Abstract
Increased worldwide trade and processing of seafood has increased the potential for species substitution on the commercial market. To detect and prevent species substitution, several deoxyribonucleic acid (DNA)-based methods have been developed that can be used to identify species in a variety of food types. For large-scale applications, such as regulatory screening, these methods must be rapid, cost-effective, reliable, and have high potential for automation. This review highlights recent technological advances in DNA-based identification methods, with a focus on seafood species identification in automated, high-throughput settings. Advances in DNA isolation methods include silica-based columns for use in high-throughput operations and magnetic bead particles for increased and targeted recovery of DNA. The three most widely used methods for seafood species identification (polymerase chain reaction [PCR] sequencing, PCR-restriction fragment length polymorphism, and species-specific PCR) will be discussed, with a focus on the incorporation of technologies such as rapid PCR cycling, microfluidic chips, and real-time PCR. Emerging methods, including DNA microarrays and next-generation sequencing will also be explored for their potential to identify seafood species on a large scale. Overall, many of the technological advances discussed here offer complementary properties that will enable species identification in a variety of settings and with a range of products.
Introduction
Seafood is an important global protein source that has shown continuous growth in world production and trade over the past 60 years. For example, worldwide imports of edible seafood products increased from 1.0 million metric tons in 1979 to 2.3 million metric tons in 2009. 1 There is also a wide variety of seafood species available on the global market, with 25 major species groups accounted for in worldwide commercial catches 2 and approximately 1700 species of commercial finfish and shellfish named by the U.S. Food and Drug Administration (FDA) in the Seafood list. 3 In their whole, unprocessed form, these species can generally be identified based on morphological indicators; however, most seafood has been processed to some extent before reaching the consumer. For instance, over half of the fresh/frozen finfish imported into the United States is processed from its original form into products such as fillets and steaks, blocks, surimi, and fish sticks. 2
Increased levels of foreign trade and seafood processing, combined with demands for certain seafood types, have increased the potential for seafood fraud. One form of seafood fraud is species substitution, in which seafood is mislabeled for the purpose of economic gain. 4 Seafood fraud can occur at any part of the seafood supply chain, ranging from large-scale multinational importers, with millions of dollars in economic impact, to individual restaurants or retail outlets. 4 In addition to economic consequences, this illegal practice can have detrimental effects in areas such as endangered species conservation, fisheries monitoring, food safety, and consumer confidence in the food supply. Some examples of seafood fraud that have been reported include farmed salmon mislabeled as wild salmon, skate wings mislabeled as scallops, Alaska pollock mislabeled as cod, and several types of seafood (e.g., rockfish, tilapia, or mahi mahi) mislabeled as red snapper. 5 Although the extent of species substitution is unclear, a 9-year survey conducted by the National Seafood Inspection Laboratory (National Marine Fisheries Service) reported that 37% of fish and 13% of other seafood products collected from randomly selected vendors was mislabeled. 6 More recently, market surveys in Europe and North America have reported seafood species substitution at levels of 15-43%,7–12 with an exceptionally high level of 75% for red snapper products. 13
To detect and prevent species substitution on the commercial market, a number of methods have been developed based on the unique protein or deoxyribonucleic acid (DNA) profiles found in different species. Protein-based methods are generally reliable for testing fresh or lightly processed seafoods, but become impractical in heavily processed foods, where proteins are degraded. 14 Although some studies have reported the use of enzyme-linked immunosorbent assays with heat-treated seafood products,15,16 this method does not work well with closely related species and requires the development of species-specific antibodies. In contrast, DNA-based methods have numerous advantages over protein-based methods, including a higher information content, greater resistance to degradation, increased specificity and sensitivity, and presence in all cell types. 14
There are three main steps in DNA-based identification: DNA isolation, polymerase chain reaction (PCR) amplification, and detection of species based on unique DNA profiles. This review will primarily discuss the PCR amplification and species identification steps, but some advances in DNA isolation techniques will also be highlighted. Some important characteristics to consider when selecting a DNA-based method for use in seafood species identification include sample processing time and costs, equipment and startup costs, reproducibility, reliability, range of target species, and ability to recover and identify DNA from processed products, complex food matrices, and mixed-species samples. Further, for large-scale screening purposes, methods that can be applied in automated and high-throughput settings are desirable. Numerous DNA-based detection methods have been used for seafood species identification, including PCR sequencing, PCR—restriction fragment length polymorphism (RFLP), species-specific PCR, random amplified polymorphic DNA (RAPD), and single-stranded conformational polymorphism (SSCP).14,17 Each of these methods has some advantages and disadvantages, as outlined in Rasmussen and Morrissey. 14 For example, PCR sequencing, PCR—RFLP, and species-specific PCR all exhibit high reproducibility. However, traditional PCR sequencing can be costly and time-consuming and cannot be used to identify multiple species in one sample. PCR—RFLP and species-specific PCR are less costly and can generally be used with mixed-species samples, but they are vulnerable to errors from intraspecies variation and they do not provide the high level of information acquired with PCR sequencing. In addition, RAPD and SSCP do not require prior knowledge of DNA sequence information, but they exhibit reduced reproducibility compared with other methods, and RAPD is vulnerable to DNA degradation.
