Abstract
Screening for ADME/Tox properties of compounds is a key step in the drug development process. Three applications defining such properties, namely Cytochrome P450 Inhibition, Serum Protein Binding and P-glycoprotein Interaction were developed and the performance of each was evaluated on Tecan's LabCD-ADMET System. The LabCD-ADMET System is an automated platform which integrates microfluidics, in the form of the centrifugally driven LabCD device; liquid handling and assay robotics, in the form of a modified GENESIS liquid handling workstation; an incubation, spinner, and detection system, the ULTRA LabCD; and platform/application-specific software. The assay approach and the data for each application are presented. Two applications, Cytochrome P450 inhibition and serum protein binding assays, have also been externally validated at leading pharmaceutical companies within Tecan's Early Access Program (EAP). The LabCD-ADMET System provides a miniaturized, integrated, and automated turnkey solution for various ADME/Tox applications.
Assay Approach
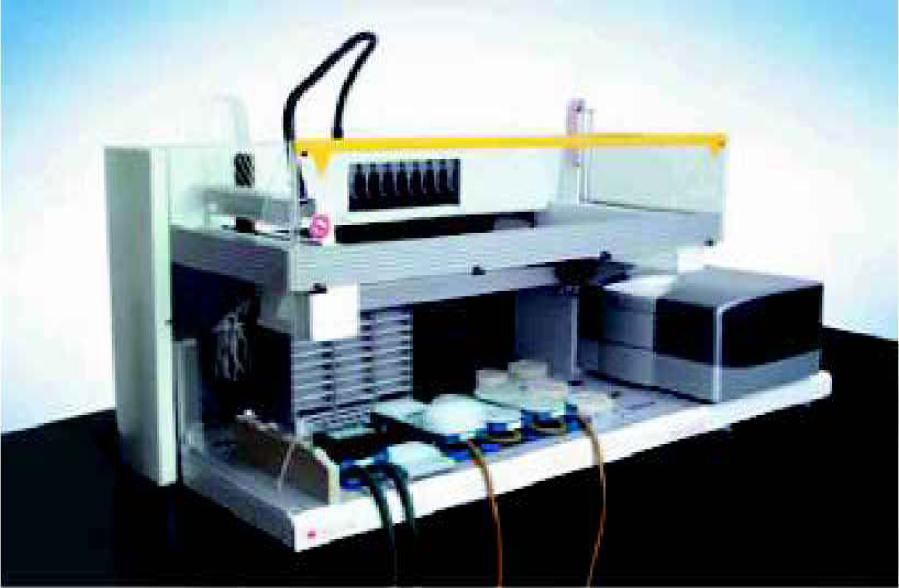
Automation: The miniaturized assays presented here use the LabCD-ADMET System (Figure 1). This system integrates the centrifugally driven microfluidic LabCD (Figure 2 & 3), an Ultra LabCD/microplate reader, and software on the Genesis robotic platform.
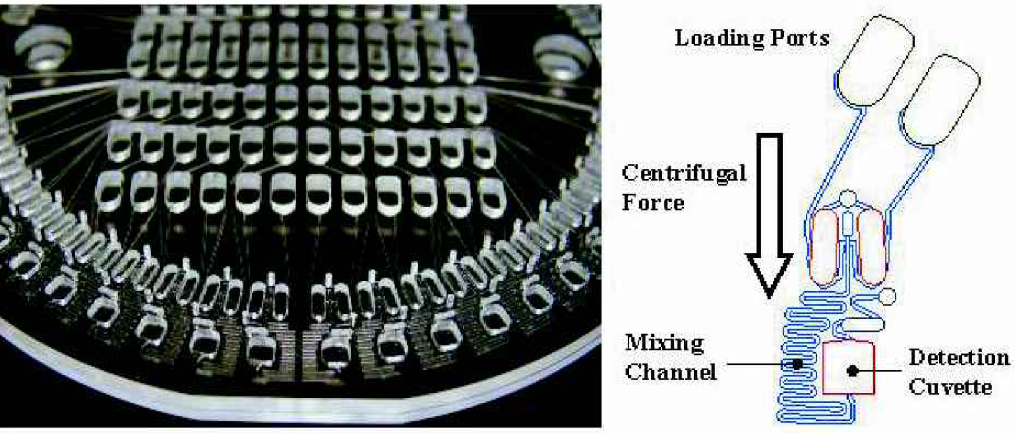


The LabCD Disc is an automation compatible microfluidic device that integrates multiple assay processes on a microscale to provide a platform technology for performing a wide range of drug discovery assays. Photograph of an EAP disc is shown at left, schematic of an individual manifold is shown at right.

