Abstract
Presented is a new approach for assaying the effects of drug candidates against protein targets. Human rhinovirus, HRV14, has been analyzed in the presence of both known and potential antiviral drug agents using enzymatically active mass spectrometer sample plates. The combination of automated mass spectrometry with proteolytic digestion provides for the rapid, high-throughput, and sensitive screening of protein-drug interactions. Multiple drug candidates can be assayed from MALDI sample plates containing 100 wells and, to further increase the number of drugs assayed, multiple drugs can be pooled into each solution (10 candidates per well, 1000 candidates per plate). Wells producing spectra indicative of drug activity can be further analyzed to discern the active compound or be tested with alternative assays.
INTRODUCTION:
The rapid growth of combinatorial libraries has created a need for faster, more accurate, and more sensitive analytical techniques capable of large-scale (high-throughput) screening. Numerous improvements in speed, sensitivity and accuracy, together with innovations in both automation and quantitation place mass spectrometry among the most powerful analytical techniques available today. Crucial to distinguishing the most active component or obtaining structure-activity relationships of compounds in a library is an efficient assay. Toward this end, electrospray ionization and matrix-assisted laser desorption/ionization have been useful for both qualitative 1 –3 and, more recently, the quantitative screening of combinatorial libraries 4 .
The combination of mass spectrometry with protease mapping 5 has recently been used to examine the binding effects of the antiviral agent WIN 52084 on the structure of the human rhinovirus (HRV14). HRV14, the causative agent of the common cold 6 , consists of a protein shell or capsid surrounding an RNA core. WIN 52084 is a member of a family of experimental drugs designed to inhibit the lifecycle of picornaviruses 7 . In these studies viral-protein digestion was dramatically inhibited by the presence of the antiviral agent WIN 52084 (figure 1), while digestion of other viral systems was not inhibited (indicating that interaction of the inhibitor with the virus HRV14, and not the enzyme, caused the inhibition). Based on these results and other evidence it is believed that binding of WIN 52084 to HRV14 inhibits capsid dynamics 6,8 .
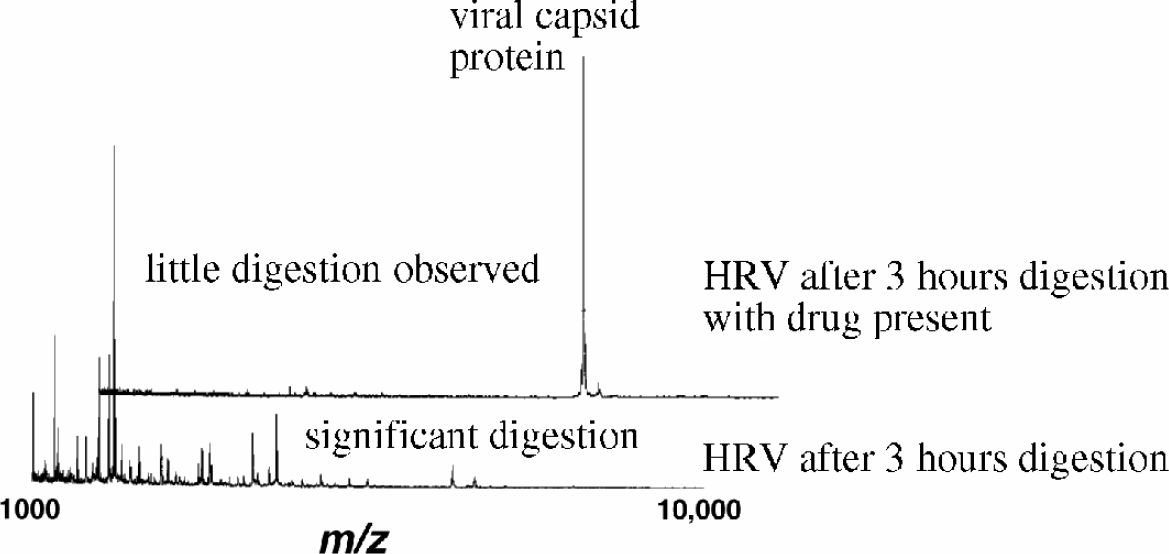
Mass spectra resulting from a 3 hour on-plate digestion of 4 femtomoles of HRV14 in the presence (top) and absence (bottom) of WIN52084. The presence of the WIN compound is found to stabilize the virus toward trypsin digestion. Inset is an electron micrograph of HRV (J. Struct. Biol., 1997, 120, 11).
