Abstract
Ahigh-throughput analytical characterization system was developed for quality control support of a central sample collection resource. This system utilizes liquid chromatography mass spectrometry with in-house developed data automation applications. Continuous operation of analytical instrumentation is accomplished by fully automating sample submission and report processing functions. Comprehensive analytical information characteristic of quality, chemical, and physical properties (e.g. relative purity, detection sensitivity, LogD) are automatically transferred to an on-line database. The application of this database for detailed quality assessment of a small sample library (ca. 24,000 compounds) is demonstrated.
Keywords
Introduction
Access to an extensive and highly diverse sample collection is critical for the discovery and development of new drug products. Biological activity screening assays of a diverse sample collection can provide valuable leads for multiple drug discovery programs.1–4 The sample collection may also be utilized for a number of synthesis or analytical research applications (e.g. selection of novel precursors, physical measurement references, or internal calibration standards).
Recognizing its importance to research operations, many pharmaceutical companies maintain a proprietary sample collection resource. With the aid of recent advances in automated parallel synthesis and sample preparation technology,4–6 it is possible for a sample collection resource to accumulate multi-million single component samples. However, regardless of the size and chemical diversity, a compound collection is of limited value if the overall quality of the collection is not controlled. Impure samples can introduce false negative or false positive results in activity screening assays. 3 6 Low quality samples applied as synthesis precursors generally result in low product yield. Also, results of analytical studies utilizing such samples can be inaccurate or inconclusive. Given the importance of the sample collection to research operations, an efficient high-throughput analytical characterization resource is needed to monitor the quality of the sample collection.
Liquid chromatography mass spectrometry (LC-MS) has become the method of choice for the characterization of compound libraries.7–9 LC methods applying high flow rates and short columns allow analysis to be performed in as little as ca. 1 minute. 10 11 Throughput has also been enhanced by the introduction of a multiplex ionization source. 12 LC-MS can also be configured with multiple detection techniques (e.g. UV absorption, light scattering, nitrogen detection) to characterize a chemically diverse compound library. 8 12
Depending on the chromatographic conditions, a single LC-MS system utilizing multiple detectors can perform analyses at a rate of ca. 60,000 to 200,000 sample per year. An analytical resource may employ multiple instruments in parallel to obtain throughput in the range of ca. 1,000,000 samples per year. However, efficient operations of such an analytical resource will require considerations to a number of issues. The LC-MS instrumentation will characterize samples at a rapid pace. Sustained operation at maximum throughput will require a mechanism to efficiently submit and queue high-density sample plates. A large quantity of report files will also be generated very rapidly. These report files contain a variety of information that maximizes the utility of the sample collection: quality control measurements (e.g. detection, average purity, signal to noise ratio), relative chemical properties (e.g. UV absorption maxima, molecular ion, detection sensitivity), and relative physical properties (e.g. LogD, solubility). This information must be efficiently extracted from the report files and formatted to be conveniently available to scientists for their various research applications.
In this report, an approach to implement LC-MS as a high-throughput quality control resource for a large sample collection is presented. This approach utilizes in-house data automation applications to control data processing functions associated with sample submission, extraction of comprehensive analytical data, and result retrieval. Application of the comprehensive result data for detailed purity assessment and support of various research applications is discussed.
Experimental
MATERIALS
Compounds, stored as 2 mM solution in DMSO, were provided by the Merck Central Repository (Rahway, NJ). The compounds were handled as 96- or 384-well sample plates. The sample plates were labeled with a barcode identifier, referencing plate map data from a sample inventory database. Automation software applications for sample submission, report processing, and search/retrieval were developed in house using Microsoft Visual Basic 6.0 (Redmond, WA) and Oracle 8.1 (Redwood Shores, CA). Automatic sample queue and data acquisition of submitted run-sheets were performed using Micromass Autolynx (Beverly, MA). ASCII report data files (*.rpt) were generated using Micromass Openlynx.
ANALYTICAL INSTRUMENTATION AND CONDITIONS
Chromatographic separation was performed on an Agilent 1100 HPLC system (Palo Alto, CA). LC effluent was characterized by a Waters ZQ mass spectrometer (Milford, MA), Agilent 1100 ultraviolet absorption diode array detector (DAD), and a Sedere (Cranbury, NJ) Sedex 75 evaporative light scattering detector (ELSD).
LC Conditions: LC separation was performed using a Metachem (Lake Forest, CA) Polaris C18A column (50 mm × 2 mm × 3 um). HPLC grade water (0.05% formic acid) and acetonitrile (0.05% formic acid) were used as mobile phase. Total analysis time was 5.5 minutes. Initial mobile phase condition was 95% aqueous phase. At time 3.5 minutes, aqueous phase was reduced to 2% for one minute. At time 4.51, mobile phase composition was returned to 95% aqueous.
DAD and ELSD Conditions: UV spectra were acquired from 220 nm to 400 nm for each chromatogram data point. LC effluent from the DAD was split to allow ca. 0.100 mL/min and 0.900 mL/min to flow to the MS and ELSD detectors, respectively. ELSD temperate and attenuation was set to 60 °C. and 9, respectively
MS Conditions: Electrospray ionization mass spectrometry was performed on a Waters Corporation ZQ quadrupole mass spectrometer. Data was acquired in both positive (ESI+) and negative (ESI−) ionization modes simultaneously. Spray and cone voltages were set at 3300 V and 35 V respectively. Interscan delay and dwell times were 0.3, 0.2 seconds respectively. Mass scan range was selected by the sample submission application, as shown in Figures 3. Various mass scan range parameters are saved as different MS method files. Based on available mass information, the software will logically select the appropriate MS method file for each sample. For compounds with mass less than 225 Da., data was acquired from 100–500 m/z (file designated as Method 1). If mass was 225 to 700 Da., mass range 200–800m/z was scanned (Method 2). For compounds with mass greater than 700 Da., mass range 600–1200m/z was scanned (Method 3).