Although all of the methods mentioned above have been applied to seafood species identification, the current review will be limited to discussing recent advances made for only the three most widely used methods (i.e., PCR sequencing, PCR—RFLP, and species-specific PCR). Emerging methods that show potential to improve automation and high-throughput aspects of seafood species identification, such as real-time PCR, microarray technology, and next-generation sequencing (NGS), will also be highlighted. To focus on recent advances in these techniques, most studies published before 2008 will not be discussed.
Advances in Current Methods
DNA Isolation
The first step in DNA-based species identification is the isolation of genomic DNA. The isolation of DNA from food matrices for use in PCR is complicated by the fact that many ingredients in food may act as PCR inhibitors. DNA quality can also be reduced by many of the conditions common to food processing, such as low pH, high temperatures, high pressure, and hydrolysis. 17 Heat-sterilized products, such as canned tuna, have been especially challenging in species identification, with maximum recoverable DNA fragment lengths of about 250—350 bp.18–20 Therefore, isolation methods with high levels of DNA recovery and purity are essential for successful PCR amplification and species identification with seafood products. This section will focus on advances in DNA isolation methods for use in high-throughput settings.
Two DNA isolation technologies that have been used in automated, high-throughput operations are silica-based DNA-binding columns 21 and magnetic particles/beads. 22 Ivanova et al. 21 described an automated DNA isolation protocol that is used in a high-throughput laboratory processing around 100,000 samples per year. This method is based on the binding of DNA to silica columns in the presence of a high concentration of chaotropic salts, and has largely replaced the more time-consuming, traditional method for DNA isolation based on phenol—chloroform extraction.21,23 Although silica-based protocols are commercially available in several kits, Ivanova et al. 21 described a procedure developed in-house that allows for significant cost-savings (U.S. $0.50/sample) when compared with commercial kits (U.S. $2.00/sample). The method may be carried out manually or automated with a robotic system. This method has been used extensively in large-scale species identification studies involving DNA sequencing of a wide range of fish species,24-26 including processed seafood products. 12
Magnetic particle technology is based on the ability to separate para- or ferromagnetic particles from chemical or biological media in the presence of a magnetic field.27,28 Spherical magnetic particles (beads) are generally prepared using a particle that is susceptible to magnetism, such as iron oxide, coated with a biological or synthetic polymer, such as agarose, silica, or polyvinyl alcohol. Selective capture of nucleic acids on the magnetic bead surfaces can be facilitated with the use of binding solutions, such as DNA-binding proteins or complementary DNA sequences. Because of the binding of nucleic acids to the surface of the magnetic beads, there is no need for the centrifugation steps used in other DNA isolation procedures, which can cause shearing and reduce DNA quality. Magnetic bead technology has been reported to improve success of DNA isolation from processed food samples, and has been used with products such as canned tuna, 29 dried and cooked bonito,30,31 and processed milkfish products. 32 A high-throughput laboratory that processes over 800 samples per week for large-scale larval fish species identification has automated their DNA isolation procedure using magnetic bead technologies combined with a liquid-handling robot. 22 The ability to use complementary DNA sequences on the surface of the magnetic beads has the potential to greatly improve DNA isolation efficiency. For example, Chang et al. 33 described the use of magnetic beads containing a DNA fragment specific to the mitochondrial D-loop to selectively isolate mitochondrial DNA (mtDNA). These beads were combined with microfluidic chip technology to develop a highly automated, rapid system that can be used to isolate mtDNA with high efficiency. A large portion of seafood species identification methods are based on mtDNA, and this method could be especially useful in seafood products with low-quality DNA.
