste container located in an easily accessible drawer at the front base of instrument. This waste container is connected to the waste manifold using a quick connect system that enables the user to remove the waste

Internal system arrangement and assembly: The consumable modules are arranged on a circular platen that allows rotation of the modules to several positions allowing for dispensing of multiple reagents, compressed air purging of the consumables, and elution of the purified samples.
The instrument construction creates a partition between the major electrical components and fluidic components. The system was designed to compartmentalize the pumps, motherboard, and touch screen on the right side of instrument, while the consumable modules, waste, and elution manifolds are located on the left compartment (Fig. 5) of the instrument. This design affords a safety measure in the unlikely event, there is a failure to contain the fluids in any of the major liquid-handling components. The current system was designed to accommodate 12 consumable modules on a rotary platen. In this format, the consumable cartridges can be positioned at various dispensing and airflow ports to enable automated preparation of any number of samples up to the full complement of consumable modules. As described, each module has the ability to operate independently of one another, which enables random access sample preparation.
The system that consists of the consumable cartridge, the consumable module, along with the system software and hardware works in concert to completely automate the sample preparation procedure. The process of sample preparation is essentially a series of commands, that when used with a particular set of reagents, produces purified samples for downstream molecular diagnostic assays. We found three parameters were critical in the effective automation of this process that include (1) temperature control, (2) adequate sample and reagent mixing, and (3) sample elution.
We found that stable temperature control was possible for all incubations required by the system (Fig. 6). In a representative example, four modules were heated while monitoring both the external block temperature and the internal fluid temperature. In this experiment, we found the first incubation at 37 °C (typically used for the lysozyme incubations) was reached in approximately 2 min, by heating the external block to 65 °C. We found no significant variation in the internal fluid temperature over a period of approximately 15 min. A second incubation at 56 °C is often required when whole blood samples are being processed. This incubation makes use of proteinase K—a proteolytic enzyme that digests many proteins contained within the sample. To reach 56 °C, the heating block temperature was increased to approximately 100 °C for a period of approximately 3 min. We found the internal fluid temperature increased rapidly to 56 °C, and was stable over a period of 15 min, the required time for effective protein digestion. The final incubation in several of the applications is the deactivation of digestive enzymes such as lysozyme and denaturing of the nucleic acids at 95 °C. To reach the required temperature, the aluminum block was heated to approximately 120 °C for a period of 1 min then lowered to 110°C for a period of 10min. Although the temperature ramp was slower, the samples did reach the required effective temperature, 95 °C, and again was stable for a period of approximately 15 min. At the conclusion of this step, a fan located on the back of the aluminum block is activated. This fan removes heat from the aluminum block by convection, and is able to rapidly cool the heating block back to room temperature in approximately 1 min. We found that the cooling of the internal fluids was also reduced to room temperature in approximately 5 min.
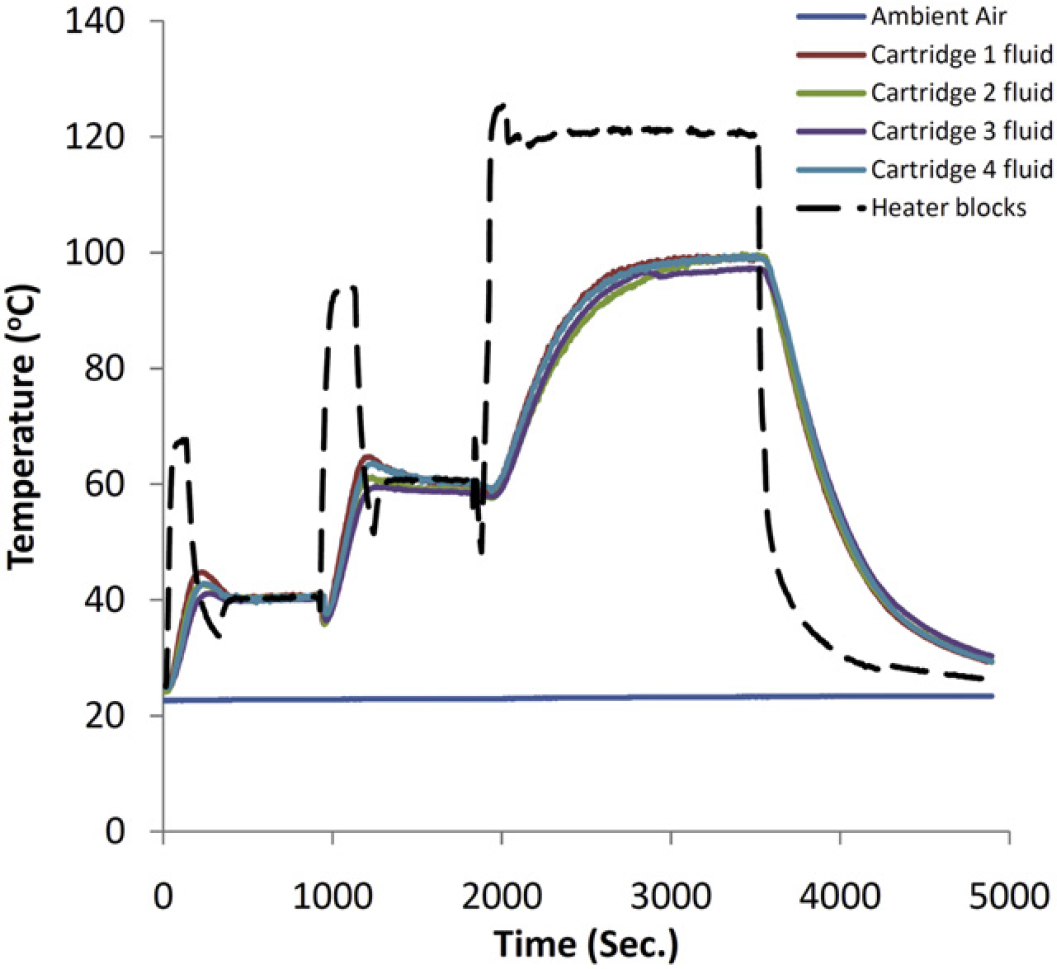
Thermal control of consumable: Traces of sleeve temperature and internal fluid temperatures are displayed. Thermal control is achieved by resistively heating an aluminum sleeve around the consumable and controlled via proportional integral derivative using thermistors for temperature feedback.