Injection molded LabCD Disc with an automation compatible microplate interface, distribution manifold for a common reagent and 48 discrete detection cuvettes.
Cytochrome P450 (Cyp) Inhibition: Miniaturized fluorescence-based assays (Figure 4) using commercially available baculovirus insect cell expressed Cyp enzymes were developed and tested on the LabCD-ADMET System.
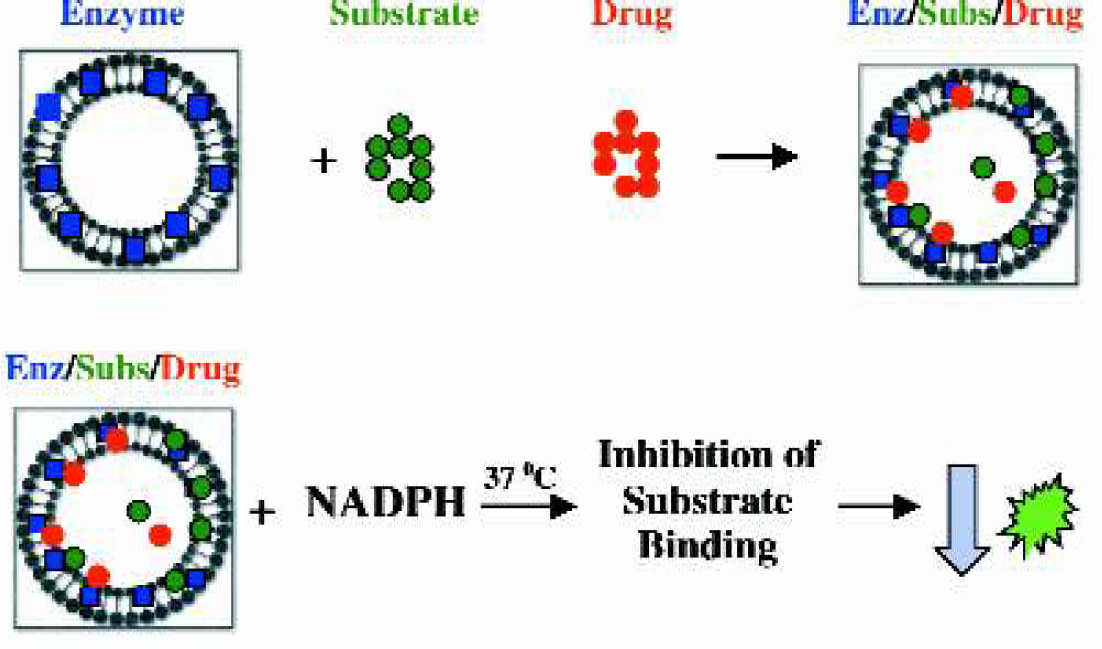
A modified assay protocol was developed with the commercially available fluorescent substrates and recombinant enzymes. The modifications include: 1) mixing of drug with enzyme/substrate mix prior to reaction initiation, 2) use of excess NADPH replacing a regenerating system, 3) use of Tecan's NSB-Block reagent to limit non-specific binding, 4) elimination of stop solution and 5) miniaturization of the assay to a final assay volume of 20ul.
Serum Protein Binding: Standard serum protein binding assays generally involve equilibrium dialysis or ultra-filtration to separate protein bound from free drug compound and LC/MS detection. An automated, high-throughput method was developed which uses drug displacement of fluorescent dyes specific for three binding sites on serum proteins: subdomains IIA and IIIA of albumin (HSA: site IIA, IIIA) and the major site of alpha-1 acid glycoprotein (a-AGP)(Figure 5).