DATA PROCESSING
After each sample analysis, peak detection and integration processes were automatically performed by the Micromass Openlynx application. Openlynx compiles integrated peak data, chromatographic and spectral data traces into a single ASCII report data file. This report data file underwent additional processing by an in-house written result processing application. Data processes performed by this application are shown in Figures 5
Detection: Selected ion chromatograms (SIC) were generated by combining (M+H)+ and (M+Na)+ ion signals for electrospray positive ionization mode (ESI+), and (M-H)- signal for negative ionization mode (ESI-). Samples were registered as ‘Found’ if the signal to noise ratio of the base peak was at least 3.
Chromatogram signal to noise: Signal to noise was calculated by taking the ratio of the absolute signal height to the root mean square average of the entire chromatogram outside the base peak region, inclusive of all other detected peak regions.
LogD Analysis: The octanol water partition coefficient was estimated by correlating analyte retention time with retention time of LogP references. LC retention times were calibrated with the following compounds: allopurnol (LogP −0.85), antipyrine (0.38), prednisone (1.46), griseofulvin (2.18), dichlorophenol (3.18), and tolnaftate (5.4). Capacity factors of LogP references were plotted vs. shake-flask LogP values. The linear curve fit formula was applied to calculate experimental LogD values using the capacity factor of a given analyte.
Results and Discussion
Depending on chromatographic conditions, the throughput of a single LC-MS instrument can range ca. 60,000 to 200,000 analyses per year. Sufficient throughput to support a large sample collection can be readily obtained by implementing several instruments in parallel. However, the operation efficiency of a high-throughput analytical system is not determined by the rate of data acquisition alone. Figures 1 shows the various processes involved in a high-throughput LC-MS analytical system: sample submission, sample analysis, report data processing, and data search/retrieval. Slowdown of any one process can significantly affect the performance of the entire analytical system. To fully capitalize the LC-MS instrument's capability to perform rapid analysis, each of these processes must be optimized to handle large amount of complex data efficiently.
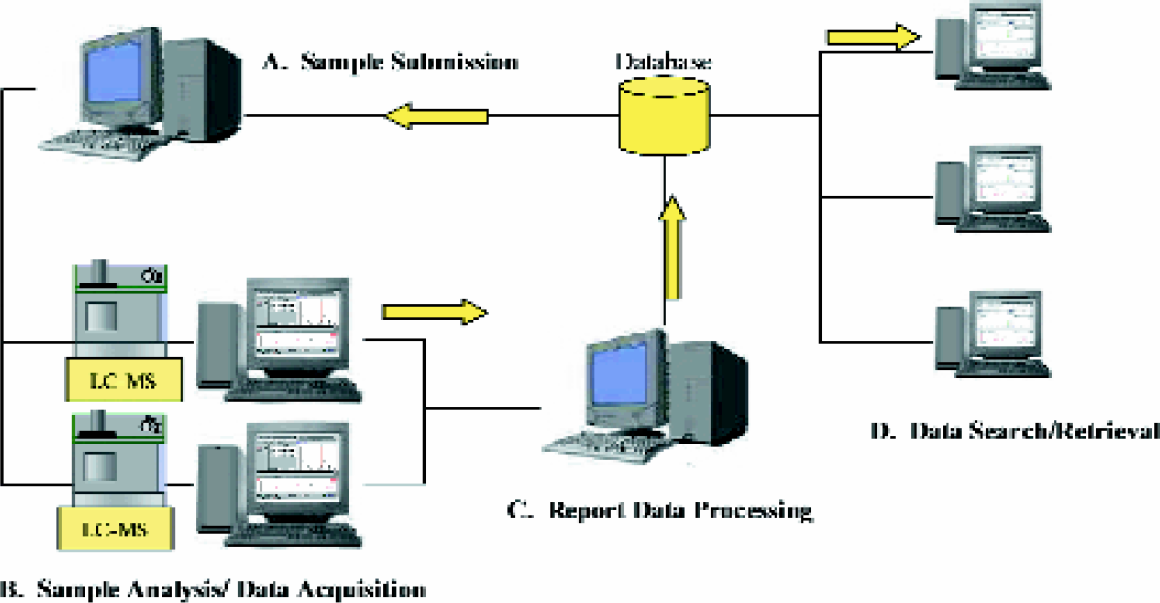
Processes involved in a high-throughput analytical system: (A) sample submission, (B) sample analysis, (C) report data processing, and (D) data search/retrieval.
SAMPLE SUBMISSION FOR ANALYSIS
In order to maintain maximum throughput and minimize downtime, high-density sample plates must be rapidly supplied to the analytical instrumentation (process Figures 1A). A graphic-user interface application was developed to automate the sample submission process in two steps. Screen captures of the application are shown in Figures 2. After the application is initiated from the home window (process Figures 2A), the user is prompted to enter the sample plate identifier (i.e. a unique barcode associated with the plate) and specify a plate location on the liquid handler (Figures 2B). In the final step, the application prompts the user to load the sample plate on the liquid handler at the designated location (Figures 2C).

