Abstract
Real-time reverse transcription PCR analysis of RNA expression in normal and diseased states is a fundamental technique used in research and screening workflows. However, traditional purification of RNA samples often represents a significant workflow bottleneck, which increases the time and automation complexity required for analysis. We describe the Cells-to-CT technology, a workflow that was designed in conjunction with real-time analysis to completely eliminate the need for traditional purification while maintaining performance relative to purified RNA. High-density sample preparation is accomplished in less than 8 min and is highly amenable to automation. We demonstrate that results are equivalent to RNA purified using traditional methods over a broad range of sample input while offering advantages when dealing with automated liquid handlers.
Keywords
Introduction
The analysis of differential RNA expression that occurs in response to spatial/temporal cues, a wide range of external stimuli, or in reaction to compound exposure is a fundamental technique used in a variety of research and screening methodologies. Real-time reverse transcription PCR (RT-PCR) analysis using TaqMan probes (Applied Biosystems, Foster City, CA) has become a key technology, which is used to analyze RNA expression levels because of the sensitivity, relative ease of use, and flexibility of the technique. Although the assembly of TaqMan RT-PCR reactions and subsequent analyses is amenable to automation and relatively high degrees of throughput, the preparation of representative RNA of sufficient quality and purity often represents a significant workflow bottleneck. Traditional techniques using extremely harsh denaturants, organic extraction, and precipitation provide RNA of extremely high quality but are extremely labor intensive and require significant hardware investments or are not amenable to automation. 1
Sample purification techniques more amenable to automation that involve binding matrices incorporated into porous fibers and placed into a 96-well plate format have been developed for high-quality RNA purification on a variety of liquid-handling equipment. 2 3 However, these formats require additional hardware, such as pumps and associated tubing to provide negative or positive pressure across the membrane, which is used to drive liquids through the binding surface. Filter-based systems rely on an ability to transiently form an appropriate seal across the plate (to maintain vacuum), are susceptible to clogging (leading to nonuniform vacuum or loss of vacuum), and are prone to aerosolization of sample, which can lead to significant cross-contamination.
More recently, a variety of sample purification matrices formed of paramagnetic particles combined with functionalized surface chemistries based on adsorptive, ionic, or pH-dependent interactions are also commercially available. Although these chemistries can be adapted to 96-well processing formats, they are not easily adapted to higher densities and still require significant time to complete. Methods based on magnetic particles require repeated addition and removal of binding, washing, and elution solutions that significantly increase method duration, consumable tip usage, and potential for cross-contamination. Traditional sample purification methods suffer from other technical hurdles, such as the copious use of disposable tips, carryover of harsh denaturants or alcohol into samples, and cross-contamination potential inherent in moving samples or liquids between plates. Most importantly, the binding and release of nucleic acids in techniques based on membrane and magnetic particles is less than 100% efficient, leading to sample loss and potentially bias in the RNA that is recovered.
The Cells-to-CT technology represents the latest evolution of technologies designed to eliminate the need for traditional purification. Enhanced features over our previous technology (Cells-to-cDNA II) include an 80% reduction in processing time, elimination of heating steps, and superior removal of genomic DNA. The Cells-to-CT technology was also optimized in conjunction with real-time RT-PCR master mixes to enable the highest signal dynamic range possible. In this study, we demonstrate how the elimination of processing steps speeds the preparation of samples on a variety of open instrument platforms and that the method is effective at high throughput densities. We also demonstrate how the ability to process directly in tissue culture plates at room temperature without the need to mix samples minimizes instrument complexity, consumables usage, and the potential for cross-contamination.
Materials and methods
Volume manipulations, including the aliquoting of cells and addition of lysis and stop solutions were performed using the Labcyte Deerac (Sunnyvale, CA) HTS noncontact dispenser in 384– and 1536-well plates. Independently controlled channels allow dispensing of reagents at any volume ranges between 50 nL and 50 μL. Addition of reagents for reverse transcription (RT) preamplification and quantitative PCR was done manually with an Eppendorf multichannel pipette or a Beckman Biomek FX (Foster City, CA).
The TaqMan Gene Expression Cells-to-CT kit and Taq-Man PreAmp Cells-to-CT kit (Life Technologies, Carlsbad, CA) come with reagents for tissue culture cell lysis/stabilization, RT, and PCR amplification. In addition, the TaqMan PreAmp kit contains reagents required for the intermediate amplification step, which extends the amount of precious sample. Figures 1 outlines the standard Cells-to-CT workflows; mixing steps were demonstrated to not be required and were eliminated from the automated methods (data not shown).
