Abstract
Two mechanical lysis devices have been developed as compact, robust components to provide rapid sample preparation for nucleic acid diagnostic systems. One such component, known as the Micro Bead-Beater™ (μBB™, BBTM, Claremont BioSolutions, Upland, CA), is a compact device that is capable of ultrarapid lysis (>90% lysis in 30 s) of micro volumes (<80 μL) ofBacillus spores in a continuous-flow format or in a disposable single-tube format. The μBB is also capable of processing much larger volumes of solutions containing spores or vegetative cells using a continuous-flow mode. A second mechanical lysis device designed as a disposable component is the microfluidic bead blender, which uses a small electric motor to spin vanes within the bead-laden solution. DNA quantification results using dsDNA-binding fluorescence dyes and real-time PCR are presented, comparing the lysis of Bacillus subtilis spores using the μBB™ with other well-known lysis techniques. Nanoscale imaging results obtained using scanning electron microscopy and transmission electron microscopy on B. subtilis spores lyzed using the μBB™ are also presented
Introduction
Genetic analysis of biological samples has been revolutionized by the ability to amplify specific sequences of nucleic acid contained in the sample. However, gaining access to these nucleic acids involves complex protocols that are typically performed in laboratories using many steps to lyse the cells and purify the nucleic acid. Typical commercial kits for DNA extraction require many manual steps including introduction of the sample, enzymes, and buffers interspersed with various mixing, incubation, centrifugation, and tube transfer steps. For example, the commonly used purification kits from Qiagen require approximately 10 pipetting steps, 3 mechanical mixing steps, 6 centrifugation steps, and 10 tube transfer steps. 1 Qiagen and others offer products to automate these preparation steps but these are typically not well-suited for emerging point-of-use diagnostic applications that require compact, automated, robust operational settings. Examples of these types of nonlaboratory settings include both indoor and outdoor environments considered for various biothreat detection systems and health care settings where rapid, low-complexity diagnosis of infectious disease is required.
Various thermal, electrical, and chemical means of lysing biological samples have been developed, characterized, and compared. 2 5 However, the focus of this report is on mechanical lysis systems that are typically required to gain access to the nucleic acid in hard-to-lyse samples, including biothreats such as spores of Bacillus anthracis. Various means of mechanical lysis have been developed using commercial instruments including the use of a French press and a variety of so-called bead beaters, disruptors, and sonicators (e.g., FastPrep Cell Disrupter 120, Qbiogene, Eschwege, Germany). Only a few of these are robust enough to lyse bacterial spores (e.g., Mini Bead-Beater, Biospec Products Inc., Bartlesville, OK) and none are readily available for integration into compact, automated diagnostic systems. 6 Most of the protocols that have been developed for hard-to-lyse samples are not suitable for routine automated use in biodefense and other applications. 6 12 We describe in this report our efforts to develop several compact mechanical lysis components that can be used to satisfy biodefense requirements for compact, rapid, and robust genetic-based pathogen detection.
The requirements for pathogen lysis in biodefense systems are mission dependent. 6 In some systems, there is a need for lysing apparatus and methods that are more efficient and/or faster than current commercial lysing instruments. The type of sample to be analyzed, the frequency of sample acquisition and analysis, the throughput of the system, and the sample volume and medium must be specified for a particular system. For stand-alone biodefense systems performing automated genetic pathogen identification, there is a clear need to minimize sample volume to maintain low reagent volumes, lower costs, minimize wash-buffer volumes, and to simplify the containment and disposal of postamplification solutions. The designs of mechanical bead-beating components are constrained by this requirement of small sample volumes such that the sample must be lyzed using a small volume of beads. In recognizing this primary requirement, we have capitalized on this small volume requirement (currently <80 μL) to develop compact mechanical oscillating/rotating systems that impart sufficient agitation for lysis while ensuring mechanical integrity for long-life operation. This is achieved primarily by imposing a vigorous shear flow within the bead-sample solution through operation at a higher frequency than commercial mechanical lysis systems. Indeed, the components presented in this report have been endurance tested to meet lifetime requirements needed for specific biodefense programs while meeting high lysis efficiency requirements.
Stand-alone, automated acquisition and analysis required in some biodefense applications can be achieved by using flow-through lysing to process many small samples over time. For example, a system could require analysis of samples taken every minute over a much longer period of time (e.g., weeks or months). The challenge in this design is that tubing must interface to the lysis vessel that must oscillate at high frequency, creating potential for mechanical fatigue. We have developed customized cartridges that adequately contain the beads using stainless steel meshes while allowing flow-through operation using standard Tygon tubing (0.02 in. internal diameter, Saint-Gobain Performance Plastics, Akron, OH) with tubing reinforcement that provides stress relief to prevent fatigue or breakage.
