Abstract
With over a decade passed since the first reported use of a Staphylococcal α-hemolysin pore to study single molecules of single-stranded DNA, research in the field of nanopores has advanced rapidly. We discuss the technological progression of nanopore-based devices from the initial use of α-hemolysin pores to the advent of solid-state nanopores to the burgeoning of organic-inorganic hybrid pores driven by the desire to achieve fast and inexpensive DNA sequencing. Additional nanopore-based efforts are also discussed that study other classes of molecules, such as proteins. We discuss the use of nanopores for protein folding and binding analysis. In addition to single-molecule analysis, we report on the introduction of nanopore arrays on thin film membranes for ultrafiltration. Owing to their reduced spatial dimensionality, such membranes offer greater control over how the pores interact with analytes thus leading to very efficient separation. With several technical hindrances yet to be overcome, the devices we report are still works in progress. The realization of these devices will enhance laboratory processes by permitting superior spatial and temporal analytical resolution at the single-molecule level resulting in laboratory capacities of great impact.
Introduction
Nanoscale transmembrane pores are found on every living cell. Their natural role is to create conduits for material to cross the cell wall and the walls of intracellular organelles. 1 These seemingly minor cellular components are of great importance to cell function and life. One class of pores allow cells to maintain different internal concentrations of solutes from the exterior environment, 2 maintaining their osmotic balance and stabilizing their volume. 3 The proper function of these pores is also of vital importance to both individual cells and the entire organism. For example, cardiac myocytes use potassium and sodium pores that are integral in keeping the rhythm of the muscle cell's contractions and consequently the heart's overall beat. 4 Pathogens use nanopores as well. Viruses can inject their genetic information via a nanopore 5 into a host's cells to further replicate, and bacteria can produce toxin pores used to lyse cells for their intracellular nutrients. These pores each have evolved over a tremendously large number of generations to have various structures and chemistries that allow them to achieve their specific tasks. It is from these evolved pores and the electrophysiological tools used to study them from which the application of both biological and solid-state nanopores as engineered tools was and is still being developed. 6
Initial experiments using nanopores as sensors used the unique nongating and inherently stable Staphylococcal α-hemolysin ion channel suspended in a planar lipid bilayer, foremost for the detection and sequencing of nucleic acids. 7 The nanopore sensing method functions similar to the working principles of a Coulter counter (Beckman Coulter Inc., Brea, CA) in which particles immersed in a conducting fluid are introduced into a microscale capillary clamped at a constant potential resulting in a partial blocking of the ionic current. 8 After recording the current, information about the particles present in the solution can be obtained by analyzing the temporary decreases in current (ionic blockades) in both the amplitude and time domain. Over a decade has passed and the driving force behind nanopore sensing research still remains rapid and low-cost genomic sequencing. The potential impact of this method to science and society is massive. 9 Access to inexpensive genomic data of individuals in both sickness and health would enable researchers to identify previously unrecognized genetic relationships to a multitude of diseases and disorders, such as the various mutations that lead to cancer. 10 Additionally, identification of genetic correlations to responses to medication may result in personalized medicine based on patients' genetic information. 11 Another impact may occur by allowing individuals access to their personal genomes, which would give them the option to take preventative measures once understanding their genetically predisposed susceptibilities. Other possible impacts also include designed or synthetic biology, in which rapid sequencing would partially enable rapid design and development of synthetic enzymes 12 or other proteins. 13 Moreover the knowledge and expertise gained from progressive works toward nanopore-based DNA sequencing have led to impacts and emergence of other potential applications that have fueled further advances in nanopore-based science.