Screen captures of the sample submission application: (A) home window, (B) plate ID and location data entry, (C) final instruction.
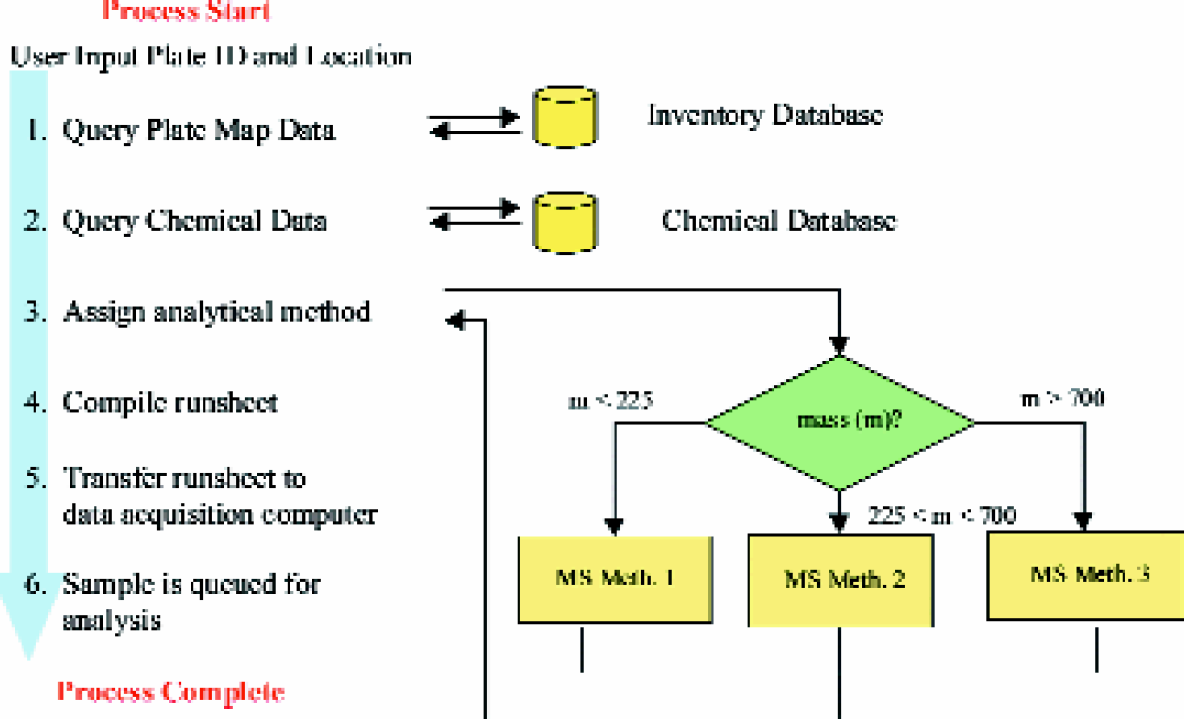
Processes performed by the sample submission application.
The sample submission application performs a number of complex procedures that are invisible to the user, as shown in Figures 3. Many of the procedures involve the compilation of parameters required by the data acquisition computer to perform a sample plate analysis. The application first queries the compound identifier (i.e. a unique number corresponding to the compound identification) and plate map coordinates from the inventory database using the plate identifier as a reference. The compound identifier, acquired from the plate map data, is then used as a reference to query molecular formula from a chemical information database. Filenames are prepared for each sample based on unique numeric sequences generated by an Oracle database. A predefined set of analytical parameter can be uniformly assigned to all samples (e.g. injection volumes, peak integration parameters), or the software can logically select certain parameters based on available chemical data. For example, mass scan range parameters are assigned based on the calculated mass of a given compound, as shown in Figures 3, step 3. If the mass of a given analyte is high, a MS scan parameter file optimized for higher molecular weight analysis will be selected by the software. More sophisticated analytical method selection logic (e.g. LC separation based on calculated LogP queried from an on-line database 14 ), adjustment of injection volume based on concentration values listed in the compound inventory database) can be readily implemented. All queried sample information and analytical parameters for each sample plate are complied into a database or runsheet file. With the runsheet data compiled, the instrument control software completely automates the data acquisition process (Figures 1B). The newly created runsheet file is detected by instrument control software (i.e. Micromass Autolynx) and placed in queue for analysis.
Manually performing the actions necessary to create and queue a runsheet is time consuming. The preparation of a runsheet for a single sample plate would require ca. 15 minutes. Manually preparing several LC-MS systems for plate analysis would require several man-hours, contributing to significant instrument downtime. The sample submission application significantly accelerates the sample submission process. The automation application allows a LC-MS system, configured with large liquid handler, to be loaded to capacity in minutes. A single operator can readily maintain continuous operation of multiple high-throughput LC-MS systems with minimum downtime.
REPORT PROCESSING
Figures 4 shows an example of the various data acquired from an LC-MS system utilizing multiple detection techniques: electrospray mass spectrometry positive ionization mode (ESI+), negative ionization mode (ESI-), UV diode array detection (DAD), and evaporative light scattering detection (ELSD). After each plate analysis, the data acquisition software examines results from each detector and creates a report file. This report file contains two types of chromatographic data: analyte specific detection traces (i.e. selected ion chromatograms), and non-specific detection traces (i.e. total ion chromatogram, DAD, and ELSD). In addition, the report contains detailed properties of the detected peaks (e.g. intensity, retention times) and spectral traces (i.e. UV and mass spectra). A variety of useful information can be extracted from this report file: quality properties (e.g. detection, relative purity, chromatographic signal to noise), relative chemical properties (e.g. MS ionization efficiency, UV absorption maxima, detection sensitivity), and physical properties (e.g. LogD).