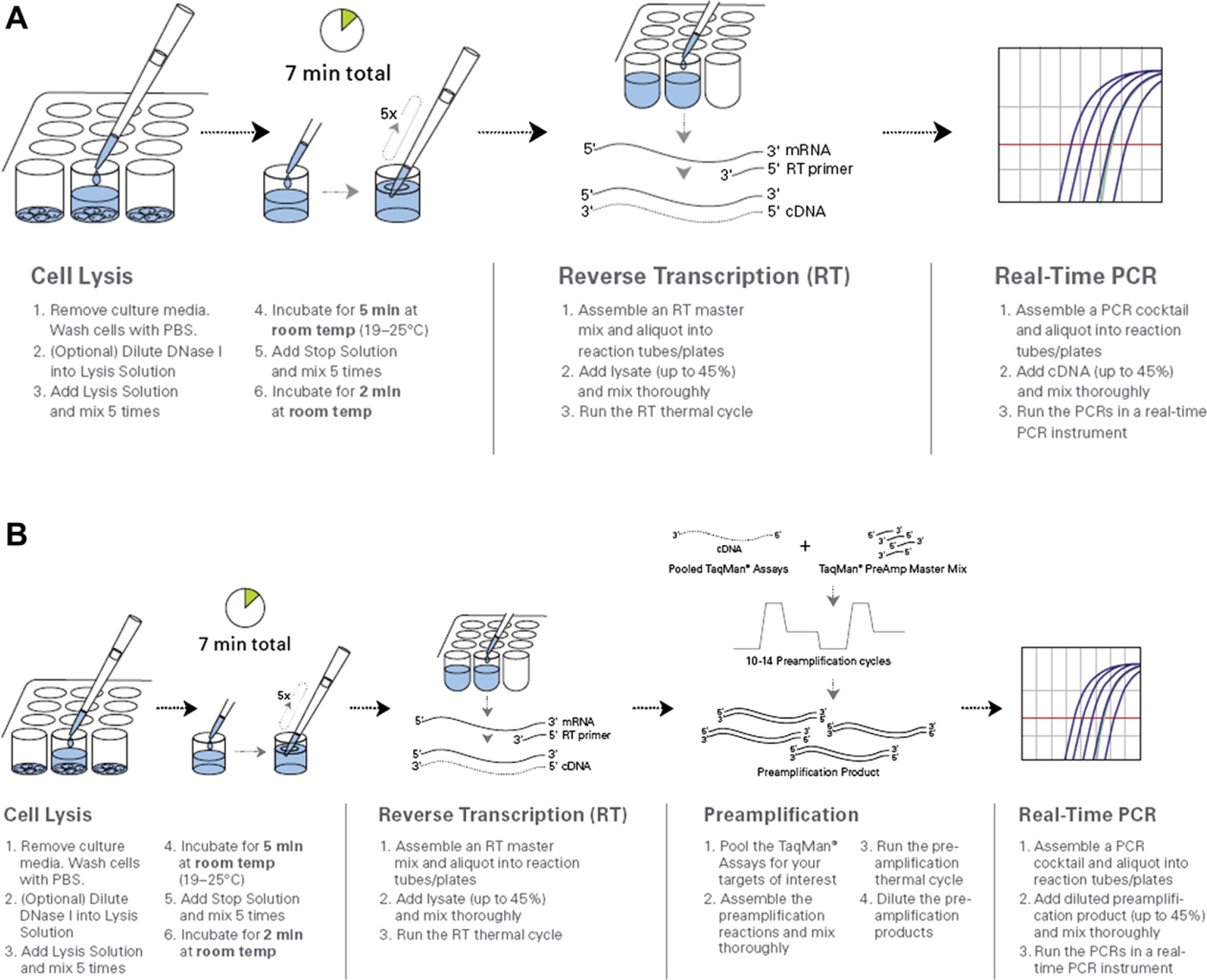
Cells-to-CT workflow. Samples are prepared for RT-PCR in a 7-minute protocol using the Cells-to-CT kit. (A) TaqMan Gene Expression Cells-to-CT workflow is further described in Materials and Methods section. (B) TaqMan PreAmp Cells-to-CT workflow is further described in Materials and Methods section.
When processing 100 or less cells in 96– or 384-well format, culture media were removed, and HeLa cells were washed once with phosphate-buffered saline. Lysis solution (8 μL) containing DNase I (added as per manufacturer's instruction) was added, and the samples were incubated for 5 min at room temperature without the recommended mixing step. Stop solution (1 μL) was added, and the samples were incubated for an additional 2 min at room temperature without the recommended mixing step. The RT reaction was carried out by adding 14 μL of the RT master mix (12.5-μL 2× RT buffer, 1.25-μL 20× enzyme mix, and 0.25-μL nuclease-free water). Samples were incubated for 1 h at 37 °C and 5 min at 95 °C as per manufacturer's instruction. The cDNA samples (2-μL aliquots in 10-μL reactions) were then analyzed by real-time analysis using primers specific for either 18S ribosomal or β-actin mRNA.
For 50 cells in 1536-well format, 2.4 μL of lysis solution (DNase I added) was added and incubated for 5 min at room temperature. Stop solution (0.3 μL) was added and incubated for an additional 2 min at room temperature. Samples were reverse transcribed by adding 5.2 μL of the RT master mix (4-μL 2× RT buffer, 0.4-μL 20× enzyme mix, and 0.8-μL nuclease-free water). Samples were incubated for 1 h at 37 °C and 5 min at 95 °C as per manufacturer's instruction. The cDNA samples were analyzed as previously mentioned.
