Abstract
Commercialization of fuel ethanol production from lignocellulosic biomass has focused on engineering the glucose-fermenting industrial yeast Saccharomyces cerevisiae to use pentose sugars. Because S. cerevisiae naturally metabolizes xylulose, one approach involves introducing xylose isomerase (XI), which catalyzes conversion of xylose to xylulose. In this study, an automated two-hybrid interaction protocol was used to find yeast genes encoding proteins that bind XI to identify potential targets for improving xylose utilization by S. cerevisiae. A pDEST32 vector re-engineered for TRP selection and containing the Gal4 binding domain fused with the Piromyces sp. E2 XI open reading frame (ORF) was used as bait with a library of LEU-selectable pOAD vectors containing the Gal4 activation domain in fusion with members of the S. cerevisiae genome ORF collection. Binding of a yeast ORF protein to XI activates two chromosomally located reporter genes in a PJ69–4 yeast strain to give selective growth. Five genes, including ADHI, were identified in the two-hybrid screen, suggesting the proteins encoded by these genes bind to XI. The effect of ADHI overexpression was examined using the pSUMOduoHisADHI vector in an automated protocol to transform eight previously identified yeast strains that showed anaerobic growth on xylose. One transformant consumed all available glucose, xylose, and arabinose during growth on wheat straw hydrolysate.
Keywords
Introduction
Plant biomass has the potential to be an inexpensive, plentiful, and renewable feedstock for biofuel production. 1 Glucose and xylose are the principal constituents of plant cell wall lignocellulose. 2 Numerous attempts have been made to engineer microorganisms capable of efficient fermentation of both glucose and xylose as a means of achieving economically feasible biomass conversion into fuel ethanol. These attempts have focused on engineering the glucose-fermenting industrial yeast Saccharomyces cerevisiae to use pentose sugars from lignocellulosic biomass by introducing the enzymes of the initial stages of xylose metabolism. 3 4
S. cerevisiae can naturally metabolize the pentose phosphate pathway intermediate xylulose. Thus, to enable this yeast to metabolize xylose, it is necessary to introduce the enzymes required to convert xylose to xylulose. One approach is to engineer S. cerevisiae to express xylose reductase and xylitol dehydrogenase, which convert xylose to xylitol and xylitol to xylulose, respectively. These steps are used by yeast strains that naturally ferment xylose, but the process gives rise to a cofactor imbalance in S. cerevisiae that results in the production of xylitol. 5 A second approach is to engineer S. cerevisiae to express xylose isomerase (XI), which catalyzes the conversion of xylose to xylulose and does not require redox cofactors. Introduction of a functional XI into S. cerevisiae allows slow metabolism of xylose by way of the endogenous xylulokinase (XKS) and the enzymes of the nonoxidative part of the pentose phosphate pathway but is not sufficient for high rates of anaerobic xylose fermentation. To ensure that flux into and through the pentose phosphate pathway is not a limiting factor, the genes encoding the enzymes involved in this pathway were overexpressed in the strain expressing Piromyces sp. E2 XI. 6 The resulting engineered strain shows a high rate of anaerobic xylose consumption; however, improvement in the rate of ethanol production is still needed for industrial applications. 7
Similar to most aerobic and anaerobic bacteria but unlike most yeasts and fungi, the anaerobic fungus Piromyces sp. strain E2 metabolizes xylose via XI and d-XKS. The Piromyces enzymes fall into the cluster comprising plant and bacterial enzymes. 8 It would be of interest to determine whether proteins in the yeast were affecting the activity of fungal XI expressed in S. cerevisiae. Very little information is available in the literature on the binding of proteins to XI. d-Xylose isomerase (d-xylose ketol-isomerase; EC 5.3.1.5) is one of the most widely used industrial enzymes. It is a soluble enzyme that catalyzes the reversible isomerization of d-xylose to d-xylulose, and, more slowly, d-glucose to d-fructose. Its activity on glucose makes it important industrially for the production of high-fructose corn syrup. The enzyme is active in multisubunit forms and all XIs require Mg2+, Mn2+, or Co2+ for catalytic activity, while Ca2+ is a strong competitive inhibitor. The active site contains two metal cations per subunit. 9 The histidine residue at position 220 is conserved in all d-XIs. The effects of replacing His-220 with Ser, Glu, Asn, and Lys on enzyme activity, thermostability, and structure were analyzed by kinetic, thermal denaturation, and crystallographic methods. All these His-220 mutants were significantly less catalytically active than the wild type and all mutants were 5–24 °C less stable than wild type because the metal binding is perturbed. 10
It has been observed that the fundamental mechanisms by which fermentation is regulated appear to be very different in the glucose-fermenting yeast S. cerevisiae and the xylose-fermenting yeast Pichia stipitis (renamed as Scheffersomyces stipitis) even though the structural genes encoding the alcohol dehydrogenase (ADH) and pyruvate decarboxylase in both show significant sequence conservation. In S. cerevisiae, glucose, rather than oxygen availability, induces high levels of fermentative ADH (ScADH1) and represses oxidative ADH (ScADH2), leading to ethanol production. In the Crabtree-negative yeast P. stipitis, it was found that oxygen availability regulates PsADH1 expression and the regulation may be mediated by heme. PsADH1 expression was about 10-fold higher under oxygen-limited conditions than under fully aerobic conditions when grown on xylose. 11 Similarly, the filamentous fungus Aspergillus nidulans ADH3 is specifically induced in response to periods of anaerobic stress, and its expression is regulated largely at the posttranscriptional level. Such mechanisms may also function in P. stipitis, because its primary response to a lowered aeration rate is to increase fermentative enzymes. The mechanism and extent of this regulation are unclear. 11 The regulation of ethanol production in Piromyces also differs from that in S. cerevisiae. Unlike other hydrogenosome-bearing anaerobes, Piromyces sp. E2 possesses an alcohol dehydrogenase E (ADHE) that combines aldehyde dehydrogenase (ALDH) and ADH activities. The final steps of carbohydrate catabolism are routed via pyruvate:formate lyase and ADHE to give formate, acetate, lactate, ethanol, and succinate. 12 The introduction of Piromyces XI into yeast may expose the fungal enzyme to proteins and possible regulators that are not present in its natural environment.
The simultaneous utilization of glucose and xylose depends on a variety of parameters that are intrinsic to the regulatory circuits in the cell. 6 In an attempt to identify potential regulatory parameters in the yeast cell that affect utilization of xylose in S. cerevisiae strains engineered to express XI, a high-throughput robotic two-hybrid screen was implemented to select yeast proteins expressed from a collection of full-length open reading frames (ORFs) representing the yeast genome that bind to XI.
Materials and Methods
Strains
Two haploid S. cerevisiae strains described previously 13 were used. They are identical except for mating type:
PJ69–4 MATa: MATa, trp1–901, leu2–3, ura3–52, his3–200, gal4Δ, gal80Δ, GAL2-ADE2, LYS::GAL1-HIS3, met2::GAL7-lacZ
PJ69–4MATalpha: MATalpha, trp1–901, leu2–3, ura3–52, his3–200, gal4Δ, gal80Δ, GAL2-ADE2, LYS::GAL1-HIS3, met2::GAL7-lacZ
Production of pDEST32 XI TRP-Selectable Haploid PJ69–4 MATalpha Bait Strain
The pDEST32 XI TRP-selectable plasmid was constructed by replacing the LEU2 gene in the pDEST32 LEU-selectable bait plasmid commercially available (Invitrogen, Carlsbad, CA) with the TRP1 gene as described previously. 13
Production of Haploid PJ69–4 MATa Strain Containing Library of pOAD Prey Plasmids with ORFs from S. cerevisiae Genome
As previously described 13 , a collection of PCR-generated ORFs predicted from the S. cerevisiae genome in pOAD LEU-selectable vectors transformed into PJ69–4 MATa haploid yeast strain and spotted in a 384-grid pattern on sixteen 86 × 128-mm Omnitrays containing CM glucose -LEU + ADE medium was provided by Dr. Stanley Fields at University of Washington, Seattle, WA. 14 15
Automated Mating of PJ69–4 MATalpha Haploid Strain to PJ69–4 MATa Haploid Strain
Mating was performed as described previously 13 except the resulting diploid strains were not spotted onto plates containing xylose medium for incubation in an anaerobic chamber, but were plated onto CM 2% glucose -TRP-LEU-HIS + 3-amino-1,2,4-triazole (3AT) medium (25 mM 3AT added to Teknova -T-L-H plate) under aerobic conditions to select for binding events between XI expressed by the pDEST32 TRP-selectable bait plasmid having the Gal4 binding domain (BD) fused in-frame to the XI bait and the yeast ORF proteins expressed by the pOAD LEU-selectable prey plasmid having the Gal4 activation domain (AD) fused in-frame to the ORF inserts based on the S. cerevisiae genome.
