Abstract
An electrowetting-on-dielectric (EWOD) microfluidic system of coplanar electrode type was designed for parallel DNA ligation of an ultra-micro volume, which was applied to droplet manipulation containing DNA, a ligation enzyme and a multi-salt reaction buffer. For environmental reasons, we applied a concept of a water–oil core–shell droplet; such droplets of volume ∼0.3 μL were generated from reservoirs between one plate with coplanar electrodes and another plate with a hydrophobic surface free of electrodes. The experimental results show that the best mixing sequence for the DNA ligation involves mixing first the insert DNA and vector DNA, and then adding the ligase solution. The waiting time was less than 5 min. In one cloning test, the total usage of reagents in an ultramicro-volume EWOD chip was 2.1 μL with no wasted volume, for comparison with 85% waste according to the standard protocol was 15 μL. The results showed also that four parallel DNA ligations were accomplished without damage to a chip and without biomaterial annulment.
Introduction
Microfluidic techniques enable efficient manipulation in complicated biomolecular procedures. Manipulating methods of many kinds have been developed; in particular, a droplet technique based on electro-wetting on dielectric (EWOD) has become popular for digital microfluidic systems. 1 –3 Not only is it practicable to manipulate minute biosamples using an EWOD chip but also EWOD-based systems have been used to develop several bioresearch devices and detectors, such as for the polymerase chain reaction (PCR), 4 for the detection of glucose 5 and for the ligation of DNA. 6 EWOD is based on varying the wettability of liquids on a dielectric solid surface by adjusting the electric potential; a thin dielectric layer or a dielectric layer of large relative permittivity allows a large change of contact angle to be obtained with a given applied voltage. 7 Droplet manipulation based on an EWOD technique enables the moving, cutting, merging, and creating of droplets. A common configuration of an EWOD chip involves two parallel plates; according to another configuration, an EWOD device with coplanar electrodes has the driving electrode and the ground electrode located on the same plate. 8 In a conventional application, aqueous droplets are manipulated directly with an EWOD device either in air or in an oil-based environment, but these actuating media have their limitations and prove inflexible for some biomolecular applications. An alternative configuration of actuation in which the droplets are enclosed in a thin layer of oil has been demonstrated in air for EWOD devices; this configuration with so-called water—oil core—shell droplets is useful for biomedical applications. 9
Besides PCR, the cloning technique is the most common tool to enable a scientist to study gene function explicitly. The process of DNA cloning is depicted in Figure 1 the fragments of DNA, containing the gene to be cloned, are inserted into a circular DNA molecule called a vector to produce a chimera or recombinant-DNA molecule. The vector DNA serves as a vehicle to transport the gene into a host cell, which is generally a bacterium, such as Escherichia coli. When the host cell divides, the vector produces numerous identical copies, not only of itself but also of the gene that it carries; the gene carried by the recombinant molecule is described as cloned. The transformed colonies are screened with hybridization, PCR, or digestion with restriction enzymes to identify those that carry the desired DNA sequence. DNA cloning should thus be based on the target DNA fragment successfully linked with vector DNA. If the clones carry a vector without insert DNA, the colonies show a blue color on an Lruia-Bertani (LB)-agar plate, whereas white colonies indicate clones carrying a plasmid containing an insert DNA. Comprising an optimized buffer system and DNA ligase, commercial DNA ligation kits are simple and highly efficient, allowing one to perform cloning procedures in vitro in only several minutes. 10 Because of this great efficiency, cloning requires only a small volume of reagent. Using a developed digital microfluidic chip, we explain here how DNA ligation with ultra-micro volume of solutions is accomplished based on an EWOD technique with coplanar electrodes.
