Abstract
The development of a tool that can provide early-stage predictive toxicology data may accelerate the identification of safer drug candidates, and thereby improve the clinical progression of drug candidates to pharmaceuticals. Such a system would require an accurate and reliable technique that is amenable to the large number of drug candidates that must be screened in the lead discovery and optimization stages of drug development. A key component of predictive toxicology is the ability to harness the metabolite-generating capacity of human cytochromes P450, which are involved in first-pass drug metabolism function of the liver. We have miniaturized P450 catalysis into a microarray format consisting of up to 11,200 isolated P450 reactions, each in 5 nL sol-gel spots, on a single functionalized glass microscope-size biochip. This dramatic scale down from more conventional 96 and 384-well plate scales (at least a 1000-fold reduction in volume) did not adversely affect P450 catalytic activity. Based on the functionality of the P450-containing microarray, we developed the metabolizing enzyme toxicology assay Chip (MetaChip), which combines high-throughput P450 catalysis with cell-based screening on a microscale platform. Proof of concept was demonstrated using anticancer prodrugs cyclophosphamide and Tegafur, as well as the analgesic acetaminophen. The MetaChip may provide a high-throughput microscale alternative to currently used

Introduction
The emergence of combinatorial methods for new compound synthesis and drug discovery has led to a dramatic increase in the number of drug candidates for
Despite the aforementioned developments, the drug discovery process remains an investment-intensive, high-risk endeavor that results in low yields of effective and safe drugs. For example, investment in pharmaceutical research and development in 2005 was approximately $40 billion, 5 which led to 20 new drugs approved by the Food and Drug Administration (FDA). The low yield is at least partly a result of the high failure rate due to toxicity of the NCE or its metabolite(s). 6 For a given pharmaceutical project, from the hundreds of thousands (in some cases millions) of compounds that are screened as potential drugs, roughly 10,000 show relevant biological activity, and only 5–10 enter clinical trials. Of these, on average, only one is ultimately approved by the FDA to become a marketable drug. 7 Thus, there is an acute need to develop high-throughput techniques that can be used in the early stages of drug development to assess the toxicity of drug candidates and their metabolites.
The human body contains a variety of enzymes that are involved in the metabolism of the myriad chemicals that comprise today's pharmaceuticals, as well as those found in foods and the environment. By far the most important drug-metabolizing enzymes are the cytochromes P450 (P450s), for which up to 58 distinct P450 genes have been identified, with most found in the liver.
8, 9
P450s play several critical roles in drug metabolism. First, P450s are directly involved in the initial clearance of drugs from the body,
10
which reduces the plasma concentration of a drug at a target site and affects the bioavailability in a manner that is difficult to predict using current technologies.
8
During this process, drug metabolites are generated, some of which are biologically active and exert the desired pharmacological effect. Often, however, drug metabolism can lead to undesirable biological consequences. A well-known example of a toxic metabolic response is the P450-catalyzed oxidation of the common analgesic acetaminophen to N-acetyl-
The advent of high-throughput screening (HTS) technologies has begun to impact P450 biocatalysis for toxicity and metabolism studies. The majority of HTS studies have focused on inhibition of P450 isoforms by specific compounds and mixtures of compounds, thus providing a window on the impact of specific drugs and their metabolites on potential toxicity. High-throughput P450 assays, which are often performed in 96 and 384-well microtiter plates and require sub-mg amounts of enzyme and inhibitor,
16
have been facilitated by the design of highly sensitive fluorogenic substrates that are tailored for specific P450 isoform reactivity.