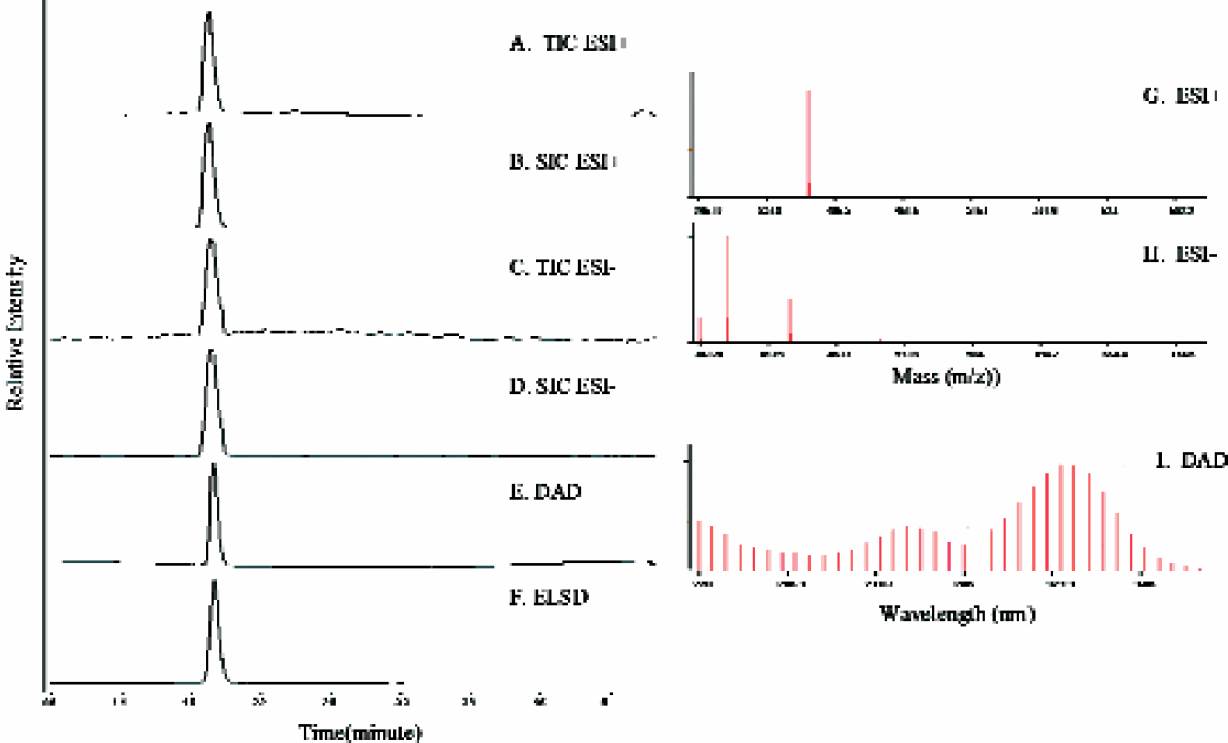
Example of acquired analytical data. Chromatographic data: (A) TIC ESI+, (B) SIC ESI+, (C) TIC ESI-, (D) SIC ESI-, (E) DAD, (F) ELSD. Spectral data traces of detected peaks: (G) ESI+, (H) ESI-, (I) DAD.
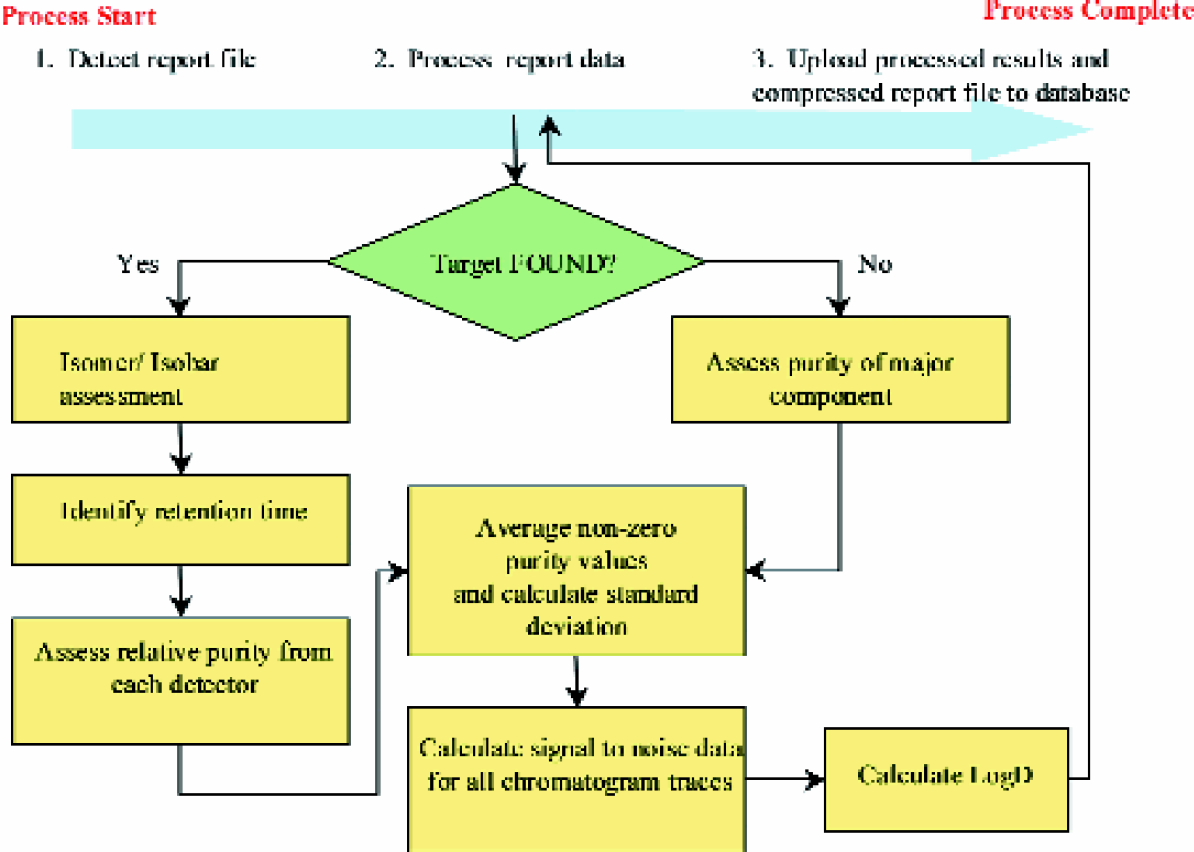
Procedures performed by the report processing application.