When processing 2500 cells, media were removed and cells were washed once with phosphate-buffered saline followed by manufacturer's instruction for the Cells-to-CT workflow.
For the 100 cell samples subjected to preamplification, 14 μL of the samples after RT reaction (Fig. 1B) were added to PreAmp master mix containing a pool of seven assays; HMBS, HPRT1, IL17RA, MYC, POLR2A, TFRC, and TGFB1 (15-μL 2× PreAmp master mix and 1-μL 2× assay mix pool). Samples were amplified for 14 cycles as per the manufacturer's instruction. PreAmp samples were diluted 1:5 with 120 μL of Tris—ethylenediaminetetraacetic acid pH 8.0. Samples were analyzed as previously mentioned.
For TaqMan gene expression analysis, TaqMan probes designed to detect targets of interest were used (Table 1). Data were collected on an Applied Biosystems 7900 instrument and analyzed using SDS software version 2.3 (Applied Biosystems, Foster City, CA). Briefly, fluorescent signal generated by the TaqMan probe assay was normalized using passive reference dye contained in each reaction, and the change in normalized fluorescence (Δ
TaqMan gene expression assays used in this study (part #4331182)

Results
Simple Workflow for High Performance Analysis of Small Numbers of Cells
The Cells-to-CT workflow was developed to eliminate RNA purification by generating samples ready for direct real-time analysis. Sample manipulation and potential loss is minimized by eliminating solid phase binding/elution by lysis of cells, performing DNase treatment, and preparing the sample for analysis directly in tissue culture plates (Fig. 1). The inclusion of a preamplification step before real-time analysis also extends small sample amounts (Fig. 1) by generating an intermediate amplification step. The simple workflow (two additions), elimination of heating, and our finding that mixing can be eliminated (data not shown) has generated a sample preparation workflow that is amenable to automated processing of cells at high levels of throughput. By closely coupling Cells-to-CT kit development for use with the specific real-time master mixes present in each kit, we have been able to demonstrate results equivalent or better than traditional RNA purification (Fig. 2). 4