PCR Sequencing
Methods that use PCR sequencing and nucleotide analysis for the differentiation of species are generally referred to as forensically informative nucleotide sequencing (FINS) 34 or DNA barcoding. 35 These are widely used procedures that involve PCR amplification and sequencing of a specific DNA fragment, followed by analysis of nucleotide variation between the target sequence and reference sequences of known species. Nucleotide variation is assessed through the calculation of genetic distances using a model such as the Kimura 2-parameter distance method, 36 and when a match is found between the target sequence and a reference sequence, species identification is possible. To ensure correct species identification, PCR-sequencing methods require a comprehensive database of reference sequences and the selection of a target DNA fragment that demonstrates high variation between species and low variation within species. The mitochondrial protein-coding genes cytochrome b (cyt b) and cytochrome c oxidase subunit I (COI) have been found to be highly suited for this purpose and have been extensively used for seafood species identification.37,38 Methods based on FINS use a variety of genetic targets, including cyt b, COI, and 16S rRNA. Recent FINS studies focused on seafood species identification have targeted small pelagic fishes, 39 snapper, rockfish and tilapia, 40 scombroids,10,41 anglerfish, 7 flatfish, 8 bivalves, 9 and salmonids. 11
The Barcode of Life (BOL) initiative is a global effort to standardize species identification using PCR sequencing and analysis of short standardized DNA sequences called “DNA barcodes” (http://www.barcoding.si.edu/). 35 For most species groups, the standard DNA barcode is a ∼ 650 bp region of the COI gene, which is relatively conserved within species, but shows sufficient variation between species to allow for differentiation. 42 An overarching goal of the BOL initiative is the development of a comprehensive reference sequence database containing DNA barcode records for all eukaryotic species, 43 and the portion of the project focused on fish is called FISH-BOL (www.fishbol.org). 38 FISH-BOL currently has DNA barcode records in place for close to 8000 species and numerous studies have been published that show high success of DNA barcoding for species identification, with 93% of freshwater species and 98% of marine species tested so far showing unambiguous species differentiation. 38 The U.S. FDA is also planning to incorporate DNA barcodes into the Regulatory Fish Encyclopedia to help detect fish species substitution on the commercial market. 26 Recent publications reporting the usefulness of DNA barcoding in species identification have targeted a variety of commercial species from the Amazon river, 44 shark and ray fins confiscated from a fishing vessel in Australia, 45 and numerous fish species in North and Central America.12, 24-26,46, 47
PCR-sequencing methods generate the highest information content of the DNA-based species identification methods and also allow for a broad range of species to be detected based on one DNA fragment; however, there are several drawbacks to these methods. For example, although technological advances have reduced both time and price of PCR sequencing, it continues to be more time-consuming and costly than alternative species identification methods, such as species-specific PCR. Furthermore, most PCR-sequencing methods use DNA fragments that are too long to be recovered from heavily processed food products, and mixed-species samples cannot be identified with sequencing unless a time-consuming and expensive cloning step is added. 17 However, these limitations are being addressed through the use of minibarcodes (discussed below)48,49 and NGS technologies (discussed in a later section). 50
Advances in DNA Barcoding.
DNA barcoding protocols must be heavily automated and high throughput to process specimens in the large volumes required by the BOL initiative. 23 In addition to the silica-based, automated DNA isolation method described above, 21 the core DNA barcoding facility uses precast agarose gels, capillary sequencing, and a well-organized data system that includes a laboratory information management system, a data management and analysis system, and a species identification engine.23,43 Despite these advances, DNA barcoding typically involves a workflow that extends over 2 days. To address this limitation, Ivanova et al. 51 recently described a procedure termed “express barcoding” that could be completed in less than 2 h. Express barcoding involves the use of several time-reducing measures, including Whatman FTA (Fast Technology for Analysis of nucleic acids; Maistone, Kent, United Kingdom) cards for DNA isolation and fast-cycling PCR and sequencing kits. This sequencing workflow was said to be faster, less costly, and more suitable for small-scale laboratories as compared with a procedure described by Applied Biosystems (Life Technologies, Carlsbad, CA), 52 which takes 4.5 h to complete. The Applied Biosystems protocol uses magnetic bead technology and calls for additional steps, such as PCR cleanup, that are not used in the express barcoding procedure. However, the express barcoding protocol does not take into account the time involved for application of samples to FTA cards, which requires the sample to dry completely before further processing. Although FTA cards are commonly used to isolate DNA from blood samples, an alternative method may be more appropriate for isolation of DNA from commercial seafood products, which often have complex sample matrices.
To overcome the limitations of DNA barcoding associated with species identification of degraded samples, the use of minibarcodes has been proposed.48,49 Minibarcodes are shorter fragments of the full-length DNA barcode and have high potential for use with samples that contain low-quality DNA, such as canned seafood, from which the full-length barcode cannot be recovered. Several minibarcode regions have been identified that allow for differentiation of a range of species48,49 and these regions have been tested in silico to differentiate commercially important salmon and trout species. 25 Minibarcoding has also been explored using advanced sequencing techniques offered by NGS, 53 which will be discussed in a subsequent section. A few studies using minibarcoding for species identification have recently appeared in the literature, targeting snakes 54 and fruitflies, 55 and this method shows high potential for use in species identification of processed seafood items.
Character-Based Systems.
Character-based analysis of DNA sequences has been proposed as an alternative to the genetic distance-based analysis traditionally used for species identification after PCR sequencing. 56 Character-based analysis relies on the identification of variable nucleotide sites that are diagnostic at the species level. These nucleotide sites are commonly referred to as either nucleotide diagnostics 57 or character attributes. 56 After PCR sequencing, nucleotides at diagnostic sites are compared among the target and reference samples to identify species. This method of sequence analysis offers complementary aspects to the distance-based approach in circumstances that require greater specificity, such as the use of PCR sequencing in a regulatory or legal setting, or differentiation of closely related species that only vary at a few nucleotides. 57 Character-based approaches to analyzing DNA barcode sequences have been reported to be successful in seafood species identification studies with sharks, 57 Cuban freshwater fish, 58 and tuna species. 59 Character-based analysis has also been applied in a high-throughput setting with a 715 bp fragment of cyt b to identify larval fish species. 22 The authors developed a character-based bioinformatics program to handle the large amount of sequence data generated, and reported an 89% success rate for this method when testing 493 samples of tuna and billfish larvae.