17, 18
In addition to P450 inhibition studies, identification of the major P450 isoforms responsible for specific drug metabolism and the kinetic susceptibility of a compound to biotransformation are also critical determinants in establishing therapeutic efficacy and the rate of
Experimental
Sol-gel microarrays were prepared by mixing 25 μL methyltrimethoxysilane (MTMOS) with 10 μL HCl (5 mM) followed by sonication for 10 min. The MTMOS-HCl sol was mixed with 5 μL CYP3A4 baculosomes (1.1 nmol P450/ml from Invitrogen (Carlsbad, CA)), 5 μL regeneration system (333 mM glucose-6-phosphate and 40 U/mL glucose-6-phosphate dehydrogenase in 100 mM potassium phosphate buffer, pH 8), and 20 μL polyvinyl alcohol (PVA, MW 10,000, Sigma (St. Louis, MO)) in distilled water (10%, v/v). Typical glass microscope slides (57 × 25 mm2 usable surface) were treated before use by spin coating an MTMOS solution (pH 7, 2 mL at 3000 rpm for 30 s) onto the glass surface. Spotting was performed using a MicroSys 5100–4SQ microarrayer (Genomic Solutions, Ann Arbor, MI) and the sols were allowed to gel for 24 h at 4 °C. P450 reactions were performed in 5 nL sol-gel spots by dispensing 25 nL of green fluorogenic substrate solution (dibenzoxymethylfluorescein (DBOMF)), containing the NADPH regeneration system, 0.25 mM NADP+, and 20% (v/v) glycerol (to retard evaporation) on top of each sol-gel spot. After 1 h reaction in the microarray chamber at 100% relative humidity, the slides were dried. Microarrays were then imaged at 532 nm using a GenePix Professional 4200A array scanner (Axon Instruments, Inc., Union City, CA). The green fluorescence intensity was quantified from the array scanner using Axon GenePix Pro 6.0 software. The overall conversion of DBOMF within 1 h was no greater than 10%; therefore, this time was used to approximate initial rate kinetics.
Cell growth was performed as described by Lee et al. 21 Briefly, MCF7 human breast cancer cells (ATCC) were grown in Dulbecco's modified Eagle's medium (DMEM) (Sigma) containing 5% heat-inactivated fetal bovine serum (FBS, Gibco, Grand Island, NY) in T-150 cell-culture flasks in a humidified CO2 incubator (ThermoForma Electron Co., Marietta, OH) maintained at 37 °C and 5% CO2 in air. A confluent layer of cells was treated with 4 mL of 0.05% trypsin-0.53 mM EDTA (Gibco) from the culture flask, and the cells were resuspended in 20 mL of FBS-supplemented DMEM. After centrifuging at 50 × g for 8 min at 4 °C, the supernatant was removed and the cells were resuspended in 40 mL of FBS-supplemented DMEM. The cell suspension (3 mL containing ca. 4 × 105 cells/mL) was then transferred to a 57 × 25 mm2 usable surface area chamber slide (Fisher Scientific, Pittsburgh, PA) and the slide was incubated for 1 day in the CO2 incubator.
Results and Discussion
We have focused on sol-gel technology for the encapsulation of proteins onto glass microarrays. Sol-gels are optically transparent, can accommodate high concentrations of protein in their 3D matrix, and provide a microenvironment conducive to favorable protein properties. In addition, sol-gels are compatible with various organic moieties, and are stable in harsh environments. 22 A wide range of biological materials has been encapsulated in sol-gel matrices without significant loss of activity, including antibodies, enzymes, DNA, RNA, and live cells. 23, 24 Scale-down of enzymatic reactions in sol-gel matrices has been reported for volumes as low as 5 μL for reactions involving glucose oxidase, peroxidase, lipase, and several proteases. 25 That particular study showed that a wide range of enzymes were functional in microfabricated wells. In the current work, we demonstrate that P450s can be incorporated into sol-gel microarrays at volumes as low as 5 nL.