For several of the applications used on this system, the methodologies require the addition of a chaotropic buffers and alcohols to facilitate the extraction of the liberated nucleic acids. During these steps, we found the most critical system aspect was the mixing of the added components with the enzyme-digested samples. To achieve effective mixing of the digested samples, we made use of a magnetic disk that when rotated creates turbulence in the sample preparation chamber, which effectively mixes the sample. We found that the rotational axis of the spinning magnetic field and the speed of the rotating field were particularly important factors in the ability to perform adequate mixing, resulting in high-efficiency nucleic acid extraction detectable by quantitative real-time (RT)-PCR (Fig. 7). In addition, the timing of mixing (e.g., during buffer addition) and the rate at which buffer addition is performed during mixing also had an impact on effective mixing. To evaluate the extraction of nucleic acids, we first performed extraction of 1.0-mL blood samples that contained Epstein Barr Viral (EBV) DNA. We found that using the benchtop mixing procedure of repeatedly aspirating the samples yielded nucleic acid extraction with approximately a 50% DNA recovery from whole blood (which is routinely achieved with the benchtop methodology). Our first attempts to mix the samples by rotating the magnetic disc on a Z-axis (which in effect spins the magnet on its end around the Z-axis) formed a pronounced vortex and a homogenous solution was visually obtained. In contrast to the control, PCR yields were significantly lower than those of the control benchtop method (Fig. 7). We further noted that increasing the duration of this mixing step using this rotational axis up to 10min did not increase the efficiency of the extracted EBV DNA. When the orientation of the rotational movement of the magnetic disk is changed to the Y-axis (which in effect spins the magnet end over end in a “flip flop” motion around the Y-axis), a homogenous solution was also visually obtained but DNA extraction efficiencies with mixing up to 5 min at 150 rpm remained lower than the bench method, although slightly improved over Z-axis mixing. We further increased the speed of the magnetic disk rotation (up to the maximum 500 rpm capable by the mixer motor). We noted that this increased mixing speed resulted in increased yields of nucleic acid compared with either the Z-axis mixing or the Y-axis mixing at a lower speeds. By increasing the duration of mixing, we found that we were able to extract EBV DNA with equivalent PCR yields compared with the benchtop protocol for the extraction of large volume (1.0-mL) whole blood samples (Fig. 7). It is important to note that throughout all of the experiments, the samples appeared homogeneous in color and consistency. We found it was critical to assess mixing performance by analyzing the downstream prepared nucleic acids by RT PCR, rather than judge the samples by appearance only. Despite the lack of differences in appearance of the mixed sample as judged by color change, significantly higher levels of amplifiable DNA were evident as judged by PCR using the Y-axis rotational mixing at high speed compared with the Z-axis or Y-axis mixing at a low speed. The mechanism at the molecular level for the optimal performance in quantitative PCR using the Y-axis high-speed mixing motion compared the low speed or alternative axis orientated mixing is unclear. We suspect the answer lies in either the conformation of the nucleic acid or, alternatively, the rupture of micelles that may be forming as a result of the detergents present in the various buffers used in the sample preparation process. We did note that single-stranded total RNA was typically extracted at a higher efficiency, which may indicate that the nucleic acid is more efficiently extracted if single stranded. Further experiments to determine the molecular mechanism at work may allow faster, higher-efficiency nucleic acid extractions.