Sealed ABGene 96-well deep-well cultures containing the resulting diploid PJ69–4 yeast strain with the pDEST32 XI TRP-selectable plasmid and the pOAD LEU-selectable plasmids with the yeast genome ORFs were pierced by the sterile stainless steel pipet tips of the liquid handler, and 20 μL were diluted in a new 384-well deep-well plate containing 100 μL CM 2% glucose -LEU-TRP + ADE liquid medium (Teknova, Hollister, CA). Three microliters of the dilution cultures were spotted in 96-well format onto the more stringent reporter medium CM 2% glucose -TRP-LEU-HIS + 3AT and incubated aerobically for 6 days at 30 °C and 100 rpm. In addition, the selected diploid strains were assessed secondarily for aerobic growth on the less stringent reporter medium CM 2% glucose -TRP-LEU-ADE plates (Teknova). For further evaluation, the selected diploid strains were plated on CM 2% xylose -LEU-TRP + ADE (Teknova) to observe for aerobic growth. The selected diploid strains were also grown on CM 2% glucose -TRP-LEU-HIS + 3AT liquid medium and used for growth analyses.
Isolation and Sequencing of Yeast ORF Inserts from pOAD Prey Plasmids
The plasmids were isolated using a QIAprep Spin Miniprep Kit (Qiagen, Valencia, CA) using the manufacturer's instruction with the following modifications. All steps were carried out in 1.5-mL microfuge tubes. Cells were pelleted using two 1-mL centrifugations of cell lysate. The resuspended cells were vortexed with beads for 1 min, cooled on ice for 30–60 s and again vortexed for 1 min. The tube was vortexed for a total of 5 min. The bottom of the spin column was tapped on a paper towel to remove any remaining PE buffer before eluting the DNA. After DNA was isolated from the yeast lysate, the resulting DNA was transformed into TOP 10 competent cells (Invitrogen, Carlsbad, CA) according to TOP 10 One Shot Invitrogen protocols and grown in 1600 μL TB AMP 50 medium (Teknova, Hollister, CA). The resulting colonies were subjected to a plasmid preparation kit procedure according to Qiagen 96-well Turbo Kit protocol. The plasmids were sequenced using the procedure described previously. 16
Binding Analysis of Mated Yeast Strains with Interacting Proteins by Growth on CM 2% Glucose -TRP-LEU-HIS + 3AT Liquid Medium
Mated yeast strains that indicated binding to XI were adjusted to an optical density at 660 nm of 0.4 and 100 μL was inoculated into 5 mL of CM 2% glucose -TRP-LEU-HIS + AT3 liquid medium and shaken at 100 rpm at 30 °C. The OD was monitored using a Beckman Coulter DU 800 UV—VIS spectrophotometer (Beckman Coulter, Inc., Fullterton, CA). The growth curves were plotted in Excel.
In Vitro ADH Activity Assay
Determination of reaction velocity was based on the method of Vallee and Hoch 17 in which rate of absorbance at 340 nm resulting from reduction of nicotinamide dinucleotide (NAD) is measured. One unit of enzyme reduces one micro-mole of NAD per minute at 25 °C under the specified conditions. The control was a solution containing 1.5 mL of 0.1 M phosphate buffer (pH 7.5), 0.5 mL 0.1% bovine serum albumin, and 1.0 mL of S. cerevisiae ADH enzymes (extracted from lysed S. cerevisiae cells, purified on gel filtration column, lyophilized; gives single band on SDS-PAGE; Worthington Biochemical Corp., Lakewood, NJ). Solutions of the ADH(L) and XI enzymes were prepared at a concentration of 1 mg/mL in 0.1 M phosphate buffer, pH 7.5 based on OD at 280 nm. ADH(L) 1 mg/mL was mixed with Streptomyces murinus XI enzyme (Sigma Aldrich, St. Louis, MO) 1 mg/mL to give ratios of ADH(L) to XI of 1:1,1:9, and 1:49, respectively. Mixtures were preincubated for 30 min before dilution and testing. Into each cuvette, 1.5 mL 0.1 M pyrophosphate buffer, 0.5 mL 2.0 M ethanol, and 1.0 mL 0.025 M NAD were pipetted and then incubated in the spectrophotometer for 3–4 min at 25 °C. At zero time, 0.1 mL of ADH(L) control or ADH(L)/XI solutions, diluted 1:2000 immediately before use, were added to the cuvette and the change in absorbance at 340 nm (ΔA 340) was recorded for 3–4 min. The ΔA 340/min was calculated from the slope of the initial linear portion of the curve. Activity (Units) was calculated using the equations:
In Vitro Analysis to Confirm ADH Binding to XI
In vitro transcription translation of Piromyces XI was performed using a RTS 100 Kit (Roche Applied Science, Indianapolis, IN) according to manufacturer's protocol and the pSUMOduo His-tagged Piromyces XI plasmid previously described. 18 The use of the bacterial modified T7 promoter in that vector was optimized by LifeSensors, Inc. (Malvern, PA). 19 The resulting 6xHis-tagged Piromyces XI was used for in vitro binding experiments with 1 mg/mL solutions of S. cerevisiae ADH(L) (Worthington Biochemical Corp., Lakewood, NJ) and 1 mg/mL preparations of purified S. murinus XI (Sigma Aldrich, St. Louis, MO). The 6xHis-tagged Piromyces XI, 10 μg/50 μL per well, was preincubated with Super-Flow nickel-charged agarose beads (Qiagen, Valencia, CA) for 4 h at 4 °C on a tumbler before addition of purified ADH(L) and S. murinus XI preparations. Mixtures of Ni-bead-bound-6xHis-tagged Piromyces XI with ADH(L), with ADH(L) plus S. murinus XI, with S. murinus XI, or with 6xHis-tagged Piromyces XI were incubated for 6 h. A set of each of these with beads alone was placed in four tubes. A third set of four tubes contained ADH(L), ADH(L) plus S. murinus XI, washed Ni-bead-bound Piromyces XI, or S. murinus XI. The resulting 12 tubes were washed with buffer prepared from 1 × PBS with Ca2+ and Mg2+ (Mediatech, Inc., Manassos, VA) modified with 1% SDS. Beads were washed four times and placed into a tumbler at 4 °C between washes for 15 min. The resulting washed bead pellets were resuspended in 20 μL of 2 × tris glycine loading buffer, 20 mL (Invitrogen, Carlsbad, CA) supplemented with 2 mL of 2-BME (Sigma Aldrich, St. Louis, MO). Samples were heated to 95 °C for 10 min, allowed to cool for several minutes, and loaded onto a 16% SDS-PAGE denaturing gel system from Novex. Gels were run for 3 h at constant 126 V and stained with Coomassie reagent (BioRad, Hercules, CA). The gels were destained with BioRad destaining solution using several changes over a period of 2 h. Photographs were taken using the AlphaImager 3400 workstation with AlphaEase FC software.