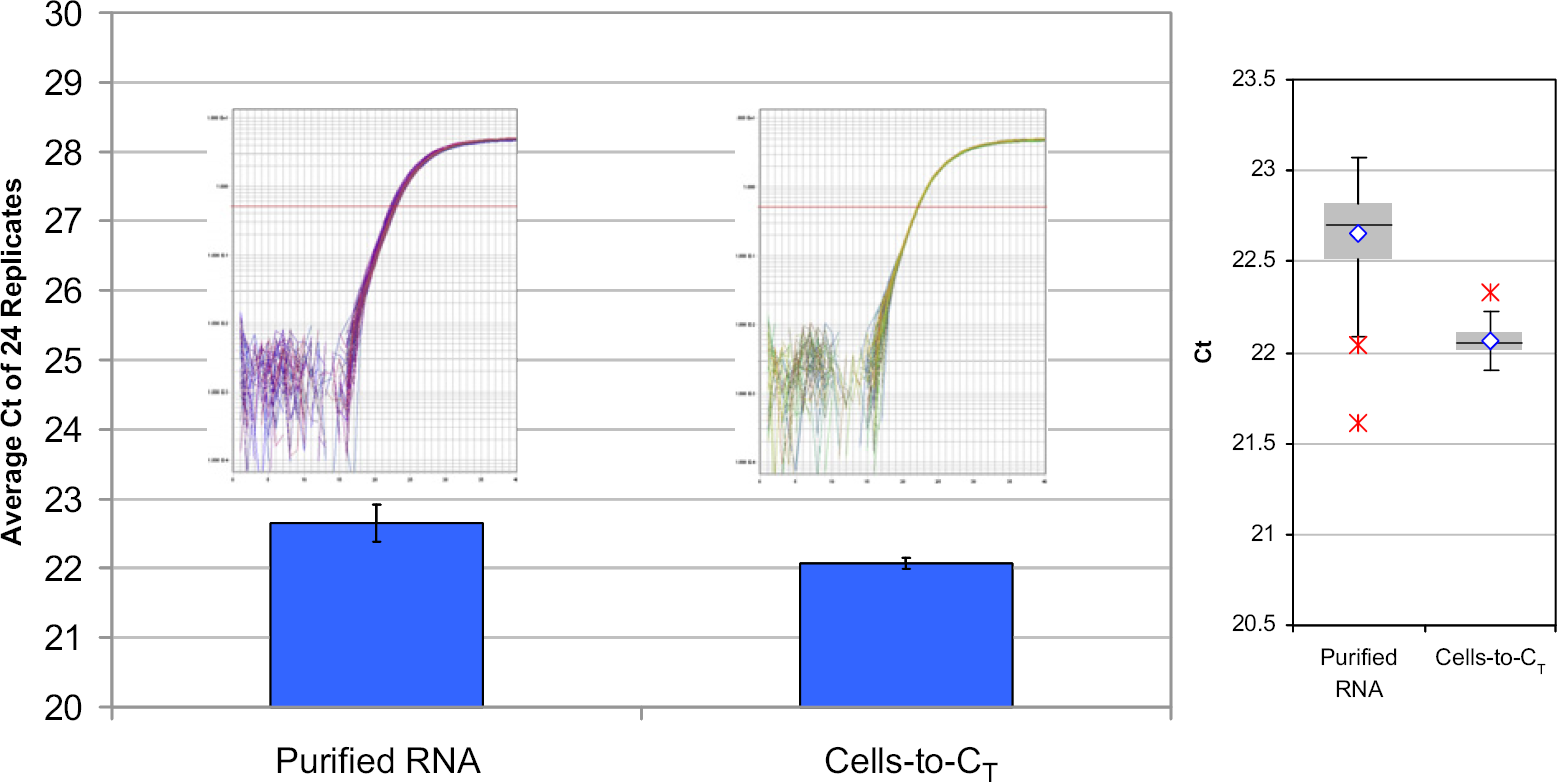
Comparison of traditional purification and Cells-to-CT kit processing of cells. Equivalent numbers (5000 cells/purification) of HeLa cells were processed using a magnetic particle-based method or Cells-to-CT. Prepared samples were analyzed using primers specific for β-actin RNA as described in Materials and Methods section. For each purification method, 24 biological replicates and three technical replicates were performed. Inset, real-time amplification plots of the data depicted graphically.
We implemented the Cells-to-CT system onto the Deerac GX low-volume liquid-handling system (Labcyte, Inc., Sunnyvale, CA) to investigate the performance capabilities of the combined system. The Deerac system is capable of addressing 96-, 384-, and 1536-well plates and can deliver reagent volumes as low as 0.05 μL using an array of eight delivery capillaries. Reagent integration was straightforward using the manufacturer's supplied software. The instrument was used for dispensing Cells-to-CT reagents and tissue culture cells. Typical sample preparation run times were approximately 7.5 min for up to 1536-well plates of cells.
To demonstrate the ability to process low numbers of cells effectively, decreasing numbers of HeLa cells were dispensed into 384-well plates using the HTS instrument, and the samples were processed using the TaqMan Gene Expression Cells-to-CT kit. Analysis of samples prepared from increasing numbers of cells demonstrated a decrease in CT values as expected (Fig. 3). Furthermore, an expected 3.3 CT decrease was observed when the number of cells processed was decreased 10-fold for both assays (compare 200 and 20 cells or 100 and 10 cells).
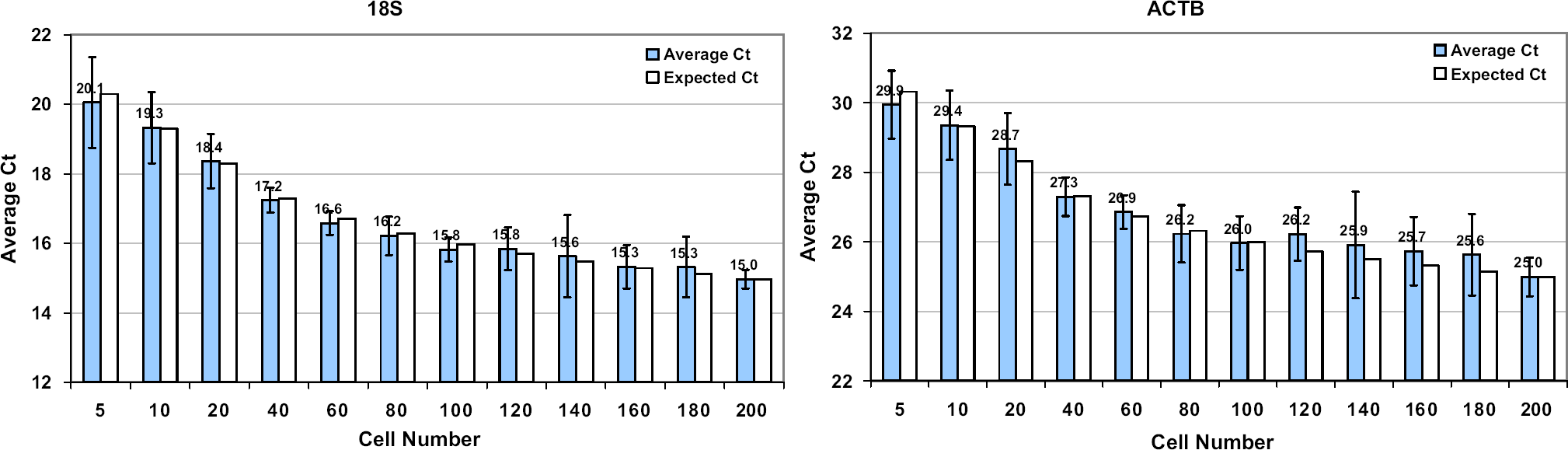
Cells-to-CT kit processing of decreasing numbers of cells. The indicated numbers of HeLa cells were prepared for RT-PCR in a 7-minute protocol using Cells-to-CT kits and analyzed using primers specific for 18S ribosomal or β-actin RNA as described in Materials and Methods section. For each bar, two biological replicates and 16 technical replicates were performed. A cell titration between five and 200 cells was performed.
