Abstract
A rapid and inexpensive DNA diagnostic platform for fingerprinting Cdn. registered, wheat varieties has been developed. Two current, real-time applications being used in Canada include the determination of purity (% contamination) of grain shipments in rail cars and the monitoring of field plots that represent a midge varietal blend. The quantification of a sample is accomplished by fingerprinting single seeds and a sample having a mixture of varieties can be assayed. The Rapid ID Technology (RIDT) platform enables high-speed, high-throughput, robotic labautomation and low-cost single nucleotide polymorphism (SNP)-DNA fingerprinting in wheat. The inexpensive seed DNA extraction method, rapid PCR amplification, and miniaturization of the Invader assay for SNP scoring are all paramount for fingerprinting millions of single seeds per year in numerous rail cars/field plots at a low cost. The cost factor is the dominant concern in any fingerprinting program and the total cost for consumables for a single marker tested is estimated at less than $0.60 per seed. The RIDT platform can also segregate closely related cultivars, an important characteristic since many varieties have a common genetic background. The results “from seed to fingerprint data” at a central lab can then be transferred electronically to any location using a laboratory information management system.
Keywords
Introduction
There is a great interest in developing an automated, robust, inexpensive, and high-throughput detection platform for the scoring of molecular markers in plants. The high-throughput expectation includes both high-sample processing and the multiplexing capability for numerous marker sites. For many cultivated crops, the most commonly used marker detection methods (amplified fragment length polymorphisms; sequence tagged sites and simple sequence repeats) have generated high-density genetic maps for regions containing economically important genes, 1 quantitative trait loci sites, 2 and fingerprints between cultivars. 3 However, these markers use a tedious, electrophoretic-based scoring platform in either gels or capillary electrophoresis (CE) and have a limited multiplexing capability. Although there are examples of fingerprinting for single genes (http://maswheat.ucdavis.edu), few exist for traits governed by multigenic loci in any actual genotyping program. 4 The difficulties of genotyping for large populations include the scaling up the number of assays, quantification, simultaneous scoring of multiple markers (multiplexing) in pyramiding schemes, and in particular, the cost per assay. Recently, the USDA-AR Regional Small Grains Genotyping Centers, using only a single marker and CE, demonstrated a need for more efficient DNA extraction technologies and marker diagnostic platforms. 5 Single-seed assays are preferred due to the ease of quantification. The percentage of contaminants or ratios of genotypes in any sample can easily be determined. However, for single-seed assays, high-speed scoring and high-throughput sample processing at a low cost are necessary.
Single nucleotide polymorphisms (SNPs) are becoming the next generation of molecular markers due to their abundant frequency and suitability for genotyping by high-throughput, automated, scoring platforms. SNPs have been discovered for a number of economically important plant species including maize, 6 wheat (http://wheat.pw.usda.gov/SNP), barley, 7 sugar beet, 8 soybean, 9 and rice. 10 SNPs are highly stable and often are found in the gene coding region, which allows for the discovery of gene-associated markers for phenotypic traits. 11 In addition, the SNP base call is independent of the plant's growing environment. There exists a plethora of SNP-scoring platforms each with its unique advantages and a detailed description of these is given by Kwok 12 and Vignal et al. 13 The seven pillars of any fingerprinting scheme are illustrated in Figure 1.

The seven important features of any fingerprinting scheme. Seeds are used, which does not require the growing of plants.
In Canada, the Federal Government mandated that Kernel Visual Distinguishability (KVD) be eliminated by year 2008 and be replaced by a farmer's declaration backed by a science-based fingerprinting mechanism. KVD is used by the wheat industry to phenotypically segregate wheat into seven to eight functionally different market classes (Fig. 2). Each class has a specific kernel size, shape, and appearance to color and grain inspectors can keep shipments segregated by class. KVD is the foundation of Canada's wheat grading system and has worked in keeping inferior varieties out of the grain pipeline. Customers specify a particular class of wheat with a minimum grade level. For example, a customer may require No.1 grade Cdn. Western Red Spring (CWRS) with a 95% purity of a mixture of up to seven CWRS varieties. For the simplest scenario, a single variety may be requested (Identity Preserved, IP, variety) such as a durum Strongfield shipment. However, KVD has now become obsolete because many varieties between classes cannot be segregated by their phenotypic appearance. There is a large genotype by environment interaction associated with the KVD traits thus reducing the reliability of KVD. With changes in farmer and customer demands, KVD is now regarded by many as a barrier to innovation as it limits the development of novel traits (hog feed and gas station wheat) and impairs the registration of these new cultivars. If a new variety with superior characteristics does not fit into KVD's appearance standard, it cannot be registered, grown or sold by farmers. For example, a low food quality, high yielding CWRW variety ideal for hog feed wheat cannot be registered because it is indistinguishable from the premium priced, high food quality CWRS varieties. Grain co-mingled at a port position can result in a downgraded vessel and represent a loss of more than $1 million. The Rapid ID Technology (RIDT) platform will ensure the consistent quality of Canada's grain exports, and the industry's exposure to accidental or intended co-mingling of varieties at a potentially significant cost will be reduced.
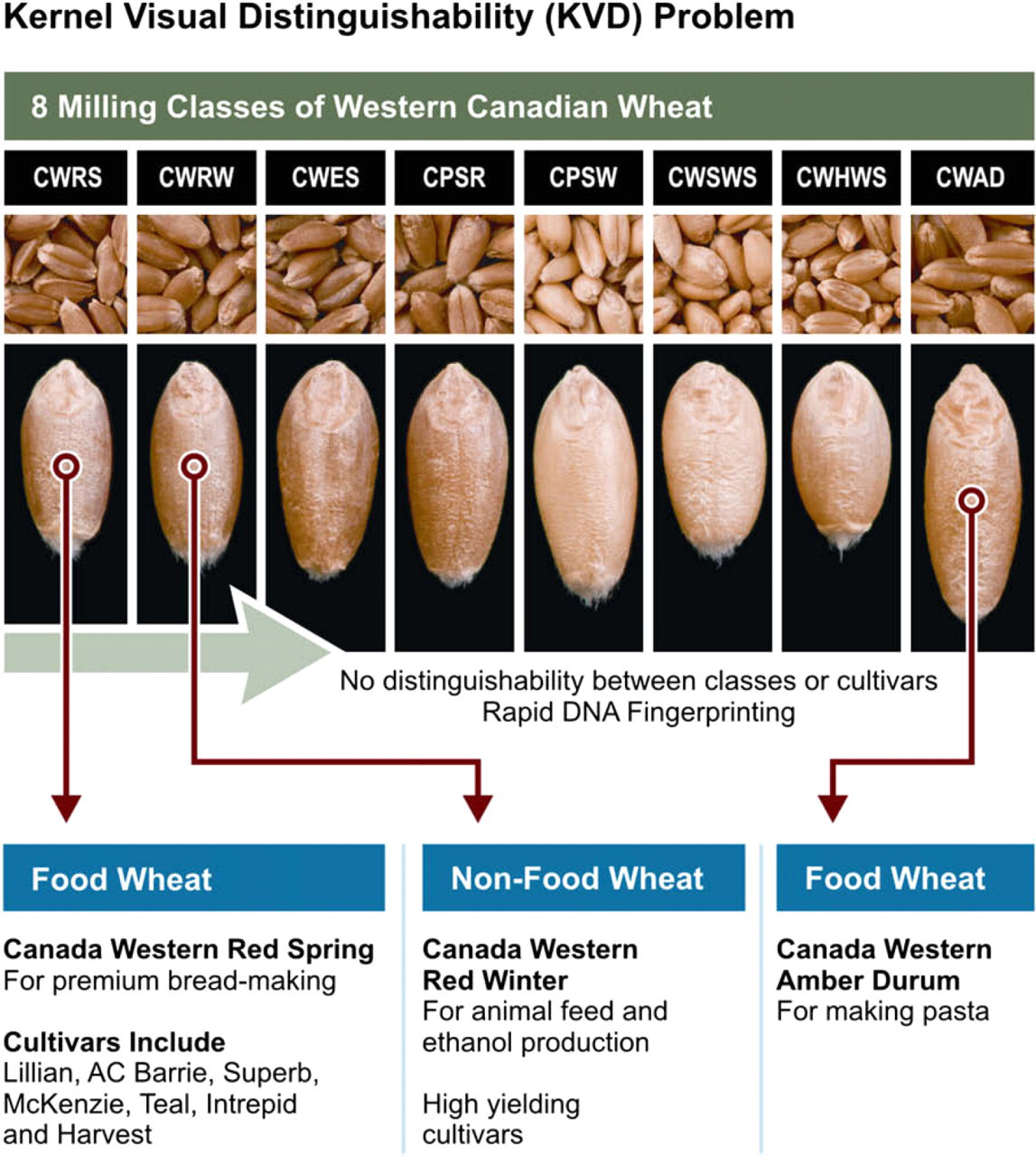
The seven to eight classes of Cdn. food wheat (CWRS, Cdn. Western Red Spring; CWRW, Red Winter; CWESRW, Extra Strong; CPSR, Cdn. Prairie Spring Red; CPSW, Cdn. Prairie Spring White; CWSWS, Cdn. Western Soft White Spring; CWCS, Cdn. Western Hard White Spring; CWAD, Cdn. Western Amber Durum). These classes are phenotypically segregated using Kernel Visual Distinguishability (KVD) by grain inspectors.