Construction of pSUMOduoHisADH1 Plasmid and Transformation into S. cerevisiae Yeast Strains Previously Engineered for Anaerobic Growth on Xylose
The pSUMOduoHisADH1 plasmid was constructed by obtaining pDONR221ADH1 plasmid from the Harvard Institute of Proteomics FLEXGene S. cerevisiae collection (HIP clone ID 114899) and using LR clonase (Invitrogen, Carlsbad, CA) as previously described 16 to clone the ADH1 insert in-frame into the pSUMOduoHis Gateway-adapted plasmid (LifeSensors, Inc., Malvern, PA). The resulting vector was used to transform TOP 10 competent cells as described previously 16 and plated onto LB AMP 50 plates (Teknova, Hollister, CA) to produce colonies that harbored pSUMOduoHisADH1. The plasmids were isolated from colonies as described previously 16 and transformed into the triple-plasmid diploid strains previously described 13 to give the quadruple-plasmid diploid strains expressing XI, XKS, an anaerobic xylose-using yeast ORF, and ADH1.
Analysis of Ethanol from Single Sugar (Glucose and Xylose) Cultures
Colonies of ADH1 transformants were picked from CM 2% glucose -URA-TRP-LEU-HIS Omnitray plates into 24-well deep-well plates containing CM 2% glucose -URA-TRP-LEU-HIS liquid medium, cells were pelleted, re-suspended in water, and adjusted to an OD at 660 nm of 0.1. An aliquot of 100 μL was used to seed 5 mL of either CM 2% glucose -URA-TRP-LEU-HIS or CM 2% xylose -URA-TRP-LEU-HIS in a 24-well deep-well plate in triplicate for each of the strains analyzed and grown for 6 days. The liquid 24-well deep-well plate cultures were analyzed in triplicate for ethanol production using gas chromatography (6890N; Agilent Technologies, Wilmington, DE) with a packed column as described previously. 20 Before injection into the GC, the samples were diluted fourfold with distilled water. The GC was equipped with an autosampler and an integrator. Ethanol production was calculated as g L-1.
Preparation of Lime Pretreated Enzymatically Saccharified Wheat Straw Hydrolysate
Quadruple-plasmid strains were compared to optimal triple-plasmid. Wheat straw was purchased from a local farmer, dried in a forced-air oven at 55 °C for 24 h, and milled in a hammer mill to pass through a 1.27-mm screen. Milled wheat straw (8.6%, w/v) and lime (0.86%, w/v) were slurried in water, mixed, and autoclaved at 121 °C for 1 h. The pH of the lime pretreated wheat straw was adjusted to 5.0 using concentrated HCl. The enzymatic hydrolysis was performed by shaking gently (100 rpm) at 45 °C using a cocktail of three commercial enzyme preparations at a dose level of 0.15 mL/g of wheat straw of each enzyme preparation for 72 h. The commercial enzyme preparations used were: Celluclast 1.5 L (cellulase), Novozymes 188 (beta-glucosidase), and Viscostar 150 L (hemicellulase). Celluclast 1.5 L and Novozymes 188 were purchased from Brentag Great Lakes, Milwaukee, WI and Viscostar 150 L was supplied by Dyadic Corp., Jupiter, FL. The activity levels of carboxymethyl cellulase, beta-glucosidase, xylanase, beta-xylosidase, and alpha-l-arabinofuranosidase in these three commercial enzyme preparations have been reported in a previous paper.
Hydrolysate Analysis for Cell Growth and for Concentrations of Glucose, Xylose, Arabinose, and Ethanol
Colonies of ADH1 transformants were picked from CM 2% glucose -URA-TRP-LEU-HIS Omnitray plates into 24-well deep-well plates containing CM 2% glucose -URA-TRP-LEU-HIS liquid medium, cells were pelleted, resuspended in water, and adjusted to an OD at 660 nm of 0.1. An aliquot of 100 μL was added to 5 mL of wheat straw hydrolysate in triplicate in a 24-well deep-well plate for each of the strains analyzed as was carried out for the single sugar substrates. Samples (1 mL) were taken after 8-day growth in the hydrolysate at 30 °C with shaking at 100 rpm and placed in triplicate onto 24-well deep-well plates containing CM 2% glucose -URA-TRP-LEU-HIS liquid medium. The plates were covered with microporous tape (Becton Dickinson, Franklin Lakes, NJ) to maintain sterility. Glucose, xylose, and ethanol were analyzed by high-pressure liquid chromatography. 22 Cell growth of the yeast strains was monitored by measuring the optical density of the appropriately diluted culture sample at 660 nm.
Results and Discussion
Automated Yeast Mating Protocol for Production of Two-Hybrid Interactor Strains to Identify S. cerevisiae ORFs Expressing Proteins That Bind to XI