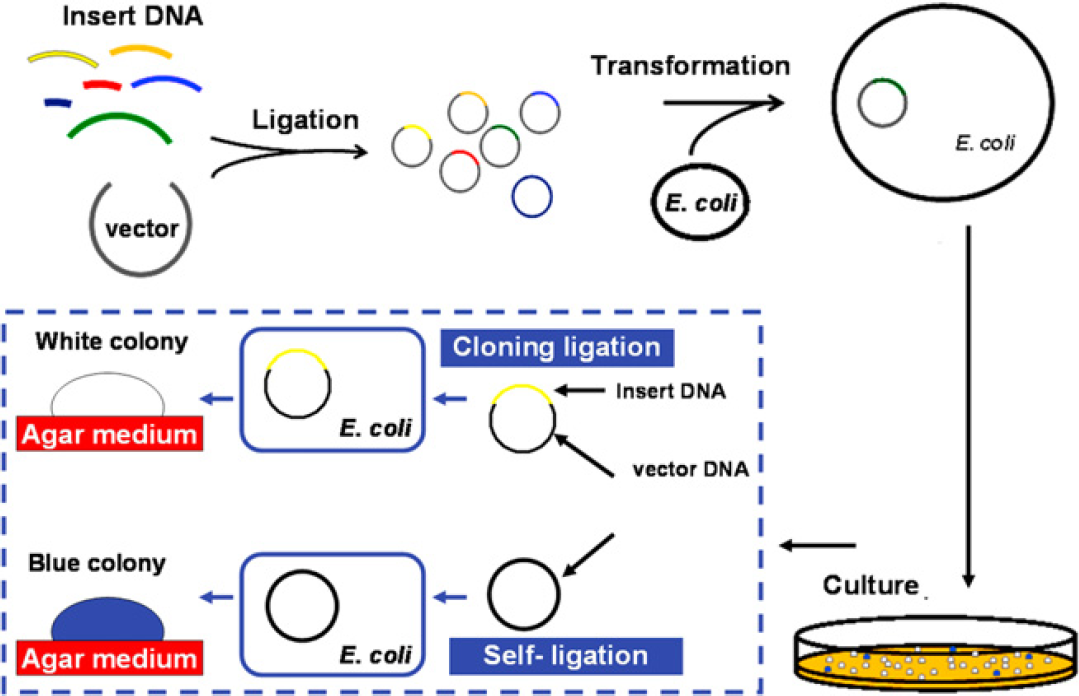
Overview of DNA ligation and cloning. The cloning has major three steps—ligation, transformation, and electrophoretic analysis. The successful ligation for a cloning purpose is defined with insert DNA linked with vector DNA and produces a white colony. If the vector ligates itself, ligation is considered also to occur, but the colony would be blue and carry no insert DNA.
Experiments
Coplanar Electrode Design and Chip Fabrication
According to the Young—Lippmann equation, materials of large relative permittivity were deposited as an insulator; pure silicon oxide or silicon nitride is a favored material in a common fabricating procedure. We propose an alternative approach, with a so-called complex dielectric layer. Based on a calculated threshold voltage required in an EWOD-based device to actuate a droplet, a complex dielectric layer is applied to decrease the actuation voltage relative to that for a pure dielectric layer. According to this procedure, a multiple dielectric layer, oxide—nitride—oxide, is deposited on an EWOD chip of coplanar electrode type, so as to decrease the applied voltage and to obtain a large contact angle change. A dielectric layer (thickness 400 nm) of oxide—nitride—oxide (thicknesses 50, 300, and 50 nm) deposited on an EWOD chip can manipulate DNA droplets smoothly with a total applied voltage 50 Vrms at 1 kHz.
To increase the flexibility of EWOD device, we have demonstrated a coplanar electrode design. The size of the control electrode is 1.5 × 1.5 mm, and the gap between subelectrodes is 20μm. A glass substrate was coated with a conducting layer of indium tin oxide (ITO) to form the coplanar electrodes. A photoresist was coated on the ITO glass; control and reservoir electrode patterns were defined with an exposure system and ITO etchant. A multiple dielectric layer (thickness 400 nm) of oxide—nitride—oxide was deposited by plasma-enhanced chemical vapor deposition to form the insulator. Teflon (1%) was coated at the end to form the hydrophobic layer. 6
Efficiency of DNA Ligation
The used biosamples in ligation process were prepared as the same steps as the one from Liu et al's. 6 The efficiency of DNA ligation was tested on varying mixing sequences and the waiting duration. A cover-free plate EWOD chip was used in this experiment; the process is shown in Figure 2. First, mix the insert and vector together at a 2:1 molar ratio, then 4-μL ligase/buffer solutions (containing 0.8-μL Fast-link DNA ligase, 1-μL 10× ligation buffer, 1-μL 10 mM ATP, and 1.2-μL double deionized water, according to the manual of Fast-Link DNA Ligation Kits, Epicentre) were dropped into the EWOD chip with a pipetman. By using the controllable program, insert DNA and ligase solution were mixed; this solution was placed on an EWOD chip at room temperature about 1, 2, 5, 10, and 20 min to vary its waiting time. The solution was then mixed with vector DNA for 5 min to continue the transformation and culture, as shown in Figure 2A. To achieve optimal mixing sequences, we mixed vector DNA and ligase solution and then added insert DNA as shown in Figure 2B, or we mixed insert DNA and vector DNA and then added ligase solution as shown in Figure 2C. We applied a 57 Vrms AC potential at 1 kHz to transport those droplets in this experiment. After the DNA ligation, the solutions were heated at 65 °C for 10 min to deactivate the ligase. Every growth of a colony on the plate implies a successfully ligated plasmid, no matter whether vector DNA ligated with insert DNA or with itself. Of the colonies of two colors, we collected and counted the white colonies to confirm the completion of ligation such that the plasmid contained both vector DNA and insert DNA.