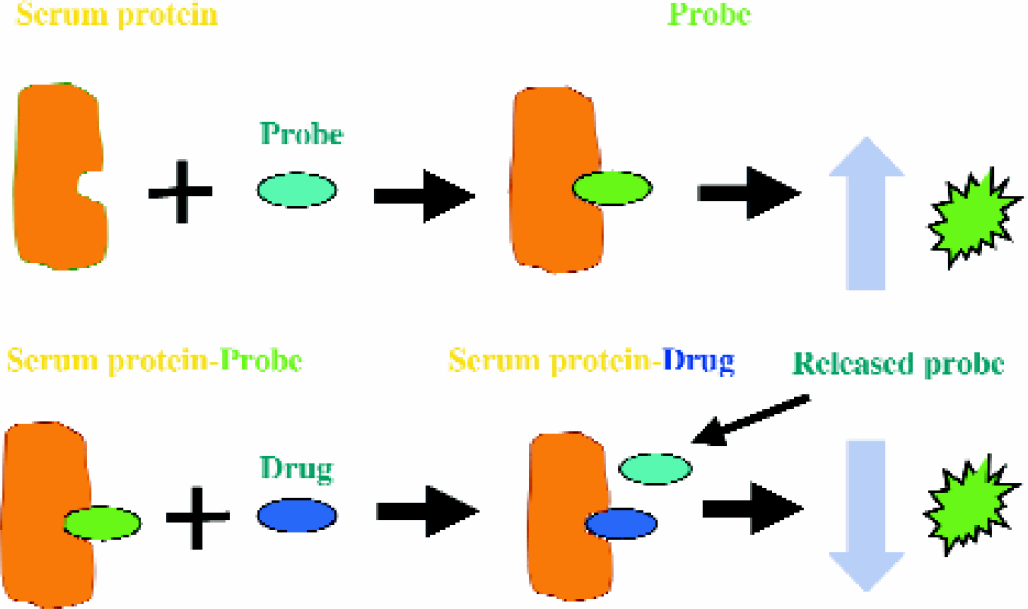
Schematic presentation of drug serum protein binding assay. A miniaturized assay using displacement of serum protein site-specific fluorescent probes was developed. The binding of the specific probe molecule to the protein site results in increased fluorescence emission of the probe molecule. This fluorescence is decreased in the presence of a drug/compound that competes with the probe for the protein binding site.
P-glycoprotein (Pgp) Interaction: Multi-Drug Resistance (MDR1)/ Pgp activity is traditionally assayed using labor-intensive cell-based approaches that determine end-points and are generally not high-throughput. In contrast to these membrane-transport approaches, assays of the Pgp-ATPase system provide a rapid screening tool with kinetic (continuous) detection for the interactions of drugs with Pgp. A novel protocol in which the Pgp-ATPase activity is enzymatically coupled to an absorbance-based detection system was developed (Figure 6) and seamlessly integrated with the LabCD system.
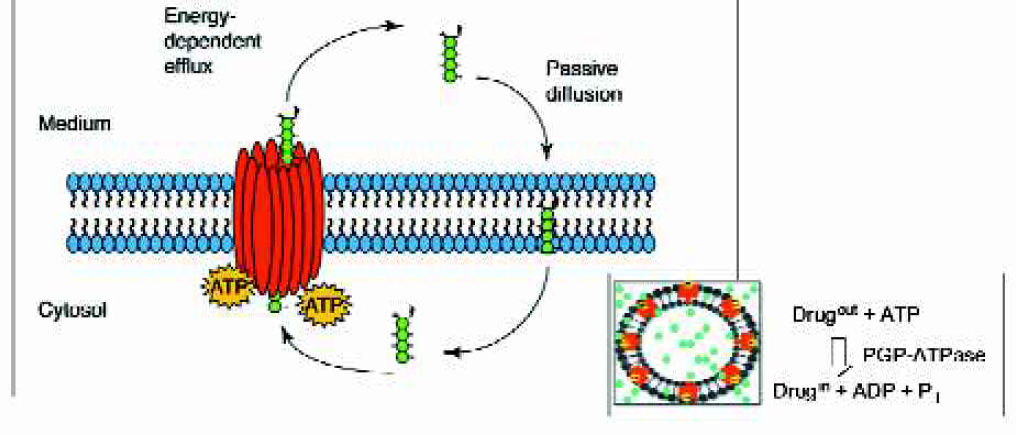
Schematic representation of Pgp Efflux Pump System. Pgp can be stimulated or inhibited by various drugs alone and in combination. The pump is energy-dependent, and is driven by ATP hydrolysis. The Pgp-ATPase assay measures free phosphate that is released during efflux of drug.