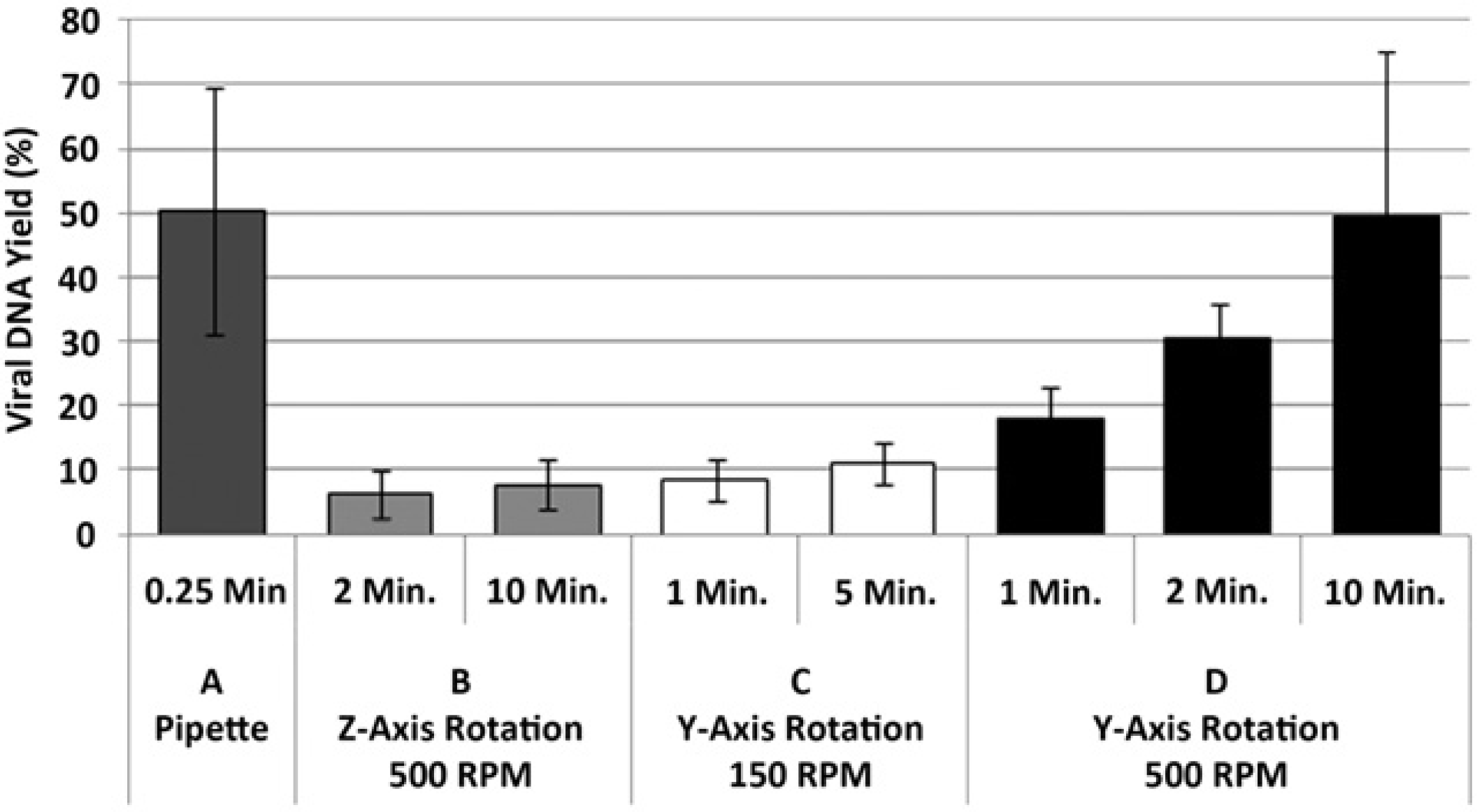
Mixing performance: DNA yields for automated and manual nucleic acid preparations with varying alcohol mixing parameters. Purified EBV DNA spike (75,000 copies) was purified from 1.0-mL whole blood, detected by quantitative PCR designed around the EBER-1 and EBER-2 genes, and percent recovery was calculated. Presented are: (A) Bench conditions using pipette mixing (
To perform this sample preparation process, the user places the consumable in a rack that is designed to contain up to 12 consumable cartridges, enough for one full load of the instrument. The samples typically are loaded and capped within a biosafety cabinet. Once loaded, the sealed consumable cartridges are transported to the instrument for loading on the consumable modules within the instrument. The user first actuates the touch screen and selects the intended method desired. The consumable device is then presented to a bar code scanner, which verifies the method selected and the loading position. The touch screen then indicates the system is ready for loading. The user opens the front access door and inserts the consumable cartridge in to the consumable module with a “snap” fit. In a similar fashion each consumable is loaded and, when complete, the user again uses the touch screen to begin the sample preparation run.
Once the preparation run is actuated, the system ejects the magnetic disks from the cartridge cap into the fluid sample (as shown in Fig. 2). The appropriate reagents are then added to the consumable and the samples are mixed using the noncontact magnetic disk apparatus. Depending on the application that is used on the system, several incubations at various temperatures may be required.
After all reagent additions, mixing, and incubations have been completed by the system, in a concerted series of actuations, the sample is pushed through the consumable to allow for capture of the nucleic acids on a material designed to extract the sample analytes from the mixture of enzymes, buffers, solvents, and contaminants. With the sample prepared for extraction, the slider component is positioned by the system into the “load” position that positions each consumable cartridge capture material beneath the “purge” port. The system raises the consumable module platen to compress the springs within the consumable modules and forms an airtight seal between the air port and the sample cap of the cartridge. With the seal pressure tight, the consumable is pressurized to force the prepared sample through the capture material as described above. Sensors within the system architecture monitor the purge pressure during this operation. At the point where these sensors detect a significant pressure drop, a valve is actuated to stop the pressurization of the consumable. In addition, the system recognizes when a sample will not flow through the capture material and notifies the user, an important feature to preserve precious samples if they are not properly loaded.