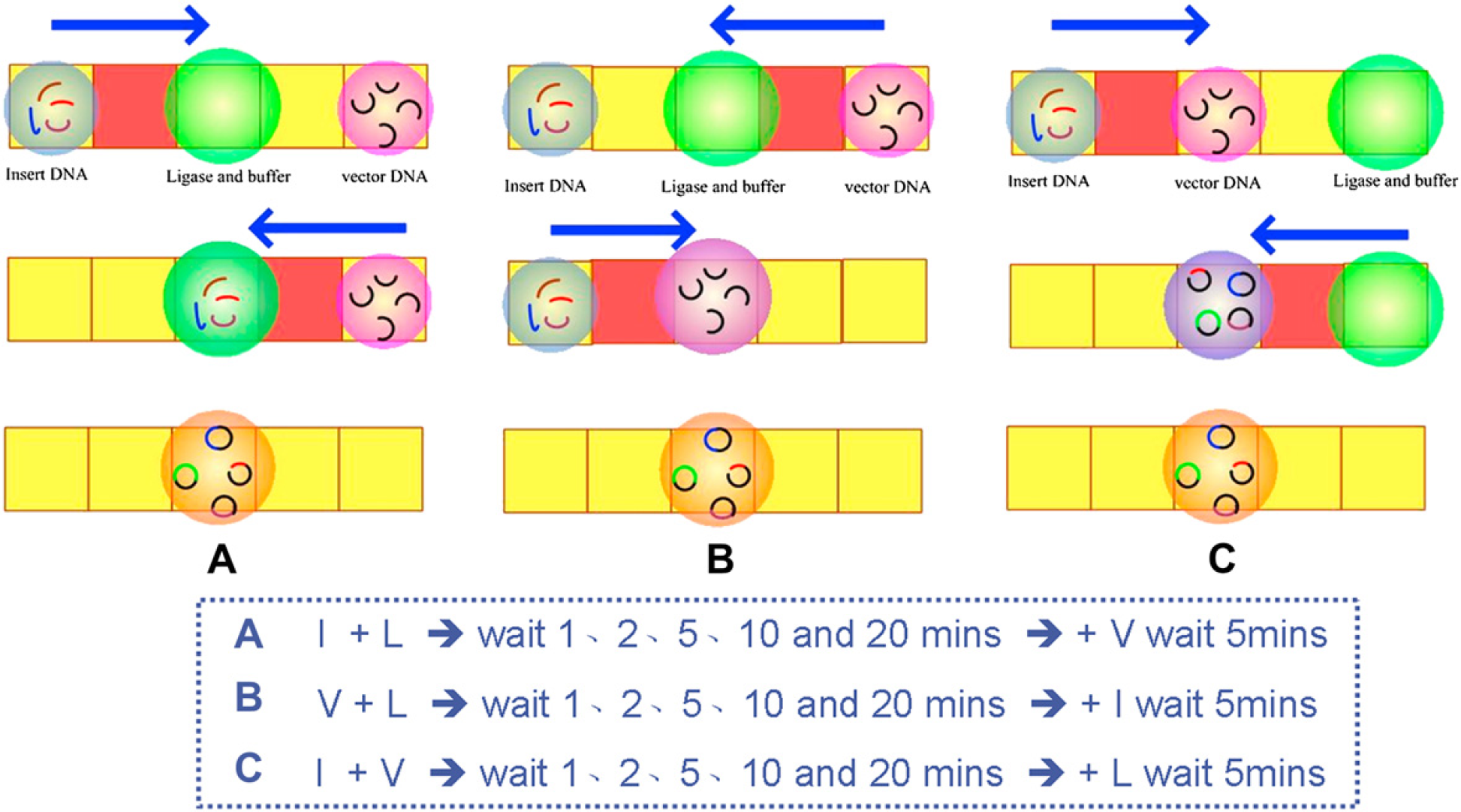
Varied mixing sequences of DNA ligation. (A) Insert DNA and ligase solution were mixed and then vector DNA was added. (B) Vector DNA and ligase solution were mixed and then insert DNA was added. (C) Insert DNA and vector DNA were mixed and then ligase solution was added.
Parallel DNA Ligation with Core—Shell Droplets
To diagnose various DNA solutions and decrease these volumes of total usages, we demonstrated an ultramicrovolume parallel DNA ligation, with an EWOD chip and total applied voltage 50 Vrms at 1 kHz for this ligation. A cartoon image of such a parallel ligation is shown in Figure 3. First, several 0.2-μL silicone oil droplets were dropped by using a pipetman. Then, each four 5-μL varied insert DNA (including 0.9-, 1.7-, 1.9-, and 0.9/1.9-kb DNA fragments), 5-μL one vector DNA, and each four 1.5-μL ligase/buffer solutions (same as above description) were dropped on the silicone oil droplets to form water—oil core—shell droplets, which would be dropped into their own reservoirs with a pipetman.