Among these emerging fields of study, protein analysis is one of the most significant. With this new application, however, come new technical challenges. For instance, proteins have a heterogeneous charge distribution along the length of their primary amino acid sequence leading to complex three-dimensional structures. Additionally, the number of amino acids making up the protein and the degree of folding result in variable protein sizes. Protein analysis using nanopores can be categorized into the following broad areas of research: structure analysis, biosensing, and binding characterization. Because current blockade events are characterized by amplitude and duration, when an event is mapped into this two-dimensional space, clusters correspond to distinct translocation events, which can be correlated to a predicted structure. Biosensing experiments are conducted by designing a pore that is adapted with a specific binding site capable of holding the analyte protein inside the pore for a characteristic period of time. Binding characterization on the other hand is performed by calculating binding rate constants using the mean interevent interval (association rate constant) and the mean event lifetime (disassociation rate constant). 14 The motivation for these studies include applications in protein engineering (de novo protein design), pharmaceutical science (drug—protein binding characterization), and biosensors (detecting disease-relevant protein biomarkers). 15
Another field wherein nanopore technology has been introduced is ultrafiltration. The ability to fabricate ordered arrays of nanopores in ultrathin membranes has made it possible to develop highly efficient molecular separation devices. 16 The selectivity of such devices can be tailored for specific applications by chemical surface modification 17 or by modifying the Debye layer around the pores using embedded electrodes. 18 The selectivity of such membranes is roughly an order of magnitude greater than conventional dialysis membranes. 16
Dna Sequencing and Analysis
In 1996, John Kasianowicz and his colleagues were the first to use a biological nanopore, Staphylococcal α-hemolysin (Fig. 1A and B) to translocate (transport through the nanopore) and characterize the length distribution of individual polynucleotides using the ionic blockade method. 7 This landmark achievement has since sparked the interest and efforts of numerous groups with hopes of developing a method to rapidly and inexpensively sequence genomes using nanopores. Simply put, it was proposed that if long single-stranded DNA (ssDNA) in ionic solution could be translocated electrophoretically through the pore, the DNA could be sequenced by recording the concurrent modulation of ions flowing through the pore. This concept relied on each nucleotide being able to block the measured ionic current through the limiting constriction of the pore by an amount according to their individual molecular size and chemical properties in a sequential fashion. Later works toward this goal demonstrated the ability to sense entire ssDNA strand characteristics, such as orientation during translocation, 7,19 ssDNA homopolymer composition, 20 –22 and differentiation of RNA block copolymers. 20
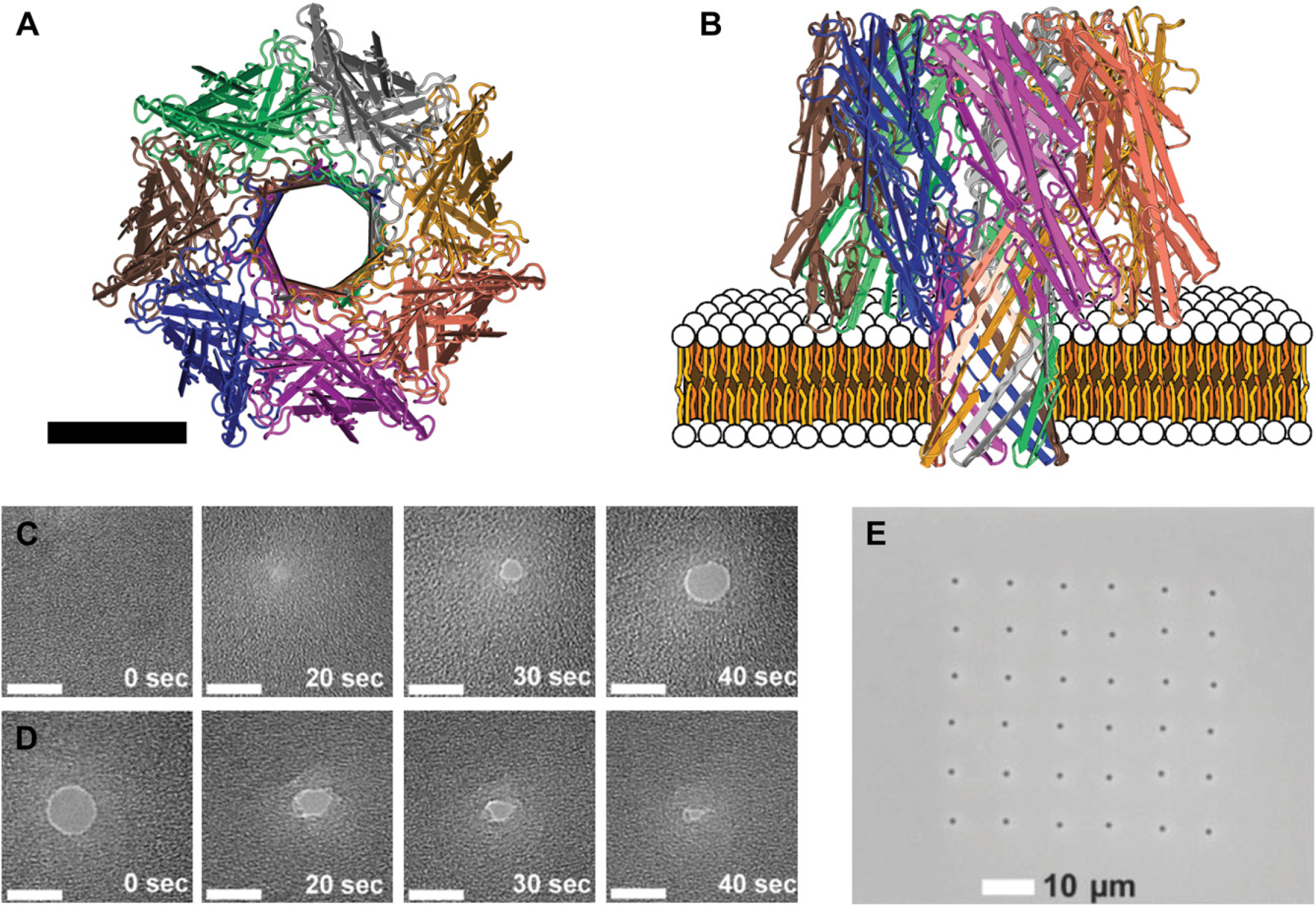
A) Axial view of a simplified representation of the α-hemolysin protein heptamer pore assembly. 24 The scale bar is 3.5 nm in length. (B) Radial view of α-hemolysin pore inserted into an illustrated bilipid membrane. A sequence of TEM images displaying the dynamics of drilling (C) and contraction (D) in the 50-nm thick silicon nitride membrane. 34 The scale bar is 5 nm. (E) A TEM image of a 6 × 6 array of 2-μm wells, each well containing a 5-nm pore 32 (C,D) Reproduced with permission from Ref. 34. Copyright (2007) IOP Publishing Ltd. (E) Reproduced with permission from Ref. 32. Copyright (2006) WILEY-VCH Verlag GmbH & Co. KGaA, Weinheim.