Although user-friendly software for character-based analysis is not currently available, Sarkar et al. 56 have reported the development of a downloadable software tool (http://www.uvm.edu/∼insarkar/CAOS/) that can identify the diagnostic nucleotide sites in a data set and can then read DNA sequences from query specimens and identify species based on variation at the predetermined sites. The current version of the program is a command-line application, but the development of a graphical online interface and visualization module is underway. Programs that readily identify diagnostic nucleotide sites can also be applied to the design of species-specific PCR assays and microarrays. 57 Although character-based approaches have demonstrated high potential for use in species identification, it is important to keep in mind that they are limited to the species pool originally used to identify diagnostic nucleotide sites and that as the species pool is widened, the diagnostic nucleotide arrangements will become increasingly complex.
PCR-RFLP
PCR—RFLP is one of the most widely used methods for species identification.17,37 This method involves the amplification of a preselected DNA fragment with universal primers, followed by digestion with restriction endonucleases, which recognize specific short sequences (four to six nucleotides) of the amplified fragment and cut the DNA at those sites. These fragments can then be separated and visualized with gel electrophoresis. The development of a PCR-RFLP method requires sequence information for the DNA fragment of interest to select appropriate restriction endonucleases that produce species-specific DNA profiles after an enzymatic digestion. The digested DNA fragments must exhibit sufficient size variation to be differentiated by electrophoresis. When designing a PCR-RFLP assay, it is important to examine sequences from multiple individuals covering a wide geographic range to account for intraspecies variability and to examine sequences from background species to avoid false positives. PCR—RFLP can be applied to processed products if a small fragment is targeted and in some cases to mixed-species samples, although the resulting DNA band patterns can be difficult to interpret. This method can be relatively time-consuming because it involves multiple post-PCR steps, including a restriction digest that is sometimes carried out overnight.60,61 Although PCR—RFLP is not considered to be a highly automated or high-throughput method, it is widely used because of its low cost, simple protocols, and limited equipment requirements. 14 Some examples of seafood species products tested in recent years with PCR—RFLP are anchovy pastes sold in Europe, 61 milkfish and carp fish balls sold in Taiwan, 32 processed puffer fish sold in Taiwan, 62 salmon and trout products sold in Europe 11 and in the United States, 63 flying fish in processed foods in Japan, 64 anglerfish sold in Europe, 7 and razor clams sold in Europe. 65
Advances in PCR—RFLP.
One of the most widely used PCR—RFLP methods for seafood species identification is based on variation within a 464 bp fragment of the cyt b gene. 37 This method has enabled identification of numerous fish species over the past decade.60,63,66-72 Although the analysis traditionally called for an overnight restriction digest, recent work has demonstrated the possibility of completing the digest in just 1 h, allowing for species identification within 8 h. 63 Furthermore, the use of commercially available precast agarose gels to analyze DNA fragments could reduce the time required for gel electrophoresis to less than 10 min. Alternatively, lab-on-a-chip technology can be used in place of gel electrophoresis to identify DNA fragment size after PCR—RFLP.73,74 This method uses microfluidic, capillary electrophoresis to separate fragments on a microchip (3 cm2) and then fragments are detected and quantified using laser-induced fluorescence. PCR—RFLP combined with lab-on-a-chip technology has been successfully carried out with numerous species, including white fish 73 and salmon, 74 and has exhibited greater sensitivity and speed compared with traditional gel-based methods. This technology has also increased the automation and high-throughput potential of PCR—RFLP because of its increased speed, ease of use, and automatic detection and quantification of DNA fragments. In April 2010, Agilent Technologies (Santa Clara, CA) announced the release of a fish species identification kit (www.agilent.com/chem/fishID) that uses PCR—RFLP and lab-on-a-chip technology combined with an RFLP pattern matching program that runs automated algorithms to identify over 50 species. According to the product brochure, the entire workflow can be completed in less than 8 h and the kit can be used with fresh/frozen fish and some types of processed fish (e.g., dried, salted, cooked, and minced samples).