A software application was developed to automate the extraction of the comprehensive analytical information (process Figures 1C). This report processing application automatically detects newly created report files and transfers the analytical information to an online Oracle database. The procedures performed by this result processing application are shown in Figures 5. After a report file for a plate is created, the application detects the file and begins to sequentially process each sample. The application first determines if signal characteristic of the compound of interest is present. This is accomplished by examining the selected ion chromatogram (SIC) from both ionization modes for peaks above a selected signal threshold. The SIC only displays signal response characteristics of a specific compound. If a base peak is detected with a signal to noise ratio greater than 3, the application registers the sample as ‘Found’ and records the properties of the selected base peak (i.e. height, area intensity, and retention time). If no such peaks are detected from either SIC traces, the instrument records the compound as ‘Not Found’. Figures 6A shows an example where an intense single peak was detected by the SIC ESI+ at ca. 1.6 minutes. The sample shown in Figures 6A would be classified as ‘Found’. In contrast, Figures 6B shows a sample where peaks were not detected on the SIC for either ESI+ or ESI-. As a result, this sample is registered as ‘Not Found’.
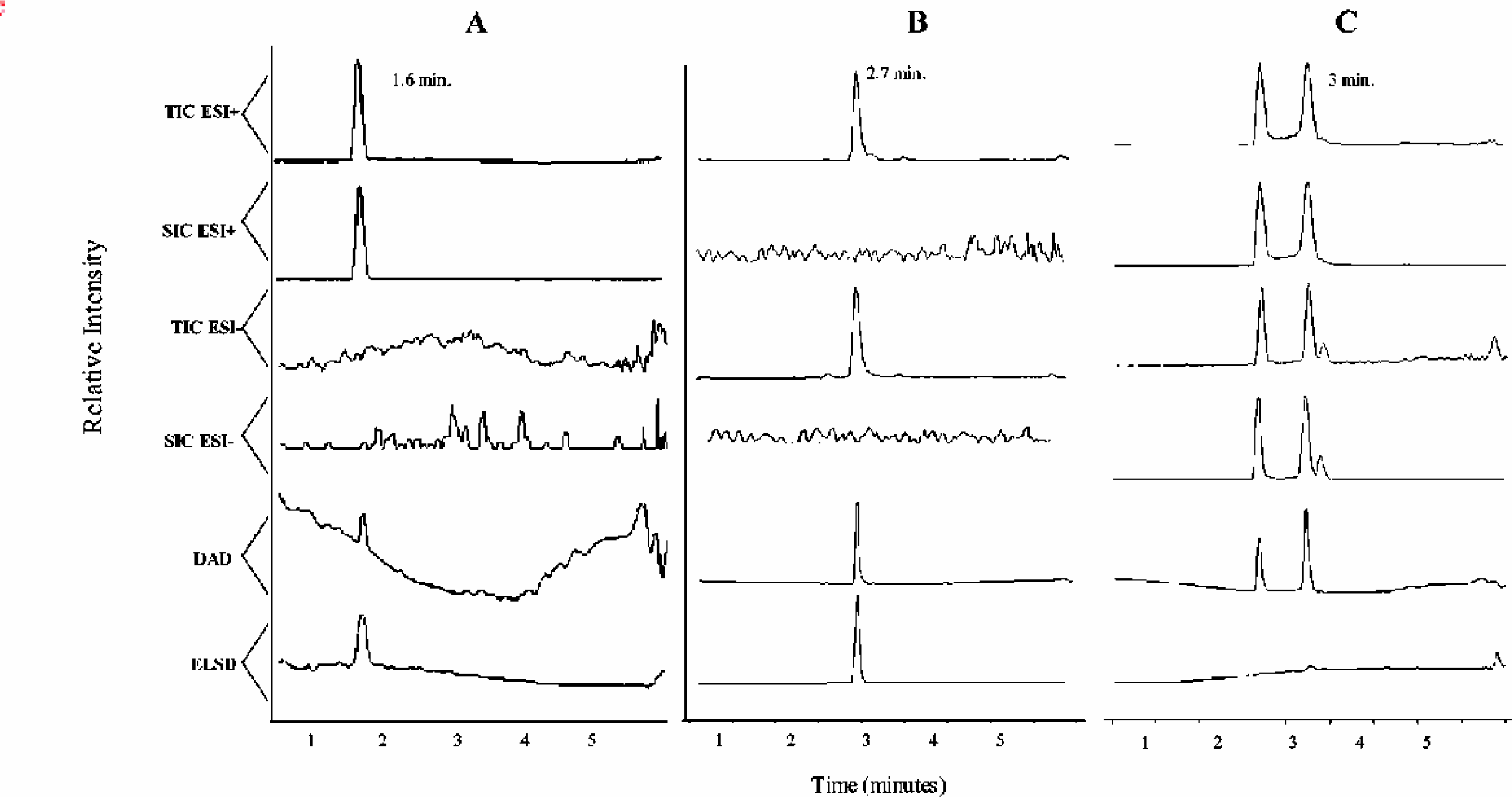
Chromatographic data for samples processed by the report processing application: (A) Found (average purity 100%), (B) Not Found (average purity 100%) and (C) Found (average purity 60%).