Another example of fingerprinting varieties by the RIDT platform is the monitoring of wheat midge varietal blends (Fig. 3). Varietal blends are a new concept not previously applied for in certified seed sales, and the Cdn. Seed Growers and Canadian Food & Inspection Agency needs information on the stability of these blends. Each blend is composed of 90% midge resistant versus 10% susceptible genotypes and this ratio must be monitored over wheat generations. The presence of susceptible genotypes will allow a sufficient number of avirulent midges to be produced in the refuge. These will mate with any rare midge harboring an avirulent-to-virulent midge gene mutation. The hybrid offspring of such matings would be avirulent and virulent midges would only occur if both parents were heterozygous for the virulence gene. This will keep the virulent midge frequency low in the blend and increase the longevity of the midge resistance gene (Sm1) effectiveness in the population. This Sm1 midge resistance gene is the only known resistance gene in western Canada and most single gene resistance populations are defeated in as few as 3 years (www.eurekalert.org/pub_releases/2008–02). The concept of using a proportion of susceptible plants among or in proximity to resistant plants to prevent or delay the development of insect resistance is now being required by law in commercially grown corn and cotton in the United States to protect the efficacy of Bt technology. 14
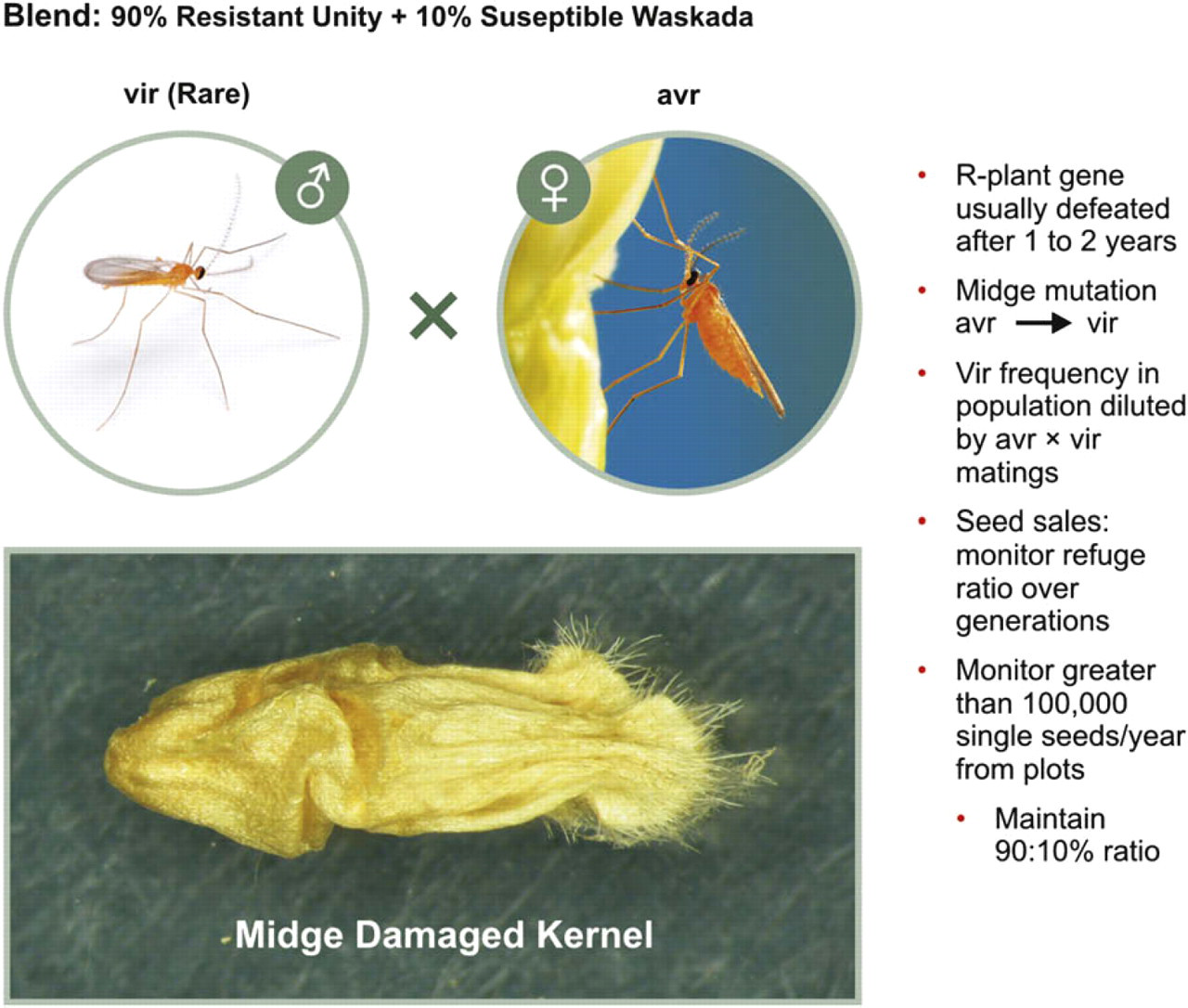
Increasing the longevity of single gene resistance lines by using a varietal blend—insect midge resistant refuges. Midge resistant plants (R-Unity; 90%) are interdispersed among midge susceptible plants (S-Waskada: 10%) to allow susceptible midges (avr) to survive on susceptible plants.