Spotting of MTMOS sols onto untreated glass microscope slides, followed by gelation, resulted in rapid detachment of the spots from the glass surface. To alleviate this problem and ensure a hemispherical spot geometry, MTMOS solution was spin coated onto the glass surface. In addition to preventing spot detachment, the hydrophobic MTMOS surface coating prevented the sol-gel spot from spreading across the surface, as shown in Figure 1. On a hydrophilic glass surface, a 1 μL MTMOS sol spot (Fig. 1(A)), which consists of ca. 60% (v/v) water, occupies a larger surface area (indicative of spreading) than on the hydrophobic MTMOS-coated slide (Fig. 1z(D)). Moreover, the MTMOS coating prevented P450 substrate solution from migrating from one spot to another. This is shown in Figure 1(B) and (E), where the substrate solution has migrated off of the sol-gel spot on an uncoated glass slide, yet remains on top of the sol-gel spot on the MTMOS-coated slide. A similar effect is observed for 5 nL spot sizes (Fig. 1(C) and (F)). For these reasons, MTMOS-coated glass slides were used throughout this work.
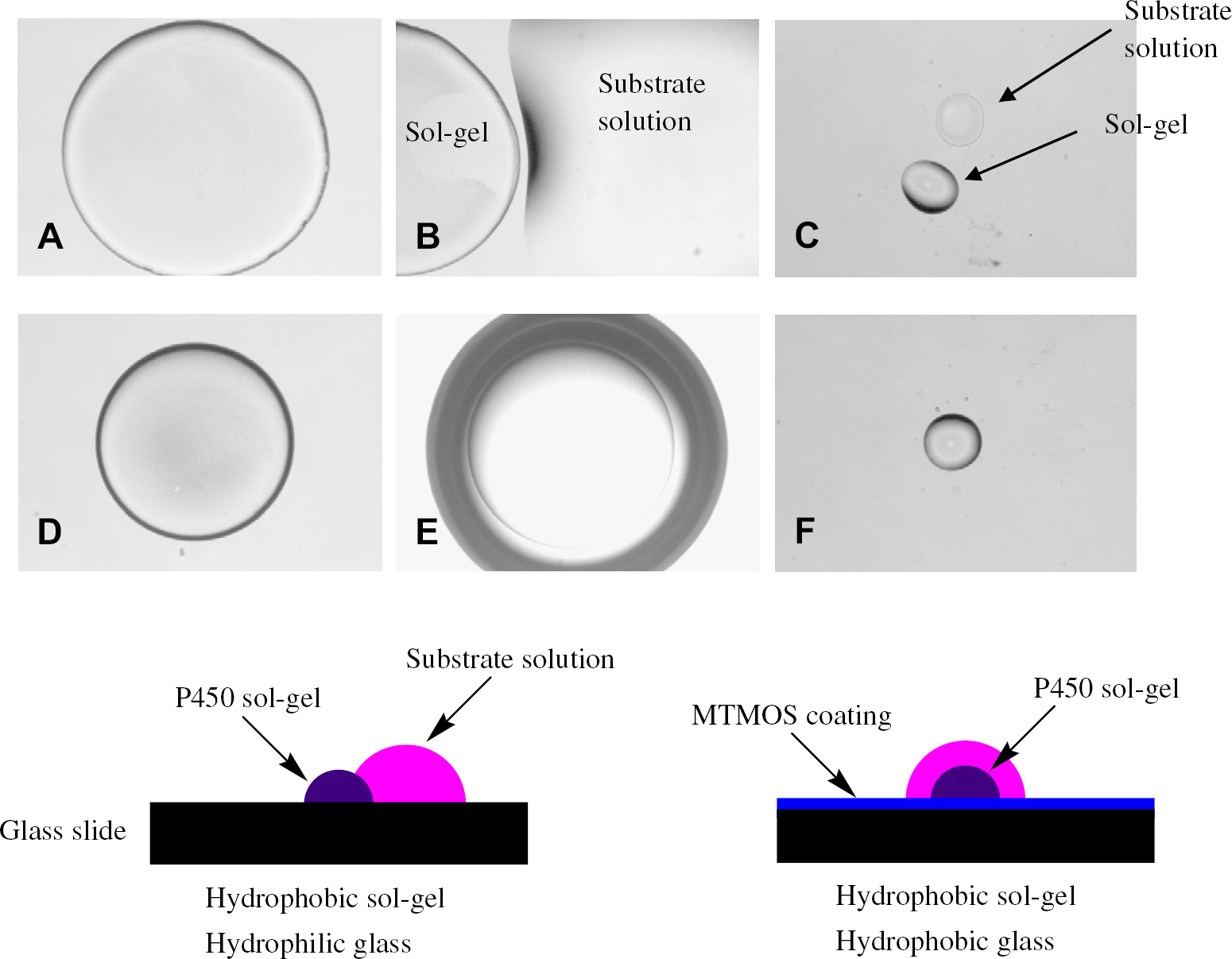
Effect of glass surface hydrophobicity on spotting of substrate solution on top of P450 sol-gel spot: (A) 1 μL of P450 sol-gel on untreated glass surface; (B) 1 μL of P450 sol-gel with 5 μL of substrate solution on untreated glass surface; (C) 5 nL of P450 sol-gel with 25 nL of substrate solution on untreated glass surface; (D) 1 μL of P450 sol-gel on MTMOS-coated glass surface; (E) 1 μL of P450 sol-gel with 5 μL of substrate solution on MTMOS-coated glass surface; and (F) 5 nL of P450 sol-gel with 25 nL of substrate solution on MTMOS-coated glass surface.