As shown in Figures 5, a series of procedures are performed only on ‘Found’ data sets. The presence of possible isomers or isobars is evaluated by counting the total number of peaks detected on the SIC that are above 25% of the base peak height. In Figures 4 and 6A, only a single peak was detected on the SIC, suggesting the presence of a single isomer. Figures 6C shows an example where two such peaks were found in both SIC ESI+ and ESI-, suggesting the presence of multiple structures with identical molecular weight. For Figures 6C data, the software would record a value of ‘2’ for each of the SIC traces.
After the data is evaluated for isomers, relative purity values are calculated from each non-specific detection chromatograms: total ion chromatograms (TIC) ESI+, TIC ESI-, UV-DAD and ELSD. The base peak retention time from the SIC data trace is used for peak reference. Due to excessive noise contributed by solvent clusters at the low mass range, purity is not calculated for TIC MS traces where the target mass is below 225. Purity is also not calculated for the UV-DAD peaks that are retained less than 0.5 minutes. The non-zero purity values from each detector traces are used to calculate the average purity value of the sample. Additional properties are also recorded (i.e. the standard deviation of purity values from different detectors, number of detectors that successfully detected the compound, total number of peaks detected for each trace, and peak intensity).
Since a signal characteristic of the sample was not detected in the SIC traces for ‘Not Found’ samples, the report processing application attempts to assess purity by using the base peak signal from each of the applied detector traces. Relative purity and retention time was recorded for each detector trace. Non-zero average purity and standard deviation (combining non-zero purity values from each detector) is recorded only if retention time data all agree within 0.1 minute.
The signal to noise ratio of the peak of interest is calculated for all chromatographic data traces. LogD values are also calculated using LC retention time for reference. After the report of each sample is processed, the data is populated into an on-line database. In addition, the entire report file for each sample (chromatographic traces and spectral data) is compressed and exported to the database as a BLOB data type.
The report processing application enables a high-throughput analytical resource to rapidly process result data. Immediately after a report file is created, the data is automatically detected and processed. The data extracted from the report file is comprehensive (i.e. detection, peak integration, indication of isomer presence, and chromatographic and spectral data traces) and made available immediately after each plate analysis. All processes are performed efficiently without user interaction. A single conventional desktop PC operating this application can handle data generated by several LC-MS instruments.
ACCESS AND APPLICATION OF THE ANALYTICAL DATABASE
Proprietary compounds stored in a central sample collection will be distributed for a number of different drug discovery applications (e.g. biological assays, synthesis, or analytical references). Each application can have different minimum purity requirements. To assess the quality of a given sample, most conventional LC-MS data systems generally record only detection and average purity values (e.g. ‘Found’, 98%). However, relative purity values may be misleading unless report chromatograms are manually reviewed. Figure 6A shows an example where the instrument integration software would calculate 100% relative purity values from the ESI+, DAD, and ELSD chromatogram traces. However, a scientist would quickly note that the traces from DAD and ELSD may not be suitable for purity assessment due to the poor signal to noise ratio. In Figures 6C, average purity would be assessed as ca. 60%. Direct visualization of the SIC chromatographic data trace would reveal that the low purity value may be accounted for by the presence of isomers. For data such as Figures 6B, where no SIC peaks were detected, purity values may not be calculated at all. For high-throughput applications, manual review of report data is prohibitively time-consuming. As a result, detection and purity values alone may not refect the true quality of a given sample. A different approach to quality assessment is necessary in dealing with large sample collections.