Species-Specific PCR
Species-specific PCR has been used for many years to identify a wide range of seafood species.37,75-77 To develop primers for species-specific PCR, the DNA sequence of the target species must be examined for diagnostic nucleotide sites that enable differentiation from background species. Species-specific primers are then designed to selectively bind and amplify the target DNA at these diagnostic sites. Several species-specific primers can be combined into a single tube to amplify multiple targets in a multiplex PCR assay, which reduces time, materials, and cost. In conventional endpoint PCR, each of the species-specific primers is designed to amplify a DNA fragment with a diagnostic size, and species can be identified with gel electrophoresis immediately after PCR. However, when species-specific primers are applied to real-time PCR (discussed below), species can be identified based on fluorescent signals that are detected as the reaction progresses, and there is no need for a post-PCR electrophoresis step. As with PCR—RFLP, design of species-specific PCR assays requires detailed sequence information for the target and background species, including multiple individuals from a wide geographic range. Reference sequence databases, such as those established by the BOL initiative (http://www.boldsystems.org) and the FishTrace Consortium (www.fishtrace.org), have provided excellent resources for the design of such assays. For example, recent studies have successfully developed species-specific multiplex PCR assays based on DNA barcodes for sharks,78,79 guitarfish, 80 and salmon and trout. 81
Species-specific multiplex PCR is advantageous and widely used for many of the same reasons that PCR—RFLP is used: it is a simple procedure, low cost, and has minimum equipment requirements. The advantages of species-specific multiplex PCR over PCR—RFLP are speed (no restriction digest step), reduced materials, and straightforward detection of species in mixed samples. However, although the same primers and restriction enzymes can be used to differentiate a wide range of species with PCR—RFLP, species identification with multiplex PCR requires the development of diagnostic primers for each target species and is more limited in scope. Numerous studies have been published in the past few years using species-specific PCR primers to identify seafood species, including dolphinfish, 82 sharks,83–85 flatfish, 86 bonitos,30,31 sturgeon, 87 oysters, 88 small pelagic fish, 89 mackerels, 90 razor clams, 91 and many more.92–96
The introduction of commercially available precast agarose gels and fast PCR-cycling technology97,98 into the above techniques could further reduce the time required for species identification with multiplex PCR to about 1—2 h after DNA isolation. Use of capillary electrophoresis with lab-on-a-chip technology, as described above with PCR—RFLP, could also improve speed and detection of multiplex PCR fragments. Another application of lab-on-a-chip technology in species identification is miniaturized, microfluidic PCR chips, which can greatly reduce PCR amplification volumes (≤3 μL) and time (<5min).99,100 A PCR chip system was recently used for multiplex PCR amplification and species detection of pathogens, 101 but this technology has not been applied to seafood species identification. Although PCR chips have high potential to benefit a wide range of PCR protocols and applications in high-throughput settings, development is still in the early stages and advances must be made to improve the ease of use and automation capability. 100
Real-Time PCR.
Another advantage of species-specific PCR is that it can be carried out in real time using sequence-specific fluorescent probes (e.g., TaqMan)102,103 or nonspecific fluorescent dyes (e.g., SYBR Green).104–106 Realtime PCR allows for the detection and quantification of target DNA fragments as the reaction progresses, eliminating the need for post-PCR electrophoresis, and greatly reducing the time required for species identification. 14 Real-time PCR is generally more sensitive and specific than conventional PCR and can readily be applied to high-throughput operations because of its minimal laboratory preparation requirements, lack of post-PCR processing steps, reduced chance of cross-contamination, and computer-generated results. Furthermore, fast PCR-cycling technology is now commercially available for real-time assays and has been used in pathogen species identification. 107 PCR chips with real-time capabilities have also been developed, 108 but have experienced limited application in species identification.
The most common type of detection method used in realtime PCR for the identification of seafood species has been TaqMan probes (Fig. 1). These probes are designed to bind at diagnostic sites found within the target DNA fragment and they release a fluorescent signal during amplification by Taq polymerase. TaqMan assays can generally include up to four different species-specific probes combined into one multiplex PCR tube and can be used for either qualitative or quantitative species identification. However, quantitative assays are difficult to standardize with food products because of diverse sample matrices and DNA quality, especially in the case of canned seafood.
102
Furthermore, quantification of mtDNA must be carried out relative to a reference gene, as mtDNA copy number varies depending on cell type and species.
109
Advances in TaqMan probe technology have allowed for increased specificity for diagnostic nucleotide sites through the incorporation of minor groove binder (MGB) moieties
110
or locked nucleic acid (LNA) bases.
111
Both MGB and LNA technologies increase the probe melting temperature and allow for the design of shorter TaqMan probes with greater specificity. MGB and LNA probes are linked to different dye sets and the choice of technology is largely dependent on the compatibility and multiplexing capabilities of these dyes in relationship to a particular realtime PCR instrument. TaqMan MGB probes have been used for identification of seafood species such as flatfish,
112
eel,
113
tuna,
109
and salmon and trout,
81
and TaqMan LNA probes have been used for identification of snappers and drum.