A successful automated strategy was previously developed for production and screening of multiplasmid diploid yeast strains by mating of two haploid yeast strains. 13 A similar protocol is used in this study, but instead of selecting for xylose utilization by plating the diploid strain on xylose medium, the diploid strain is used in a two-hybrid setting to identify proteins that bind to XI (Fig. 1). The mated yeast cells were plated onto CM 2% glucose -TRP-LEU-HIS + 3AT medium to select for binding events between XI expressed by the pDEST32 TRP-selectable bait plasmid having the Gal4 BD fused in-frame to XI and the yeast ORF proteins expressed by the pOAD LEU-selectable prey plasmid having the Gal4 AD fused in-frame to the ORF inserts based on the S. cerevisiae genome. CM 2% glucose -TRP-LEU-HIS + 3AT medium was used as the primary screen because the HIS reporter gene gave more stringent selection than the ADE reporter gene and growth on CM glucose -TRP-LEU-ADE plates. Six yeast ORFs were identified by growth on the selective medium CM 2% glucose -TRP-LEU—HIS + 3AT as expressing proteins that bound to XI. The ORF coding for ADH1 was selected twice, so the screen yielded five unique ORFs expressing proteins that bound to XI. A list of the yeast genes encoding proteins that bind to XI and the Saccharomyces GenomeDatabase gene ID, nomenclature, and function are given in Table 1. Four of these yeast proteins, ADH1, CSM4, APM1, and RNR1, have been ascribed a function, whereas one with locus tag YOR342C does not have an assigned function. The latter is designated XIB1 (XI-binding) protein in this study. In S. cerevisiae, the ADH1 gene encodes ADH, the major enzyme responsible for converting acetaldehyde to ethanol. The enzyme functions as a tetramer of four identical subunits with each subunit containing two zinc ions, one is essential for catalysis and the other is important for the structure of the protein. 23 CSM4 protein mediates chromosome dynamics and recombination during yeast meiosis. 24 25 It is required for accurate chromosome segregation during meiosis. APM1 is the Mu1-like medium subunit of the clathrin-associated protein complex AP-1. Clathrin-associated protein complexes are believed to interact with the cytoplasmic tails of membrane proteins, leading to their selection and concentration. The AP-1 complex interacts directly with clathrin. It has a late Golgi sorting function in S. cerevisiae. 26 RNR1 is part of the ribonucleotide-diphosphate reductase complex that catalyzes reduction of ribonucleotides to the corresponding deoxyribonucleotides, the rate-limiting step in dNTP synthesis. RNR1 activity is regulated by complex checkpoints to ensure high fidelity of DNA replication during normal cell-cycle progression and of repair in response to genotoxic stress. 27
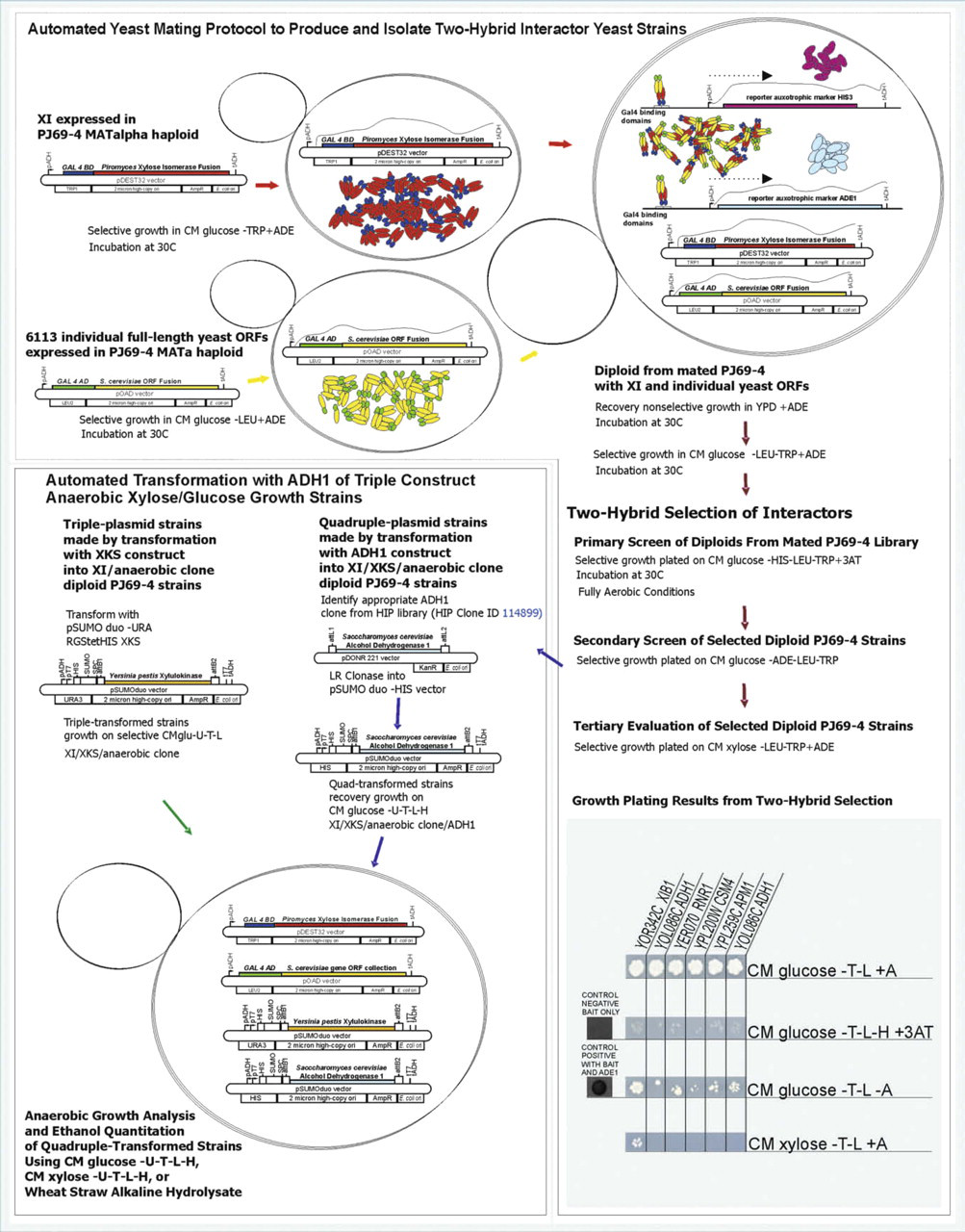
Schematic of automated yeast mating protocol (top and right) for two-hybrid analysis to identify yeast open reading frames that express proteins binding to XI and outline of the automated transformation protocol (bottom left) to produce yeast strains overexpressing ADH1 (an XI-binding protein identified in the two-hybrid screen).
Identification of five yeast genes encoding proteins that bind to xylose isomerase (XI)
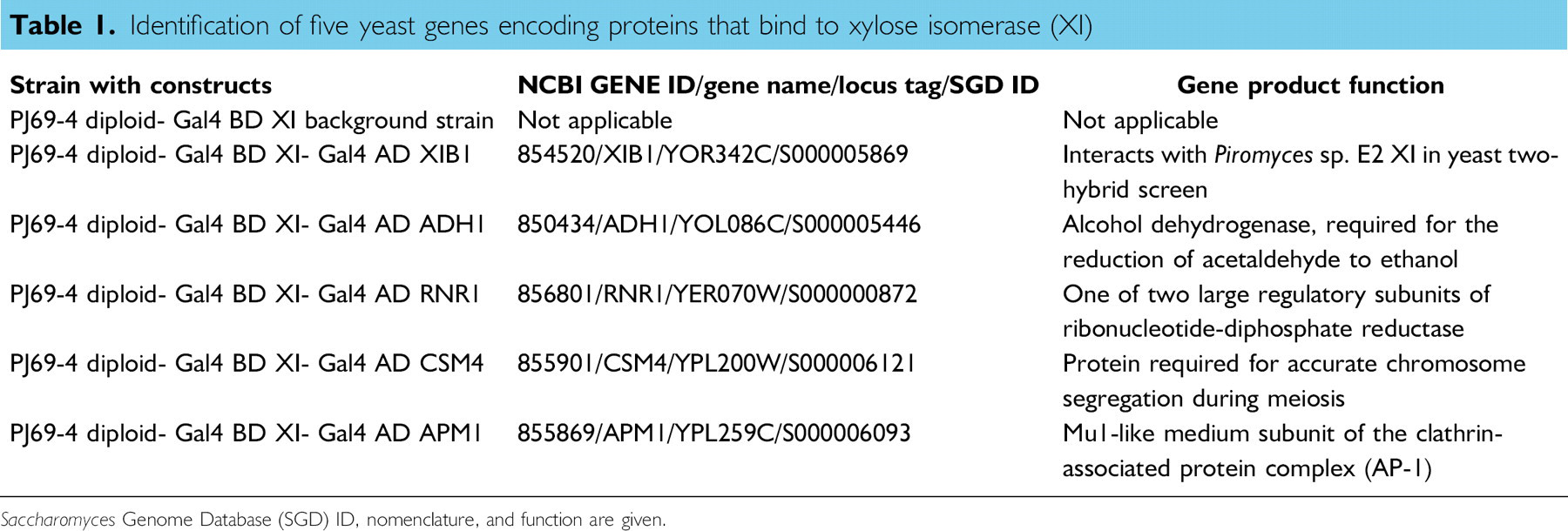

Saccharomyces Genome Database (SGD) ID, nomenclature, and function are given.
A secondary screen was performed using the less stringent ADE reporter gene. When the selected diploid strains were plated on CM 2% glucose -TRP-LEU-ADE plates, the same strains that grew on the plates with the more stringent medium, CM 2% glucose -TRP-LEU-HIS + 3AT, demonstrated strong growth, confirming the interaction between XI and the yeast ORF proteins.
In an additional evaluation to determine if growth on xylose could occur with strains containing XI and the XI-binding ORFs in the absence of XKS, the colonies were plated on CM 2% xylose -LEU-TRP + ADE. One of the strains identified as expressing an XI-binding protein also showed aerobic growth on xylose. This strain contained the yeast ORF with locus tag YOR342C, which has not yet been ascribed a function and is designated XIB1 in this study.
Binding Analysis of Mated Strains Using Growth on Reporter Selective Medium as Indication of Strong Interaction with XI
The results of the analysis using growth on CM 2% glucose -TRP-LEU-HIS + 3AT liquid medium to select the diploid S. cerevisiae strains with ORFs expressing proteins that bind XI are depicted in Figure 2. No other strains showed growth on this medium. The strains containing ORFs encoding yeast proteins identified as ADH1 and CSM4 showed slightly better aerobic growth on CM 2% glucose -TRP-LEU-HIS + 3AT, but all strains demonstrated a good rate of growth for two-hybrid interactor strains. ADH1 is the enzyme in S. cerevisiae responsible for converting acetaldehyde to ethanol. S. cerevisiae does not naturally express XI, while Piromyces does not express the same ADH as S. cerevisiae. Unlike other hydrogenosome-bearing anaerobes, Piromyces sp. E2 possesses an ADHE that combines ALDH and ADH activities. 12 It could be speculated that an interaction between S. cerevisiae ADH and Piromyces XI may affect the rate of xylose utilization by S. cerevisiae. The doubling times derived from these growth curves are also shown in Figure 2 (graph insert). The strain with ADH1 had the shortest doubling time, suggesting it has the strongest interaction with XI.