This protein mapping approach provides a rapid and sensitive mass-based assay for screening libraries of potential antiviral agents. Those compounds which show anti-viral potential (very little to no viral-protein digestion), can be further tested for activity or modified to improve their activity. The conventional assay means of screening for antiviral drug activity is the plaque assay, an assay that requires significantly more time (days) than does the mass mapping approach 10 . Ultimately, plaque assays must be performed to confirm antiviral activity, yet the protein mass mapping method acts as a preliminary screening procedure which can significantly reduce the number of drugs that must undergo the plaque assay. As an automated mass spectrometric method for screening libraries of antiviral agents, protein mass mapping offers a novel and viable alternative.
EXPERIMENTAL:
HRV14 was prepared as previously described 10 to a final concentration of 3mg/ml in 10mM TRIS buffer at pH 7.6. On-plate trypsin digests were performed at 25°C in a high humidity environment. Aliquots of 1.0 μL of the digest solutions containing 25uM drugs (10 drugs per digest solution for a 250 μM total drug concentration), 20mM Tris(1%DMSO), and 4nM HRV14 at pH 7.8 were deposited on trypsin-active plates (Intrinsic Bioprobes Inc., Tempe, Arizona). Digests were allowed to proceed for 1 hr before being terminated by the addition of 1.0 μL of 3,5-Dimethoxy-4-hydroxy cinnamic acid (Aldrich) in a saturated solution of acetonitrile/water 0.25% TFA (50:50).
Automated MALDI-MS mass analysis was conducted using a PerSeptive Biosystems Voyager Elite equipped with delayed extraction and a nitrogen laser. External mass calibration typically was accurate to 0.05% and allowed unequivocal assignment of most proteolytic fragments. The identity of trypsin released fragments was determined by the Protein Analysis Worksheet (PAWS, MacIntosh version 6.0b2, copyright © 1995, Dr. Ronald Beavis) available on the internet.
RESULTS AND DISCUSSION:
Three 96-well plates, with each well containing an analytical solution of 10 drug candidates (for a total of ∼2880 unique drug candidates), HRV14, and the digestion buffer, were loaded onto trypsin-activated MALDI mass spectrometer sample plates (figure 2) and were allowed to digest for one hour prior to mass analysis. As a control each sample plate also contained wells for the digestion of HRV14 in the presence and absence of WIN 52084. Liquid sample handling robotics may ultimately be used to mix and load samples onto mass spectrometer plates, further increasing the number of drugs assayed.