114
Real-time polymerase chain reaction (PCR) using TaqMan probes. The probe contains a reporter (R) fluorophore and a quencher (Q) fluorophore. When the probe is intact, the quencher fluorophore prevents the reporter fluorophore from emitting fluorescence. Probes are designed to hybridize with a complementary sequence on the target deoxyribonucleic acid (DNA) fragment. Following DNA denaturation, the TaqMan probe hybridizes to the target DNA and then during primer extension, Taq polymerase separates the reporter fluorophore from the quencher. The result is emission of a specific fluorescent signal that can be detected and quantified. Reprinted with permission from John Wiley and Sons: Comprehensive Reviews in Food Science and Food Safety,
37
copyright 2008 Institute of Food Technologists.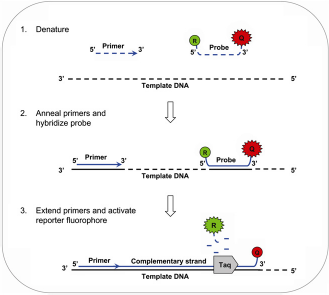
Emerging Methods
DNA Microarrays
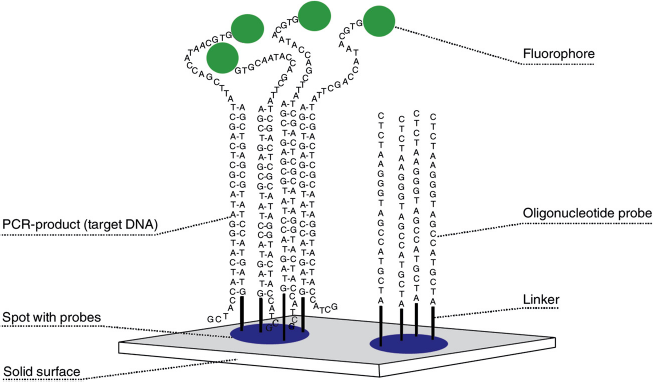
An emerging DNA-based method that is well suited for automated, high-throughput operations is DNA microarray technology (Fig. 2), which enables the screening of a large number of samples simultaneously. DNA microarrays, also known as DNA chips, contain between ten and tens of thousands of different oligonucleotide probes that are immobilized on the surface of a glass slide or microscopic beads.115,116 To identify species with a DNA microarray, the target DNA fragment is labeled with a fluorophore during PCR amplification with universal primers, and then the labeled amplicon is applied to a microarray, where it will hybridize with any probes that exhibit a complementary nucleotide sequence.
117
Following a series of washing steps, any amplicons that have hybridized to the probes can be detected and quantified by their fluorescent labels. Even a DNA microarray containing as few as 25 probes may prove useful as a high-throughput diagnostic tool, because of the low sample volume requirements, ability to analyze mixed-species samples and sensitivity of the assay.
116
The development of a DNA microarray requires extensive sequence information to design probes with high specificity, and all probes must be empirically tested to avoid false positive or negative identifications.
118
Although DNA microarrays have already reached widespread use as a high-throughput technology in areas such as gene expression analysis and single nucleotide polymorphism (SNP) genotyping, they have only recently been applied to seafood species identification.
118
Example of a deoxyribonucleic acid (DNA) microarray. The target DNA is labeled with a fluorophore during polymerase chain reaction (PCR) amplification. The labeled amplicons are then exposed to oligonucleotide probes that are immobilized on a solid surface. Following hybridization between probes and target DNA, the levels of fluorophore are detected with a fluorescence scanner and used to identify and quantify the level of target DNA at each spot. DNA microarrays can contain between ten and tens of thousands of different oligonucleotide probes. Reprinted with kind permission from Springer Science+Business Media: The Future of Fisheries Science, Trends in Fishery Genetics, 2009, pp. 453-493, Kochzius, M., Fig. 24.4.
118
In 2004, the first microarray for food and animal feed testing was launched in France by the biological diagnostics company bioMerieux (Marcy l'Etoile, France; www.biomerieux.com). This product was called the FoodExpert-ID system and used a DNA chip produced by Affymetrix GeneChip (Santa Clara, CA) technology containing 80,000 oligonucleotide probes based on variation in the cyt b gene. The FoodExpert-ID system was reported to enable simultaneous identification of over 30 vertebrate species, including 15 species of fish, 5 bird species, and 12 mammals, in both raw and processed products. Since its launch, the FoodExpert-ID system has been tested and compared with real-time PCR for its ability to screen animal feeds and meat mixtures. 119 The microarray was reported to show good performance with many of the samples and the results generally agreed with real-time PCR. However, the microarray did exhibit some cases of reduced specificity (i.e., false positive results) and sensitivity, and the relative amount of each species in an admixture could not be determined. Chisholm et al. 119 reported that the analysis required about 8 h to complete and could be useful as a general screen to determine the type of species present in a food sample.
Two additional microarrays have been developed that enable identification of fish species.117,120 Kochzius et al. 117 developed a prototype DNA microarray targeting 11 commercially important European fish species. They designed oligonucleotide probes based on variation within the mitochondrial 16S rDNA gene. The authors reported promising results and are using this microarray design as the basis for creation of the “Fish Chip,” which will enable identification of 50 species that are important in fisheries research and monitoring. Teletchea et al. 120 designed probes based on variation in the cyt b gene to identify 71 commercial and/or endangered vertebrate species in food and forensic samples, including 26 ray-finned fishes and 2 sharks. They used an additional primer set specific for smaller targets to enable identification in heavily processed foods. Ten food samples were tested with this method and species were identified in products such as smoked salmon and canned mackerel, with a sensitivity of about 10% in fresh samples. According to the authors, the microarray method was easy to use, accurate, and could be completed within 2 days. Teletchea et al. 120 plan to incorporate COI probes into future microarrays as a complement to cyt b. The BOL database will be a valuable tool for the development of COI microarrays. Algorithms for the design of COI oligonucleotides that could be used in DNA array technology have been developed 121 and used for a high-density membrane-based COI oligonucleotide array to detect Penicillium species. 122 Although over 60% of oligonucleotides designed in silico exhibited low specificity during testing, 76 probes designed with the algorithms provided useful group-specific or species-specific information.