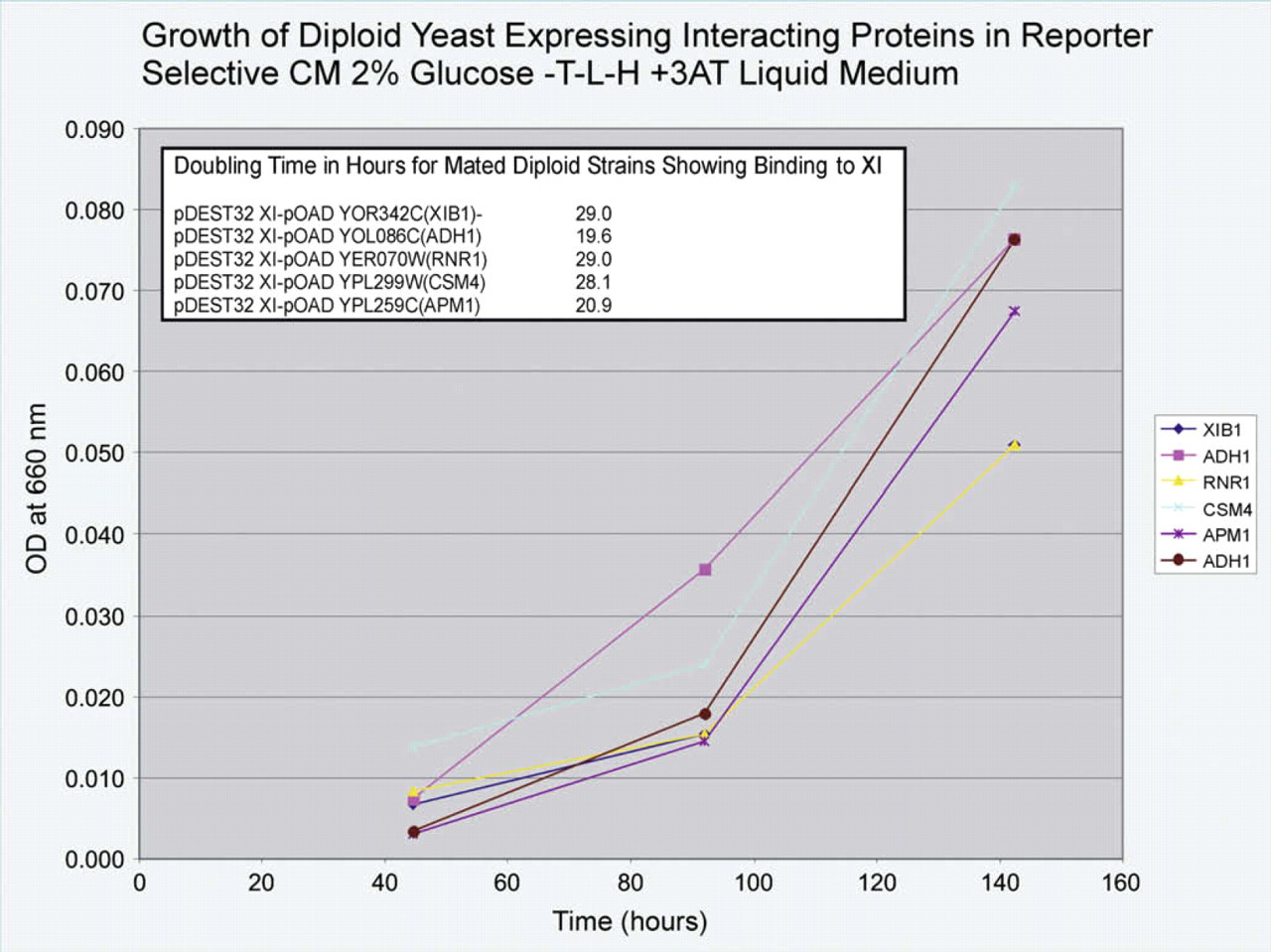
Binding analysis for proteins that bind XI showing growth curves of the Saccharomyces cerevisiae mated diploid strains with interacting proteins obtained from the two-hybrid screen in reporter selective CM 2% glucose -TRP-LEU-HIS + 3AT medium. Doubling times (n = 3) for these strains derived from the growth curves are shown in the table insert. Equation used to calculate doubling time: ln 2/{[ln(A/A 0)]/Δt}. Terms: ln = natural log; A 0 = optical density at t 0; A = optical density at t; Δt = t – t 0. Note: Two curves are shown for ADH1 because it was selected twice.
In Vitro Confirmation of ADH Binding to XI
In vitro binding confirmed that ADH purified from S. cerevisiae binds to Piromyces XI expressed in vitro from the pSUMOduoHis Piromyces XI plasmid. The His-tagged Piromyces XI is bound to SuperFlow nickel beads and then incubated with a purified lyophilized preparation of yeast ADH. The SDS-PAGE analysis of this mixture after thorough washing with 1 × PBS with Ca2+ and Mg2+ modified with 1% SDS (Fig. 3, Lane 3) shows bands for ADH at 35.0 kD, for HisXI at 44.9 kD, and for SUMOHisXI at 61.5, indicating that because ADH has no His tag to bind to the Ni beads, it was bound to the XI. ADH also binds Piromyces XI in the presence of S. murinus XI because the ADH band is visible after thorough washing with PBS containing SDS in the lane with His-tagged Piromyces XI bound to SuperFlow nickel beads and incubated with purified yeast ADH and S. murinus XI (Fig. 3, Lane 4).
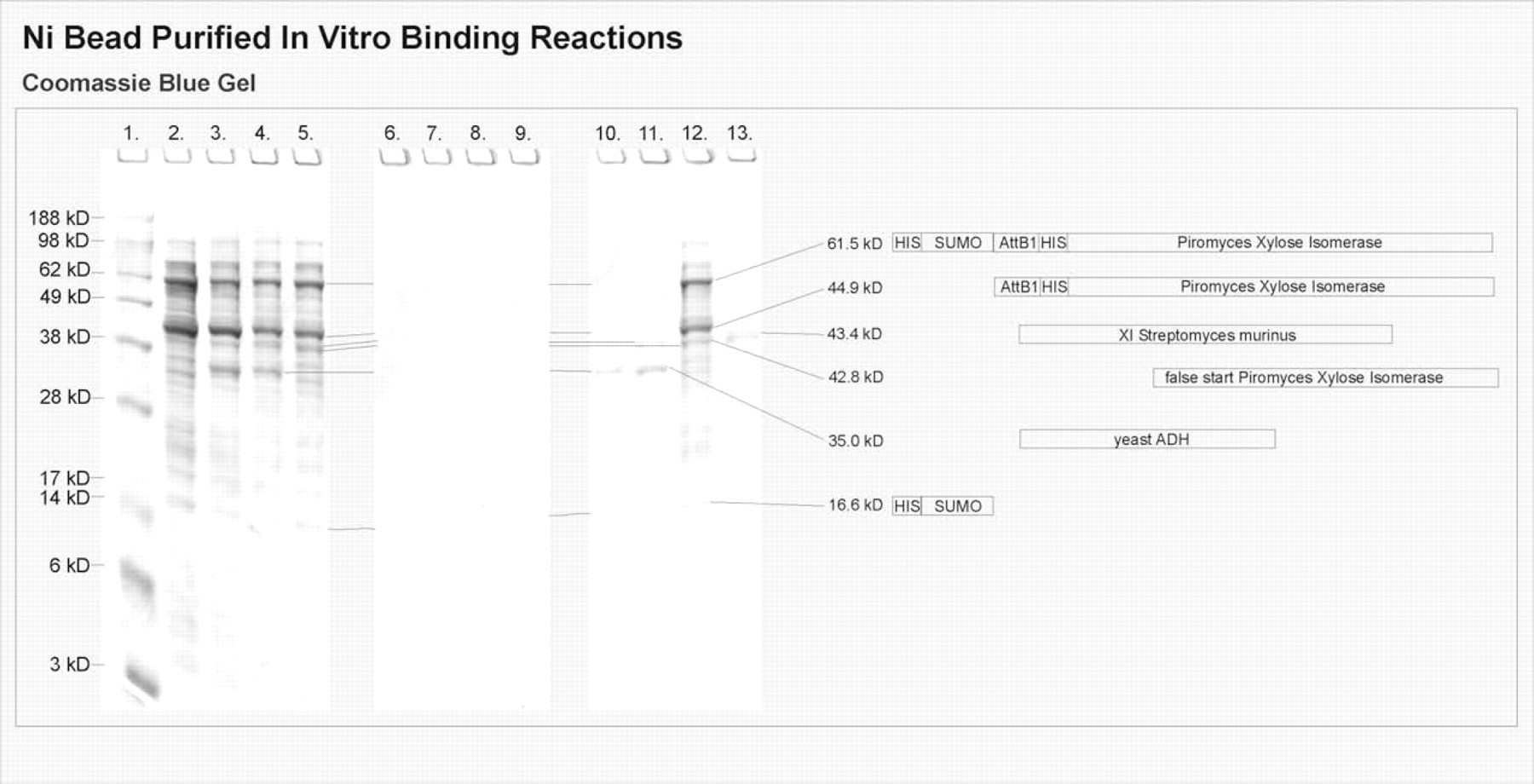
Gel showing results of in vitro nickel bead experiment to confirm ADH binding to XI using Piromyces XI expressed in vitro. Lane 1: Markers See Blue Plus II; Lane 2: Ni-bead-6xHis Piromyces XI; Lane 3: Ni-bead-6xHis Piromyces XI and Saccharomyces cerevisiae ADH(L); Lane 4: Ni-Bead-6xHis Piromyces XI, ADH(L), and Streptomyces murinus XI; Lane 5: Ni-bead-6xHis Piromyces XI and S. murinus XI; Lane 6: Ni beads; Lane 7: Ni beads, ADH(L); Lane 8: Ni beads, ADH(L), and S. murinus XI; Lane 9: Ni beads, S. murinus XI; Lane 10: ADH(L); Lane 11: ADH(L) and S. murinus XI; Lane 12: Washed Ni-bead-6xHis Piromyces XI; Lane 13: S. murinus XI.