Principle of Operation
The basis of mechanical lysis is to generate confined periodic flow fields with angular accelerations that provide strong vortical flow fields and large shear rates between beads in the liquid solution. We describe here our qualitative observations that led us to the development of the lysis systems. Two prototype mechanical systems were initially developed to characterize lysis in oscillating flow fields. The first of these systems moved the sample back and forth with a stroke of approximately 1 cm along a rail using a linkage driven by an electric motor. The second mechanism created an arc motion, also with a stroke of approximately 1 cm (2.5 cm radius of gyration with near 20° peak-to-peak angular amplitude). Using the PCR and PicoGreen assays described below, it was determined that oscillation of the tube along a single linear axis, even at high frequencies (100–200 Hz) resulted in very little lysis of biological samples while oscillation in an arc motion generated lysis superior to commercially available bead beaters. High-speed movies at 4000 frames/s (FASTCAM-PCI, Photron USA Inc., San Diego, CA) using an optically clear 200-μL polypropylene tube were taken to visualize the bead motion. Although only crude quantitative analysis of bead velocities, shear, and flow-field characteristics was performed, it was qualitatively very clear that linear motions result in a periodic concentration of the beads followed by expansion of the beads away from each other but that there is little relative motion of beads that is not along the axis of motion. In contrast, within a cartridge that moves on an arc, the beads are seen to compress to high density just as a strong swirl is induced. With optimization, this principle has been used to develop very effective lysis systems.
Based on these principles, the first system that was designed, developed, and tested at Keck Graduate Institute in support of several biodefense programs required compact, unattended, and robust lysis of over 10,000 samples over a period of several weeks. A device was developed that met these requirements and it was named the Micro Bead-Beater, or μBB and is now sold by Claremont BioSolutions, Inc. (Upland, CA) as a commercial product. A graphical layout of the system is shown in Figure 1 and a photograph of the top part of the device is shown in the upper part of Figure 2 during oscillation. In the simplest mode of operation, a single 200-μL polypropylene tube filled with beads and sample solution can be inserted into a tube holder and the sample lyzed by oscillation of the unit for a duration of tens of seconds. Flow-through lysing operation typically involves using a syringe pump or peristaltic pump to provide sample solution at a rate of 10s to 100s of microliters per minute through a cartridge containing beads. Depending on the type of downstream assay being performed, a variety of additional sample preparation steps may be required. A variety of sample types have been pumped through the bead beater to date and in some cases downstream size exclusion chromatographic separation has been used to remove protein and cell debris before performing DNA analysis. One study involved the collection of airborne biological samples collected into deionized water, requiring filters to remove airborne dust.
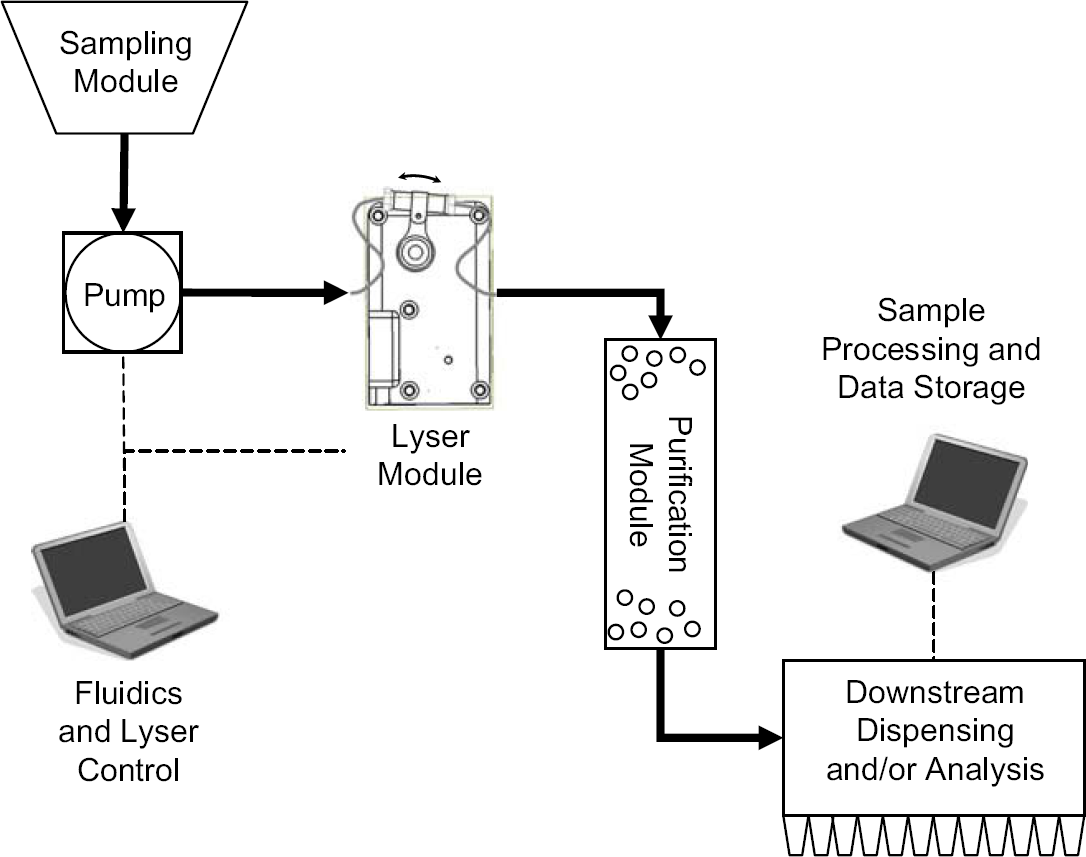
Automated mechanical sample lysis system: the Micro Bead-Beater or μBB. Operation with either continuous flow or with a stopped-flow control may be used.