Experimental Details
Wheat SNP Database
A key element of the RIDT platform was the construction of the wheat SNP database of 96 genotypes by 200 SNP sites (Table 1). This database is updated frequently by the addition of new cultivars or wheat lines. In general, two 3′-end cDNA libraries were constructed from two wheat lines (FHB No. 37 and Thatcher) and individual clones were sequenced. From these sequences, oligonucleotide primer pairs were designed, PCR was performed on a diverse set of wheat genotypes, the amplicons sequenced and compared, and SNPs were discovered.
Example of a portion of Cereal Research Centre wheat single nucleotide polymorphism database a
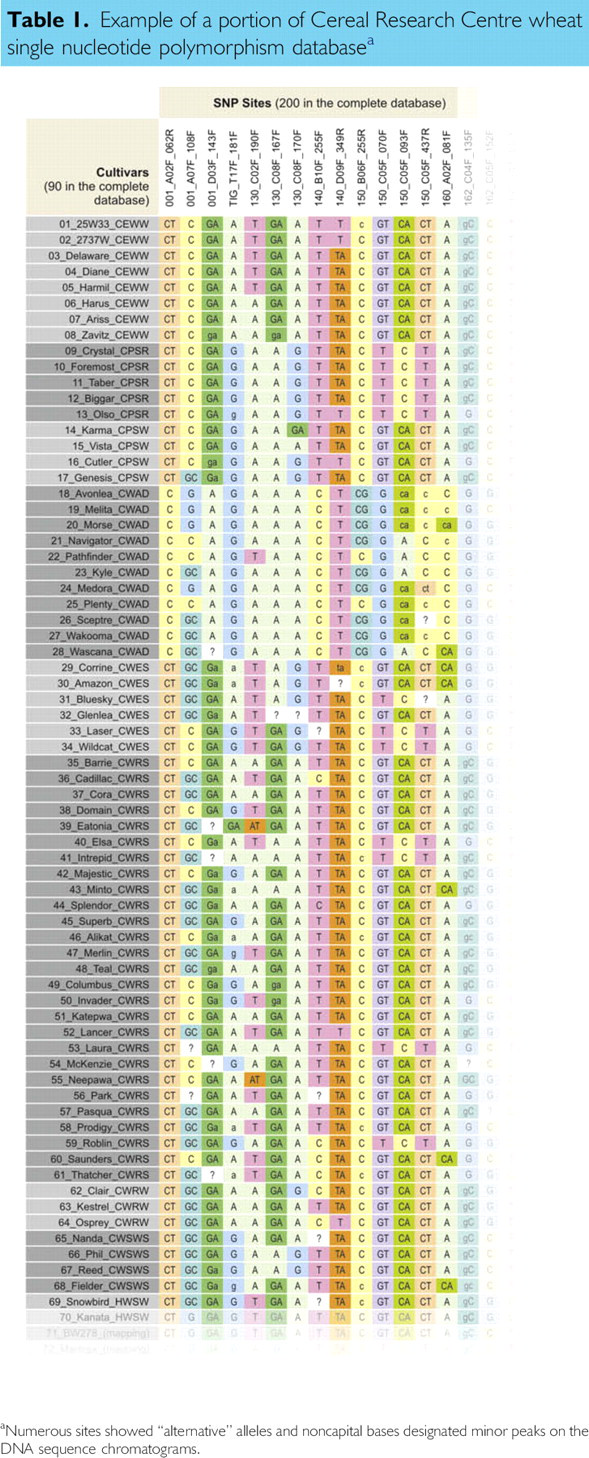
Numerous sites showed “alternative” alleles and noncapital bases designated minor peaks on the DNA sequence chromatograms
The cDNA sequences were assembled into contigs using the CAP3 assembly program 15 with the criteria of 95% homology over a 500 bp region. Singleton sequences that were singly represented and not part of any multimembered contig were selected. This enhanced the selection of single copy genes. Primer pairs were designed using DNAMAN Sequence Analysis Software (Lynnon Corp., Quebec, Canada) from the more polymorphic 3′-end untranslated region of these cDNA genes. Only the primer pairs that showed a single, clean amplification product from Thatcher genomic DNA were selected for further analysis.
These selected primer pairs were used for amplification of the 96 wheat cultivars/lines and the PCR products sequenced. The sequences were aligned using the Multiple Alignment program of the DNAMAN sequence analysis software. By comparative DNA sequence alignment, putative SNPs between the genotypes were identified. These base variations were checked with the DNA sequence chromatograms for confirmation.
An independent SNP detection method was used to cull SNP base calling errors from the putative SNP population. The Single Base Extension (SBE) method was done for SNP validation. The PCR product of the gene containing the putative SNP(s) was used as the template for this assay. SBE primers were designed with their 3′-ends immediately adjacent to the putative SNP position. The SBE reaction was done using the ABI Prism SnaPshot Multiplex kit (Applied Biosystems, Foster City, CA) and SBE fragments monitored in the ABI3100 Genetic Analyzer.
From the population of putative SNPs found by comparative sequence alignment, 45% were true SNPs as validated by the SBE assay.
Wheat is a polyploid species with three (hexaploid; 2n = 6X = 42 chromosomes; AABBDD) or two (durum tetraploid; 2n = 4X = 28 chromosomes; AABB) homoeologous genomes. These homoeologous genomes originate from different wheat species. Near-identical gene sequences originating from homoeologous genomes can be amplified resulting in multiple near-identical amplification products. The sequence data demonstrated that a single SNP site may show two alternative SNP bases at the same position within the amplicons generated from homoeologous genomes for a single genotype. A single SNP site was defined in this report as the base position in the PCR-generated amplicon(s) and this base(s) was polymorphic between genotypes. This site can then be used to fingerprint wheat cultivars/lines (Table 1).
Invader Genotyping Assay for SNP Base Calls
Most molecular marker analysis used today are time consuming, labor intensive, and cost prohibiting. Many involve electrophoretic separation by gels or CE thereby preventing truly high-throughput analysis of thousand of samples. 16 The first large-scale genotyping method used “gene chips,” 17 which now can yield more than 10,000–20,000 genotypes per microscope slide. However, the design and manufacture of microarrays are quite expensive and each one is not reusable (one slide per sample).
The invasive cleavage reaction (Third Wave Technologies, Madison WI) offers a simple, elegant, and specific method for genotyping SNPs using PCR products as templates. This reaction is based on cleavage of a unique secondary structure formed by the hybridization of a DNA target sequence to both an Invader oligonucleotide and primary probe that overlaps by one base (Fig. 4). The unpaired region at the 5′-end of the primary probe forms a flap that is a substrate for cleavage by a structure-specific 5′ endonuclease. Only in the presence of a perfect match between the primary probe and the template is the flap cleaved to drive a universal secondary cleavage reaction with a fluorescence resonance energy transfer (FRET) label. Signal is detected with a conventional fluorescence microtiter plate reader.