Automated Transformation of Three-Plasmid Yeast Strains That Showed Anaerobic Growth on Xylose to Give Four-Plasmid Strains That Overexpress ADH1
The triple-construct strains engineered with plasmids expressing XI, XKS, and the yeast ORFs that gave anaerobic growth on xylose 13 were transformed with pSUMOduoHis ADH1 using an automated transformation protocol (Fig. 1). The steps on the robotic work cell and the plates used for producing the ADH1-transformed strains are depicted in Figure 4. The triple-plasmid yeast strains expressing the yeast gene ORFs PIP2 and CDC7 were not able to be transformed and no significant colonies survived. On the plates for triple-plasmid strains with yeast ORFs MAK5, IMG2, VPS9, COX10, ALE1, and MMS4, trans-formant colonies were seen and propagated.

Outline of the yeast transformation protocol depicting steps on the robotic work cell and plates required for producing ADH1-transformed strains.
Growth Analysis of Quadruple-Plasmid Strains Overexpressing ADH1
Growth curves in CM 2% glucose -TRP-LEU-URA-HIS liquid medium or in CM 2% xylose -TRP-LEU-URA-HIS liquid medium for the ADH1-transformed strains with XI, XKS, yeast ORFs for anaerobic growth on xylose (IMG2, MAK5, VPS9, COX10 ALE1, and MMS4), and ADH1 are presented in Figure 5. The doubling times derived from these curves are shown in the table on the graph. The doubling times for the strains with XI, XKS, IMG2, MAK5, VPS9, COX10, ALE1, or MMS4 and overexpressing ADH1 on glucose were 69.83, 65.77, 61.90, 58.90, 76.11, and 65.73 h and on xylose were 103.05, 103.42, 98.11, 88.59, 169.72, and 115.17 h, respectively (Fig. 5 insert). These times are much slower than the doubling times for the strains without ADH determined previously 13 in CM 2% glucose -TRP-LEU-URA liquid medium or in CM 2% xylose -TRP-LEU-URA liquid medium. The doubling times for the strains with XI, XKS, and IMG2, MAK5, VPS9, COX10, ALE1, or MMS4 (without ADH1 overexpression) grown on glucose were 3.44, 5.39, 4.95, 4.11, 5.72, and 3.89 h, and grown on xylose were 11.79, 9.75, 17.02, 20.08, 17.44, and 13.62 h, respectively. 13 The strain carrying plasmids encoding XI, XKS, COX10, and ADH1 grew fastest overall on both xylose and glucose single sugars. However, without ADH the strain with COX10 grown on xylose had the slowest doubling time. The xylose consumption decreased and the glucose consumption increased for the quadruple-plasmid strains when they were grown in trough plates with baffles as the strains in the previous study were, suggesting the relative aeration will shift glucose/xylose consumption (data not shown). COX10, farnesyl transferase, participates in the farnesylation of heme b, which is necessary for formation of heme a, the prosthetic group required for cytochrome oxidase assembly and activity. Heme a is a unique heme compound present only in mitochondrial cytochrome c oxidase, which is the terminal enzyme of the mitochondrial respiratory chain and catalyzes transfer of electrons from cyto-chrome c to molecular oxygen. 28
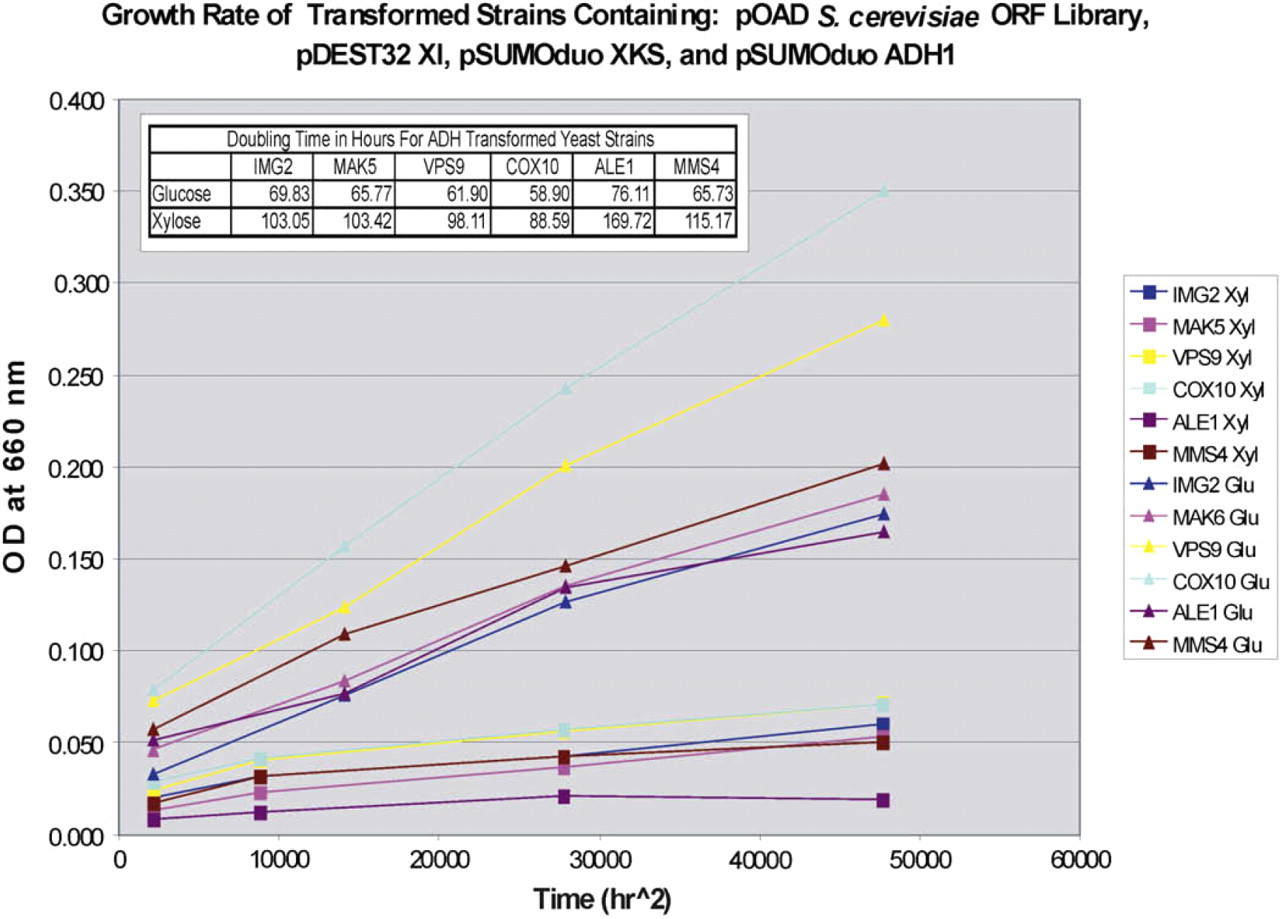
Growth curves of quadruple-plasmid strains overexpressing ADH1 in xylose and glucose medium based on optical density at 660 nm with doubling times indicated (insert table). Strains were grown in 24-well deep-well plates at 30 °C with 100 rpm shaking in CM 2% glucose -TRP-LEU-URA-HIS liquid medium or in CM 2% xylose -TRP-LEU-URA-HIS liquid medium.