Hindrances to genomic sequencing using this initial concept do exist. For instance the α-hemolysin nanopore's sensing zone (volume in which translocating material is detectable by the measured pore current) was later found to occupy its entire 5-nm long stem, 23 which is much longer then the base-to-base distance (3.4 Å) of DNA. Because of this, decreases in the measured ionic current through the pore during ssDNA translocation include the effects of the many nucleotides within the stem. 24 More recently, it has been revealed that there are three recognition sites or axial positions within the stem where a nucleotide will have greater effect on the overall ionic flow than other positions leading to further complexity. 25 Another well-known technical hurdle is the average time (∼1μs) under typical experimental conditions at which free translocating nucleotides will remain at such recognition sites. This is well below the time needed to take reliable measurements of the individual nucleotide's restricted electrolyte flow. One promising effort to remedy these shortcomings uses a mutated α-hemolysin pore along with a molecular adapter to temporarily bind individual nucleotides during translocation, increasing their time at the recognition site and allowing for the identification of the base. 26 It is envisioned that by fixing a controllable exonuclease near the entrance of such a pore, the sequence of long strands of nucleic acids can be recorded a single nucleotide at a time as they are cleaved. 27 Many technical hurdles exist in achieving this objective at satisfactory detection rates for genomic sequencing as well. These include integration and alignment of the exonuclease to the pore and control of the exonuclease's cleaving rate. 27 To date, no group has been able to develop a nanopore-based experimental setup that has the temporal or electrical resolution to rapidly characterize ssDNA at the single-nucleotide level.
In addition to the fundamental limitations detailed above, the inherent noise, instability, and limited bandwidth of the patch clamp/Staphylococcal α-hemolysin nanopore experimental setup 28 are also known hindrances to the rapid and continuous data acquisition needed for genomic sequencing. The experimental setup of Kasianowicz et al.'s seminal work, consisting of the α-hemolysin toxin reconstituted in a planar bilipid membrane (Fig. 1B) separating two aqueous compartments is presently still the most widely used. The planar bilayer is applied using one of the two traditional methods 29,30 on a ∼100-μm diameter aperture through a ∼25-μm thick Teflon septum. The septum' hydrophobic surface chemistry encourages the deposited lipids' fatty acid tails to orient toward the septum resulting in the formation of local lipid monolayers on both sides of the septum and a suspended bilayer across its aperture. Teflon's chemical stability, high resistivity, ability to form high resistance seals with lipid layers, and low permittivity prevent excess noise during current measurement. Nevertheless, this well-established 28 bilayer support system's notorious sensitivity to physical vibration, temperature, electrical noise due to capacitance across the bilayer, and propensity to rupture has led to searches for its improvement or replacement. Toward this, Mayer et al. 31 showed that by fabricating apertures reaching diameters as low as 2μm using soft-lithographic molds and amorphous Teflon AF lead to decreases in membrane capacitance and noise and increases in stability (multiple hour lifetimes) and breakdown voltage (∼150 mV). The difficulty in fabricating the Teflon AF-based apertures has led many leading groups to continue using the larger diameter standard Teflon septums.
An alternative to bilayer-suspended pores arrived with the advent of solid-state pores (Fig. 1C, D) and pore arrays 32 (Fig. 1E). The first solid-state nanopores were drilled in micro-fabricated silicon nitride or silicon dioxide freestanding films supported by silicon frame chips using argon ion-beam sputtering 33 and later commercially available transmission electron microscopes. 34,35 Methods of creating individual nanopores in glass capillaries 36 and track-etched polymers 37 also exist, yet are not as widespread as the use of silicon-based platforms that promise to allow the easy integration of pores with other nanoelectromechanical/microelectromechanical devices. Further advantages of solid-state nanopores include fixed pore position (as opposed to a diffusing protein pore within a lipid bilayer), controllable dimensions (both pore diameter and thickness) (Fig. 1C, D), longer lifetime, increased temperature and ionic strength ranges of operation, easily modifiable surface properties, 38 –40 a single recognition site, and much higher breakdown potentials. These characteristics have allowed the translocation of larger molecules, such as dsDNA, 41,42 the discrimination of double stranded from single-stranded nucleic acids, 43,44 experiments under nonphysiological conditions, 45,46 and optical detection using multiple fixed pores in parallel, 32 which could not have been accomplished using the earlier protein pore/bilayer system.