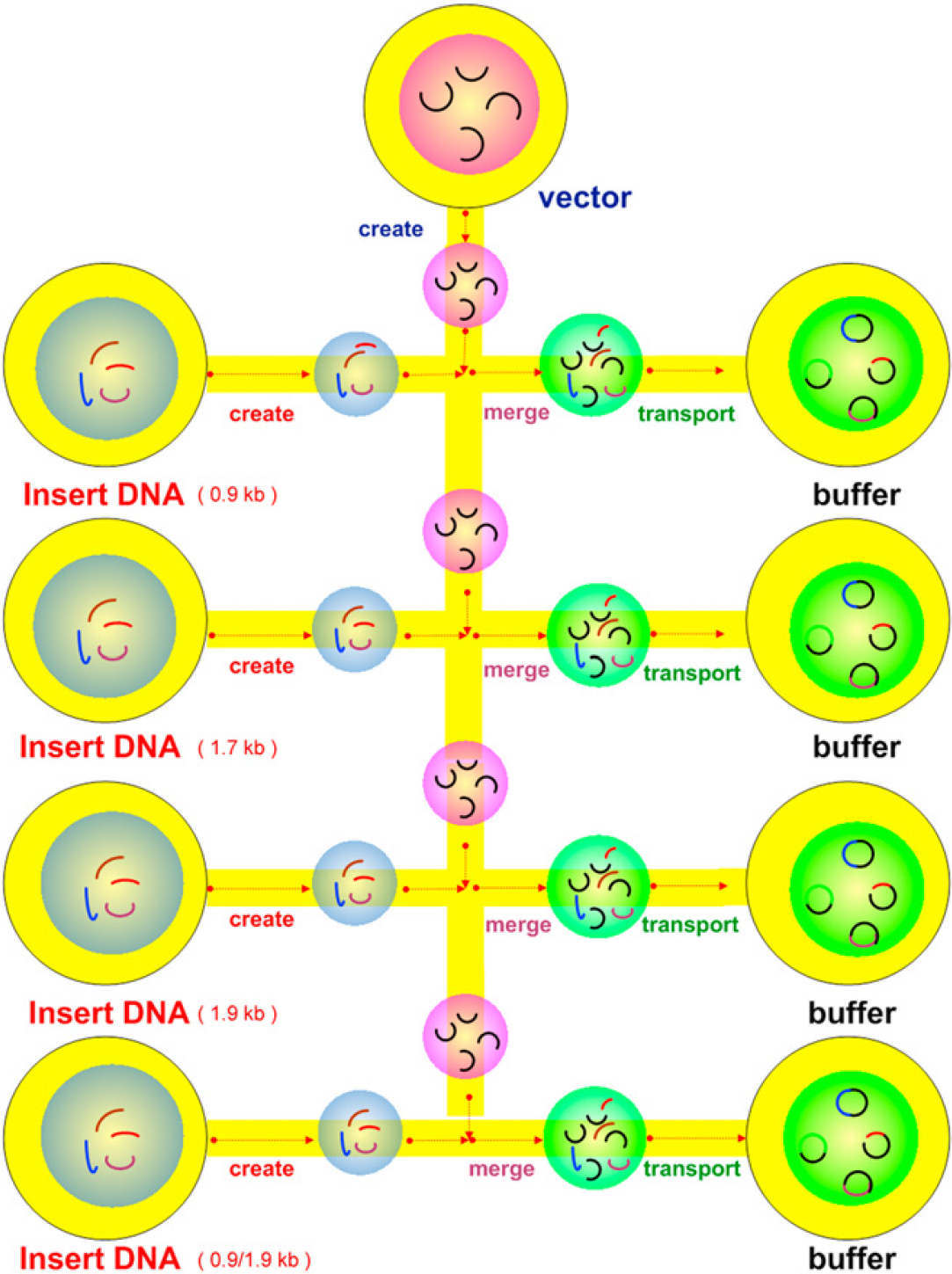
Cartoon images of four parallel DNA ligations. Vector DNA and insert DNA were created from their respective reservoirs; droplets were transferred and merged with ligase/buffer solution.
A function generator (NI PXI-5402) supplied an AC potential signal (3.5 Vpp at 1 kHz, sine wave); the signal was amplified (50 Vrms, A.A. Lab. System Ltd, A-303). We used software (LabVIEW 8.0, National Instruments, Austin, TX) to plan the droplet transport routes and to output the signal to switch relays to turn on or off through a relay module (NI PXI-2569). An EWOD chip was connected to the controllable circuit with peripheral component interconnect slots; the AC potential (50 Vrms) was applied to the EWOD chip, and then the droplet motion of the DNA ligation was planned at the human—machine interface. We carefully observed the droplet motion with an optical microscope.
The cover-glass plate was then covered with spacers (thickness ∼ 100μm) formed from one piece of tape (double sided). Second, four drops of vector DNA and four drops of insert DNA were created, one by one, from their own reservoirs. All core—shell droplets in this ligation were formed from biomolecular solutions and the silicone oil (octamethyl-trisiloxane, viscosity 5cSt, Sigma—Aldrich). Afterward, the four combinations of insert DNA and vector DNA were mixed simultaneously and then transported to their reaction buffer reservoirs. Each DNA solution was mixed with ligase buffer solution (1.5μL). The entire mixtures were then incubated on a chip at room temperature about 5 min to link the cohesive end of the insert DNA and the vector DNA with the ligase. After this DNA ligation, the solutions were heated at 65 °C for 10 min to deactivate the ligase. These mixtures were then ready for transformation.