The stability of the sol-gel spots on the MTMOS-coated glass slides prompted us to examine the reactivity of a typical human P450 in a high-density microarray. To that end, the MTMOS sol solution containing CYP3A4 was printed onto the MTMOS-coated slide using the microarrayer and then allowed to gel for 24 h at 4 °C. P450 reactions were performed in 2800-spot arrays consisting of 35 × 80 spots (5 nL each, spot-to-spot (center-to-center) distance of 700 μm). CYP3A4 was active on the 2800-spot microarray, as shown in Figure 2 using DBOMF as a fluorogenic substrate, which was varied in concentration from 0.1 to 10 μM. The enzyme followed Michaelis-Menten kinetics with an apparent kcat/Km of 5.5 ± 0.3(× 104) M−1min−1. Corresponding values of CYP3A4 reactions in 1 μL sol-gel spots and in 1 μL aqueous solution (both on the slide format) were 2.0 ± 0.2(× 104) and 4.6 ± 0.4(× 104) M−1min−1, respectively. Thus, CYP3A4 retained its native activity in the sol-gel microarray.
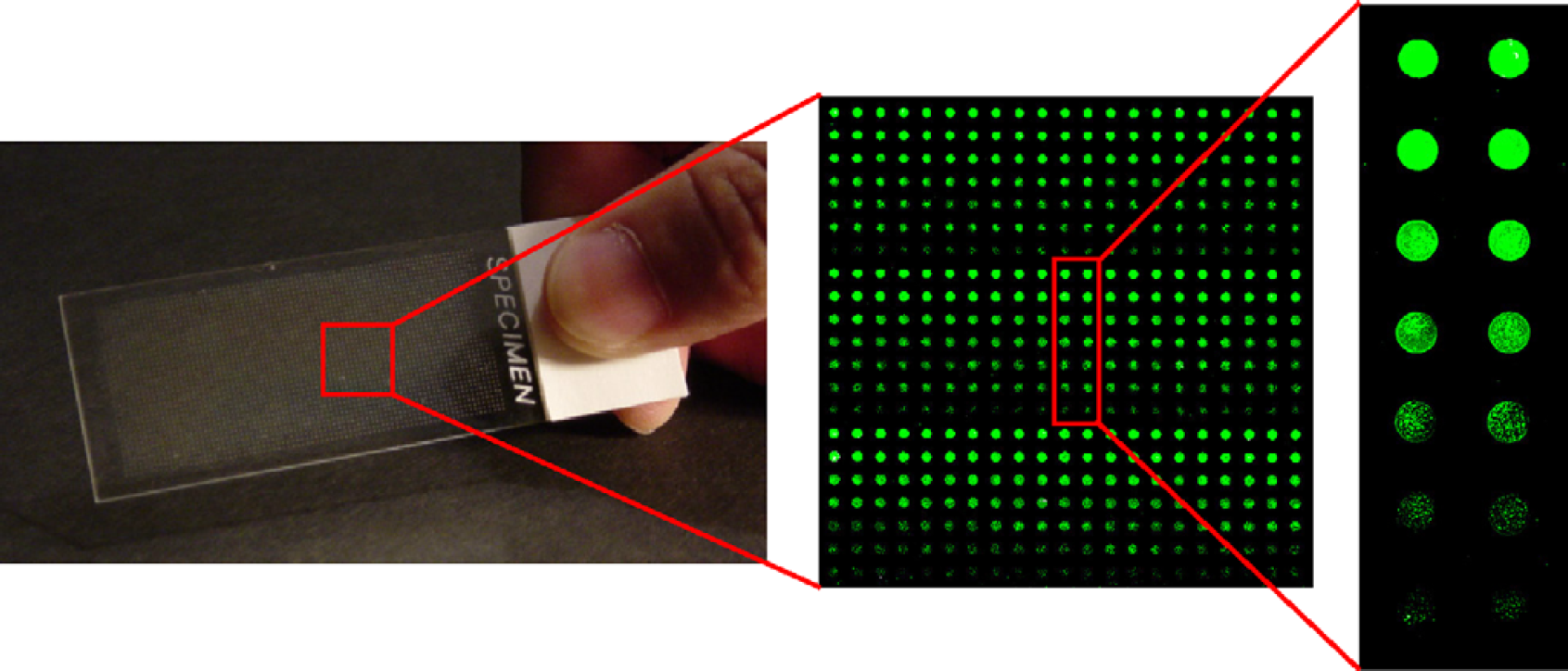

CYP3A4 catalysis in 5-nL sol-gel spots (35 × 80 spot array, 700 μm spot-to-spot distance, 200 μm diameter). Reactions were performed for 1 h. The enlarged regions show scanned images of chip sections, with higher fluorescence intensities corresponding to higher substrate concentrations and faster reaction rates. See text for experimental details.
Having established that CYP3A4 was active on a 2800-spot microarray, we proceeded to further miniaturize the microarray by shrinking the spot-to-spot distance to 350 μm, while maintaining the 5 nL spot volume. This yielded a microarray that contained 11,200 spots (Fig. 3). To test the reactivity of the high-density array, the slide was immersed in a 1-mL solution of DBOMF (10 μM) for < 1 min and then removed. After 1 h reaction, the slide was washed twice with phosphate buffer solution (pH 8) and imaged at 532 nm using the microarray scanner. The microarray visually showed very high uniformity with 10 μM DBOMF as substrate. A random sampling of 40 spots revealed that the reactivity of CYP3A4 was also uniform (average reaction rate of 15.4nM/min with a standard deviation of 1.4nM/min). A magnified image of this microarray showed no apparent spot-to-spot contact.