Although solid-state pores offer many advantages over the traditional protein pores, the physics associated with translocation of nucleic acids through solid-state nanopores is still not completely understood. As opposed to the translocation of long ssDNA in the fixed diameter α-hemolysin pore that have mean translocation times linearly related to their length at constant applied voltage and temperature, 47 translocations times in solid-state pores vary with pore dimensions, and have faster mean translocation times with broader distributions. 48 The physics associated with solid-state pore translocation and ionic conductance is the subject of a great deal current experimental 49 –52 and theoretical 53 –55 study. Additionally it is well known that solid-state pores have a similar fundamental sensing zone limitation as the α-hemolysin-based system. Although solid-state pores have only one recognition site, the thinnest nanopore membrane reported thus far, ∼ 5 nm, 33 is again much longer than the base-to-base distance (3.4 Å) of DNA. Significant resistance is also created directly outside of the pore's volume to an approximated diameter in length away. 56,57,53 Because of this, decreases in the measured ionic current through the pore during ssDNA translocation include the effects of the many nucleotides both within and directly outside the pore. In addition, solid-state pores have been found to have greater capacitance, and other noise issues related to surface phenomena as compared with biological pores. 58 –61
Alternatives to the traditional ionic blockade detection technique are driven by the ease of integrating other microfabricated architectures on the solid-state pore chip. Proposed alternatives include detection using capacitance, 62,63 massively parallel single-molecule fluorescence, 64 or transverse tunneling. 65 Further microfabrication-based solutions include using chip integrated electrodes to manipulate local electric fields for DNA translocation control 66 and deposition of various chemistries to decrease noise by changing the chip's surface properties. 40,60 A drawback to these possible solutions is the lack of reproducibility or atomic level accuracy of a genetically controlled protein pore as even subtle changes in a pores size can alter translocation dynamics significantly. 50
A promising method to use the advantageous properties of both classes of pores and eliminate their inherent disadvantages is suspending lipid bilayers and biological pores across solid-state pores. Similar to the traditional method of suspending a lipid bilayer across Teflon apertures, solidstate pore membranes with hydrophobically chemically modified surface chemistry will encourage deposited lipids' fatty acid tails to orient toward the membrane. Applying a suitable amount of lipid near the membrane via the Mueller“painting” method 30 will result in the formation of local lipid monolayers on both sides of the membrane and a suspended bilayer across the pore. Using chemically modified ∼ 150-nm diameter conical glass pores White et al. 67 were able to achieve 800-mV breakdown voltages, 2 week lifetimes, and 70 Gω membrane/bilayer resistances, far exceeding the properties of traditionally suspended lipid bilayers. These glass capillary-based pores on the other hand are limited in their throughput due to their single pore restricted fabrication method. 36 To remedy this, Han et al. 68 suspended bilayers across hydrophobically silanized silicon nitride nanopore arrays, and later Studer et al. 69 detailed the suspension of various organic pores in similar silicon nitride arrays as a function of pore size and total pore area. Although Studer et al. were successful in suspending organic pores in an array allowing parallel throughput, the membrane/bilayer system's properties were not at par with White et al.'s single glass hybrid pore's properties and additional work in this area is still needed. It is also necessary to consider that further decrease of the size of the supporting apertures greatly increases the lag time needed to insert the biological pore into a bilayer, especially for oligomer pores, such as α-hemolysin, which self-assemble from protein monomer in the bilayer. 69 Rapid insertion methods have been developed to stem pre-experimental setup delay using microscale septum-suspended bilayers and plastic probes tipped with agarose gel containing adsorbed preoligomerized α-hemolysin pores. 70 Also, glass probes that adsorb and transfer oligomerized protein pores directly from bacterial colonies, 71 have been investigated; however, oligomerized pore insertion into nanopore or nanopore array still remains an unresolved matter. Overall the hybrid pore scheme seems to provide the most immediate promise for low-cost high-speed genomic sequencing by combining the strengths of both organic and solid-state pore technologies while significantly decreasing or eliminating their individual present weaknesses.