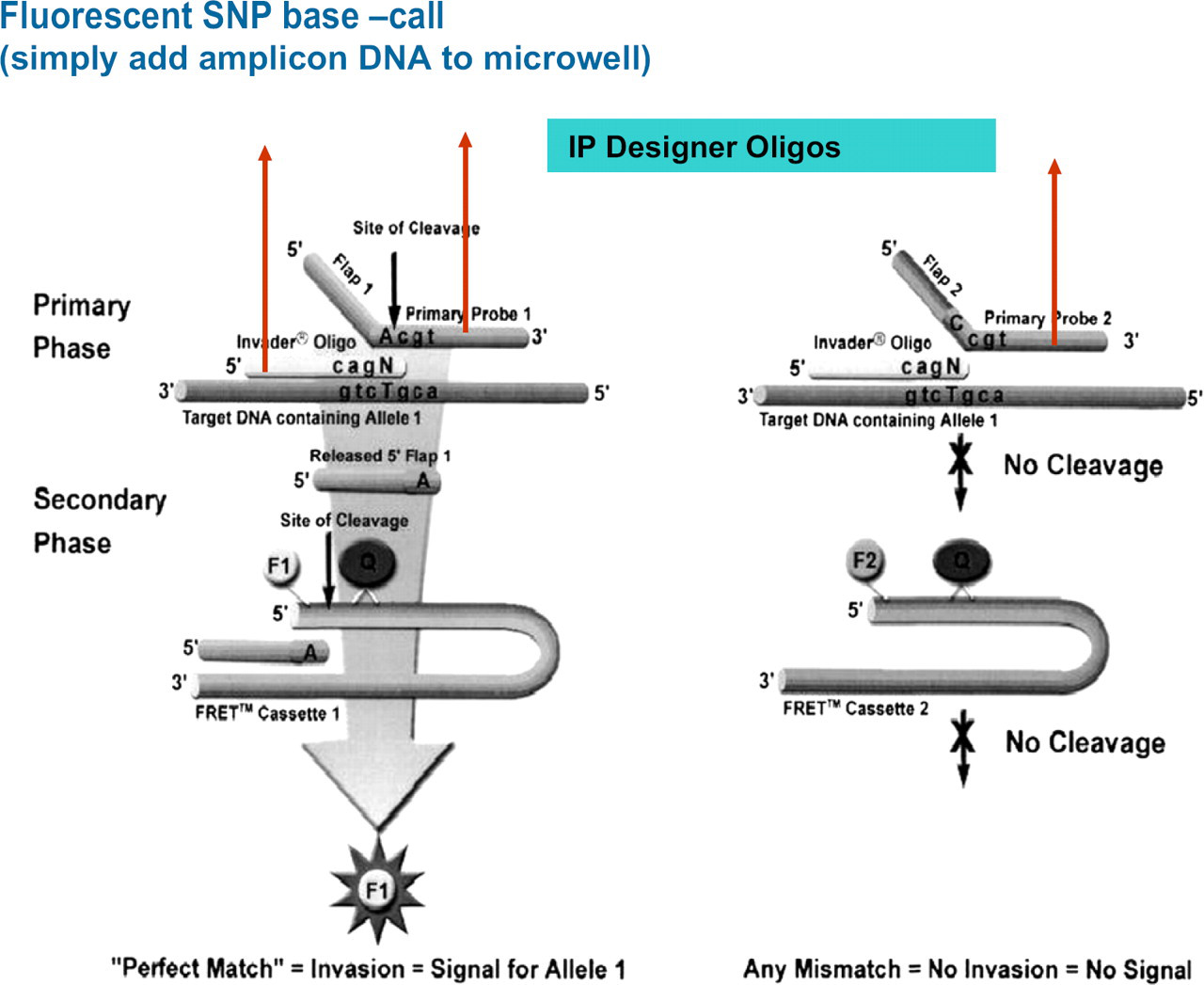
Third Wave Technology Genotyping assay. The intellectual property (IP) designer oligonucleotides are shown and are site specific.
The PCR-Invader technology has several features that suit its use for high-throughput genotyping. 18 19 The assay is performed entirely in a microtiter format with no electro-phoresis or purification steps. The reporter system can be dried down to the surface of the microtiter plate wells with only the addition of the PCR template required. The use of two oligonucleotides enhances the specificity for the intended target sequence and only “perfect matches” give a signal. This results in a higher level of specificity as compared to the commonly used hybridization schemes. The biplex reaction allows for single-base discrimination of alternative alleles present in polyploid genomes. Finally, the platform can easily be adapted to a labautomation robotic format. One disadvantage with the Invader technology is that only two alleles are interrogated in a single reaction well (biplex reaction). The development of more fluorescent dyes will allow greater multiplexing of loci for this stage.
RIDT Seed DNA Extraction System Coupled with Fast Thermocycling and Miniaturization
Quantification down to the 1.5% impurity level or ratios of resistant versus susceptible lines in a midge blend is a key prerequisite of the RIDT platform (Fig. 5). Although real-time PCR with the TaqMan-nuclease assays using numerous seeds in a single, bulked, group sample are widely used for quantitative measurements, variations during the PCR cycle can be incurred from several sources. 20 In particular, the SNP detection of the minor allele frequency can be suppressed. 21 Also, quantification on a mixture of up to seven different wheat varieties (CWRS class) is required for the Cdn. wheat transportation system. The RIDT platform uses the high-speed single-seed assays for the quantification.
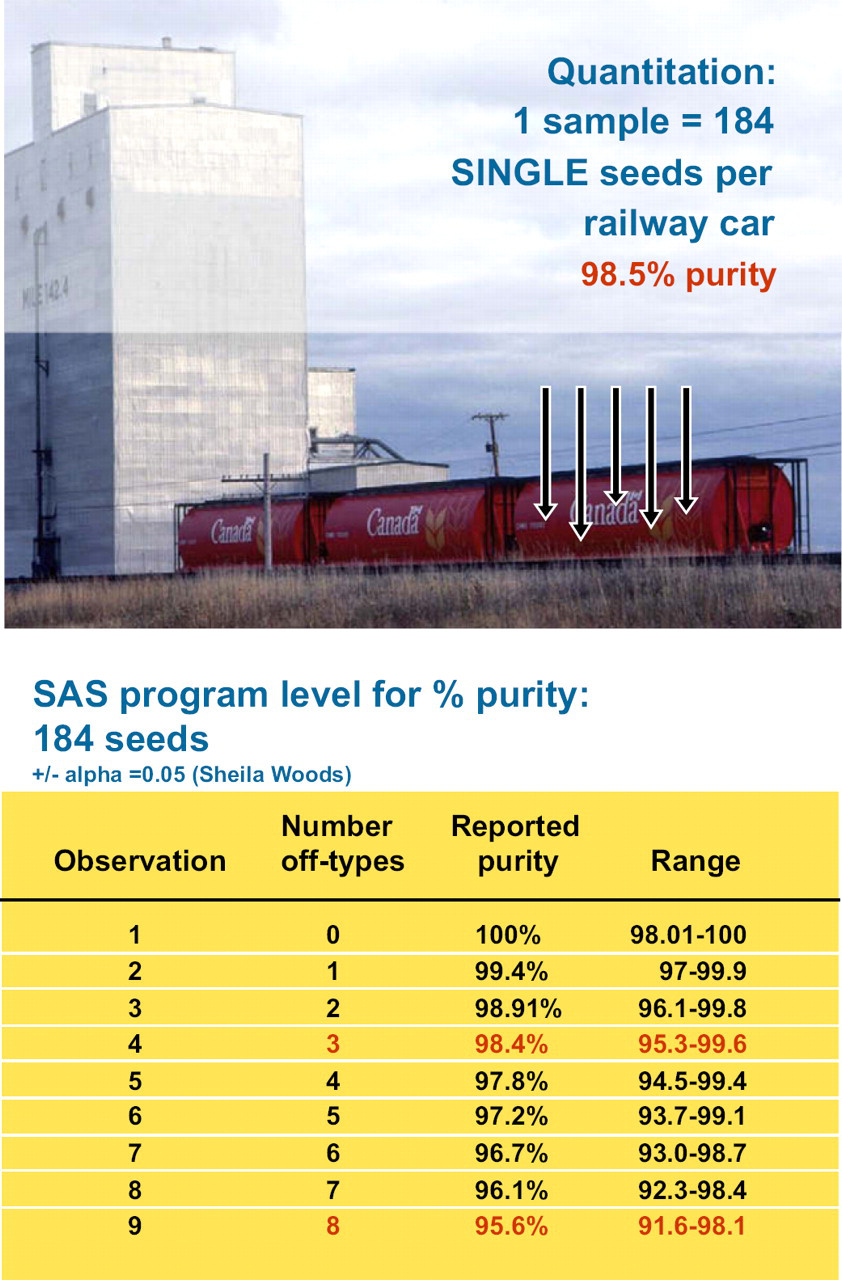

Seed sampling from a rail car. The two-sided SAS program gives the % purity with confidence intervals for a 184 seed sample.