In Vitro Analysis of ADH Activity in Presence of XI
No indication of ADH activity decreasing with increasing amounts of XI was observed (Table 2). Solutions containing ratios of purified ADH to XI of 1:1, 1:9, and 49:1 were examined in a biochemical assay to determine the activity of ADH in the presence and absence of XI. No change in the activity of ADH enzyme was observed in the presence of increasing XI (Table 2). Conversely, high levels of ADH1 expression may affect activity of XI and/or XKS because the rate of growth on xylose dramatically decreases with ADH1 overexpression.
Biochemical assay to measure alcohol dehydrogenase activity in presence of increasing concentrations of xylose isomerase (XI)
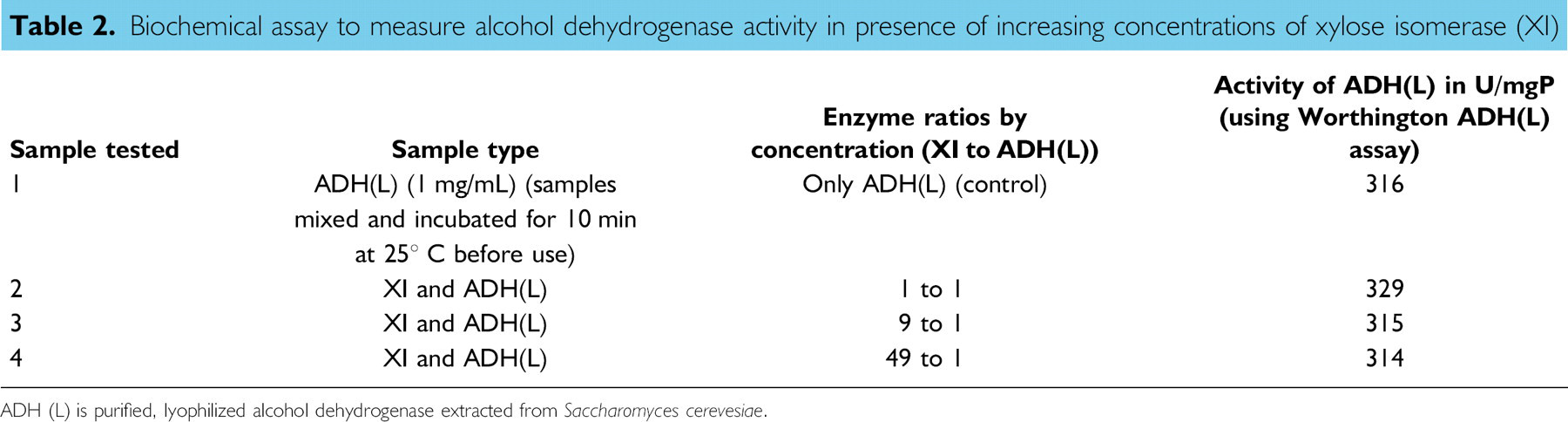

ADH (L) is purified, lyophilized alcohol dehydrogenase extracted from Saccharomyces cerevesiae.
Ethanol Production from Quadruple-Plasmid Strains Overexpressing ADH1 from Single Sugar Cultures
The amount of ethanol produced from single sugar use measured by gas chromatography is presented in Figure 6. Strains were grown on CM 2% glucose -URA-TRP-LEU-HIS in a 24-well deep-well plate for 6 days. The strain expressing XI, XKS, the anaerobic xylose utilization yeast protein COX10, and overexpressing ADH1 produced the greatest amount of ethanol when grown on glucose, 0.570 g/L, among the five quadruple-plasmid strains overexpressing ADH1. The COX10 strain also had the shortest doubling time grown on glucose. No ethanol was produced by any strain when the strains were grown on CM 2% xylose -URA-TRP-LEU-HIS in a 24-well deep-well plate for 6 days (data not shown).
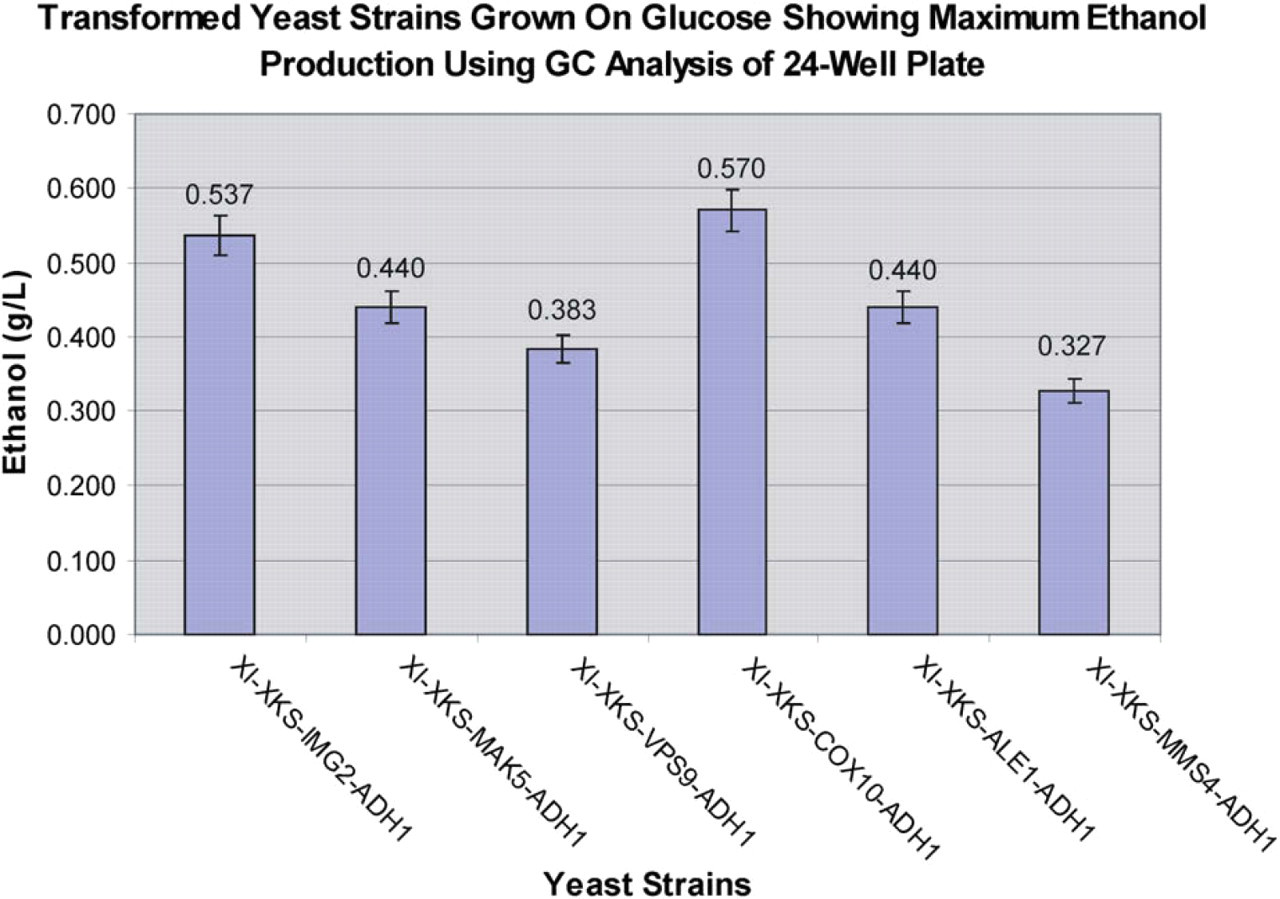
Ethanol production from quadruple-plasmid diploid yeast strains grown in CM 2% glucose -URA-TRP-LEU-HIS liquid medium in a 24-well deep-well plate (n = 3) for each of the strains analyzed. Cultures were incubated for 6 days at 30 °C with shaking at 100 rpm in a volume of 3 mL. Error bars represent standard deviations.