Protein Analysis
Proteins are biopolymers that can be composed of a sequence of more than 20 types of amino acids varying in charge and polarity. 72,73 The sequence of amino acids is responsible for how a protein folds into its three-dimensional shape that is critical to the protein's function. When a protein is synthesized, local segments of the protein begin to fold into secondary structures, such as α-helices and β-sheets, which help define the overall three-dimensional conformation of the protein called the tertiary structure. 73 The relationship between the amino acid sequence, structure, and function is demonstrated by considering the disease sickle cell anemia, which is caused by a substitution of a single amino acid in the protein hemoglobin found in red blood cells. This small alteration causes the protein to aggregate, changing the shape and elasticity of the blood cell resulting in decreased functionality. 74 Although sensitive to structural alterations, healthy proteins are incredibly robust at performing their complex functions. A classic example is the molecular motor protein assembly used by bacteria, responsible for rotating their flagella to propel themselves through their environment. 13 From an engineering perspective, the sophisticated molecular machine-like functionality of proteins or protein assemblies is desirable in nanoscale systems. In the future, protein engineers will be able to more predictably alter the properties of proteins and create new proteins with a prescribed function. 13 This de novo design approach requires understanding of the relationships between amino acid sequence, three-dimensional structure, and function.
The current method for studying protein folding using nanopores involves inducing a structural change (i.e., unfolding) in the protein either during or before nanopore translocation and correlating the ionic current blockade characteristics to predicted folding states. 75 The method for inducing structural change in a protein is complex and involves altering the balance of chemical, electrostatic, and thermodynamic forces that act on the protein. The two predominant experimental techniques to unfold proteins include using denaturants (guanidium chloride and urea) and using a pore smaller than the protein such that the protein is deformed due to the steric hindrance caused by the pore (nanopore-induced unfolding). 75,76 A third mechanism has been recently suggested which uses the electric field within the pore to pull positive and negative amino acids in opposite directions thereby inducing protein unfolding. 76 Although it is hypothesized by researchers that these unfolding mechanisms can shed light onto novel unfolding pathways, a consensus on the best method to unfold a protein has yet to be reached. It has been realized however, both nanopore-induced unfolding and electric field-induced unfolding have physiological significance to the processes that occur in cells. 15,76 The type of information gained from current work includes the discrimination between folded and denatured proteins as well as the detection of single-point mutations in a protein. 75,77 Future works are likely to integrate nanopores with other single-molecule tools, such as fluorescence, optical tweezers, and atomic force microscopy enabling further study of native and engineered proteins. The current methods of protein unfolding provide complimentary information about different folding transitions and the pathways to forming the final folded state. Hence, no one method alone is fundamentally superior. 76
Binding is one of the many manners in which a protein may partly or completely accomplish its prescribed function in a biological system. For instance, proteins bind to segments of DNA to control gene expression, enzymes bind to proteins to catalyze reactions, and ligand—receptor binding is responsible for numerous types of cell-to-cell communication. As such, protein binding characterization is a significant area of study and has helped identify pathological cellular pathways. 78,79 Perturbed protein kinase-mediated signaling pathways, for example, have been linked to cancer, diabetes, and inflammation. 78 Working to restore proper signaling, protein kinase inhibitors have emerged as a promising cancer treatment. The rapid screening of kinase inhibitors, using the measured binding constants as an indicator of drug efficacy, is one potential application of nanopores. 14 Binding interactions have also been measured between proteins, 14 a protein and DNA, 80 and a protein and a disaccharide. 81 The protein binding kinetics were measured using α-hemolysin as a site for protein binding or as a steric barrier to force proteins from their binding partners. 14,22,80 These techniques provide ways to screen binding affinities of biological and small-molecule therapeutic agents more quickly than traditional microwell assays. 82 Additionally, nanopores reveal information potentially hidden by bulk protein measurements. 80 To make these devices truly high throughput and practical for routine use, nanopore arrays or hybrid arrays, as described previously, hold promise for future applications.