We next investigated the interplate variability observed using the HTS instrument with Cells-to-CT kit sample preparation. A fixed number of cells (100 cells/well) were dispensed into duplicate 384-well plates, and the samples were prepared using the Cells-to-CT kit method. Technical duplicates were analyzed for each sample by real-time analysis using primers specific for either 18S ribosomal or β-actin mRNA (Fig. 4, panel A). The results from both targets demonstrate a high degree of consistency within a plate (indicated by low standard deviations) and between plates (indicated by similar CT values). Box plots for the data confirm the consistency of the data (Fig. 4, panel B). The ability to prepare samples for real-time analysis in 1536-well plates was also demonstrated. Equal sample volumes containing 50 cells were dispensed into 1536-well plates, and the plates were processed using the Cells-to-CT kit procedure. Analysis using probes specific for β-actin demonstrated that a high degree of reproducibility was possible (Fig. 4, panel C), with a mean
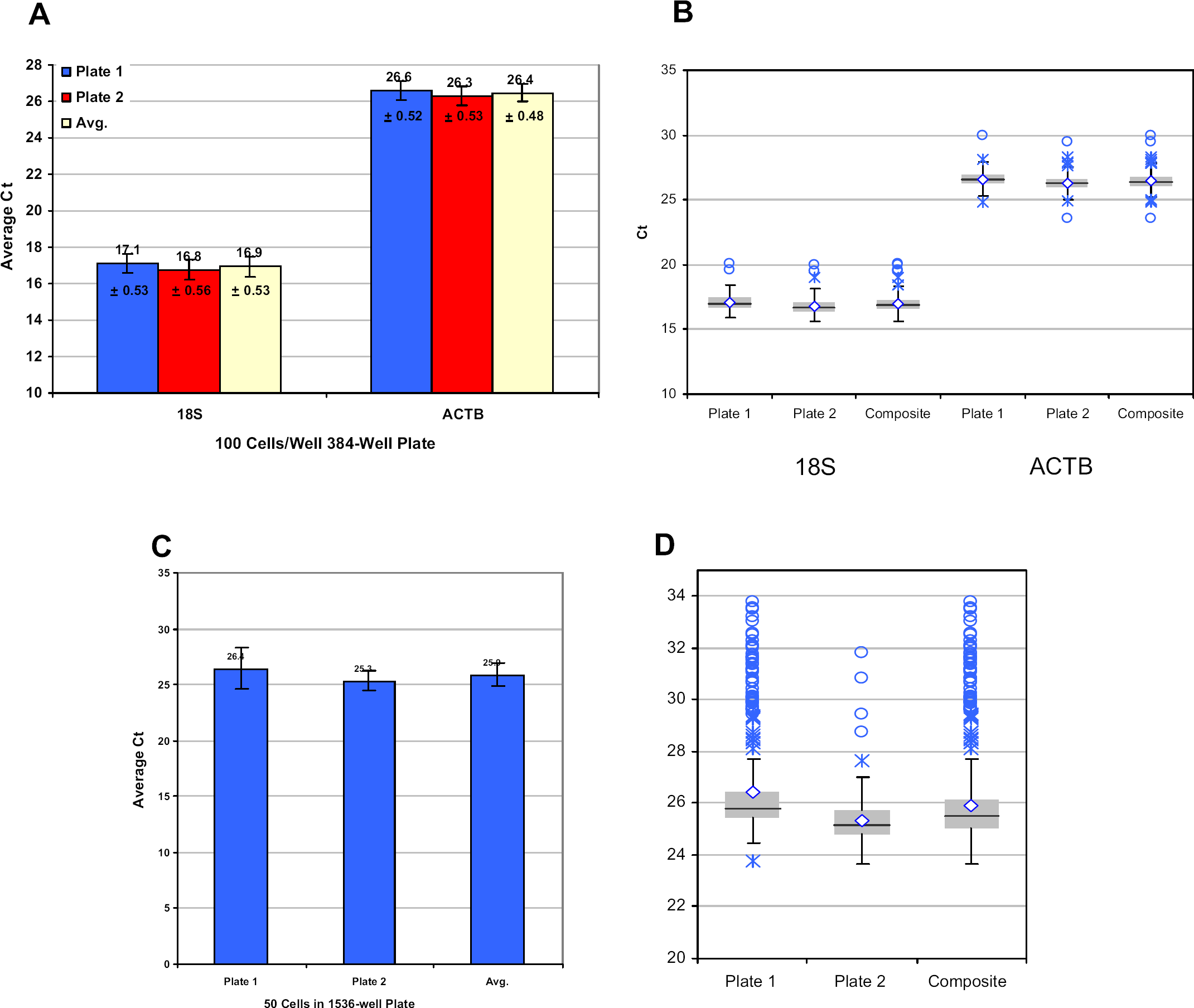
Cells-to-CT kit consistency. (A) A constant volume containing 100 HeLa cells was dispensed into two 384-well plates and prepared for RT-PCR in a 7-minute protocol using the Cells-to-CT kit. Prepared samples were analyzed using primers specific for 18S ribosomal or β-actin RNA as described in Materials and Methods section. Each bar represents the average of 384 biological replicates with two technical replicates performed per sample. (B) Box plot for data used in panel A. (C) A constant volume containing 50 HeLa cells was dispensed into wells of two 1536-well plates and prepared for RT-PCR in a 7-minute protocol using the Cells-to-CT kit. Prepared samples were analyzed using primers specific for β-actin RNA as described in Materials and Methods section. Bars representing plate averages of 384 biological replicates (one technical replicate) and the average across the two 1536-well plates tested are presented. Average CT values and standard deviations from the mean are presented. (D) Box plot for data used in panel C.