Transformation
For use in the transformation, the entire ligation mixture was mixed with chemically competent cells (20 mL) and incubated (on ice, 5 min) before being suddenly switched into a water bath (42 °C, 45 s) to prevent a thermal shock effect. The competent cells were recovered in a super optimal broth with glucose medium (100μL, about 1 h, 37 °C, horizontal shaking at 225 rpm). The cells were finally plated on an LB-agar plate that contained ampicillin (100μg mL−1) and incubated (37 °C, overnight).
Culture and Insert DNA Check
After incubation (16–18 h), four colonies were selected from the LB-agar plate, incubated in LB broth medium (1 mL) with ampicillin (100μg mL−1, 37 °C, overnight), then subjected to shaking at 100 rpm. All cells were harvested in a centrifuge (13,000 rpm, about 1 min), and the plasmid was extracted with a commercial kit (Gene-Spin 1–4–3 DNA extraction kit, Protech Inc.). To verify the correct length of vector DNA and insert DNA, we digested the plasmid with HindIII (same conditions as above). These DNA fragments were separated on electrophoresis with agarose gel (1%, in 1 × TAE buffer) containing ethidium bromide (1 mg mL−1).
Results and Discussion
Efficiency Test of DNA Ligation
The efficiency of DNA ligation with varied mixing sequences and waiting time was tested before the parallel DNA ligation of ultra-micro volume. The efficient DNA ligation with a free-cover coplanar EWOD chip grew many colonies. The results of varied mixing sequence and waiting time for DNA ligation are shown in Figure 4. Every colony grown on the plate contains a ligated plasmid no matter whether the vector ligated with insert DNA or with itself. These clones carrying a vector without insert DNA show a blue colony on the LB plate, whereas the white colonies implied clones carrying a plasmid containing insert DNA through successful ligation as shown in Figure 1. According to these results, the third mixing sequence (Fig. 2C) attained the greatest efficiency of the DNA ligation sequence. Also, more white colonies were obtained, which showed that the clone carried a plasmid containing insert DNA through ligation for which the waiting time was less than 5 min.
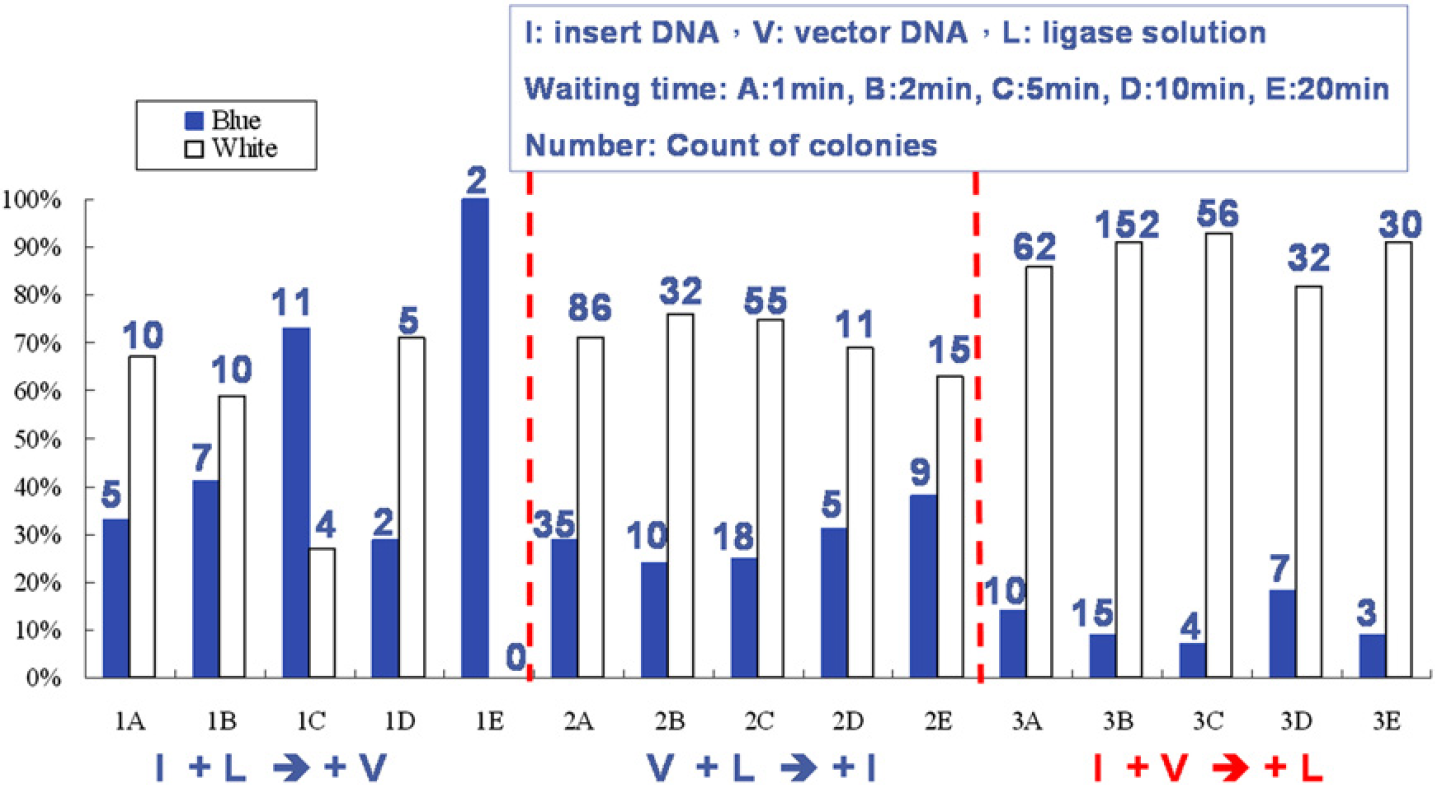
Results of varied mixing sequences and waiting time for DNA ligation. The blue rectangles represent that these clones carry a vector without insert DNA on the LB plate, whereas the white rectangles denote clones carrying a plasmid containing insert DNA through ligation. (The numbers above the rectangles indicate the count of colonies on the LB plate.)