The main body of the bead beater is a rectangular aluminum box containing the mechanical components and is about the size of a bar of soap (1.5 × 2.25 × 3.75 in.). It uses a four-bar linkage design driven by a brushless DC motor to oscillate either off-the-shelf 200-μL PCR tubes or custom flow-through cartridges at frequencies of 100–250 Hz back and forth along an arc of 1 cm length with a radius of curvature of 25 mm. The flow-through cartridge shown in the bottom of Figure 2 is an easily removable PEEK cartridge filled with 100-μm diameter zirconium-silica beads that are contained between stainless steel meshes at the inlet and exit.

Photograph of Micro Bead-Beater in motion (top) with disposable cartridge, tubing, and bracket assembly (bottom). Attachment of a standard 200-μL polypropylene PCR tube requires a different bracket (not shown).
The second system described in this report meets an entirely different set of requirements with disposability and small volume lysis of bacterial spores of primary importance. Two versions of the microfluidic bead blender are shown in Figure 3. The first image shows a partially assembled version of the bead blender with the tubing inlet and exit at 90° angles. The electric motor is 4 mm in diameter and the motor-impellor rotation axis is the same as that of the exit tube. The custom vaned-impeller of Nylon 66 is attached to the motor and is conical in shape with a taper from a maximum vane diameter of 4 mm at sample entry location to 1 mm diameter at the sample exit. The tubes fit into entrance and exit ports, which are milled into acrylic along with chambers for housing the impellor and flow channels. The second image shows six bead blenders with the inlet and exit ports along a single axis (horizontal) and the motor/impellor at 90° orientation (vertical from top). The impellers used in the bead blenders in the second image are laser machined from acrylic.

Microfluidic bead blenders. Early prototype showing a six-vaned conical impeller mounted to a small electric motor before insertion and sealing of the motor/impeller assembly into the orifice at the top of the rectangular body. U.S. 10-cent coin shown for size reference (top). The entrance and exit of the sample may be through either tube. Six fully assembled bead blenders. U.S. 1-cent coin shown for size reference (bottom).
Experimental Methods
At Keck Graduate Institute (Claremont, CA) we have developed a series of experiments which were designed to systematically compare three different bead-beating strategies as rapid methods to lyse Bacillus subtilis spores. KGI has developed a protocol for the acquisition and treatment of B. subtilis spores that is used as a model for B. anthracis, obviating the need to perform lysis experimentation in bio-safety level 2 (BSL-2) or more restrictive laboratories. The specific lysis methods used for comparison were as follows: the μBB and two commercial-off-the-shelf technologies; (1) traditional bead beating using the industry-standard Biospec Mini Bead-Beater and (2) lysis by sonication with beads in a 100-μL Cepheid SmartCycler PCR tube (Cepheid, Inc., Sunnyvale, CA) and a sonication horn (VibraCell VCX-130, Sonics & Materials, Inc., Newtown, CT) placed on the exterior against the tube wall with a uniform 3 N force. A summary of the major parameters of each experimental system is presented in Table 1. Note that power supplies constitute a major portion of the mass budget and that the μBB requires much lower power than the other methods. In these experiments, we used a large 2A DC power supply with mass of 1240 g but this could easily be replaced with four AA batteries weighing 95 g.