A challenge faced in using small numbers of cells is the ability to detect less abundant messages. The TaqMan Pre-Amp Cells-to-CT kit is specifically designed to extend material from valuable or small amounts of sample, and we wanted to demonstrate the use of this kit in 384-well formats. Samples containing 100 cells were processed using Cells-to-CT kits, and aliquots of the resulting materials were used for preamplification of a 7-target assay pool before real-time analysis. Analysis of technical replicates performed in two separate 384-well plates confirms that targets were easily detected with low plate-to-plate variation using a procedure that extends the amount of usable sample up to 64-fold (Fig. 5).
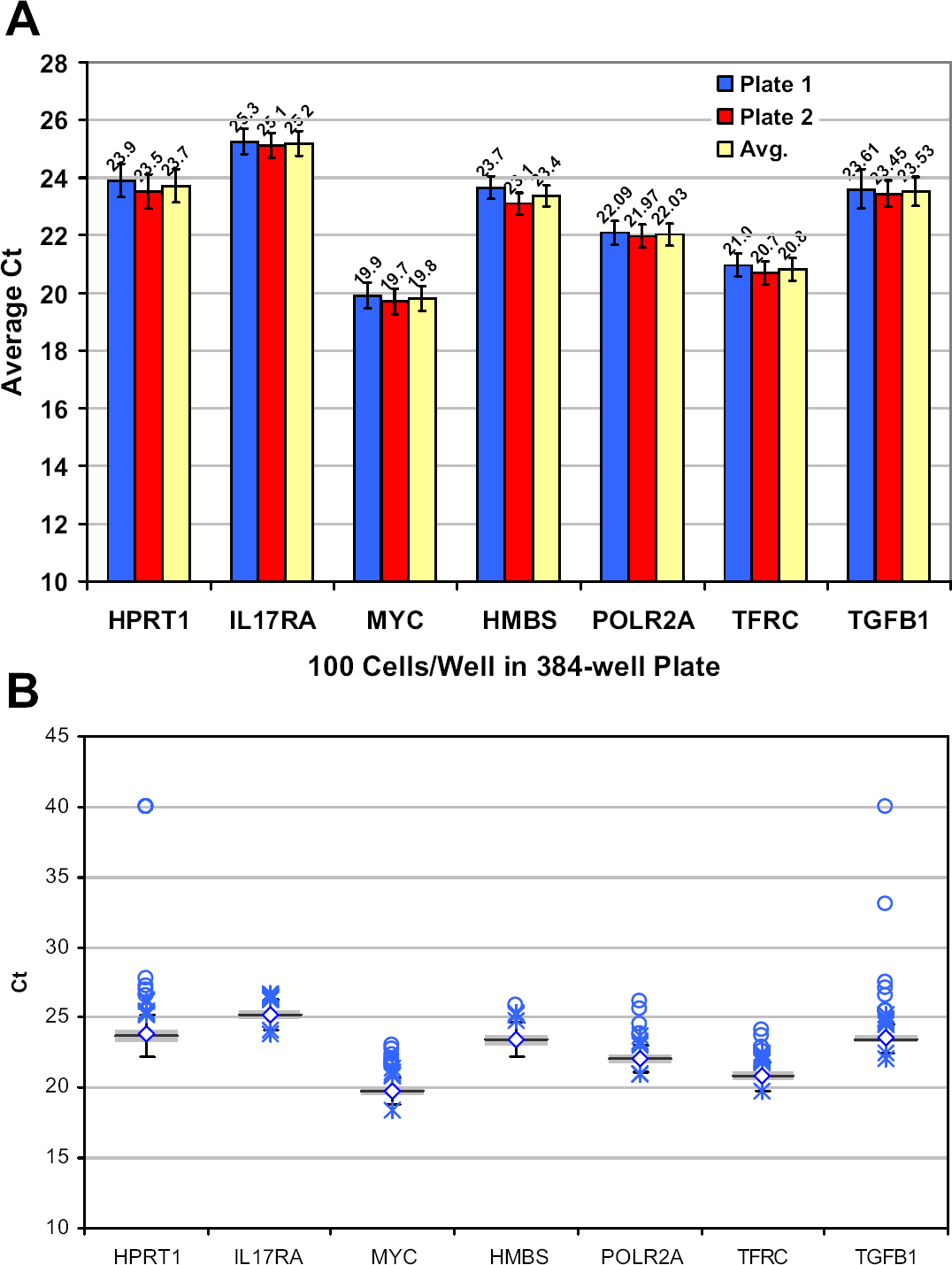
Preamplification of samples. (A) A constant volume containing 100 HeLa cells was dispensed into a 384-well plate, and the samples were prepared using Cells-to-CT kits. Prepared samples were preamplified using pooled primers for the indicated targets and analyzed using real-time PCR for each specific target as described in Materials and Methods section. The averaged
To minimize the possibility of cross-contamination, all Cells-to-CT kit sample preparation takes place in a single well with no transfer of sample or movement of liquids from processing wells. We used the ability of the HTS instrument to selectively distribute the cells and the reagent according to a predesigned plate map to add cells to specific wells of a 384-well plate and subsequently perform real-time analysis to confirm that cross-contamination was not detectable after preparing samples using the Cells-to-CT kit procedure. Figures 6 demonstrates that consistent real-time signal was obtained in wells that contained cells (average