Transmission of Core—Shell Droplets on an EWOD Chip with Coplanar Electrodes
Four drops of insert1 DNA (5μL, encapsulated with 1-μL silicone oil shells) with various length, one drop of vector DNA (5μL, encapsulated with 1-μL silicone oil shells), and four drops of enzyme buffer solutions (1.5μL, encapsulated with 0.5-μL silicone oil shells) were previously pipetted into reservoirs and covered with a glass plate, as a preparation for further ligation steps. Some sequential images of four parallel ligation processes are shown in Figure 5. First, a core—shell vector DNA droplet (0.μL) was created from its own reservoir (picture not shown), the core—shell insert DNA droplet (0.3μL) was then created from its own reservoir shown in Figure 5A—D, and the insert DNA and the vector DNA were mixed as shown in Figure 5E—F. Afterward, the DNA mixture (0.6μL) was transported to the ligase/buffer reaction reservoir (1.5μL) for the ligation reaction as shown in Figure 5G—I. Another three DNA ligation processes were manipulated concurrently with the controlling program. In these four parallel DNA ligations, the total volume of each reaction was 2.1μL. In this driven process, we observed that the droplets were generated, transported, and mixed smoothly. The surface of the chip was not damaged and no bubbles appeared after the entire process. Even when the concentration of a DNA droplet attained 250 ngμL−1, it was still driven smoothly without electrolysis.
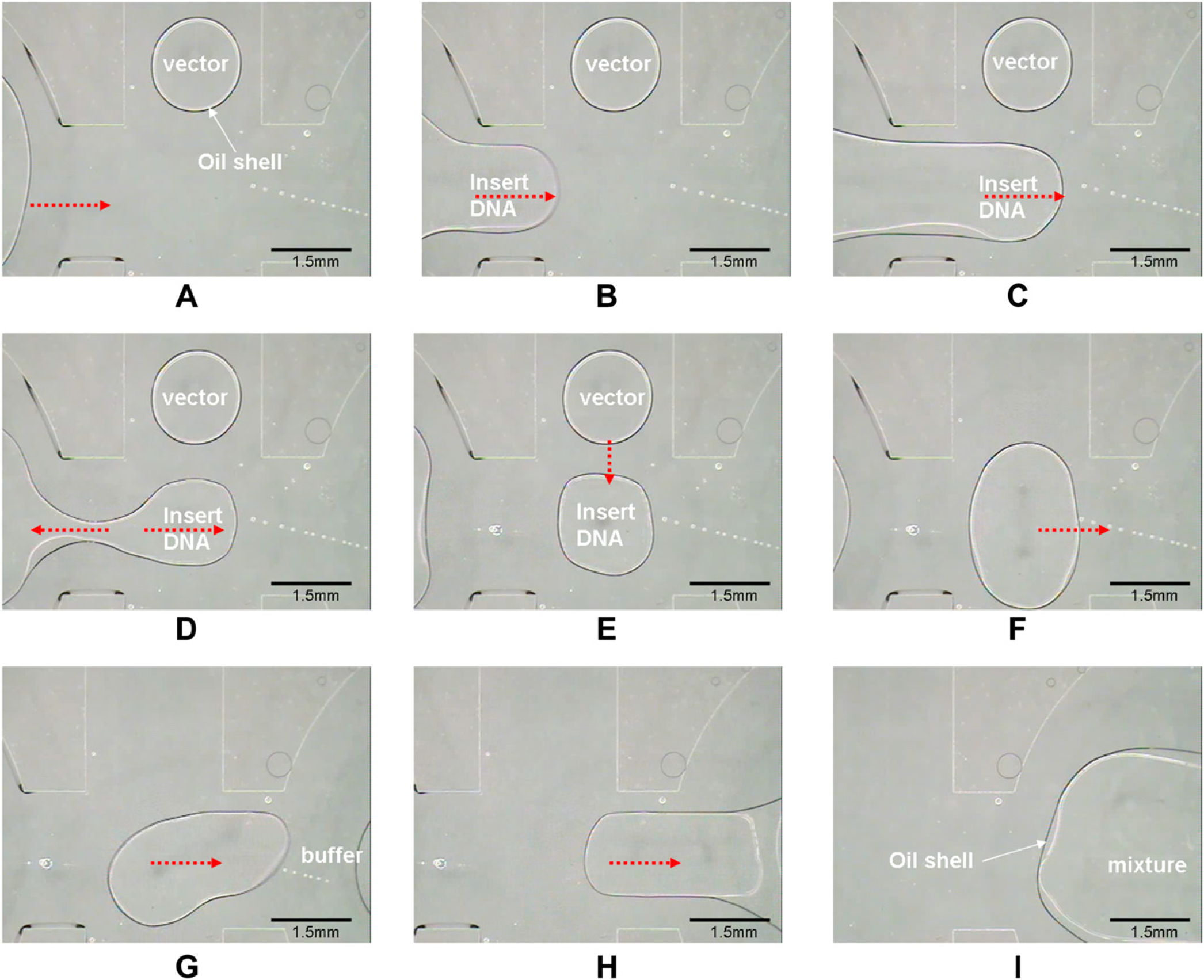
Sequential images of ligation. Video sequence (left-to-right and top-to-bottom) depicting (A)—(D) a core—shell droplet of insert DNA (0.3μL) created from a reservoir. (E)—(F) Insert DNA and vector DNA being mixed. (G)—(I) A DNA droplet being transported to the ligase/buffer reaction reservoir for ligation.
To compare the total usage of reagents according to various methods, we applied two other methods for this ligation operation—a method recommended from a commercial kit and another involving a cover-free coplanar EWOD chip. The former and latter methods left about 85% and 80%, respectively, of the total volume of reagent after ligation