A related application that uses protein binding is the use of nanopores as protein biosensors. Protein binding is often described as a“lock and key” interaction 83 referring to the complimentary morphology of the binding sites. The specificity implied by the above metaphor is evidence for proteins' valuable molecular recognition capability, which can be used for biosensing and detection applications within nanopore systems. Biosensing applications are not immediately concerned with the kinetics of binding but rather the detection of specific targets, such as disease-relevant biomarkers. Nanopores have an advantage over other detection techniques, such as immunoassays, due to their superior sensitivity and lack of costly labeling. 84,85 Biosensing experiments vary in design and detection scheme. Initially, α-hemolysin-based biosensing was limited to proteins smaller than the pore's recognition site (1.5 nm). 23 This limitation was overcome by using a polymer (polyethylene glycol) chain for protein detection. 86 The polymer chain could either stay in the trans chamber, allowing current to flow freely, or span the pore's length leaving the binding site in the cis chamber and thereby partially blocking ionic current. When the analyte protein would bind to the polymer chain, the chain would be fixed in one of these two states allowing for the presence of the protein to be detected. A second scheme places a binding element very close to the opening of the pore using either covalent chemistry 82 or by incorporating the binding element into the α-hemolysin genetic code (Fig. 2A). 14 Using this binding site configuration, the protein analyte binds close to the pore entrance and blocks ion flow through either steric or electrostatic interactions. A third detection scheme is based on measuring a reduction of translocation events on adding an analyte-specific binding molecule, which creates a complex larger than the pore's diameter. 22 For example, this detection scheme has been demonstrated by detecting the prion protein responsible for Creutzfeldt—Jakob disease. 85 Because more than one biomarker may be important to diagnosing disease, further development of multianalyte nanopore analytical systems is one of the future works awaiting this field. 22
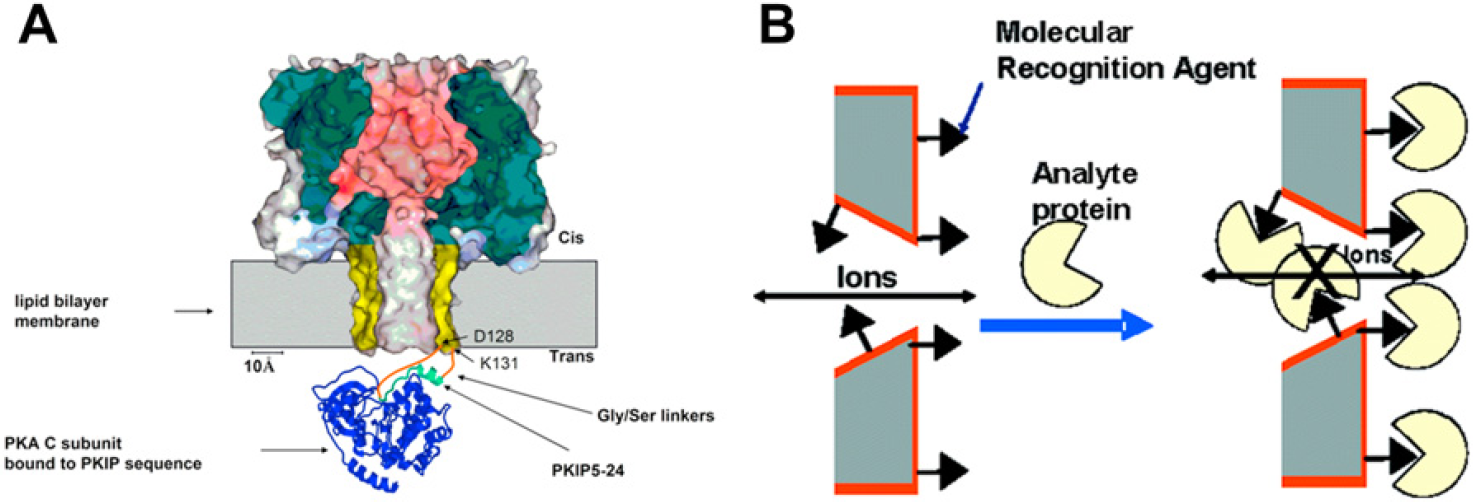
The biological pore, α-hemolysin, is shown on the left and has been modified to contain a protein binding site on the trans side of the lipid bilayer. This binding site is used to transiently hold a protein analyte near the pore's ion-conducting channel which partially blocks the flow of ions through the pore. This allows the protein analyte to be detected in solution with great specificity and probe the binding kinetics of the protein. The solid-state nanopore on the right, made from a gold-coated polymer membrane, demonstrates permanent protein binding to a molecular recognition agent attached to the pore and membrane surface. Once the protein analyte has bound sufficiently to these binding sites, the protein partially occludes the pore and hinders the flow of ions through the pore. (A) Reproduced with permission from Ref. 14. Copyright (2006) Wiley-VCH Verlag GmbH & Co. KGaA. (B) Reproduced with permission from Ref. 90. Copyright (2005) ACS.