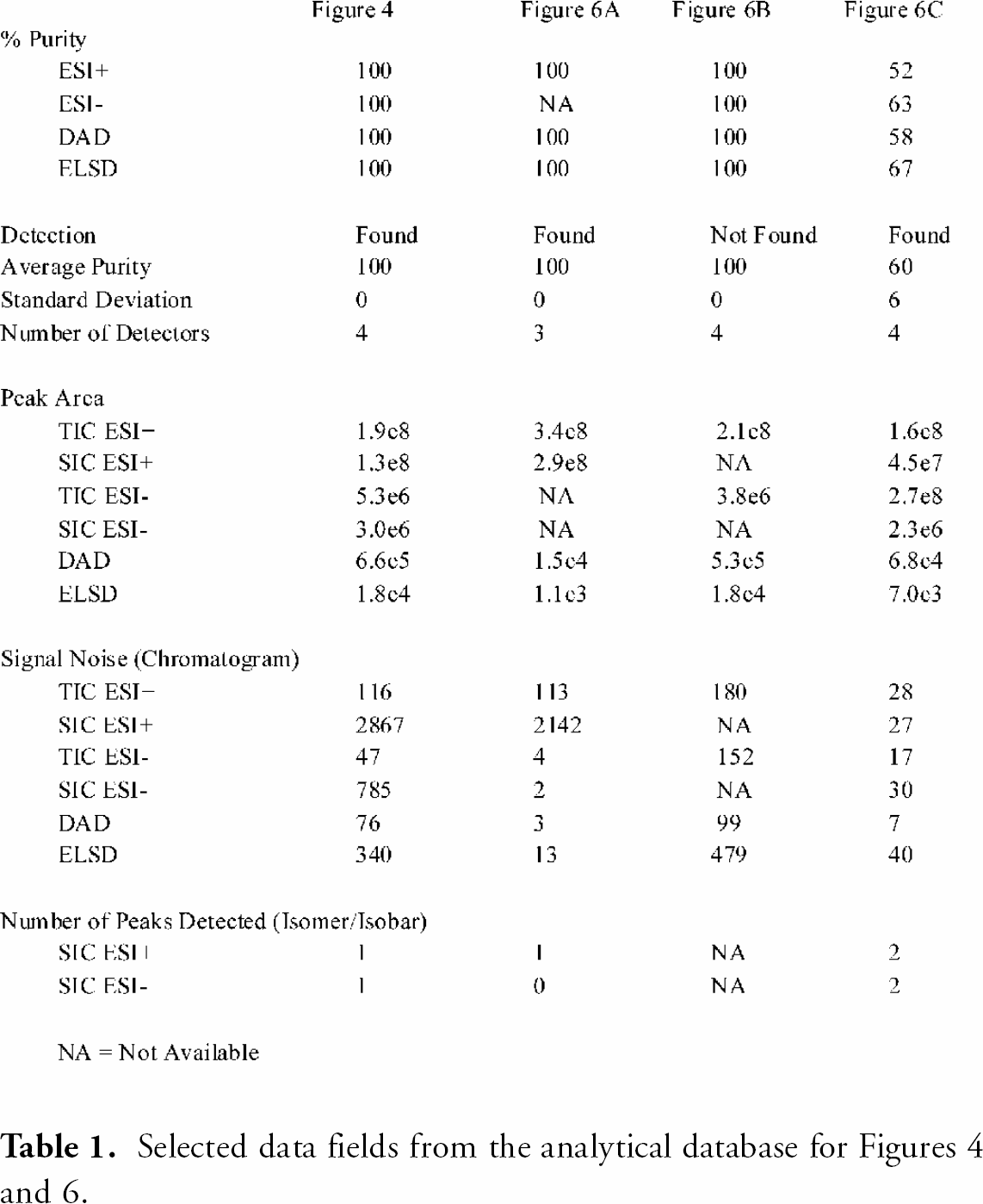
The analytical database populated by the report processing application contains a number of data fields that reflect sample quality (e.g. Found, average purity, number of detectors, standard deviation, isomer indication, signal to noise). By performing a search that cross-references multiple data fields, it is possible to accurately assess sample quality of a large collection without direct visualization of the report data. Table 1 lists selected data fields from the analytical result database for chromatograms shown in Figures 4 and Figures 6. As shown in Figures 4, high quality samples will exhibit high average purity, minimum standard deviation, high peak area and high signal to noise values. Samples that potentially contain a mixture of isomers or isobars, as shown in Figures 6C, can be readily identified and sorted by referencing the number of SIC peak count data field (i.e. isomer/isobar assessment). If a sample is classified as ‘Not Found’, but contains a single peak of high intensity from multiple detectors (e.g. Figures 6B), a chemist may choose to retain that compound for further analysis.
An example of applying this quality assessment approach to a small sample library is shown in Table 2A. Of the total 24,801 samples, 17,622 had an average purity between 75 to 100%, as shown in Table 2A. Table 2B shows a sub-search of the 17,622 samples using various result criteria. Approximately 6% of this set did not have a SIC peak in either ESI+ or ESI-, and 10% of the samples had an isomer/isobar impurity. Searches using the signal to noise data fields will eliminate samples that were poorly detected. The majority of the sample set (89–92%) yielded a signal/noise response of 50 or greater from either ESI+ or ESI- ionization modes. Approximately 71% and 90% of the sample set had a DAD or ELSD signal/noise response of 10 or greater, respectively. Performing a search combining all the criteria would reveal samples of the highest quality (ca. 57% of the 75–100% sample set). Research applications with maximum purity requirements (e.g. follow-up screening assays, organic precursors) would utilize a similar search protocol. Applications with lower purity requirements (e.g. initial lead screening assay) may choose to limit search criteria or change the parameter values (e.g. lower signal to noise, or increase average purity range).