In these experiments, we used both real-time PCR and intercalating dsDNA-binding dye to assess the lysis efficiency of DNase-treated B. subtilis spores that were placed at concentrations of 10 6 spores/μL in deionized water. Lysis efficiency was determined through the use of the Quant-it PicoGreen dsDNA Assay Kit (Invitrogen Corp., Carlsbad, CA) per the manufacturer's specifications. The real-time PCR experiments were performed using a LightCycler (Roche Applied Science, Indianapolis, IN) and the FastStart DNA Master SYBR Green I reagents. The primers used to specifically identify B. subtilis were 5'-CCGGGCGGTT ATGTATATTTA-3' (forward) and 5′-GTTGAAAGAGT AAGAAGTGACA-3′, producing an amplicon of 467 bp at location 54883 of the sequence. In PCR experiments, the crossing-threshold values (Ct values) were obtained by determining the thermal cycle number during real-time amplification for which the fluorescence level first exceeded 10% of the maximum signal. The PCR protocol used for this study consisted of an activation step of 95°C for 10 min, followed by cycles consisting of 95°C for 5 s, 61 °C for 12 s, and 72 °C for 23 s. Amplicon fidelity was confirmed through a standard melt curve analysis performed on the LightCycler following PCR, ranging from 37 to 95 °C at a rate of 0.1 °C/s.
The μBB™ experiments used standard 200-μL polypropylene PCR tubes packed with 300 mg of 100-μm diameter zirconium-silica beads (Biospec Products, Inc., Bartlesville, OK), whereas the 2-mL tube used in the Biospec Mini Bead-Beater was packed with 3 g of the beads and the 100-μL SmartCycler tube contained 100 mg of the beads.
Bead-Beating Lysis Performance Results
A preliminary experiment was performed to characterize the dependency of the real-time PCR crossing-threshold (Ct) cycle-time values on the concentration of DNA, using both purified genomic DNA and using the solution obtained directly upon lysing spores. As seen in Figure 4 (top), a pattern of increasing Ct values is observed when a dilution series of standard B. subtilis genomic DNA (ATCC, Manassas, VA) is performed. A similar dilution series for bead-beaten samples was obtained for DNase-treated B. subtilis spores, which were bead beaten for 3 min in the Biospec Mini Bead-Beater and then immediately diluted to 10 4 , 10 3 , and 10 2 spores/μL solutions. As seen in Figure 4 (bottom), the data show a statistically significant increase in Ct value (an indication that there is less DNA available) for each 1:10 dilution of spores, ranging from approximately 29 cycles for 10 5 spores/μL to about 37 cycles for 10 2 spores/μL. These two graphs are generally consistent, considering that a single B. subtilis genome is approximately 5 femtograms (fg), which means that 500 fg DNA should correspond to approximately 10 2 lyzed spores and that 500 pg DNA should correspond to approximately 10 5 lyzed spores. In both graphs, the Ct values are seen to increase by approximately nine cycles for three orders of magnitude decrease in concentration of either genomic DNA or bead-beaten spores.