Once the sample has been captured, several wash buffers are dispensed into the consumable cartridge to remove any remaining unwanted contaminants on the capture material. Depending on the method selected, the number of sequential washes varies significantly, which is typically dependent on the turbidity and protein load of the sample being prepared. The final step in the purification process before the elution is removal of the remaining fluids contained within the capture material after the washes. To accomplish this, the system conducts air through the capture material while the consumable is heated to facilitate drying and removal of any additional contaminants.
With the loading and purification of the sample complete the system prepares for the elution of the sample. In this step, the consumables are rotated on the platen to the elution manifold. Once positioned beneath the elution manifold, the slider once again positions the capture material orifice beneath the elution port on the consumable base. With the consumable in the elution position, the system then compresses the consumable module against the elution manifold to form a pressure tight seal between the elution port of the consumables and the ports located on the elution manifold. The system dispenses the thermally controlled elution buffer through the capture tip (also thermally controlled) to elute the purified sample into a microcentrifuge tube located beneath the capture tip. This is an important system step, as reproducible elution volumes are critical for robust downstream quantitative molecular diagnostic assays. To provide reproducible elution volume, the system is capable of delivering elution buffer at a desired temperature, at various flow rates, and at varying increments. The system also allows for pressure release between elution positions in the event a clogged consumable is present, it will not affect the elution profiles of the remainder of consumables.
We found that using this process, reproducible elution volumes were achievable (Fig. 8). When a volume of 200μL was delivered to a consumable cartridge at each position of the automated system, we found that there was an approximate dead volume of 20 μL, as judged by the delivery of approximately 180 μL to each position. We found, over greater than 100 runs, that the elution volume had an average coefficient of variation (CV) of approximately 6.0%. We are continuing to optimize this scripted operation to reduce the CV to less than 5.0%. The data indicate that while there was variation in the delivery of the elution volume to each position, the elution volume for each sample at each position was reproducible and predictable.