Average purity results for the characterization of a 24,801 sample library

Sub-query of the 75–100% purity range set with additional result criteria

The analytical database also contains information that reflects physical and chemical properties (e.g. absolute signal response, LogD, LC retention time, UV and MS spectra traces). The availability of this information can improve how compounds are searched and selected for various research applications. A comparison of peak intensity with various signal to noise ratio data fields will identify the detection technique most suited to monitor a given compound. LogD or retention time data can aid separation chemists in identifying the ideal chromatographic conditions for a given class of compound. Chemists performing quantitative pharmacokinetics studies can cross-reference signal intensity area with retention time to identify the most suitable internal standard for a specific analyte. This database can also provide valuable information on novel compounds. By cross-reference structural similarity data, chemists may be able to predict the retention and detection properties (e.g. relative sensitivity) of newly synthesized products.
Access and detailed searches of the comprehensive analytical data can be performed using a number of different applications (process Figures 1D). Figures 7A-C, shows screen captures from a custom data browsing tool that allows users to search, review result data, and visualize chromatographic or spectral data. A Web-based application was also developed, as shown in Figures 7D. With the aid of more sophisticated search applications, it is possible to cross-reference the analytical result data with other on-line database (e.g. structural information, biological assay data).
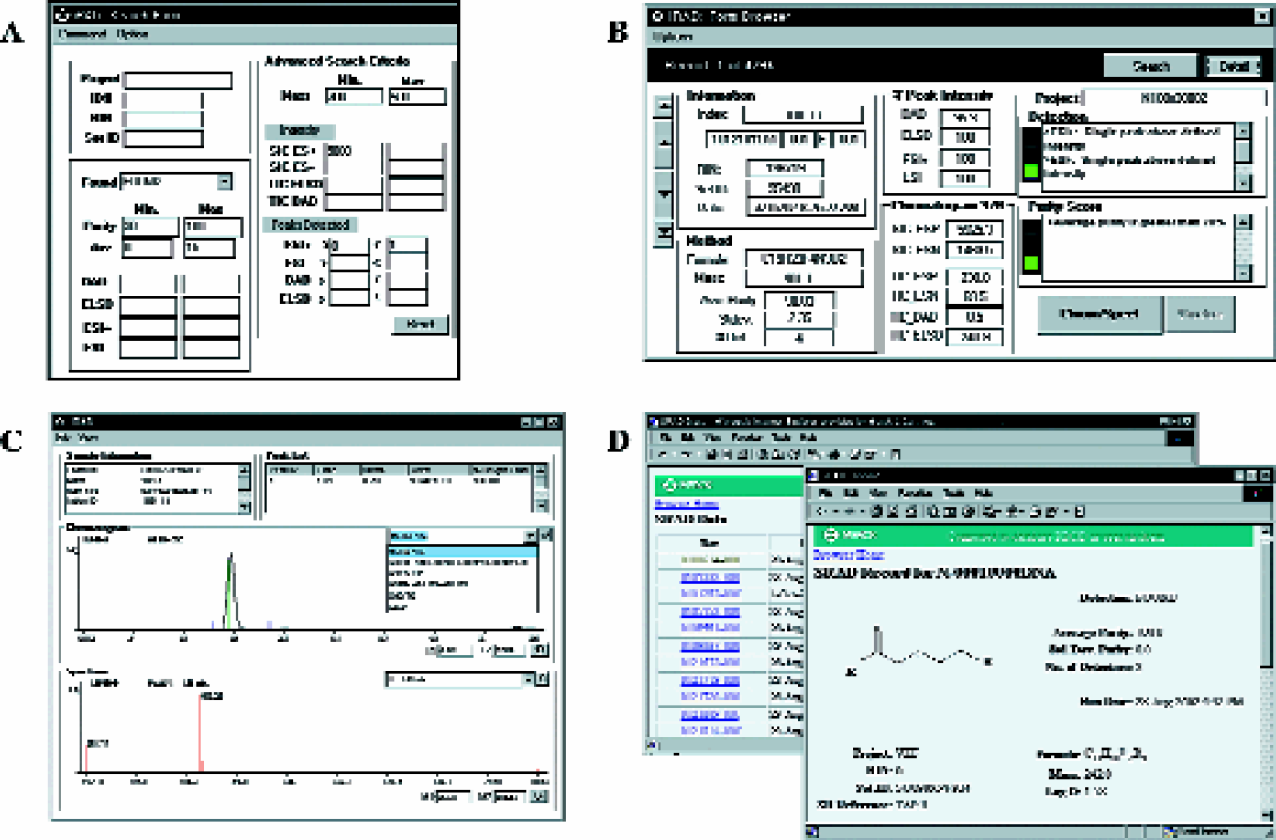
Screen captures of the data search and browsing application (A) search window, (B) result summary, (C) chromatogram and spectral data viewer, (D) Web data access.
Summary
A high-throughput LC-MS analytical characterization system was developed. This system was configured with in-house data automation applications to efficiently support a large sample collection resource. Sample submission, report processing, and search/retrieval procedures for high-density sample plate analysis were fully automated. High-throughput operations can be sustained while minimizing instrument downtime and manpower requirements. This entire system can be readily scaled to manage multiple LC-MS systems and accommodate demands of a large sample collection resource.
Comprehensive analytical information characteristic of quality, chemical, and physical properties are automatically extracted and populated into a relational database. A client side data search and retrieval application was developed to allow result property query over a broad number of samples or visualization of a specific sample result. The availability of comprehensive analytical information allows scientists to efficiently perform detailed quality assessment of large collection of compounds. The data will also be useful in the selection of compounds with ideal physical or chemical properties for various research applications.