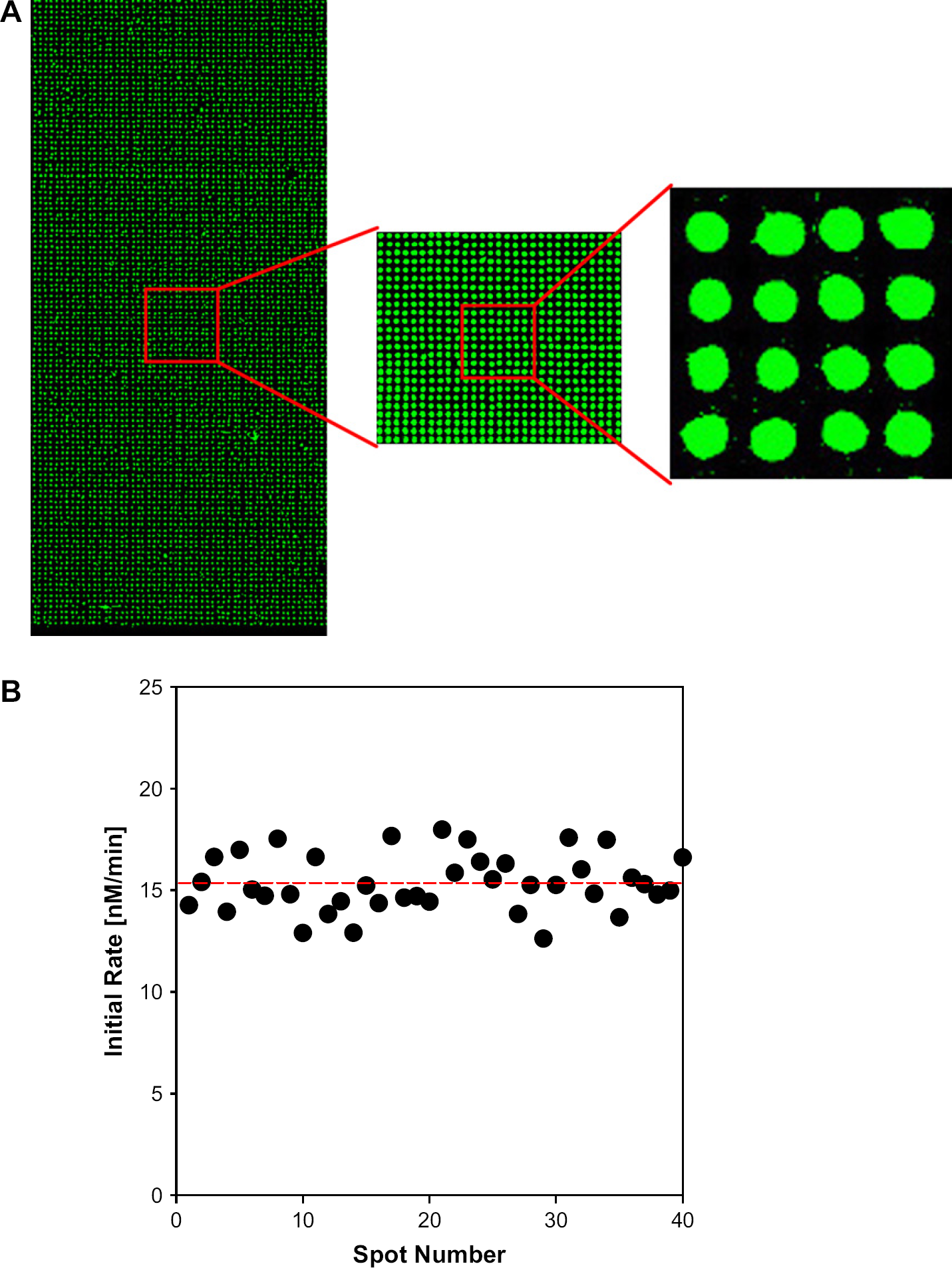
High-density P450 spot array (see text for experimental details): (A) Scanned images of 5-nL CYP3A4-encapsulated sol-gel spots after reaction with 10 μM of green substrate for 1 h (70 × 160 spot array, 350 μm spot-to-spot distance, 200 μm diameter) and (B) initial reaction rates from randomly selected CYP3A4 sol-gel spots.
The ability of P450s to function within sol-gel spots in a microarray led us to design a metabolizing enzyme assay chip, as previously described in Lee et al. 21 This chip, called the metabolizing enzyme toxicology assay chip (MetaChip), is based on the combination of a biocatalytic event (the in situ generation of a drug-candidate metabolite(s)) with the cell-based screening of the metabolite(s). As a result, P450-generated drug-candidate metabolites can be generated and screened against human cell lines on a single microscale platform, which remains useful even if the metabolites are unstable. The MetaChip involves two key components, as depicted in the schematic of Figure 4. A P450-containing sol-gel microarray comprises one slide, while a human cell monolayer, housed in a chamber slide, comprises a second slide. Following addition of a solution of a compound to the sol-gel spots using the microarrayer, the cell monolayer slide is stamped onto the sol-gel array, and the slides are maintained in the stamping position for 6 h to afford sufficient time for P450-generated metabolites to form and diffuse into the cell monolayer. After this stamping period, the cell monolayer slide is removed, the cells are then incubated for 18 h at 37 °C, and finally the cells are stained with live-dead stain to determine the percentage of dead cells using the microarray scanner.