The preparation of complex biological samples for high-throughput analysis remains a significant bottleneck. The most limiting factor in analyzing large plant populations on the molecular level is the extraction of DNA from the thousands of individual plants. This extraction of DNA is time consuming, labor intensive, requires the use of many chemicals and steps, and expensive. The removal of interfering species such as salts, detergents, and PCR inhibitors and the concentration of the dilute DNA samples when using a small amount of starting material are required. The vast majority of plant DNA extraction methods reported have been done on leaf tissue. This requires the growing of plants, which is a waste of greenhouse space, staff resources, and time. On the contrary, there are few reports on seed DNA extraction methods since it is more difficult due to the seed's flour/starch and phenolics, which inhibits PCR amplification of DNA. 22 23
For the RIDT Seed DNA Extraction System-TM (Figs. 6 and 7), single seeds are dispensed into 96-well microplates (one seed/well). A prototype robot capable of dispensing up to 10,000 of single seeds per day is required. There is no need for extensive maceration, cutting, or crushing that results in a “gluey pulp” of starch/flour. This inhibits subsequent liquid-handling manipulations and the starch polysaccharide inhibits the PCR. The procedure requires no time-consuming ethanol precipitation of DNA, centrifugation, and washing of the DNA precipitate. The genomic PCR is done directly on the crude DNA filtrate after plate centrifugation. This PCR is critical for this procedure and the PCR conditions may vary for each unique SNP site. The efficiency of PCR and “single band versus smear” results from genomic amplification are easily affected by subtle variations in reaction conditions due to the exponentially amplifying nature of PCR. Therefore, the major factors affecting the reproducibility and comparability of PCR must be considered and controlled to establish reliable PCR-based assays. These factors include the quality and quantity of components such as template DNA, Taq polymerase used, chemical components, primer oligonucleotides, and the performance of the deployed thermal cycler. A substantial portion of commercial thermal cyclers has been clearly demonstrated to insufficiently perform for rapid PCR proto-cols. 24 We routinely test multiple primer pairs for each SNP locus 25 and use the Applied Biosystems Verti Fast 96W Thermal Cycler (Ontario, Canada) with six independent temperature blocks to determine the optimal temperatures. The minimum cost of $1.50 per sample is prohibitive for large genotyping populations. Our estimated cost for the DNA extraction method used here is less than $0.01/sample and thousands of samples can be processed simultaneously.
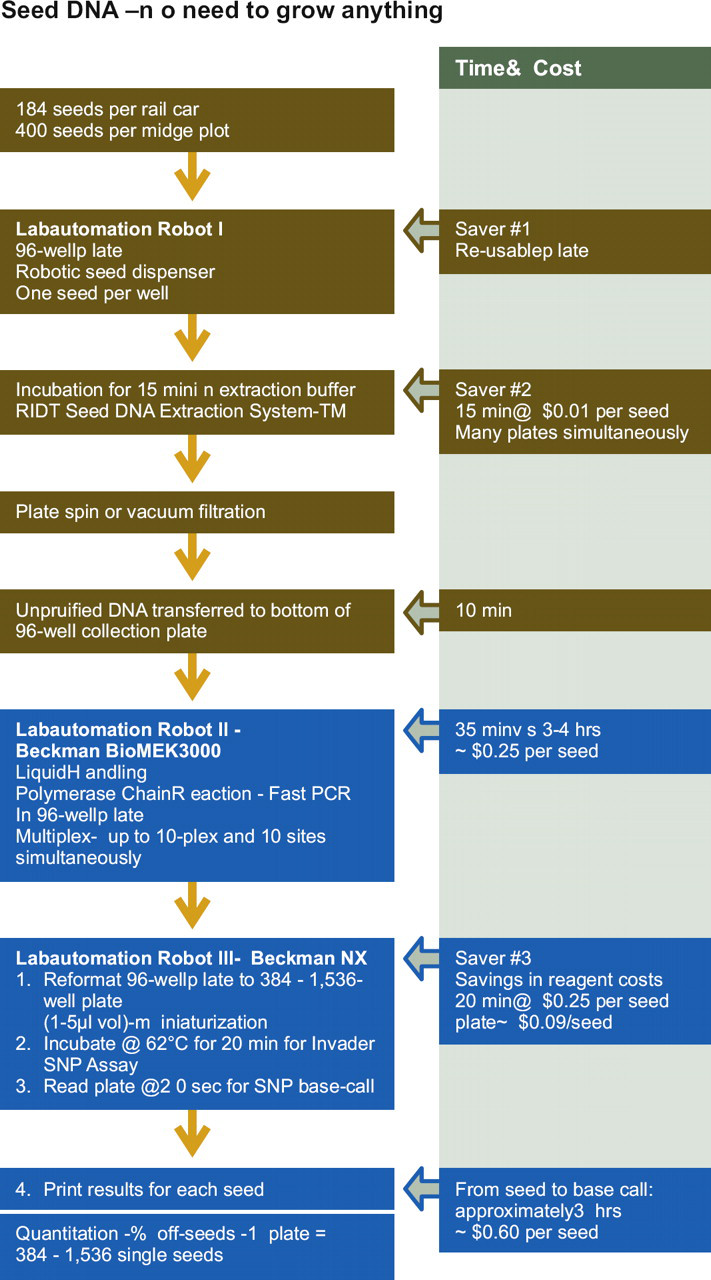
Rapid ID Technology (RIDT) seed DNA extraction system coupled with the Invader single nucleotide polymorphism (SNP) Genotyping assay. The approximate cost per seed and timelines are given.