Most of the benefits that solid-state pores hold for DNA sequencing are similarly true for protein analysis. For example, fixed pore position makes interrogating protein translocations using secondary single-molecule tools much simpler. Controllable pore dimensions allow for the study of proteins irrespective of their molecular size and can be used to unfold proteins to varying degrees. Lacking the need to prepare lipid bilayers will facilitate the incorporation of such pores into ready to use lab-on-a-chip devices. Finally, much higher breakdown potentials allow probing protein unfolding at greater forces using higher strength electric fields. In spite of these advantages, the design of solid-state nanopores specifically for protein binding has been challenging due to the inability to functionalize the pore with a single reactive group allowing for single-molecule binding. 84 As previously mentioned, current chemical surface modification techniques of solid-state pores do not compare to the single-site mutagenesis of α-hemolysin. 87 –-89 The goal of this research is to have proteins bind transiently such that during each binding event the rate constants can be recorded. Although recent attempts have demonstrated the ability to bind proteins to the pore surface, the binding involved many proteins due to the uncontrolled attachment of binding sites. As a result, the protein binding affinities (i.e., rate constants) could not be obtained using this experimental setup (Fig. 2B). 90 Protein folding studies on the other hand have been accomplished using solid-state nanopores. Talaga and Li were able to detect three different folding states of β-lactoglobulin. 76 In their experiment, points in which the protein would stop moving (i.e., stall) inside the pore were used to distinguish the folding state of a protein and differentiate between proteins. The mechanism for stalling was suggested to be due to a net zero charge of the amino acids residing inside the pore. 76 Therefore, the thickness of the solid-state membrane is a critical component of designing such an experiment because this determines where and how many protein stalling events occur. In summary, solid-state nanopores have an advantage in studying protein folding as their geometry and surface chemistry can easily be modified. On the other hand, the current technical challenge of adapting a pore with a single binding site is hindering solid-state nanopore protein binding advancements. In the immediate future, research is likely to focus on the folding characteristics of different proteins in integrated single-molecule systems providing proof-of concept techniques to be used by proteomic researchers.
Ultrafine Molecular Sieving
Recently, nanopore arrays as platforms for biomolecule separation have been investigated 16,91 owing to their inherent advantages, such as better separation efficiency and higher throughput. 92 –97 While conventional separation techniques, such as gel electrophoresis use an isotropic matrix with randomly distributed pores, the geometry of micro-and nanofluidic systems can be controlled precisely thus providing finer control over how macromolecules interact with such systems. 93,95 Scaling lengths down to the nanometer regime also greatly enhances the surface area to volume ratio, thus making surface governed phenomenon especially dominant. This makes it possible to achieve more efficient anisotropic separation that is not possible at the macroscale. 95 Owing to such advantages, several micro-and nanofluidic-based separation devices have been developed and extensive reviews on the subject exist. 93 –95 The ultimate aim of all such devices is to allow for faster and more efficient separation as compared with currently used techniques. Similarly, nanopore array membranes that separate macromolecules in much the same way as conventional dialysis or ultrafiltration membranes, have been reported to have better resolution and higher throughput. 16 It is also possible to chemically modify the surface of such membranes to tailor them for specific applications. 98 Although such nanopore-based membranes seem like a promising platform for efficient molecular separation, certain practical limitations, such as membrane fragility, and the ability to integrate such membranes in high throughput continuous flow devices, need to be overcome.
Recently, Striemer et al. 16 have reported the fabrication of robust, porous nanocrystalline silicon (pnc-Si) membranes using standard microfabrication techniques. Their membranes have average pore sizes ranging from 5 to 25 nm and are only 15-nm thick (Fig. 3A, B). Despite their ultrathin nature, these membranes are relatively robust, withstanding up to an atmosphere of differential pressure. Using such membranes, they demonstrated for the first time separation of bovine serum albumin (molecular weight [MW] = 67 kDa) and immunoglobulin-γ (MW = 150 kDa), two proteins that only have a twofold difference in their MW, using steric effects. This is a significant improvement over conventional dialysis membranes that can only resolve proteins with at least a 10-fold difference. Moreover, they achieved separation rates an order of magnitude faster than conventional dialysis membranes. 16 In an earlier study, Tong et al. 99 fabricated an ordered array of extremely uniform 25-nm pores on a robust 10-nm thick silicon nitride membrane. Since then, various different protocols to fabricate nanopore arrays on silicon-based membranes were developed 100,101 and all these can be used for macromolecule separation as well. Although in the case of Striemer et al., separation was governed by diffusion; it should be possible to separate molecules under the effect of an electric field or, depending on membrane robustness, hydrostatic pressure to increase throughput.