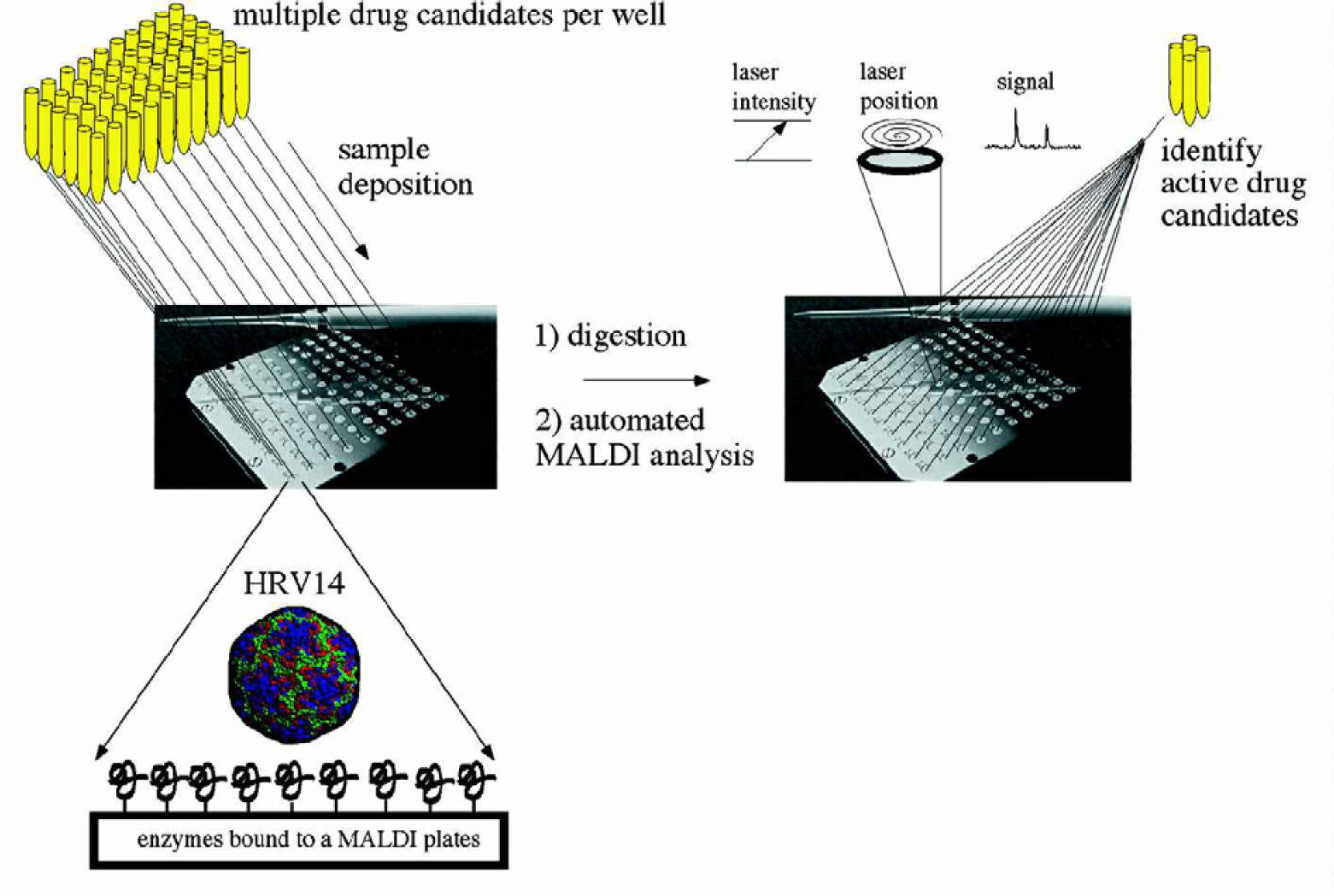
Drug efficacy is determined by assaying the digestion of protein(s) in the presence of multiple drug candidates. Protein digests were performed on the enzyme-activated MALDI sample plate. Addition of MALDI matrix terminates the digestion and automated MALDI analysis follows. Analytical solutions resulting in spectra indicative of antiviral drug activity (digestion is none or significantly retarded) are further analyzed for the active compound. Those compounds that show activity can then be used to synthesize new analogues in an attempt to improve their activity.
While the digests in the presence of the WIN compound were inhibited, the digests in the presence of other drug candidates were not inhibited, indicating no antiviral activity for this particular candidate library.