Despite the high-throughput advantages offered by DNA microarrays, there are several limitations to this method that have prevented their widespread use in species identification up to now. The development of microarrays is time-consuming, requiring the design and empirical testing of all probes, and sample analysis is relatively slow and costly compared with other DNA-based methods for species identification. 17 However, DNA microarrays have proven to be effective for processing samples on a large scale and they will likely experience increased use in future efforts to monitor seafood species substitution in world trade, especially if the technology becomes less expensive, as predicted. 123 There have also been some interesting advances that might improve the potential for microarrays in this field, such as the combining of PCR amplification with microarray technology.124–126 For example, one study reported the use of immobilized primers in a multiplex PCR amplification carried out on a microarray platform, with results detected in real time using SYBR Green. 126 This method was described for the identification and quantification of pathogenic viruses and was said to be one of the most promising approaches to pathogen detection. 126
NGS Technologies
NGS methods use advanced techniques to generate large volumes of sequence data simultaneously and relatively inexpensively. 127 Compared with traditional, “first-generation” DNA sequencing based on the dideoxy chain termination technique, 128 NGS relies on immobilization of fragmented DNA templates on a solid support system. 127 The spatially separated, immobilized fragments can then be amplified simultaneously by PCR and subjected to massively parallel DNA sequencing. There are several platforms available for NGS, including the Roche/454 Life Sciences (Indianapolis, IN), Illumina/Solexa Genome Analyzer (San Diego, CA), and the Applied Biosystems/SOLiD System, each with its own unique enzyme system, sequencing chemistry, hardware, and software engineering.127,129,130 Sequencing reads obtained with NGS technologies and the total sequencing output are different from one platform to another. The Roche/ 454 platform can generate from several hundred thousand to 1 million reads of 400—500 bp DNA fragments per run (www.454.com). In addition, Illumina/Solexa and Applied Biosystems/SOLiD platforms can generate tens of millions of short reads (25—35 bp) per run. The short read length associated with some NGS technologies is the major limitation of these platforms, especially with regard to sequence library construction. For this reason, traditional first-generation sequencing remains the preferred technology when longer sequence reads (700 bp) are desired. However, NGS technologies have already reached widespread use in DNA sequencing research, with over 700 scientific publications using Roche/454 sequencing since its release in 2005 (www.454.com). Some of the applications of NGS include metagenomics, gene expression, and ancient genome sequencing. 129 NGS has also proven useful for the discovery of SNPs that can be used to develop microarrays. 131 In a similar manner, NGS has the potential to be used for identification of diagnostic nucleotide sites that can be targeted in species-specific PCR assays or for species identification using direct sequencing of diagnostic fragments.
Pyrosequencing.
Among the available NGS platforms, Roche/ 454 sequencing offers the greatest potential for use in species identification because of its speed and capacity to generate longer read lengths.
127
Roche/454 uses pyrosequencing technology (sequencing by synthesis, Fig. 3), which is based on the real-time detection of pyrophosphate (PPi) molecules released during the incorporation of nucleotides by DNA polymerase.
132
After it is released, PPi initiates a series of enzymatic reactions that leads to the production of light by the firefly enzyme luciferase. Pyrosequencing was commercialized by the Biosystems unit of Biotage AB (Uppsala, Sweden; acquired by Qiagen [Hilden, Germany] in 2008; www.pyrosequencing.com) and the technology was later licensed to Roche/454 for high-throughput use with NGS. The pyrosequencing instruments now offered through Qiagen are available in automated, low-to-medium throughput settings and generate shorter read lengths (50—60 bp) than the Roche/454. Previous versions of these systems (Biotage PSQ 96 series) were used for species identification in several studies, with demonstrated applications in pathogen identification133-135 and identification of animals for forensics purposes.136,137 More recently, the Qiagen PyroMark series of instruments has become available for pyrosequencing of samples in 24- to 96-well formats, with specific applications in microbial species identification and quantification (www.pyrosequencing.com).