Method
Cyp Inhibition: A modified assay protocol was used with commercially available fluorescent substrates and baculovirus-insect cell expressed Cyp isoforms. A proprietary reagent termed ‘NSB-block’ was developed and used to limit any non-specific binding. The influence of drug concentration on product formation was used to calculate IC50 values. The performance of the LabCD-ADMET System (final assay volume of 0.02 ml) was compared to the conventional 96-well microplate (final assay volume of 0.2 ml) method. Assays on both platforms were examined for twenty drugs, eight fluorescent substrates, and seven CYP isoforms (CYP3A4, 2D6, 2C9, 1A2, 2C19, 2C8 and 2A6) during the validation performed at Tecan Boston. These assays on the LabCD system were also externally validated at five leading pharmaceutical companies within Tecan's Early Access Program (EAP).
Serum Protein Binding: An automated protocol using drug displacement of fluorescent dyes specific for three binding sites on serum proteins was developed. Subdomains IIA and IIIA of human serum albumin (HSA) and the major site for basic/acidic ligands of human alpha-1 acid glycoprotein (AGP) are the sites probed. Miniaturized (0.02 ml) assays of the binding of >30 drugs to these three sites were performed on the LabCD System and the apparent affinities (Kd) were determined. These assays on the LabCD were also externally validated at three leading pharmaceutical companies within Tecan's EAP.
Multi-Drug Resistance (MDR1)/ P-glycoprotein (Pgp): Miniaturized assays (final assay volume of 0.02 ml) employing an enzymatically coupled absorbance-based detection system with commercially available baculovirus-insect cell expressed Pgp were performed with 13 drugs. Direct stimulation and inhibition of verapamil-stimulated Pgp-ATPase activity by these 13 drugs was measured in the continuous format.
Results
Cyp Inhibition: Typical dose responses (Figure 7) obtained during the testing are shown. Miniaturized assays on the LabCD System (0.02 ml) were performed as well as assays on 96-well microplates (0.2 ml). Correlation analysis comparing the IC50's obtained with the LabCD System to those of 96-well microplates during Tecan's internal validation had an r-squared of >0.95 for more than fifty different combinations of drugs, substrates, and Cyp isoforms (Figure 8a). The correlation statistics from the EAP Sites around the world also result in an r-squared value and slope of >0.9 (Figure 8b).
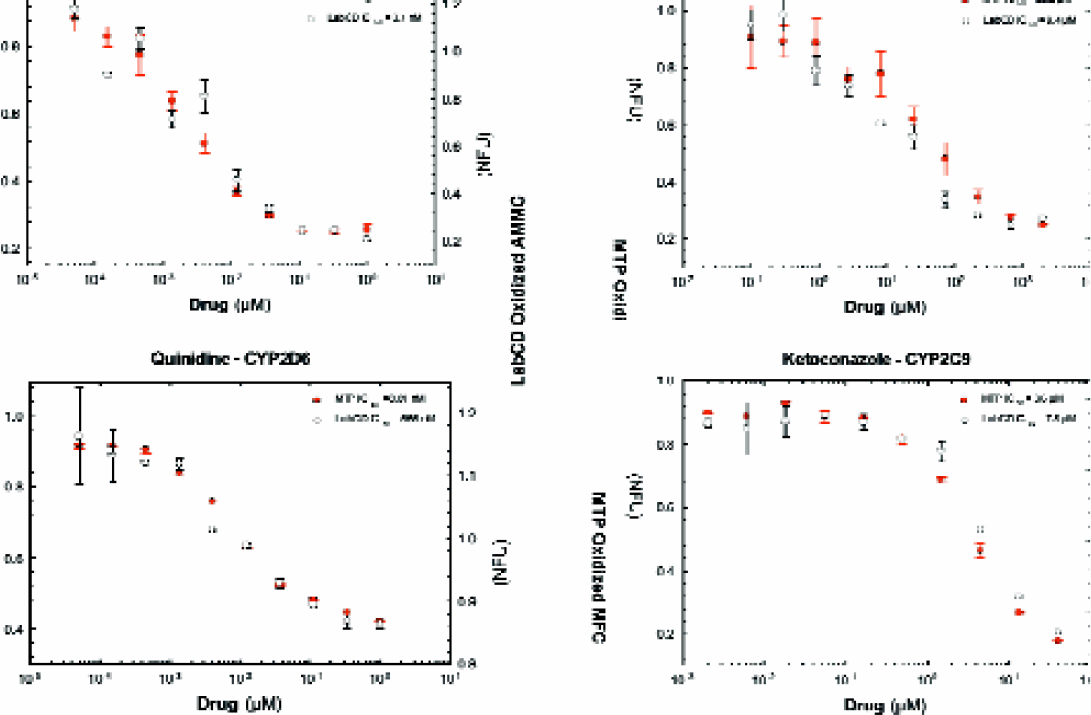
Comparison of IC Curves for CYP 3A4, 2D6, and 5O 2C9 on LabCD System and 96-well microplates (MTP).