was completed. A coplanar EWOD chip with a hydrophobic shelter in this work produced microdroplets of volume only 0.3μL, which might be further diminished: such a volume is too small to be measured with a general microinjection pipetman. In this manner, an ultra-micro volume of DNA ligation decreased the required amount of ligation reagent.
Analysis of the Insert DNA Length
The pUC19 plasmid (HindIII cloning-ready) was used as a vector and four DNA samples (containing 0.9-, 1.7-, 1.9-, and 0.9/1.9-kb DNA fragments) were used as insert DNA. A commercial kit (Fast-Link DNA Ligation and Screening Kit, Epicentre) and chemically competent cells were also used in the cloning process. After ligation, cloning and culture, the plasmid from four parallel DNA ligation trails were purified and cut with HindIII. After incubation (16–18 h), more than 100 white colonies were obtained. Every growth of colony on the plate implies a ligated plasmid no matter whether the vector DNA ligated with insert DNA or with itself. The white colonies were collected to confirm the completion of ligation such that the plasmid contained both vector DNA and insert DNA. After extraction of plasmid and digestion with restriction enzyme, the electrophoresis results are shown in Figure 6. Every colony has a vector DNA fragment that correctly shows a band about 2.6 kb by comparison with the lambda/HindIII-digested DNA marker on the left side (label“M”) of the gel. Lanes 1 to 3 show the 0.9-, 1.7-, and 1.9-kb insert DNA, respectively, and lane 4 shows both 0.9-and 1.9-kb insert DNA. These insert DNA band positions matched correctly the lambda/HindIII-digested DNA marker, which was a source of insert DNA for ligation. These data showed that an EWOD chip of coplanar electrode type with a hydrophobic shelter is usable to accomplish four parallel ligations with a small volume. Furthermore, we found no damage to either the biosamples or the chip itself caused during the entire process. Based on these results, an EWOD chip of coplanar electrode type becomes an effective tool to deliver diverse biosamples, such as DNA, enzyme, and salt solution without problem.