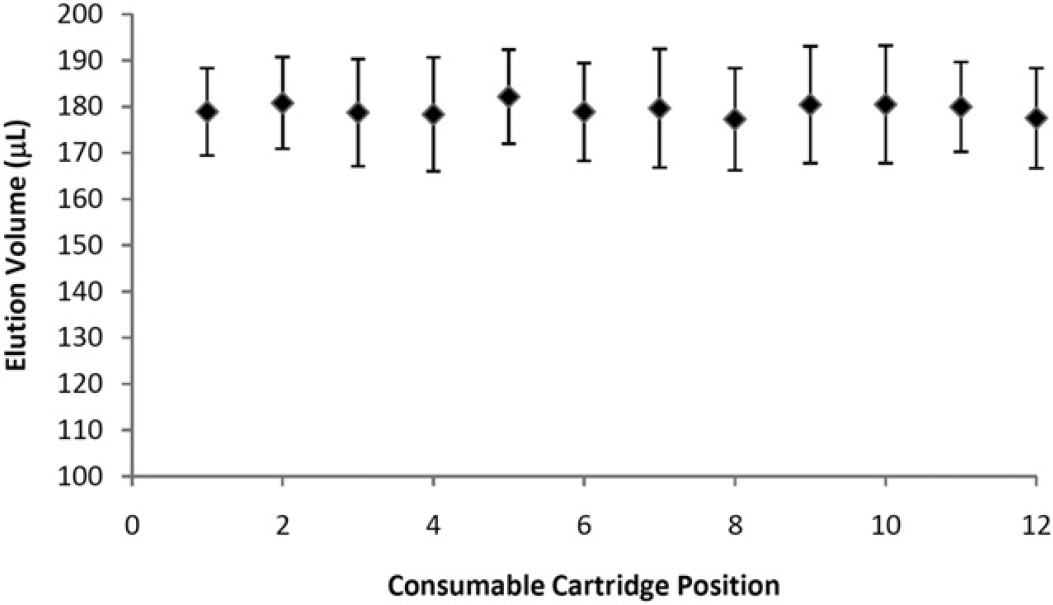
Elution profile: Repeated elutions performed on a fully populated 12 consumable prototype system. A total elution volume of 200μL was delivered to each location, and dispensed in two 100-μL dispenses. Note that the actual volume collected at each location is approximately 20 μL less than dispensed to each location. This is because of backpressure generated at the capture material, which is released between consumables to maintain consistency. The elution manifold is fed by one syringe pump and flows are directed to each consumable via solenoid valves. Only one position is dispensed to at any one time.
Having first individually demonstrated the ability to effectively incubate samples at a variety of temperatures, mix the samples with various reagents, and elute purified nucleic acids in a reproducible volume, we subsequently demonstrated the ability to perform completely automated sample preparations from both whole blood and water. This process can include several steps to perform temperature-controlled enzyme-mediated lysis of the intact organisms and digestion of proteins within the sample matrix and, if required, denaturation of these enzymes at high temperature (Fig. 9). Various temperature-based incubations are used depending on the method selected by the user, which are included as automated scripts on the instrument. The duration of the process is variable depending on the selected method and is dependent on the ability of the target organism (gram-positive bacteria vs RNA virus or free DNA), and the complexity of the sample matrix (i.e., whole blood vs water or cell free media). To demonstrate the flexibility of the automated sample preparation platform, three target analytes were selected for processing, including viral DNA from Epstein Barr virus, rat liver total RNA, and bacterial DNA from
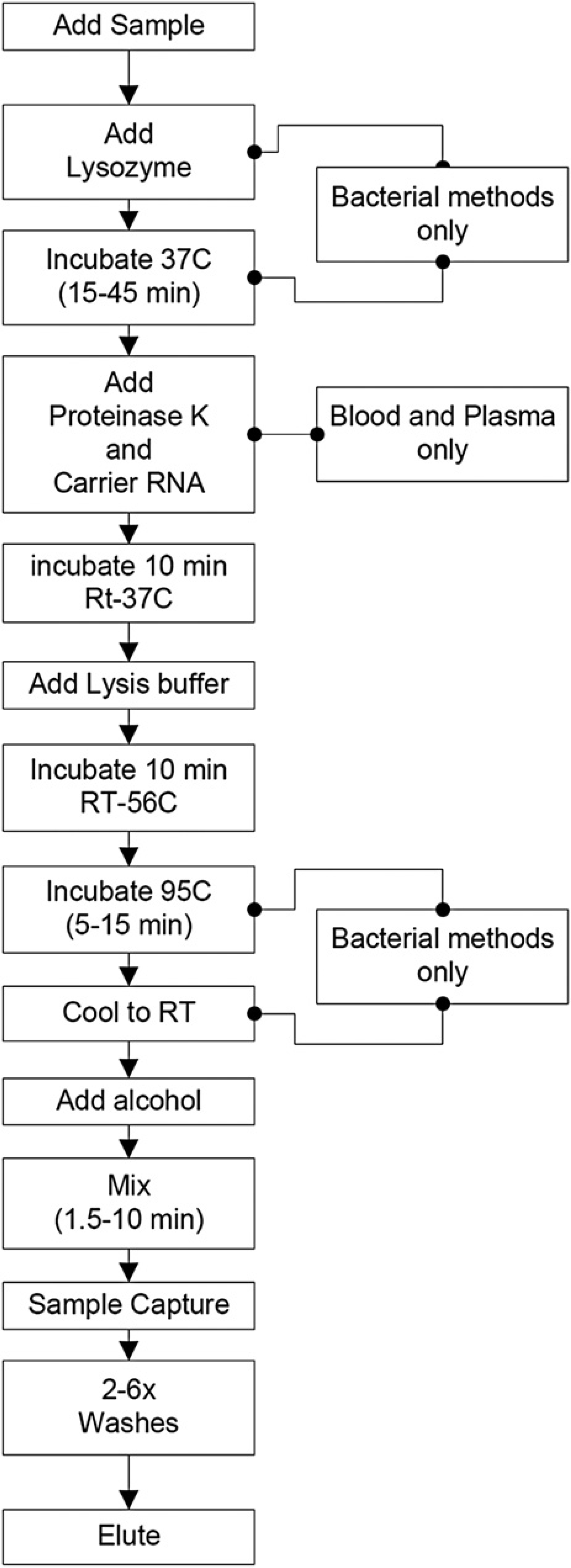
Process flow for automated sample preparation: Automation of the sample preparation process was achieved by breaking each step down into a series of operations and time frames. Several steps of the process are common to all methods, whereas additional steps, such as lysozyme and proteinase K incubations or heating steps are only required for methods that include bacterial lysis and/or whole blood or plasma processing.

Automated nucleic acid extraction using the Xisyl consumable and hardware system. In (A), samples (
