Abstract
Major changes in production and distribution practices in the global food industry are prompting increased scientific efforts to identify, understand, and control the transmission of microbial contaminants. Through the development of advanced diagnostic tools, significant inroads in microbial source tracking are making important contributions to the safety of the food supply.
For decades, conventional or phenotypic tests involving culture-based and biochemical approaches have served as the primary method of identification for unknown microorganisms. Because they require unique growth requirements, phenotypic tests can sometimes be difficult to perform and interpretation can be highly subjective. These factors, combined with several other limitations—such as long incubation times, poor reproducibility, and requisite expertise to identify microorganisms microscopically—are key reasons why phenotypic methods have largely fallen out of favor in the food testing industry.
In the 1990s, the advent of molecular biology and polymerase chain reaction (PCR) ushered in a new era of genetics-based technology for the detection and identification of bacteria, yeasts, and molds. PCR requires DNA as the starting material and is used to amplify a specific DNA sequence (“DNA target”). With every cycle of PCR, the number of copies of the DNA target doubles; thus, very little DNA is needed as the starting template. More recently, detection using PCR and gel electrophoresis has been replaced by real-time PCR and the use of fluorogenic dyes, enabling visualization of detection and shorter times for results.
As the technology surrounding molecular biology continues to advance, microbial genome sequence databases are expanding. Considering the objectivity provided by DNA sequence information, gene sequencing has emerged as the “gold standard” for microbial identification. Given the limitations of phenotypic identification, DNA-based testing provides more definitive results and eliminates the subjective interpretations that are typical in traditional biochemical methods.
Great Unknown
Bacteria, yeasts, and molds are diverse in nature, and knowing whether or not an unknown microorganism, present in a food product or a processing environment, is a pathogen, spoilage microorganism, or toxin producer is necessary to avoid recalls, minimize wasted product, and solve shelf-life problems. Research studies have shown that microbial identification by gene sequencing provides fast, specific, and reliable results.
The gene sequencing process involves sequencing a specific region of the genome. For identification purposes, the ribosomal DNA sequences often serve as the target for sequencing. These regions are stable and relatively conserved, subject to a low rate of polymorphism and thus are able to reasonably characterize microorganisms at genus and species level.
A number of gene sequencing systems are commercially available in the food industry, including the MicroSeq Microbial Identification System by Applied Biosystems (Foster City, CA). 1 3 The MicroSeq system analyzes the ribosomal RNA gene sequence of an unknown bacterium, yeast, or mold and matches it to sequence of a known organism within the extensive MicroSeq database. MicroSeq uses PCR to amplify a region of the 16S rRNA gene of bacterial isolates and the D2 region of the large subunit rRNA gene of fungal isolates. Sequencing is performed via capillary electro-phoresis, which separates dye-terminator sequencing products. MicroSeq software captures and analyzes the data, and the resulting sequence is matched against a known sequence for identification.
The benefits of identification by gene sequencing are many, including shorter turnaround time for results, higher specificity and reliability, and higher reproducibility of results. Gene sequencing technology is especially advantageous in the identification of molds. Some molds, particularly the Ascomycota (e.g., Eupenicillium), may require up to 3–4 weeks for colonies to develop, followed by biochemical tests. Gene sequencing considerably speeds up identification. Moreover, the relative ease of microbial genome sequencing due to advancements in technology allows creation of more extensive sequence databases and more accurate identifications.
Genus and Species Level
Although ribosomal DNA-based gene sequencing provides identification of an unknown microorganism down to the genus and species level, at times it is important to know the particular strain of a given genus species. Molecular sub-typing, also referred to as strain typing and fingerprinting, provides differentiation beyond the species or subspecies level. Common molecular subtyping methods include pulsed field gel electrophoresis (PFGE), ribotyping, repetitive element-PCR (rep-PCR), and DNA-sequence—based methods such as multilocus sequence typing.
PFGE provides strain characterization through whole genome digestion with a restriction enzyme to generate the DNA fingerprint. PFGE has long been considered the “gold standard” for subtyping of bacterial pathogens as it offers high discriminatory power. However, the technique has many disadvantages: it is time consuming; requires a high level of technical skill; can be difficult to interpret; and approved protocols have been developed for only a few microorganisms of interest. The presence of conserved noncoding repetitive DNA sequences found interspersed in the chromosomes of microorganisms allows opportunities for DNA fingerprinting by PCR. 6 These “rep-elements” have been found in nearly all bacteria, mold, and yeasts studied to date. 4 5 Rep-PCR primers target these rep-elements, and the resulting amplicons can be separated by gel electrophoresis for visualization of the banding pattern or DNA fingerprint (Fig. 1).