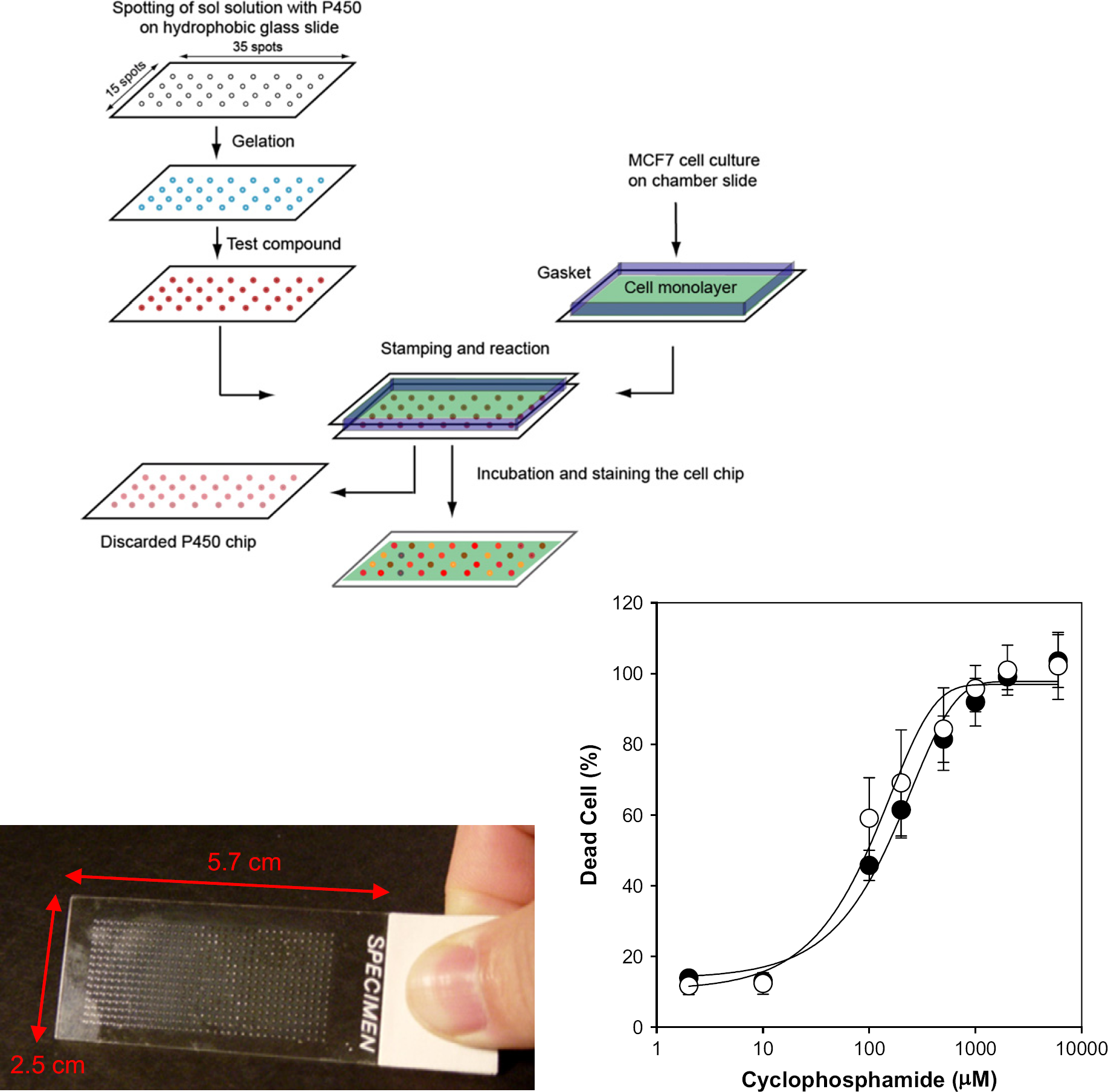
Schematic of the MetaChip platform for metabolite toxicology assays. Compounds are spotted onto the P450-containing sol-gels. Simultaneously, a cell monolayer is prepared and the two slides are contacted and incubated. A picture of a 525-spot MetaChip is shown. Results for CP metabolism are shown; the two curves include CYP3A4 (•) and CYP2B6 (○) containing sol-gels.