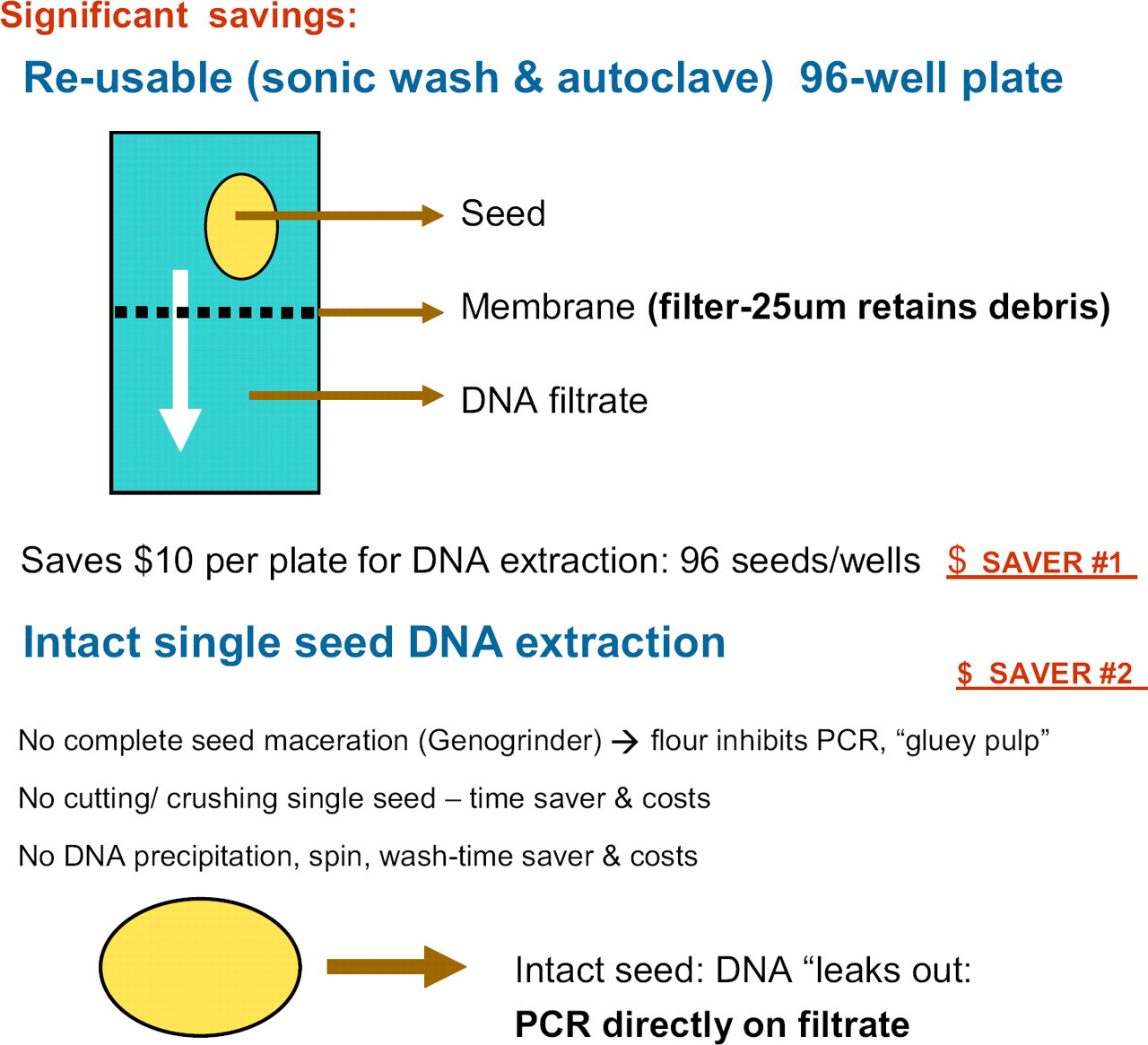
A simplified view of the reusable 96-well plate for seed DNA extraction.
The DNA filtrate in the 96-well collection plate is transferred into a 96-well PCR plate by the Biomek 3000 liquid-handling robot (Beckman Coulter Inc., Fullerton, CA). This robot also has tube to plate capability for dispensing the PCR reagents. The Applied Biosystem 9800 Fast PCR System reduces the reaction time to about 35 min and reagent costs are cut in half. After the PCR, the Beckman NX robot reformats four 96-well plates into a 384-well plate. Reformatting is also possible into one-quarter of a 1536-well plate. This miniaturization of the Invader reaction volume (1–5 μL) greatly reduces reagent costs. The Invader reagents (Invader, WT, MUT oligos; FRET core kit; enzyme/MgCl2) are premixed, added to each microwell, and subsequently the diluted (1:20) PCR amplicon DNA added. The Invader SNP assay is done for 20 min at 62 °C and the plate read by a standard fluorescence plate reader.
Applications
The RIDT platform and associated kits have a wide variety of uses in plants. One current real-time application being used in Canada is the fingerprinting of wheat varieties being shipped in rail cars and determining the contamination (% impurity) of these shipments. Another is the monitoring of field plots that represent a midge blend of 90% resistant versus 10% susceptible plants. Two other possible applications are its use in Marker Assisted Selection programs and Genetically Modified Organisms testing of plant seeds.
Varietal Fingerprinting (Cdn. Western Amber Durum, Durum)