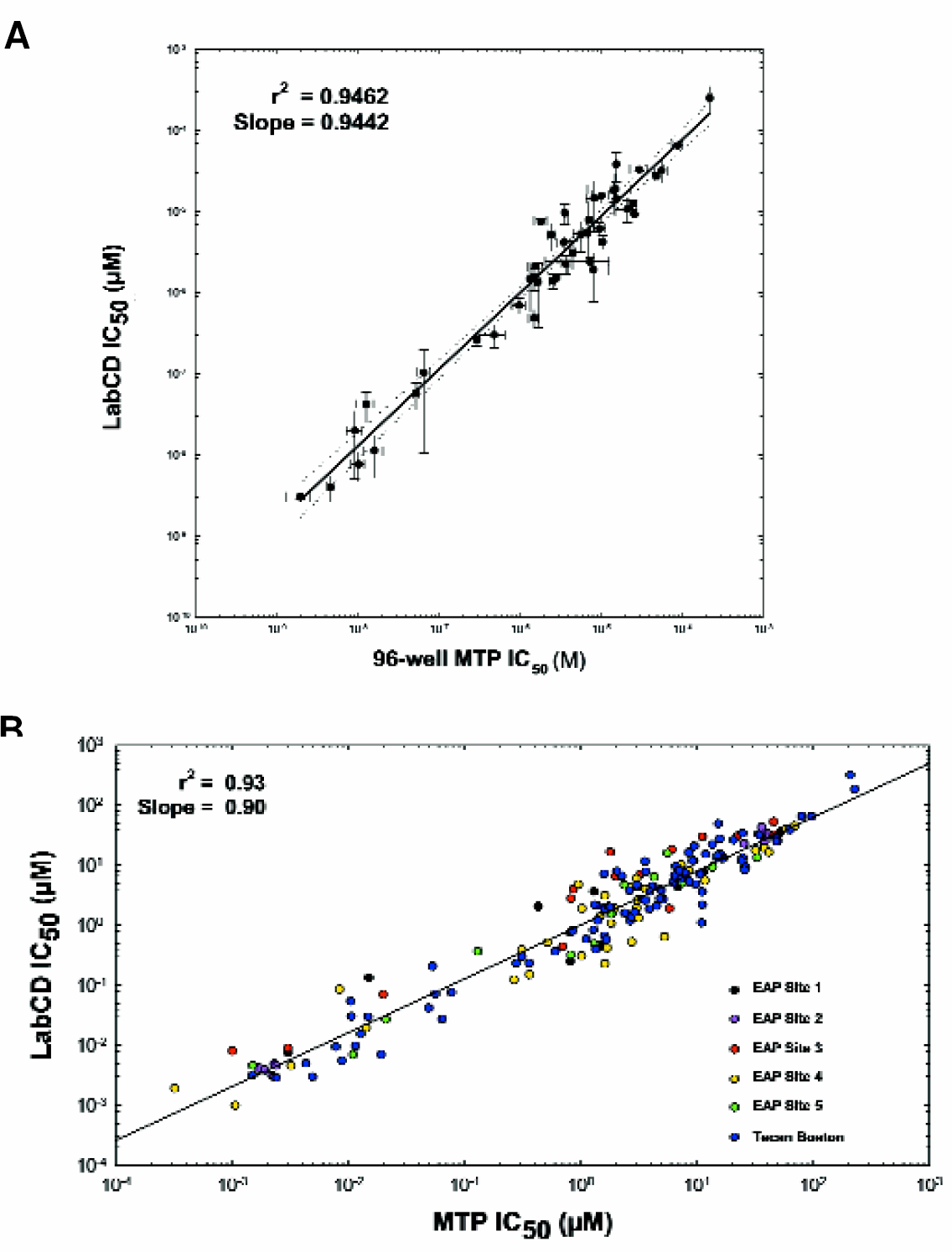
(A) Correlation plot of IC5O values for CYP450 inhibition on LabCD System and 96-well microplates (MTP) for internally tested compounds. (B) Correlation plot of IC5O values for CYP450 inhibition on LabCD System and 96-well microplates (MTP) for internal tests compared with EAP site tests.