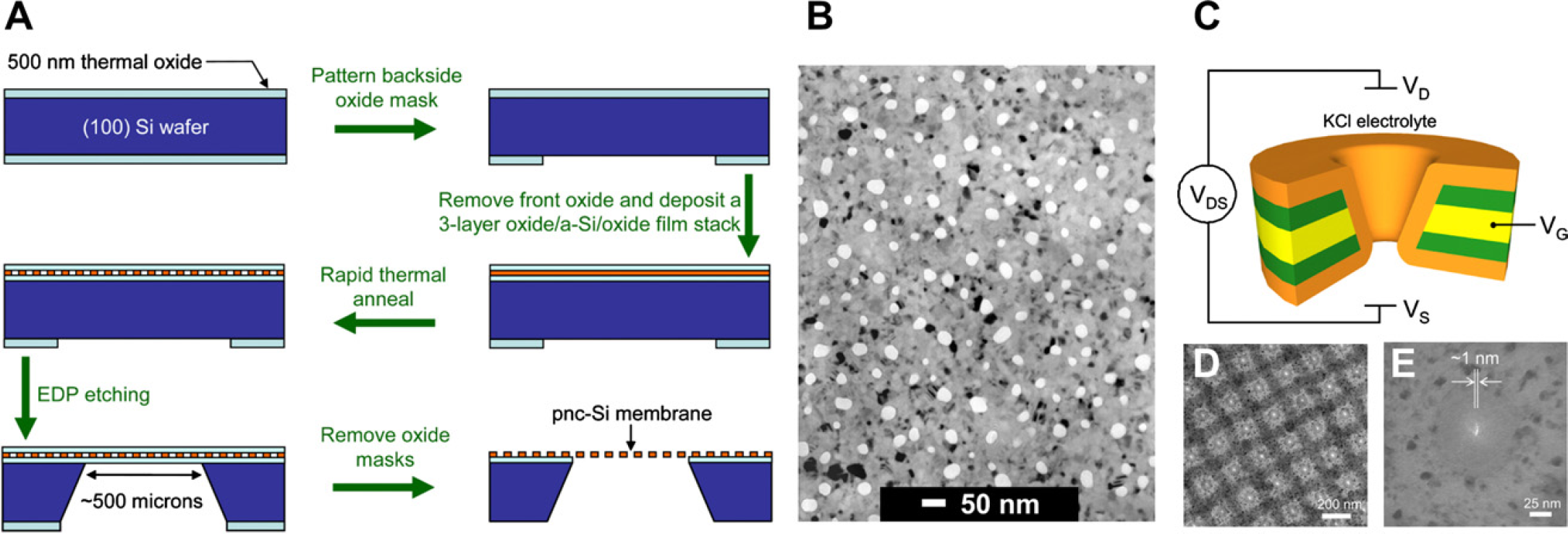
A) The fabrication process for the pnc-Si nanosieves. 16 (B) TEM image of the nanosieves shows pores with a narrow size distribution in the 15-nm thick pnc-Si membrane. (C) Schematic of the setup used by Nam et al. to manipulate the surface charge of the nanopore membrane. 18 (D) and (E) TEM images of the nanopore array and an individual nanopore, respectively. Figures A and B Figures C-E Reprinted with permission from 18. Copyright (2009) ACS. Figures A and B, reprinted by permission from Macmillan Publishers Ltd. Nature (Ref 16), Copyright (2007). Figures C-E reprinted with permission from Ref 18. Copyright (2007).
Another critical parameter that can greatly influence the separation through nanopore arrays is the surface charge of the membrane. 102 Instead of using the nanopore membrane as a passive filter, Nam et al. have fabricated a nanofluidic device using an electrode-embedded nanopore array that makes it possible to actively control the surface charge of the nanopore membrane (Fig. 3C-E). 18 Like a conventional field effect transistor (FET), their device consists of source and drain electrodes, to establish an electric field across the membrane and a gate electrode to finely tune the permeability of positively and negatively charged ions through it. Using this setup, it is possible to control the concentration of different ions within the nanopores and even completely exclude one type of ions. 103 Similar setups, called flow FETs, have been used in the past in conjunction with capillary electrophoresis 104 and microchannels 105 to achieve better molecular separation. These principles could be applied to nanopore arrays as well to carry out charge-based separation through them.
It is possible to enhance the selectivity of nanopore arrays by chemically functionalizing their surface as well. 17 Although currently no such work has been reported with ultrathin nanopore array membranes, Dai et al. 98 functionalized commercially available nanoporous aluminum-coated silicon membranes with antibodies and detected fluorescently labeled antigens in concentrations as low as 0.02 ng/mL. Moreover, chemical modification of individual nanopores 106 and other microfluidics channels 107 is actively being studied. These techniques could directly be applied to nanopore arrays as well to tailor them for specific separation or analytical applications.
The use of nanopore array-based membranes for separation is still in its infancy with only a few applications reported so far. Nonetheless, such systems have shown potential as efficient molecular sieving platforms in initial experimental results. Further, significant developments that have been made in molecular separation using other micro-and nanofluidic systems, such as chemical functionalization and surface charge manipulation, can be adapted to nanopore array membranes as well to enhance their functionality.
Conclusions
This review outlines some of the progress made and challenges waiting ahead toward the goal of nanopore-based DNA sequencing at single nucleotide resolution and the emergence of nanopore-based protein characterization and nanoscale sieving. Each of the applications presented are not working technologies but works in progress toward exciting new and high impact bioanalytical methods or complements of such. Although much progress has been made since the pioneering work by Kasianowicz et al. the ultimate aim of rapid DNA sequencing is yet to be realized. Overcoming the hurdles toward achieving this goal has given new insights that have inspired the use of nanopores for different applications, such as ultrafiltration and protein analysis and will continue to into the future.
Competing Interests Statement: The authors certify that they have no relevant financial interests in this manuscript.