An IP durum variety, Strongfield, has a 20% higher protein content or gluten strength than other Cdn. durum varieties and this quality is an important prerequisite for good spaghetti making. Strongfield quickly became one of the dominant varieties grown in Western Canada, along with Navigator, Avonlea, and Kyle. Other durum varieties comprised less than 8%, in total, of the acreage (Table 2). The goal was to fingerprint and segregate Strongfield from all other varieties grown in Western Canada and determine the % purity of Strongfield-containing rail cars destined to the port of Vancouver, BC, Canada.
Single nucleotide polymorphism (SNP) database for fingerprinting Identity Preserved Strongfield cultivar
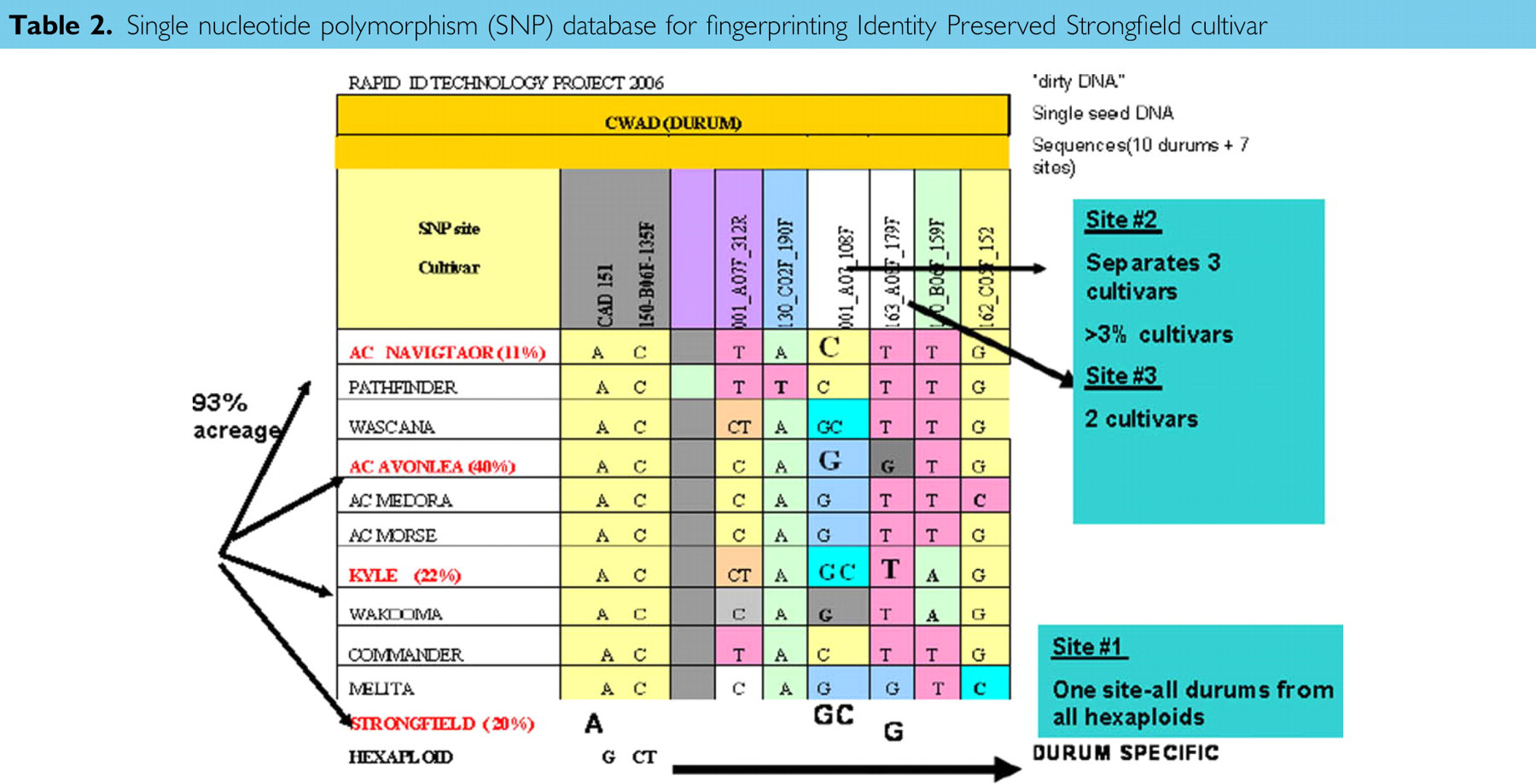
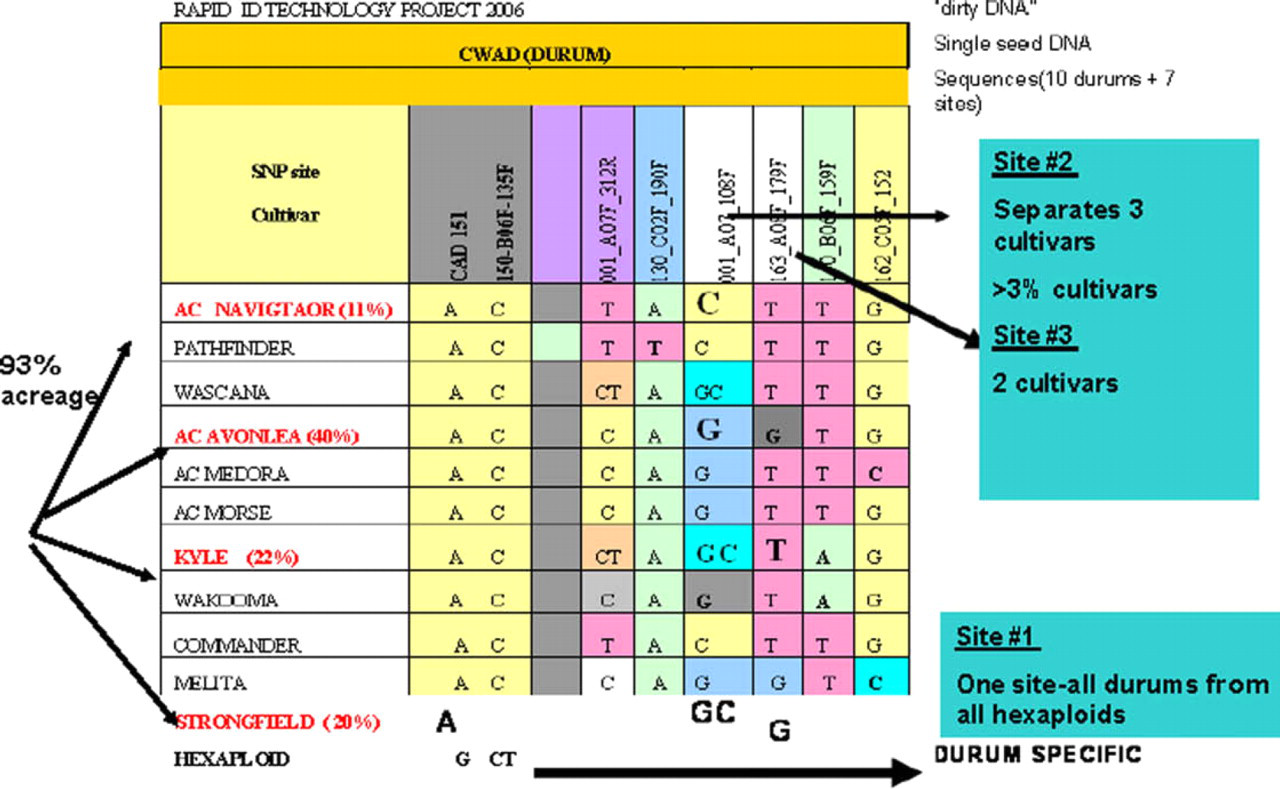
Using the RIDT seed DNA extraction system coupled with fast thermocycling, the subsequent Invader genotyping assay was done in 384-well microplates. A single SNP marker segregated 21 hexaploid from 10 durum (tetraploid) varieties (Table 2, Fig. 8, site 1). Two additional SNP sites can fingerprint the dominantly grown Navigator, Avonlea, Kyle, and Strongfield varieties (Fig. 8). This also demonstrated that the SNP markers can distinguish between closely related cultivars, an important characteristic. Strongfield is genetically related to both Kyle and Avonlea, Strongfield having the latter two as parents in a breeding program.
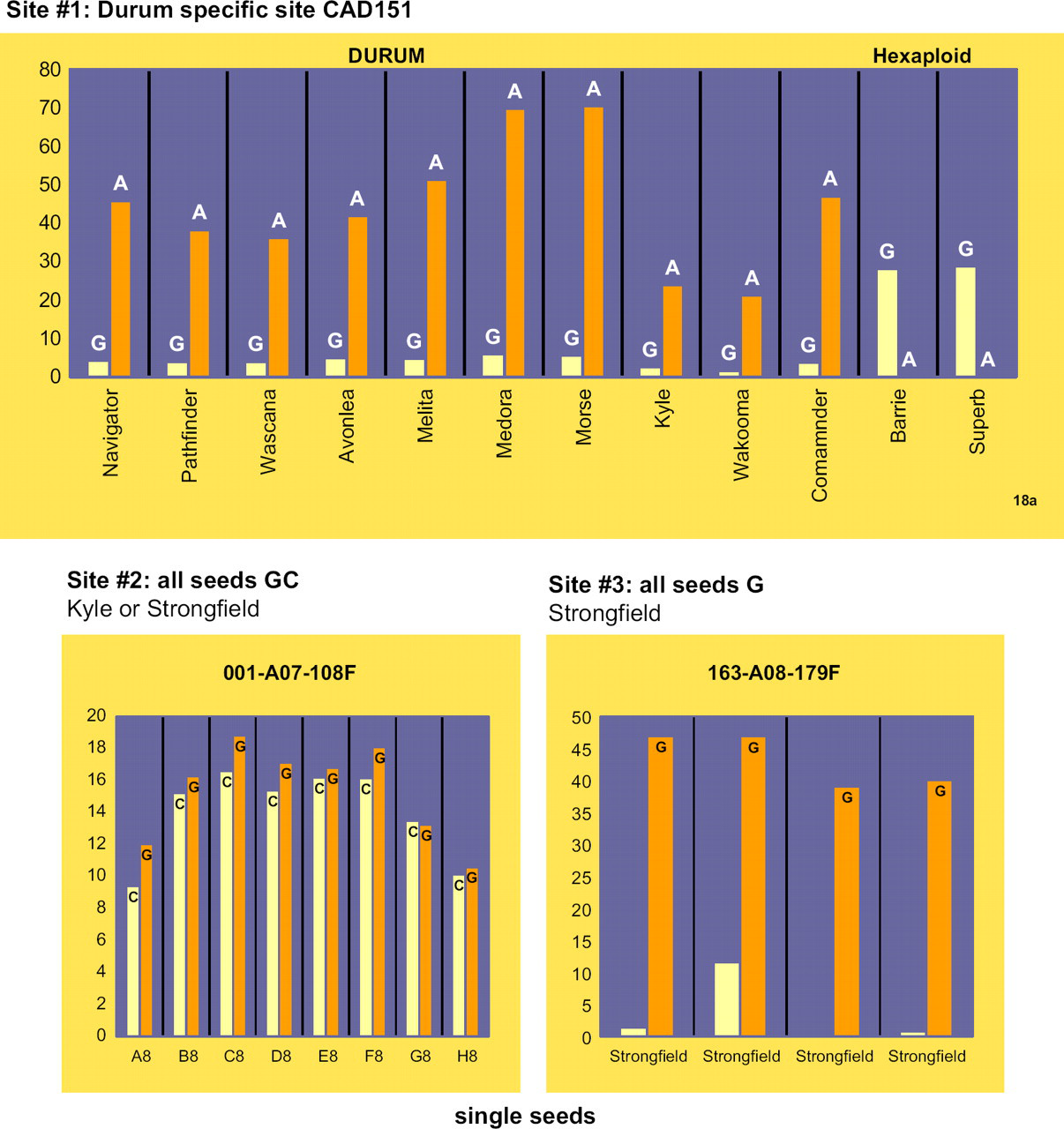
Invader single nucleotide polymorphism (SNP) assay for single-seed Strongfield fingerprinting. The minor peaks for sites #1 and #3 resulted from the control (mismatched/MUT) oligos used. Site #2 showed the alternative alleles (G/C).