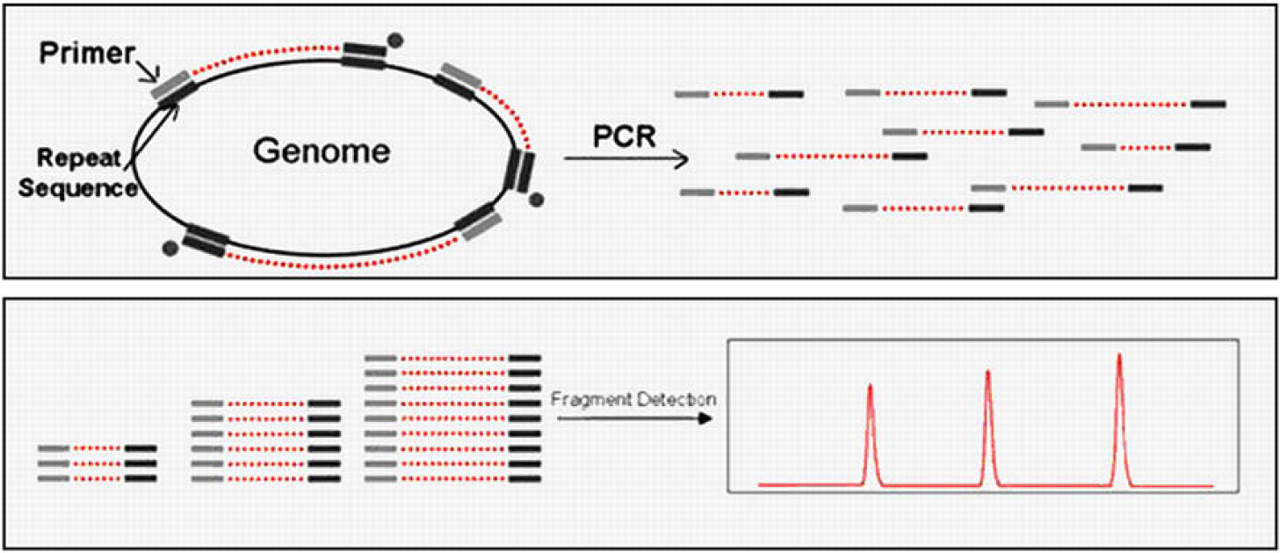
Repetitive element-PCR diagram.
At the forefront of this technology is the DiversiLab System, an automated rep-PCR system for strain typing developed by Bacterial Barcodes, Inc. (Athens, GA) The system uses kits that include rep-PCR primers to work with a specific genus and use a microfluidic chip to separate and detect amplicons. The automated rep-PCR process uses web-based software for data analysis. An excellent tool for strain comparison, the DiversiLab System is relatively easy to use, highly reproducible, and offers quick turnaround for results. The entire DiversiLab process takes approximately 4 h for one chip, or 13 samples.
Representative Fingerprints
Determining a microorganism's DNA fingerprint allows for strain-to-strain comparison. Discrimination between strains can be performed only if the strains have been typed through the same methodology (i.e., a ribotype pattern cannot be compared to a rep-PCR fingerprint). Strain typing provides valuable information and can be used in many ways including microbial source tracking.
The concept behind microbial source tracking in a food production or processing environment is to track the source and spread of the bacterial contamination throughout the food production system, from checking incoming raw materials to testing finished product. Sampling at different points of the production chain will allow a food producer to identify contamination sources and sites for cross-contamination that may lead to an undesirable outcome (e.g., product spoilage or recall due to presence of a pathogen). Potential data applications from microbial source tracking include devising plans for sampling or sanitization and/or disinfection. In addition to providing a robust and easy-to-use workflow for strain comparison, the DiversiLab System allows the user to build a user-specific isolate fingerprint library.
Gene sequencing provides information on the genus and species of a microorganism, thus allowing identification of which microorganism may be causing spoilage issues and product formulation problems that result in a decrease in the shelf-life of the product and ultimately, economic loss. 1 6