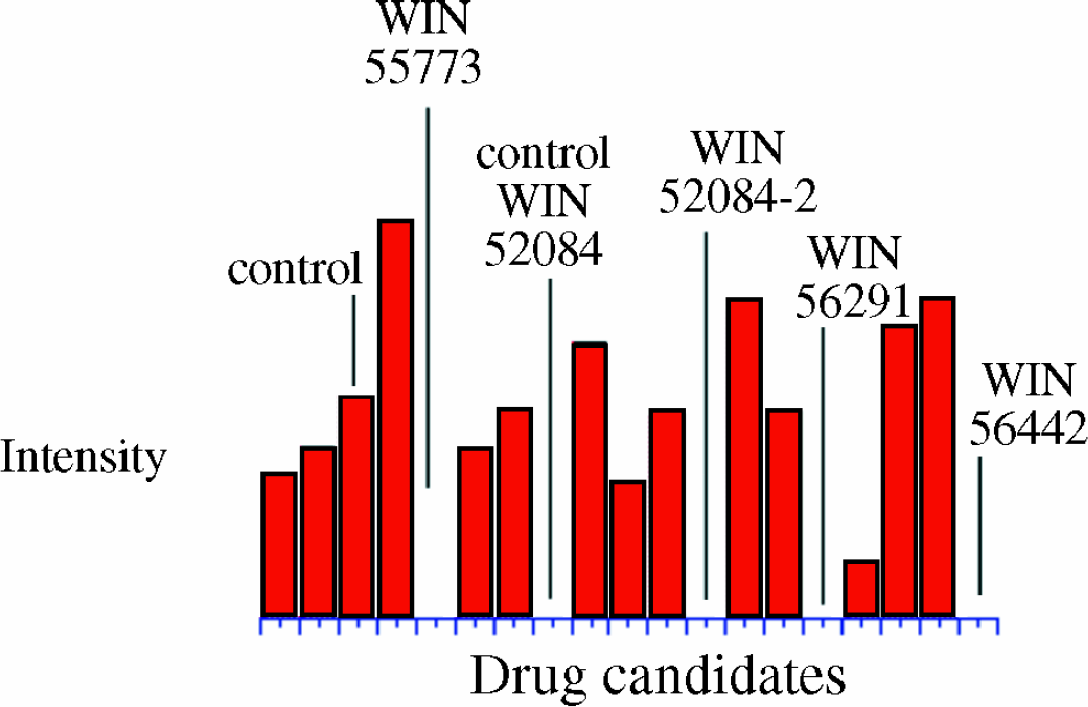
Bar graph representing the amount of digestion per sample well (only 15 sample wells are represented). The y-axis is the sum of the intensities of 5 of the most commonly observed HRV14 trypsin digestion fragments. Also shown is the extent of digestion for the other WIN compounds analyzed. In all cases the WIN drugs significantly inhibit the digestion of HRV14.
Potential limitations to this method for drug analysis include matrix-drug interactions and limited drug solubility. As a result of matrix-drug interactions the quality of some of the mass spectral data was reduced. For instance, broad peaks (poor resolution) and a low signal-to-noise ratio were found to be due to the presence of the drugs interfering with matrix crystallization. Of all the mass spectra obtained about 1% of the spectra was affected by the drugs being tested. This problem was overcome by reducing the concentration of the drug candidates.
Currently the use of MALDI protein mass mapping as a screening tool requires that a system undergo changes (with respect to proteolytic cleavage) upon interaction with a potential drug candidate. The ability to compare the effectiveness of drug candidates with known drugs would provide an ideal scenario, however without known inhibitors available, it is still possible to screen using protein mass maps. With WIN 52084 the inhibition of digestion represents one way a positive “hit” could be recognized, however with other protein systems a “hit” may be indicated by the generation of a new set of fragments occurring or even a significant increase in the rate of digestion. This assay relies on the hypothesis that mass maps are a reflection of the conformational state of the protein or protein complex and that any significant conformational perturbations, due to the binding of a drug molecule, will be represented by a new mass map. Overall even in the early stages of this assay's development, we believe this approach could be generally applicable to a large variety of protein-drug interactions.
ACKNOWLEDGMENTS
The authors gratefully acknowledge the editorial assistance of Jennifer Boydston and funding from the National Institutes of Health grant 1 R01 GM55775-01A1.
Jay Chiang