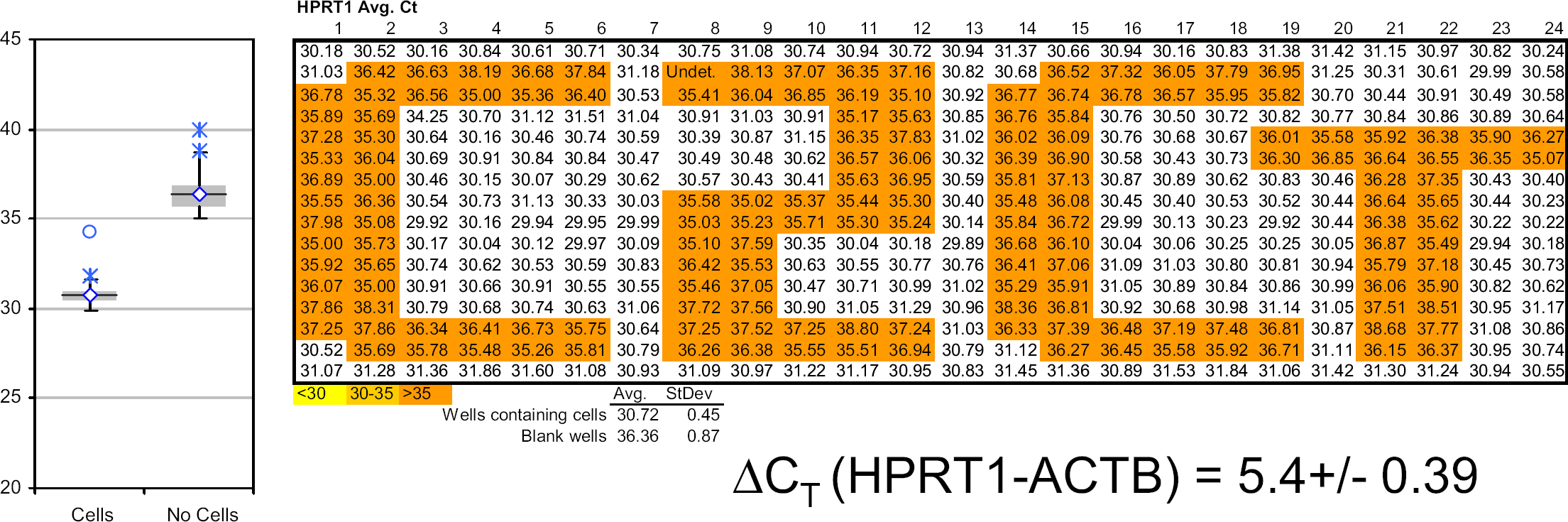
Undetectable cross-contamination. A constant volume containing 2500 HeLa cells was dispensed into specific wells of a 384-well plate, and the entire plate was prepared using Cells-to-CT kits. Prepared samples were analyzed using real-time PCR for the HPRT target and β-actin target (not shown) as described in Materials and Methods section. A box plot of the data set and the averaged Δ
Conclusions
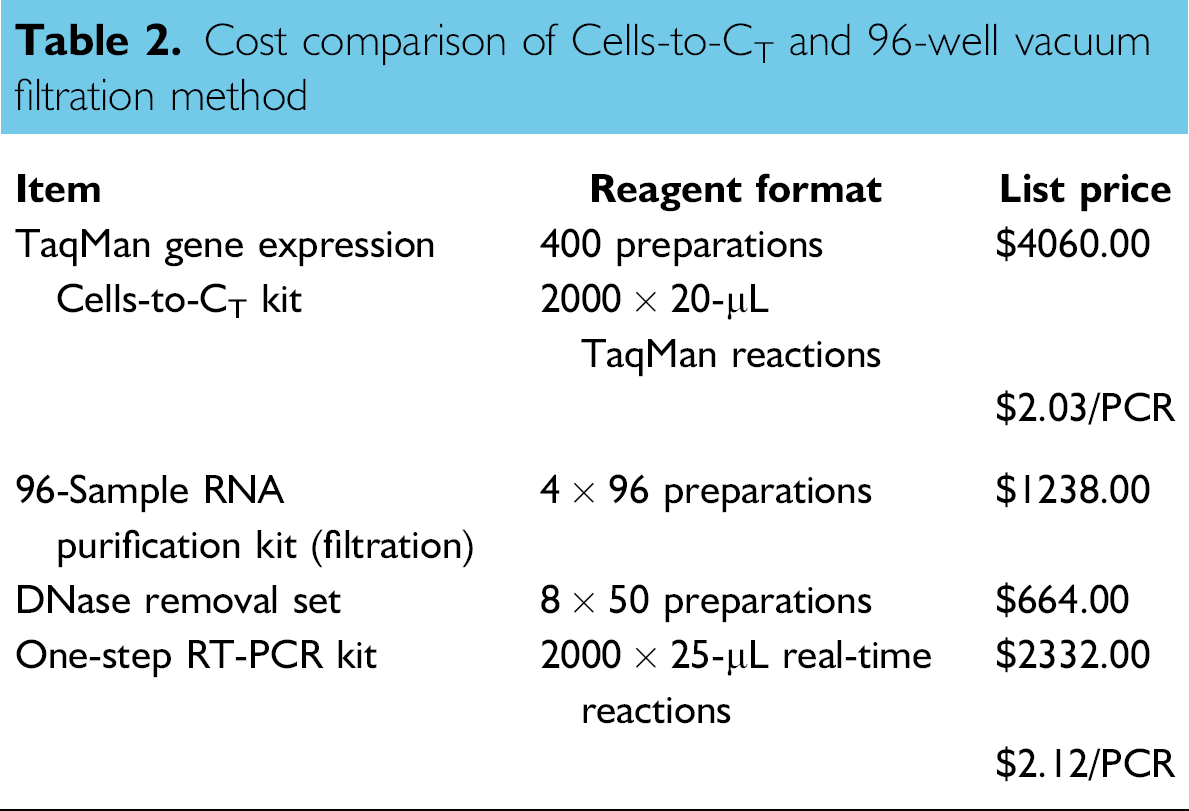
The ability to rapidly prepare high-quality RNA from cell culture samples for subsequent real-time analysis is often a critical bottleneck in many high throughput workflows. The Cells-to-CT technology prepares samples for direct realtime analysis without the need for traditional purification and has been shown to provide equivalent or better performance across a wide variety of cell lines and amplification targets. 4 The Cells-to-CT workflow is also cost effective when compared with other methods (Table 2) and is designed to be more environmentally friendly by eliminating the use of guanidine salts and alcohol and reducing plastic waste by greater than 95% (by weight) relative to traditional filtration methods. We demonstrate here that the Cells-to-CT technology is easily applied to high-density expression analysis and can be performed in 96-, 384-, and 1536-well plates in a sample preparation procedure that is accomplished in about 8 min.
Cost comparison of Cells-to-CT and 96-well vacuum filtration method

We observed an approximate three
The Cells-to-CT workflow allows for simple noncontact dispensing of liquids in a format that is easily adapted to rapid processing of high-density plates. The analysis of cells dispensed into 384-well plates demonstrated consistent performance within a plate and across plates. Although the Cells-to-CT is validated for use with as few as 10 cells, 4 our analysis of cells in 1536-well plates highlighted that careful preparation of cells becomes increasingly important as the numbers of cells decrease. Analysis of 50 cells demonstrated excellent reproducibility in one 1536-well plate but was suboptimal in a second plate (Fig. 4, panel D). We attribute this to incomplete dispersion of the cells during preparation, and this highlights the special cell-handing needs that are required as fewer cells are processed. The ability to couple Cells-to-CT kit sample preparation and the TaqMan PreAmp technology is a valuable method for extending the amount and signal from valuable samples, and we successfully demonstrated the performance of these coupled technologies in a 384-well format. The demonstrated ability to rapidly and reproducibly prepare small numbers of cells at high plate densities coupled with preamplification methods provides a high-performance workflow ideal for working with precious samples or in applications, such as compound screening.
Acknowledgments