In a first set of experiments, 1.0-mL horse whole blood samples were spiked with both EBV DNA (40,000 copies) and rat total RNA (2 μg). These samples were aliquoted into consumable cartridges, and placed on to the instrument for sample preparation, purification, and extraction without any preprocessing. The first step in this protocol was the additional of Proteinase K and carrier nucleic acid. The samples were mixed and incubated for 10min at room temperature. A 2.0 mL volume of lysis buffer was added to each sample, mixed, and incubated for 10min at room temperature. On completion of this incubation, 2.2 mL of alcohol was added to the sample and mixed for a period of 10min. Samples were purged through the capture matrix, in this case a silica fiber matrix to extract the viral DNA and total RNA. For all of the methods described including the extraction of viral DNA, RNA, and bacterial genomic DNA, the samples were cooled to room temperature (approximately 22–26 °C) before the addition of alcohol and remained at this temperature through the sample capture and wash steps. No effort was made to cool the alcohol below room temperature, because of the difficulty in maintaining controlled temperature throughout the fluidic delivery. Once extracted, several wash buffers are dispensed into the cartridge and subsequently purged to purify the sample. Specifically, two volumes (0.25 mL) of wash 1 containing a combination of chaotropic salts and alcohol and four volumes (0.25 mL) of wash 2 containing alcohol were dispensed into the consumable cartridge and subsequently purged from the consumable through the capture material. Once removal of the unwanted contaminants was completed, the capture material was purged with air at an elevated temperature to dry the capture material. The slider was automatically moved into the elution position and heated elution buffer was dispensed on to the capture material allowing sample elution into a microcentrifuge collection vial. The contents of this tube were analyzed using RT PCR on an ABI 7500 Fast RT PCR system. EBV DNA was detected and quantified using the following primers and probe designed to detect the EBER-1 and EBER-2 genes: F: 5′-AAA CCT CAG GAC CTA CGC TGC-3′, R: 5′-AGA CAC CGT CCT CAC CAC-3′ and a dual-labeled probe 5′-/56-FAM/TAG AGG TTT TGC TAG GGA GGA GAC GTG TG/3BHQ_1/-3′. Each reaction contained 10μL of sample, which was added to 15 μL of PCR master mix containing 12.5 μL of 2× Taqman fast universal master mix, 0.5 μL of 13.3 μM EBV Taqman probe, and 1.0 μL each of 10μM EBV Forward and Reverse primers for a final volume of 25 μL for amplification on a 7500 fast RT PCR machine (ABI, Foster City, CA). The protocol included a 20 s activation step at 95 °C followed by 40 cycles of 95 °C for 15 s and 60 °C for 30 s. Standard curves for EBV DNA were generated from dilutions of EBV type 1 quantitated viral DNA (Advanced Biotechnologies, Columbia, MD). Extraction efficiency was calculated by quantifying the genes for EBER-1 and EBER-2 on the EBV DNA standard curves.