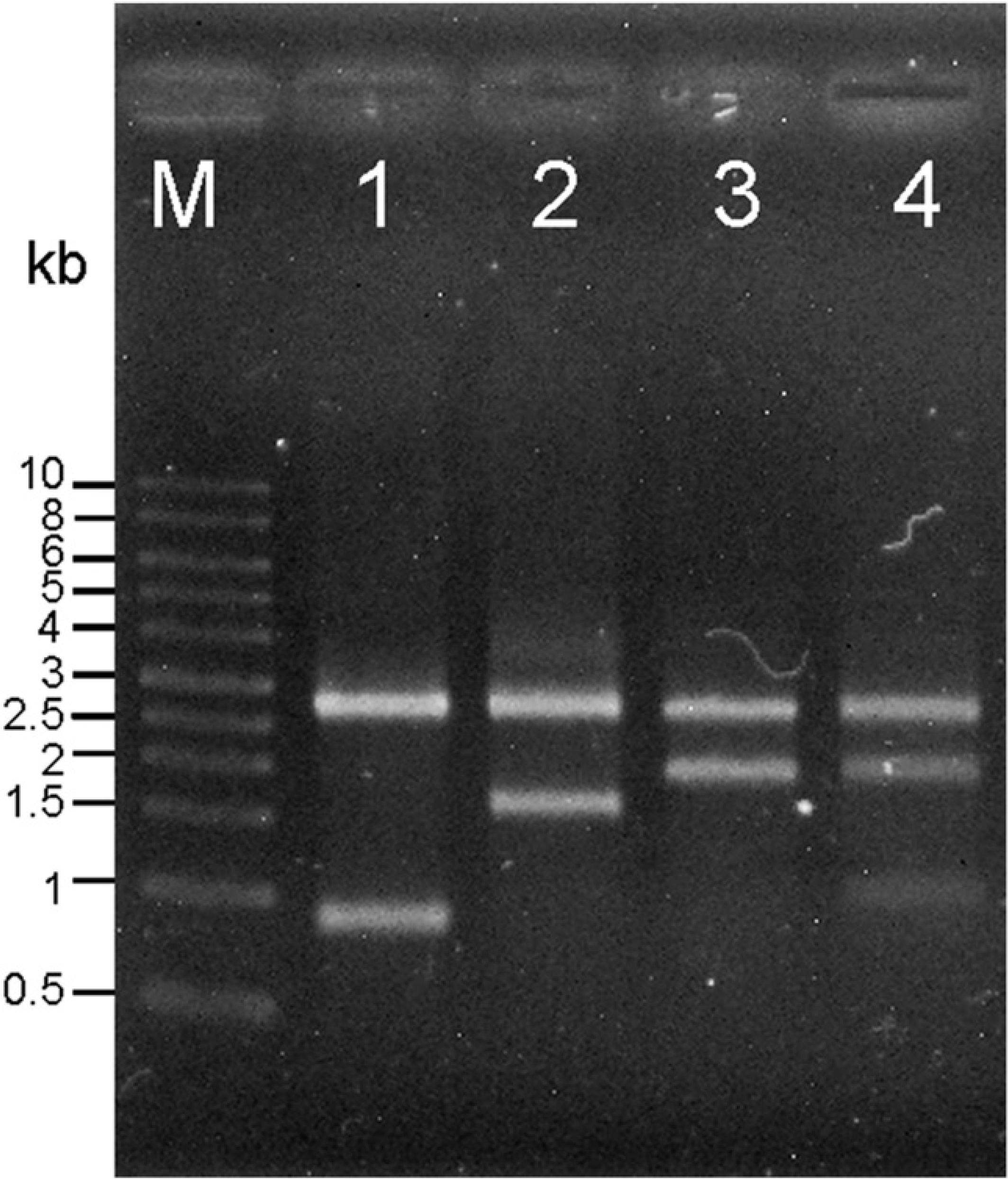
Verification of insert DNA length using HindIII cutting. All plasmids from four trials contain the pUC19 vector (2.6 kb) and insert DNA. Lanes 1 to 3 show the insert DNA 0.9, 1.7, and 1.9 kb, respectively and lane 4 shows both 0.9-and 1.9-kb insert DNA. (Lane M is the lambda/HindIII DNA marker.)
Conclusion
In this work, we propose a core—shell droplet and EWOD digital microfluidic platform with coplanar electrode types, which can accomplish four independent DNA ligation of ultra-micro volume simultaneously. Using a 400-nm complex dielectric layer and a core—shell configuration, we decreased the applied voltage to 50 Vrms at 1 kHz to actuate biomedical regents. Furthermore, this reaction platform minimized the usage of total reagents by creating ultramicro-volume droplets from reservoirs. In one cloning, an ultramicro-volume EWOD chip achieved a ligation mixture of volume 2.1μL with no waste. For comparison, with a standard protocol and a freecover coplanar EWOD chip, about 85% and 80%, respectively, of DNA and enzyme/buffer volume were wasted. Therefore, in four independent clonings, the DNA and enzyme/buffer usage volume (2.1μL × 4 = 8.4μL) was less than a standard protocol (15μL × 4 = 60μL) about seven times. In the future, via integrating other cloning functions, such as thermal control and cell incubating areas, coplanar EWOD digital microfluidics might be designed and improved to become an efficient DNA cloning system to construct an artificial library or expression library in the post-genome era.
Acknowledgments
The National Science Council, Taiwan (grant NSC 96-2221-E-007-128-MY3), provided financial support.
Competing Interests Statement: The authors certify that they have no relevant financial interests in this manuscript.