Ethanol Production and Consumption of Glucose, Xylose, and Arabinose from Wheat Straw Hydrolysate
Production of ethanol and the consumption of glucose, xylose, and arabinose from wheat straw alkaline hydrolysate in a 24-well deep-well plate for the ADH1-transformed quadruple-plasmid strains are presented in Figure 7. The MAK5, XI, XKS triple-plasmid strain (not transformed with ADH1) was used as a control. It may be that the MAK5 strain does not completely represent all of the three-plasmid strains, because a direct comparison of the MAK5 three-plasmid strain to the MAK5 four-plasmid strain indicates that of the parameters measured, only ethanol production and xylose consumption are altered in the four-plasmid strain. That may not be the case for the other four-plasmid strains. All of the four-plasmid strains except one showed complete consumption of available glucose, slight utilization of xylose, and no consumption of arabinose. The exception was the COX10, XI, XKS, ADH1 strain, which used all of the available xylose and most of the arabinose and all of the glucose. It might have been expected that overexpression of ADH1 would increase ethanol production; however, the strain not transformed with ADH1 produced more ethanol than the ADH1-transformed strains. The COX10 strain with the shortest doubling time on glucose (58.90 h) did not produce as much ethanol as the IMG2 or ALE1 strains that had the longest doubling times (69.83 and 76.11 h, respectively). The highest cell densities at the final endpoint (OD 1.10) were in the MAK5 triple-plasmid strain and the MMS4 quadruple-plasmid strain (Fig. 7). It appears that ADH1 overexpression impacts the growth as well as the ethanol production. Xylose utilization and arabinose consumption appear affected primarily in the COX10 strain.
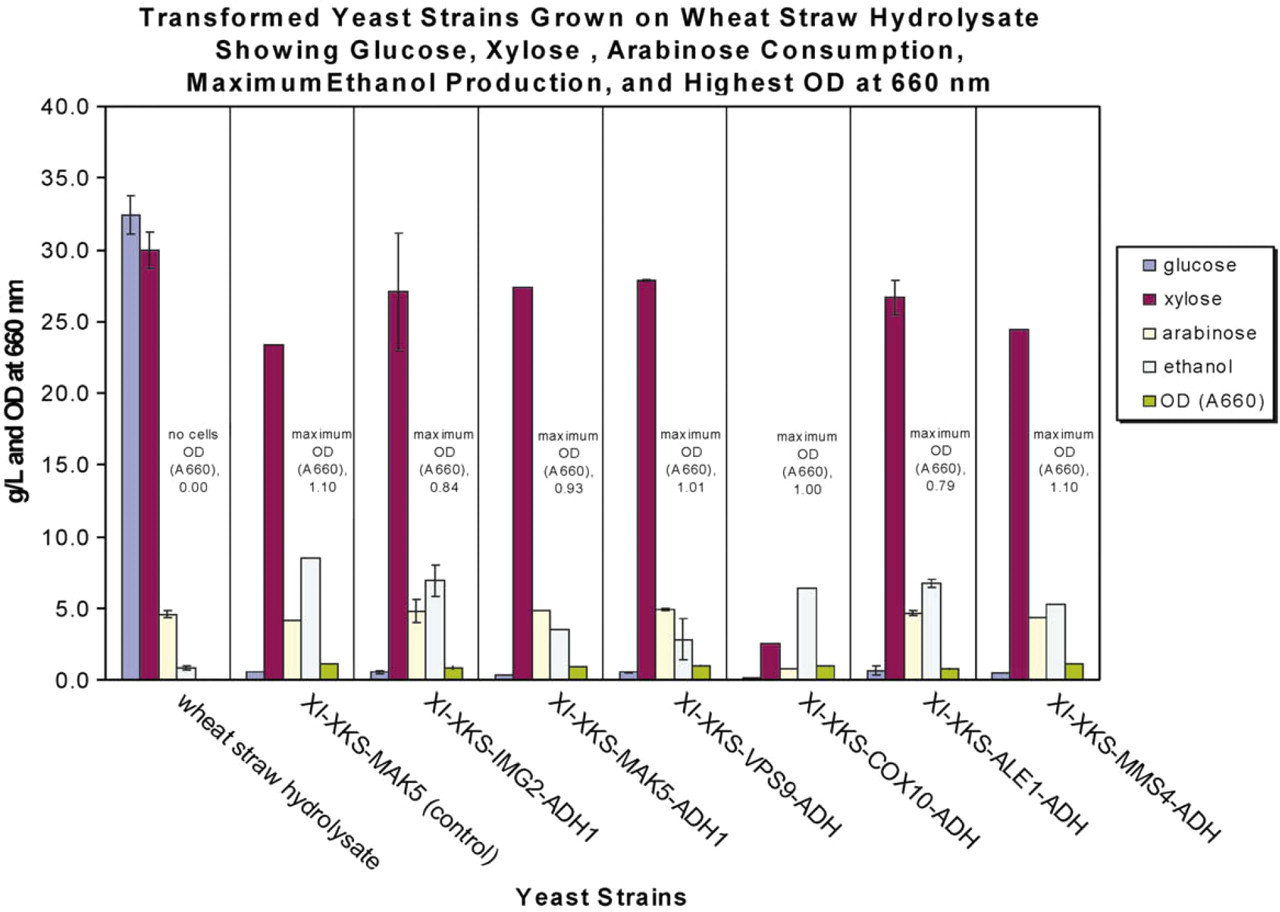
Maximum amounts of ethanol from quadruple-plasmid diploid yeast strains that gave optimum growth on wheat straw alkaline hydrolysate. The glucose, xylose, and arabinose levels are also shown after 8-day growth at 30 °C with shaking at 100 rpm in 5 mL of wheat straw hydrolysate in a 24-well deep-well plate (n = 3). Optical density at 600 nm is also indicated. Control is the triple-plasmid Saccharomyces cerevisiae diploid strain engineered to express XI, XKS, and an anaerobic xylose utilization gene (MAK5). Error bars represent standard deviations.
Conclusion
Five yeast ORFs were identified that encoded proteins that bound to XI, including ADH1, CSM4, APM1, and RNR1, which have known functions in the yeast cell, and one with locus tag YOR342C, which does not have an assigned function, using an automated two-hybrid interaction protocol. The four proteins with assigned functions are all enzymes that are critically important to the growth of the yeast cell. These may serve as potential targets for improving xylose utilization by S. cerevisiae. The effect of ADH1 overexpression was examined using an automated protocol to transform previously identified yeast strains that showed anaerobic growth on xylose. One of the transformants grew on wheat straw hydrolysate and consumed all available glucose, xylose, and arabinose. This strain will be used for further optimization of S. cerevisiae for cellulosic fuel ethanol production. Strains containing the other XI-binding proteins will also be used for further investigation.
Acknowledgments
We thank Karen Hughes for critical reading and formatting of the manuscript. We also thank Nathane Orwig for performing the sequencing work. We acknowledge the technical assistance of Francesca L. Scalpati, Thomas R. Ryan, David M. Skrincosky, and Von Worthington at Worthington Biochemical Corporation, Lakewood, NJ in performing the in vitro analysis of ADH activity. The gift of the library of pOAD prey plasmids for the yeast two-hybrid screen from Stanley Fields at University of Washington, Seattle, WA is greatly appreciated.