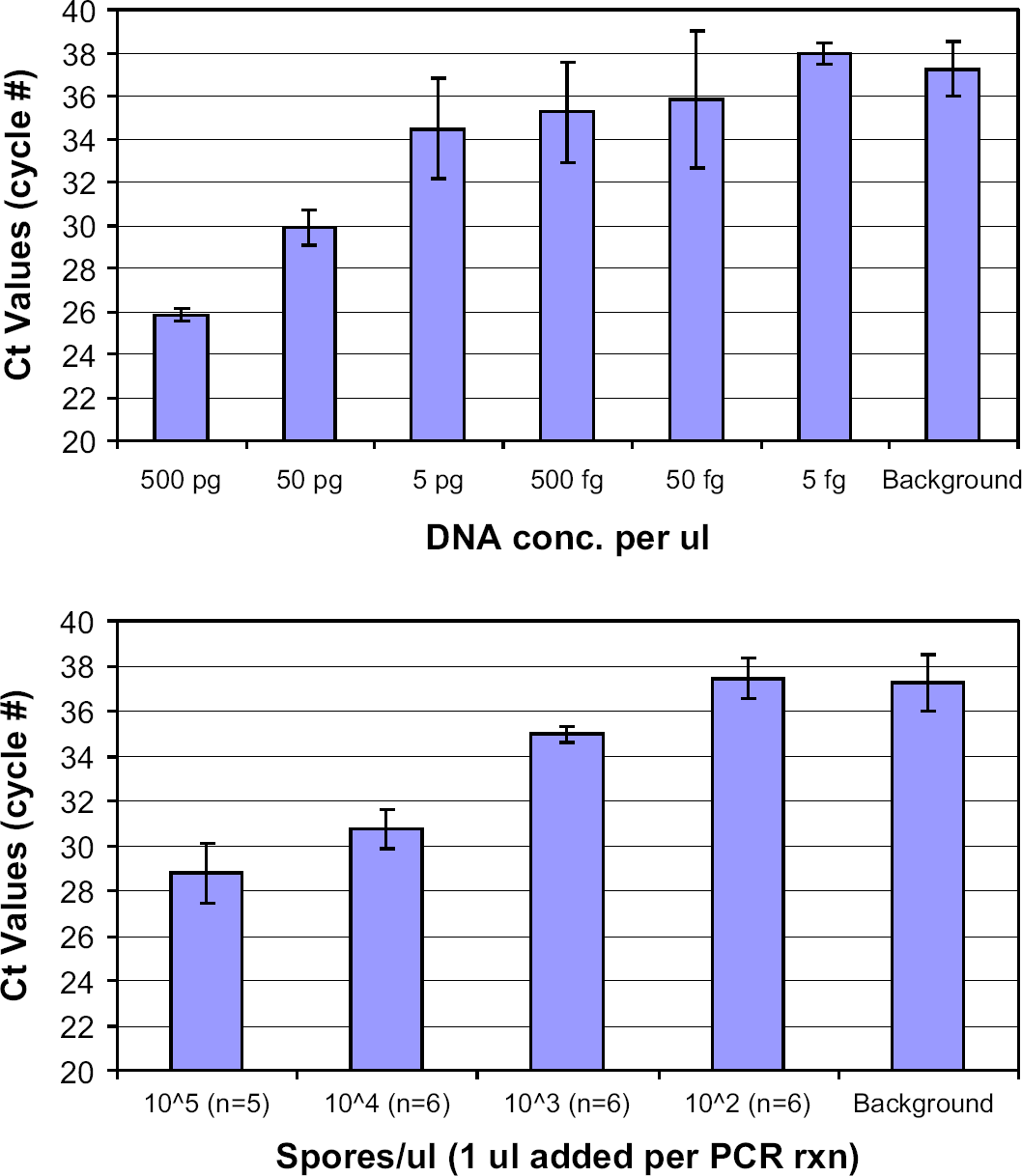
Dependence of PCR crossing-threshold values for ATCC genomic DNA performed with three replicates of each concentration (top) and for Bacillus subtilis spores (bottom) after 3 min of bead beating in the Biospec Mini Bead-Beater as a function of initial concentrations.
With this basic PCR information, we proceeded to compare the three systems described in Table 1. The characteristic performance of the μBB and the Biospec instrument is shown in Figure 5, which plots the PicoGreen assay fluorescence as a function of the duration of bead beating. This fluorescence is used as a measure of lysis efficiency because the dye binds preferentially to dsDNA that is liberated upon lysis. As seen in the figure, the PicoGreen results obtained for the 3-min duration are taken as 100%, such that all other values represent the lysis relative to this condition. The results clearly show that the lysis in the μBB device occurs much more quickly than the Biospec instrument. The data consistently show that it takes less than approximately one-third of the time to expose an equal amount of dsDNA in the μBB than in the Biospec Mini Bead-Beater. The μBB data indicates that approximately 50% lysis is accomplished in 15 s and 90% lysis in 30 s of bead beating. The PicoGreen results show performance of the Cepheid tube sonication method as being comparable to, or less efficient than, the Biospec results. However, this result is misleading in determining lysis efficiency for this method; a point that becomes clear upon review of the PCR results presented in the following paragraph.