Rat total RNA was also analyzed using RT PCR on an ABI 7500 Fast thermocycler for detection of the Gusb transcript and was quantitated using a commercial predesigned primer and probe set (rat-Gusb) from Applied Biosystems (Foster City, CA). Each reaction contained 1.0 μL of sample, which was added to 9.0 μL of PCR master mix containing 5 μL of 2× Taqman fast universal master mix, 0.100 μL 20 U/μL Rnase inhibitor, 0.125 μL of 50 U/μL MultiScribe Reverse transcriptase, 0.5 μL of 20× Gusb primer/probe mixed with FAM, and 3.275 μL nuclease free water for a final volume of 10μL for amplification on a 7500 fast RT PCR system (ABI, Foster City, CA). The protocol included a 30 min reverse transcriptase step at 48 °C, a 20 s activation step at 95 °C followed by 40 cycles of 95 °C for 3 s and 60 °C for 30 s. Standard curves for Gusb RNA were created from dilutions of rat liver total RNA. Extraction efficiency was calculated by quantifying Gusb presence on the total RNA standard curves.
In these experiments, we found that the extraction of EBV DNA and rat total RNA from whole blood was approximately 33% and 55% efficient, respectively. We found from the extraction of EBV DNA, there was a degree of PCR inhibition found for all purified samples regardless of purification method used. Despite this inhibition, the automated process on the prototype system (Fig. 10A) outperformed the manual bench method, which was statistically significant (
In a second set of experiments, bacterial DNA was extracted from tap water and whole blood samples. Bacterial genomic DNA can be extracted from 101–108 cells in either tap water or whole blood using our standard bench methods (data not shown). To enable reproducible calculation of yield, a range of cell inputs, as calculated by absorbance or O.D., were spiked into either 1.0-mL tap water (
Summary
The overarching goal of our efforts has been focused on a systems wide approach to completely automate the process of nucleic acid sample preparation. Through a combined engineering and biochemistry integration effort, we have demonstrated the ability to effectively prepare samples for downstream RT PCR analysis using a prototype system. The data indicate that viral DNA, total RNA, and genomic DNA from gram (−) and gram (+) bacteria can be extracted from complex samples such as whole blood. The system automates the process of enzyme addition, reagent and solvent addition, component mixing, analyte capture and purification, and additionally provides the ability to perform thermo-controlled enzymatic digestion of the proteins present in the sample. From a systems wide approach, using a random access processing mode, the instrument and consumable enables the user to simply load a sample into the consumable device, load the consumable on the instrument, and activate the instrument to prepare nucleic acids for downstream molecular analysis in a completely automated process.
Footnotes
Acknowledgments
The authors would like to acknowledge the contributions of Mr. Christian Miller, and Dr. Lars Majlof in the optimization of the system electronics and firmware. They further acknowledge the experimental assistance from Ms. Pallavi Shah, Ms. Lena Emadi, and Mr. Deshane Campbell. They thank Mr. Tom Stadfelt, Mr. Shalendra “Slim” Maharaj, Fabian van de Graaf, and Michael DeRenzi for their engineering support during the design, fabrication, and testing of the Xisyl Sample preparation system. They would also like to thank Mr. John Roche for his assistance with the design control aspects of the system development. Arcxis Biotechnologies is a small business corporation, which owns the rights to the Xisyl automated sample preparation system.