Pyrosequencing using a Roche/454 platform: (A) the deoxyribonucleic acid (DNA) template is immobilized using emulsion polymerase chain reaction (emPCR). The reaction mixture is encapsulated into single aqueous droplets using an oil—aqueous emulsion. PCR amplification takes place within these droplets to create beads containing thousands of copies of the same template sequence. (B) DNA-amplified template beads are deposited into PicoTiterPlate (PTP) wells and additional beads coupled with sulphurylase and luciferase are added. The fiber-optic slide is mounted in a flow chamber, enabling the delivery of sequencing reagents to the bead-packed wells. In this example, the 2 -deoxyribonucleoside triphosphate (dNTP) cytosine (C) is shown flowing across the PTP wells. The incorporation of a dNTP by DNA polymerase causes the release of inorganic pyrophosphate (PPi), which initiates a cascade of reactions involving the conversion of adenosine-5 -phosphosulfate (APS) to adenosine triphosphate (ATP) and the production of light by luciferase. (C) The light generated by the enzymatic cascade is recorded as a series of peaks called a flowgram. Adapted by permission from Macmillan Publishers Ltd: Nature Reviews Genetics,
127
copyright 2010.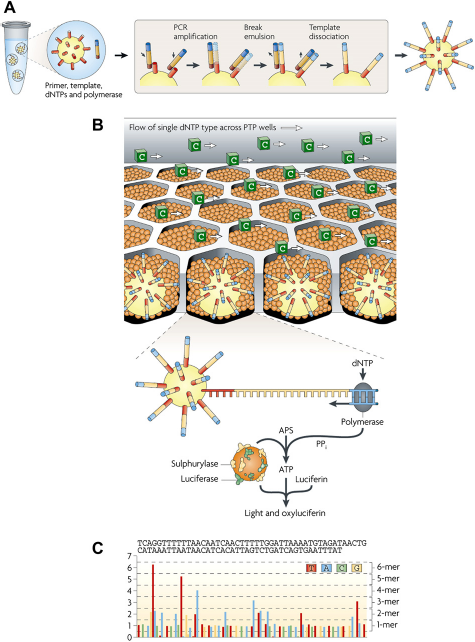
Pyrosequencing is advantageous in its flexibility, accuracy, and automation potential. 136 Furthermore, recent advances in this technology have allowed for simultaneous analysis of DNA from multiple individuals through the use of tagged primers.138,139 For example, Binladen et al. 138 demonstrated the ability to simultaneously sequence fragments of 16S rDNA from 13 different mammal species using a 454 pyrosequencing platform with tagged PCR primers. Despite these advances, pyrosequencing has not yet been applied to seafood species identification and it continues to have a few limitations. For example, stretches of homopolymeric DNA can be challenging to sequence with this method, because the intensity of the chemiluminescent signal does not allow accurate detection of a stretch of more than three identical nucleotides. 140 However, this is generally only a problem if there is no reference sequence to which the pyrosequenced fragments can be aligned.
Pyrosequencing technology could readily be applied to seafood species identification, especially when using a comprehensive sequence database, such as BOLD or FishTrace, combined with short sequences known to exhibit diagnostic variation, such as minibarcodes.48,49 This type of species identification might be carried out in low-to-medium throughput settings in a similar manner as microbial species identification with the PyroMark system or on a larger scale with Roche/ 454 pyrosequencing. Indeed, Roche/454 pyrosequencing of minibarcodes was recently demonstrated to be effective for accurate identification of Lepidoptera (butterflies and moths) species and the protocol was reported to be simple, rapid, and less costly per sequence, considering the deep sequencing coverage, when compared with traditional sequencing. 53 Although traditional sequencing remains the method of choice for obtaining full-length DNA barcodes, Roche/454 pyrose-quencing may prove complementary in the case of degraded 53 or mixed-species samples. 50 Overall, pyrosequencing shows high potential for use in automated, high-throughput identification of seafood species in different product types and for the development of sequence information necessary for the design of species-specific PCR assays or microarrays.
Conclusions
As demonstrated in this review, DNA-based techniques for species identification are rapidly evolving. The three most commonly used methods, PCR sequencing, PCR—RFLP, and species-specific PCR, are all experiencing ongoing technological advances leading to increased speed and automation potential. Microfluidic chip systems have enabled the miniaturization of procedures such as PCR and electrophoresis, resulting in lower sample volumes, increased speed, and increased capacity to accurately identify and quantify species. Emerging techniques for species identification, such as micro-arrays and NGS technologies have provided extremely high-throughput, automated platforms for DNA analysis that could be applied to regulatory screening of seafood products or large-scale sequencing operations. Many of the technological advances discussed here have already been applied and proven to be successful in pathogen species identification, and could readily be modified for use in the identification of seafood species. Many of these methods exhibit complementary properties that, when combined, allow for detection in a range of settings and with a variety of sample types. For example, PCR sequencing and NGS technologies not only enable high-throughput species identification, but also provide important sequence information for the design of species-specific PCR assays and microarrays. Ultimately, the choice of method(s) is dependent on a combination of factors, including the desired scale of automation, speed, and throughput; the range and degree of processing of target species; and the required level of information content and specificity. Overall, the technological advances described here represent important contributions to the field of DNA-based species identification and demonstrate high potential for high-throughput, automated detection of seafood species on the commercial market.
Footnotes
Competing Interests Statement: The authors certify that they have no relevant financial interests in this article.
Acknowledgments
The authors would like to thank the Association for Laboratory Automation (ALA) for the opportunity to submit this article as part of the 2010 ALA Young Scientist Poster Award. Thanks also to Shadi Shokralla, Ph.D., for edits on the NGS Technologies section. This work was supported by the Oregon Innovation Council through the Oregon Economic Development Department.