Comparison of experimental lysis systems

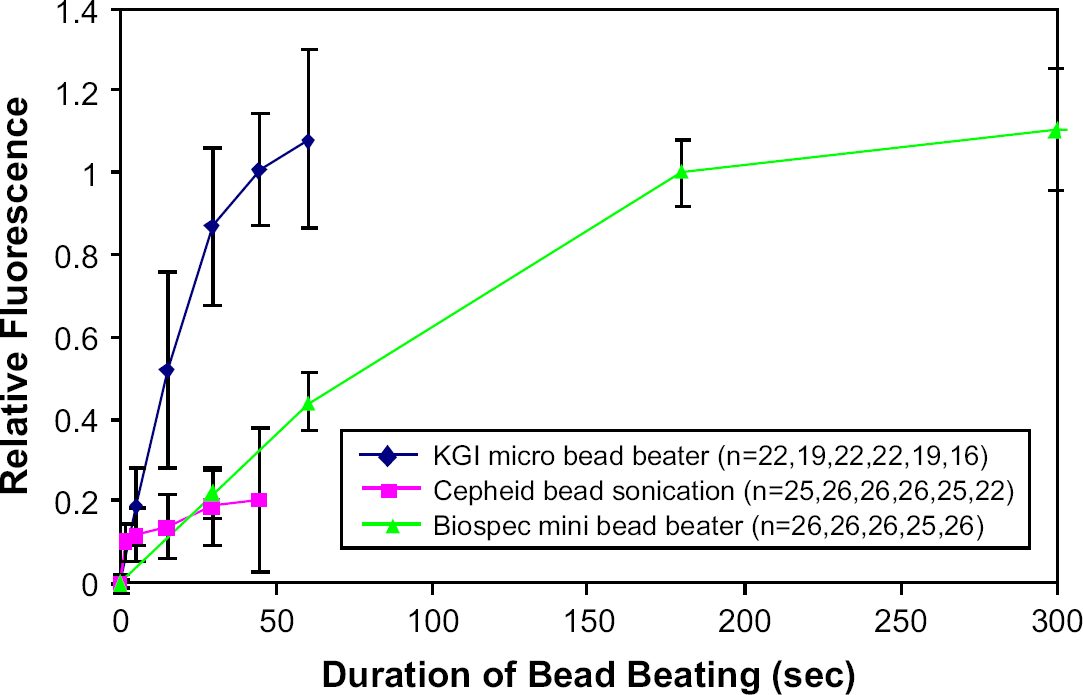
Dependence of lysis efficiency, as determined by the PicoGreen assay, as a function of duration of bead beading (time points are 0, 5, 15, 30, 45, 60, 180, and 300 s).
In applying the real-time PCR assay to the bead-beaten spore samples, we use low Ct values as a measure of efficient lysis because availability of larger amounts of DNA for PCR amplification should be the measure of efficiency for nucleic acid diagnostic systems. As shown in Figure 6, we observe that the μBB again performs much faster lysis than the Biospec, providing lower Ct values for any given duration of bead beating. It is not clear why the 60-s duration for the μBB yields a higher Ct value than the 45-s duration, but we speculate that this may be caused by rupture or shearing of the DNA into such short fragments that the number of contiguous primer-hybridization sites is reduced.
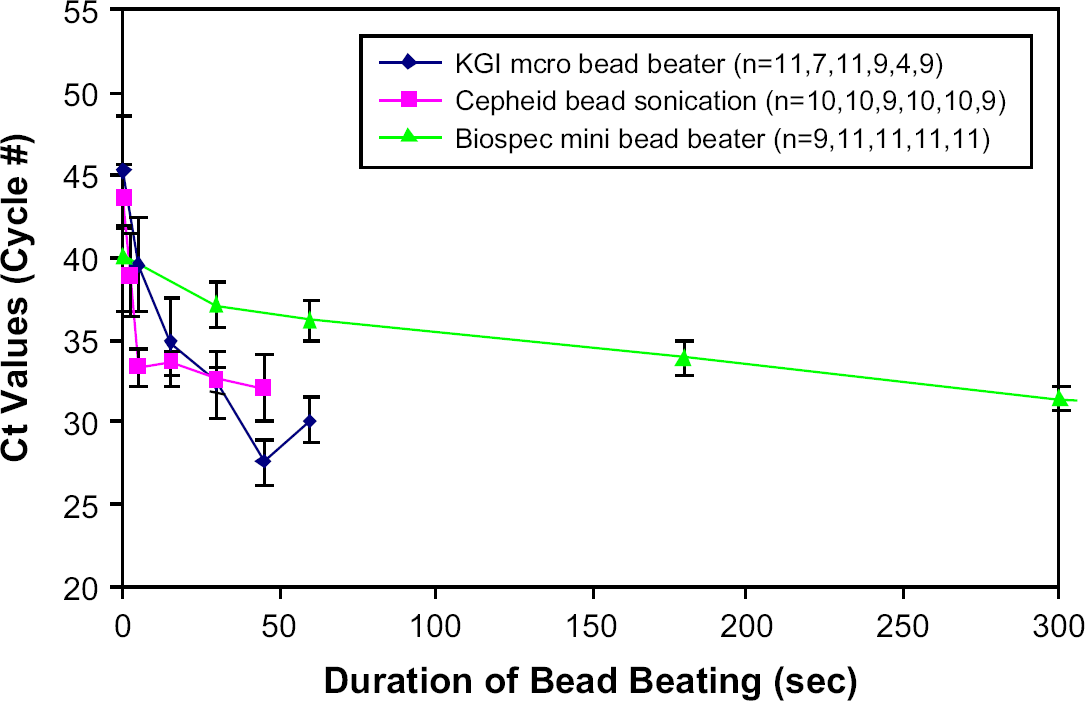
Dependence of lysis efficiency, as determined by the real-time PCR assay, as a function of duration of bead beading (time points are 2.5, 5, 15, 30, 45, 60, 180, and 300 s).