A prototype MetaChip consisted of 525 spots (15 × 35 sol-gel array (30 nL/spot) containing CYP3A4, the most common human P450. The microscope slide was pretreated with MTMOS, as described above. A monolayer of MCF7 breast cancer cells was used as the screening target. CP, a well-known anticancer compound, was chosen as a model compound to demonstrate the MetaChip concept. CP is nontoxic until it is metabolized by CYP3A4 and CYP2B6 in the liver. 12, 26 A 60-nL solution of CP was spotted onto each sol-gel spot of the array, followed by stamping of the cell monolayer onto the array (Fig. 4). Following separation of the two stamped slides, the cell monolayer was incubated, stained, and scanned. In the absence of P450 a lawn of green cells was observed, indicating viable MCF7 cells.
The sensitivity of the MetaChip was evaluated by varying the CP concentration (0–2mM). CYP3A4 and a second P450 isoform, CYP2B6, were used to compare the MetaChip with solution-phase incubations in 96-well plates. Dose-response curves for the CYP3A4 and CYP2B6-catalyzed oxidation of CP are shown in Figure 4. The average calculated LD50 value for CP was 174 ± 21 for CYP3A4 and CYP2B6. These values were within 20% of those obtained in solution phase in 96-well plates. Hence, the MetaChip produced a cytotoxic response profile close to that of more conventional and lower-throughput solution-phase P450 incubations. Demonstration of the MetaChip concept was extended beyond CP to include Tegafur, which yields the potent anticancer compound 5-fluorouracil upon P450-catalyzed oxidation, and the common analgesic acetaminophen, which is oxidized to the cytotoxic
Conclusions
A high-throughput, chip-based format provides distinct advantages over more conventional well plates, including ease of washing, fully contained components on the chip, and the ability to couple in situ metabolite synthesis and cytotoxicity screening. We have developed a high-content biochip that contains isolated human P450s encapsulated in a sol-gel matrix that maintains high P450 activity. Up to 11,200 isolated P450 reactions were performed on a single chip. In the presence of cell monolayers stamped onto the P450 chip (the MetaChip approach), we have performed 525 cell screening assays per chip. We are currently expanding the MetaChip platform to include high-throughput P450 inhibition analysis, thereby providing an additional toxicity endpoint for the MetaChip platform. We envision that the MetaChip concept can ultimately comprise the entire repertoire of human P450s, including tailored mixtures of P450s, to provide predictive toxicity data for different populations of individuals. Such a capability would be critical for the realization of drug toxicity testing for personalized medicine.
Finally, the chip-based platform described herein is amenable to laboratory automation, including chip fabrication, drug candidate spotting, and pharmacological screening, e.g., P450 inhibition and cytotoxicity of drug candidates and their P450-catalyzed metabolites. To that end, we have begun to develop the MetaReader, which is expected to serve as a benchtop analyzer that carries out all steps necessary to use the MetaChip. In addition to enabling the user to obtain full service of the MetaChip platform, the automated platform built into the MetaReader is expected to yield rapid and reproducible analysis of MetaChip results, thus providing human toxicology information at unprecedented speed and ease.
Acknowledgments
This work was supported by the National Institute of Environmental Health and Safety (ES012619). We thank Sumitra Sukumaran for helping us with the figures.