Out of the 10 rail cars initially tested, 6 contained the expected 98–100% Strongfield grain. One car, claimed by the Cdn. Wheat Board (CWB) to have 100% Avonlea grain, only showed a 91% Avonlea purity by the RIDT test. The RIDT-SNP profile of three cars did not fit the CWB's claim of varieties present in these cars. This demonstrated that discrepancies can occur and an independent verification platform such as DNA fingerprinting is necessary.
Monitoring Wheat Midge Blends
A single antibiotic resistance gene (Sm1) against the midge fly has been introgressed into spring wheat from winter wheat by breeders. The wheat midge can cause significant damage to wheat crops by seed yield losses and downgrading of the kernels from CWRS#1 to #3. A midge blend comprising of 90% midge resistant: 10% susceptible will be used to protect the longevity of the single Sm1 gene's resistance in the field. The efficacy of using a blend to prolong the field resistance has been shown in Bt cotton and corn (www.eurekalert.org/pub_releases/2008).
Four genetically distinct blends, each grown in different environments under different soil conditions, climate, and midge pressure, will be monitored for their resistant versus susceptible plant ratios. In total, 12,800 single seeds will be fingerprinted using the RIDT platform (4 refuges x 100 seeds per plot x 4 replicas x 8 locations). This will be done for three successive wheat generations. This will evaluate whether an interdispersed refuge can maintain the 90:10 ratio through successive generations. An example of the Good-eve versus Intrepid blend after the first generation is shown in Fig. 9. A 92:8 ratio was observed, thus little or no drift was observed for this location and these genotypes. High rates of drift significantly different from the 90:10 ratio in subsequent generations would show the appropriate generation for the blending of resistant and susceptible varieties. Also, farmers can test their farm-saved seed and may need to use certified seed on occasion. This midge control is an example of biological control through environmentally friendly means by not using damaging pesticides that kill the midge's natural predators and will save the agriculture industry millions of dollars.
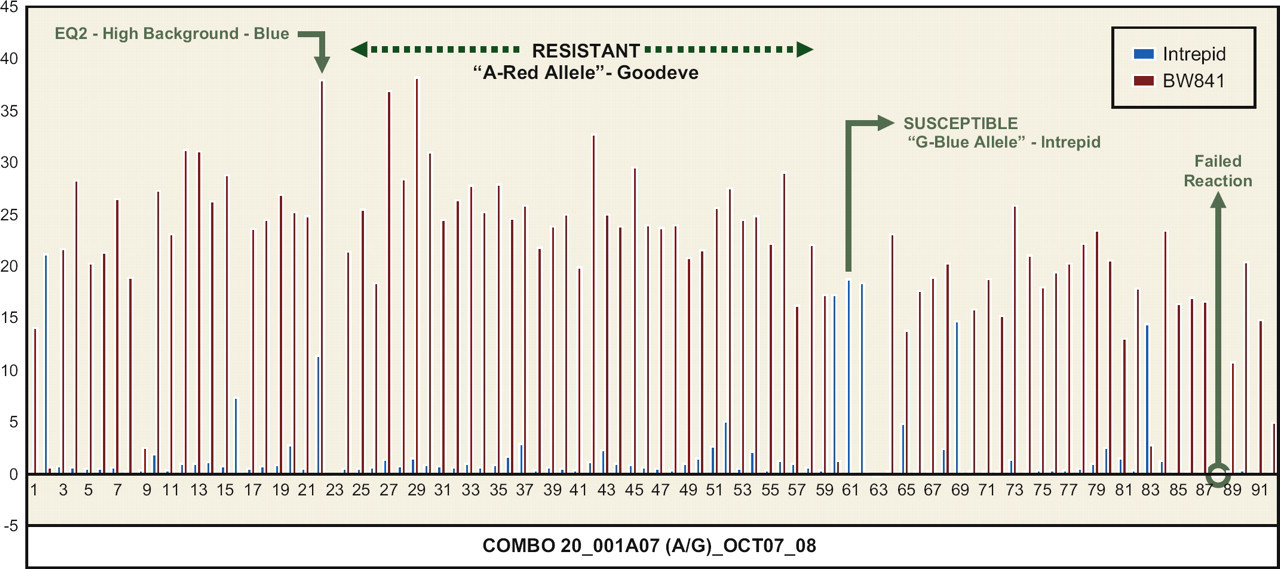
First generation midge blend #3-Melfort_plot 4–28: unequivocal scoring.
Conclusion and Outlook
We have demonstrated that the RIDT strategy can be applied to any large-scale fingerprinting platform with labautomation. RIDT can fingerprint and segregate IP wheat cultivars in rail cars and monitor the genotypes of wheat midge blends in field plots. With a high degree of labautomation and miniaturization, the cost factor can be reduced significantly. High-speed single-seed assays are used for the quantitative aspect of this genotyping protocol.
One future aim is achieving a very high throughput with the goal of fingerprinting millions of single-seed samples per year. Segregating food wheat from non-food wheat in rail car shipments and monitoring the approximately 85,000 Prairie wheat farms for compliance to the wheat midge refuge strategy is a primary goal. The combination of Third Wave Hologic (TWT Inc., Madison, WI) Invader-Plus genotyping system and the array tape-based automation technology (Douglas Scientific Inc.) can potentially achieve this objective. Tape-based system uses no microplates or thermal cylers and the PCR component is done in a water-tank bath with up to 186,000 samples on the tape roll. TWT's Invader-Plus SNP genotyping allows for both the PCR component and SNP detection to be done in the same well and both assays completed in hours.
Acknowledgment
This work was funded by a consortium of Government and Industry partners, which included the CWB, Ottawa (MII grant), Cdn. Seed Growers Association (CSG), and Cereal Research Centre (CRC, Winnipeg).