Regarding the Cepheid tube sonication experiments, in contrast to the PicoGreen assay results for this method, the PCR assay suggests that this method yields very good lysis results that are similar to those of the μBB. Indeed, with a few as 5 s of sonication in the Cepheid tube, good lysis efficiency indicated by a Ct value of less than 35 cycles is obtained which is better than the 3-min Biospec value. With increased duration of sonication, however, only modest decreases in Ct value are observed in contrast to the μBB that apparently continues to liberate more PCR-ready DNA. Atomic force microscope images (not shown) of postlysis supernatant show strands of spaghetti-like DNA strung between circular blobs about 20 nm in size that are thought to be ribosomes. In the images of the sonicated spore solutions, there are very few intact ribosomes, whereas in the μBB beaten spore solutions there are many intact ribosomes. With the high power and high frequency of the sonication method, we speculate that rapid access (∼5 s) to the PCR primer-binding sites is provided and that further sonication exposes more primer-binding sites while simultaneously shearing some primer-binding sites, with the result that the lysis efficiency never reaches the level attained by the slightly more gentle μBB after 45 s of bead beating.
The results of bead-beating methods using the two assays as measures of lysis efficiency are supported by direct imaging of bead-beaten spores using electron microscopy. Characteristic scanning electron microscope images of B. subtilis spores, with 6000 × magnification, lyzed for various durations in the μBB are shown in Figure 7. Intact spores are 1–1.25 μm long and 0.5–0.75 μm wide. The time course shows typical spores shaped as oblate spheroid or cylindrical bodies for the first 5 s, followed by the appearance of some hollow or C-shaped bodies after 10–15 s. By 30–45 s, most of the spores appear hollow. The amount of spore debris, defined as pieces substantially smaller than 1 μm in size, increases with each snapshot. In Figure 8, a clump of four or five spores is seen in a transmission electron microscope image with the spore on the left clearly bent into a C shape. DNA coated with ribosomes and other DNA-binding protein is observed to be released on the left and top of the image. These images clearly show the effect of mechanical disruption on the integrity of the spore coats and make clear that access to DNA is provided for both the PicoGreen dye and for the primers, polymerase, and deoxyribonucleotide triphosphates for the PCR assay.

Lysis time course of Bacillus subtilis spores beaten in the Micro Bead-Beater visualized using a Hitachi S520 scanning electron microscope. Magnification 6000x. Spores are 1–1.25 μm long and 0.5–0.75 μm wide.

Bacillus subtilis spores beaten in the Mini Bead-Beater for 3 min visualized using a Hitachi H600 transmission electron microscopy. Magnification 25,000x. Spores are 1–1.25 μm long and 0.5–0.75 μm wide.
Microfluidic Bead-Blender Lysis Results
The same assays used to characterize bead-beating performance above were used to characterize the performance of the bead blenders. The bead blenders are designed to hold less than 50 μL of solution and they are used in either a stopped-flow mode or in a continuous flow-through mode. We have used the continuous flow-through mode to lyse 1 mL of sample in ∼90 s. The Biospec 100-μm zirconium-silica beads are contained in the bead-blender cartridge such that they are present in the impellor region as well as upstream and downstream of the impellor. The motors operate at 4 V DC and consume 400 mW of power at approximately 40,000 RPM (1/3 of maximum unloaded speed of 120,000 RPM). The PicoGreen assay results are shown in Figure 9. The results demonstrate 50% lysis in less than 30 s and 75% lysis in 45 s of bead blending.
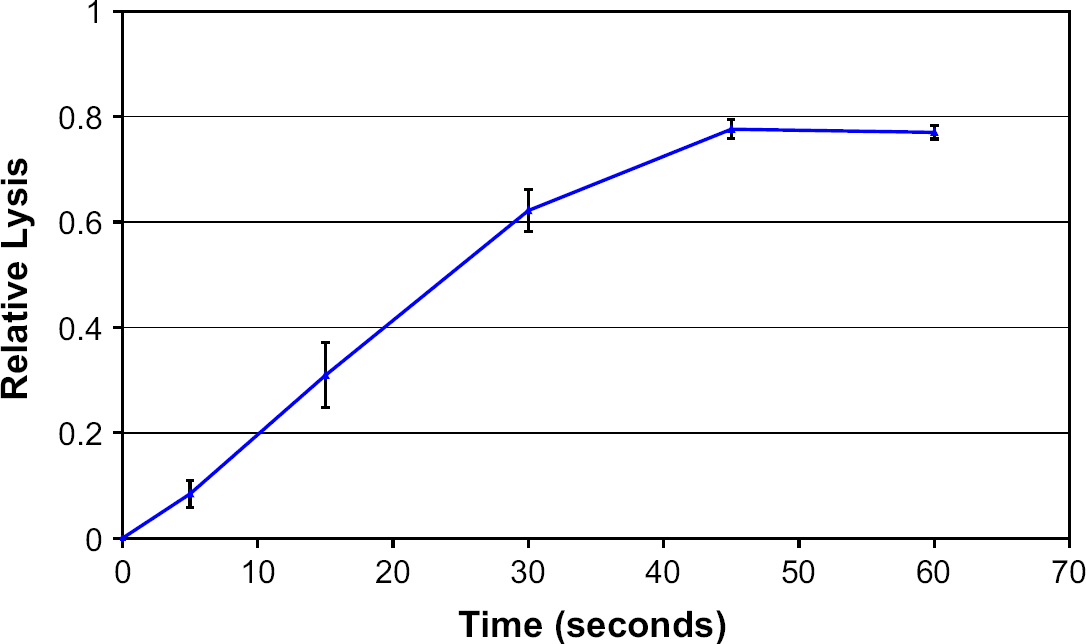
Dependence of lysis efficiency of bead blender, as determined by the PicoGreen assay, as a function of duration of bead beating. Fraction lyzed is relative to the 3-min Biospec value. Samples were run in the blender at 4 V at 100 mA with 180-mg beads and 50 μL of spore solution.
Real-time PCR results also demonstrate rapid lysis in the bead-blender system. The system can be adjusted to attain a desired lysis efficiency by changing the flow chamber volume, the continuous flow rate of solution through the device, and the rotational speed of the impellor. The characteristic residence time of beads within the impellor region is given by the chamber volume divided by the flow rate; lysis efficiency is observed to increase with this residence time for a particular impellor speed. However, the best results seen to date have resulted from a stopped-flow method to increase the residence time of the spores within the chamber. Typically, a spore solution stopped for 45 s within the blender chamber containing 100 mg of beads and operated at 6 V (∼50,000 RPM in solution) will achieve a real-time PCR Ct value equal to, or slightly lower than, the 3-min Biospec Mini Bead-Beater result. Finally, we note that scanning electron microscopy (SEM) images of the bead-blended samples show similar mechanical rupture as observed for the μBB™
Conclusions
We have described two novel devices that can be used as components to provide sample preparation for point-of-use integrated nucleic acid diagnostic systems. The μBB™ and the microfluidic bead blender were characterized using a PicoGreen dsDNA-binding fluorescence dye assay and a realtime PCR assay. Results consistently showed rapid lysis (<1 min) in small volume (∼50–100-μL) samples using these devices. SEM and transmission electron microscopy images of spores bead beaten for various durations showed clear effects of mechanical disruption of the spore coats and exposure of DNA. The small size and low power requirements of these devices make them good alternatives to the use of sonication or commercial bead-beaters for rapid lysis of hard-to-lyse biological samples.
Acknowledgments
We would like to thank Ionian Technologies, San Diego, CA and Dr. Anne Hultgren of The Department of Homeland Security Science & Technology Directorate (DHS—S&T) for their support of portions of this work through TTA2 Rapid Automated Biological Identification System Program (HSHQPA–04–9–004).