Serum protein binding: Typical dose responses (see Figure 9) obtained during the testing are shown. Apparent affinities and specificity results of the 30 drugs for the three binding sites were reasonably consistent with the published data. Fractional occupancy was within a half log in all cases (Figure 10). Results were also very consistent across several sites, giving an r-squared of 0.87. (Figure 11)
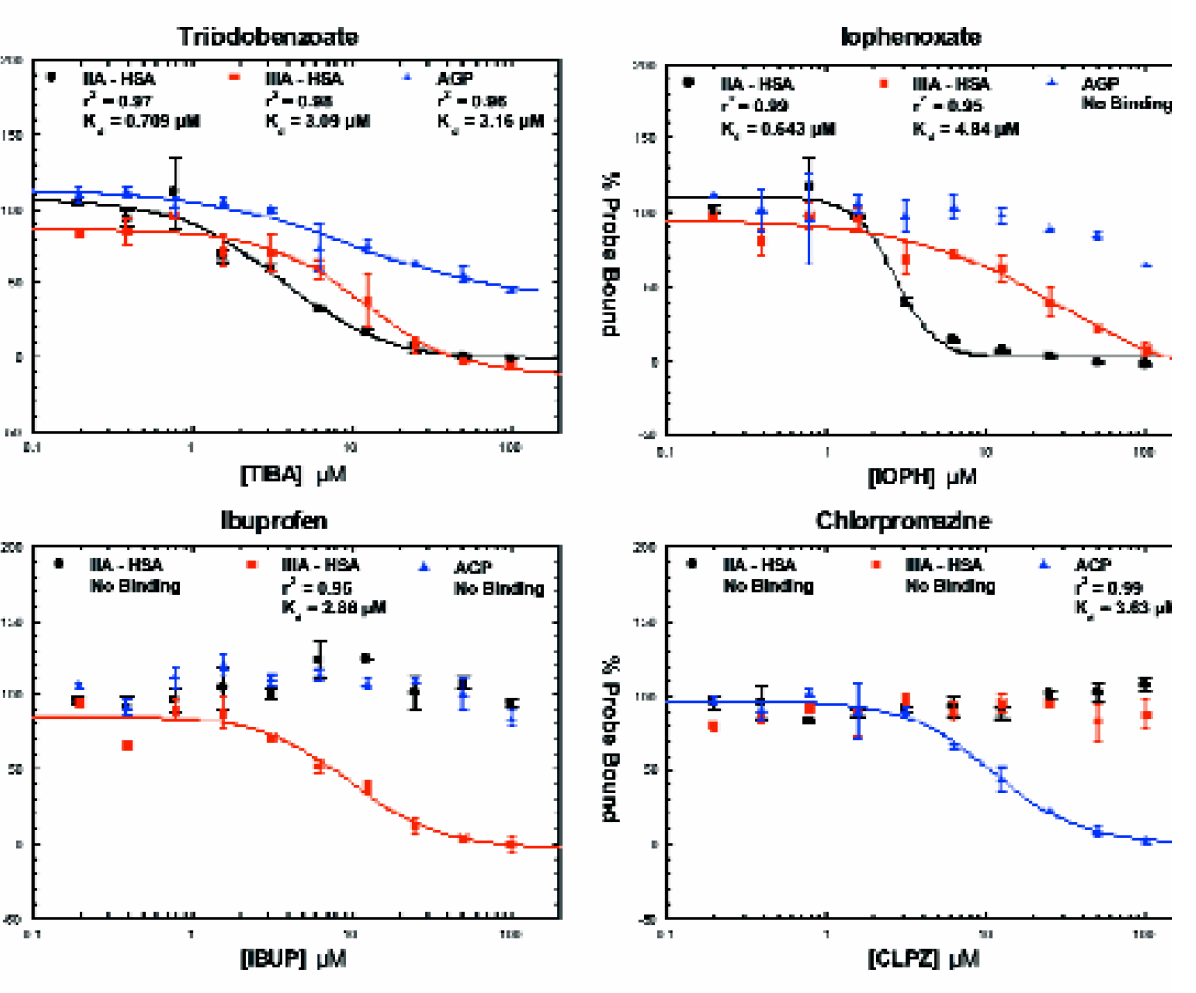
Drug competition curves for binding of Triiodobenzoate (TIBA), Iophenoxate (IOPH), Ibuprofen (IBUP), and Chlorpromazine (CLPZ) to Three Probe/Protein Combinations on LabCD System.
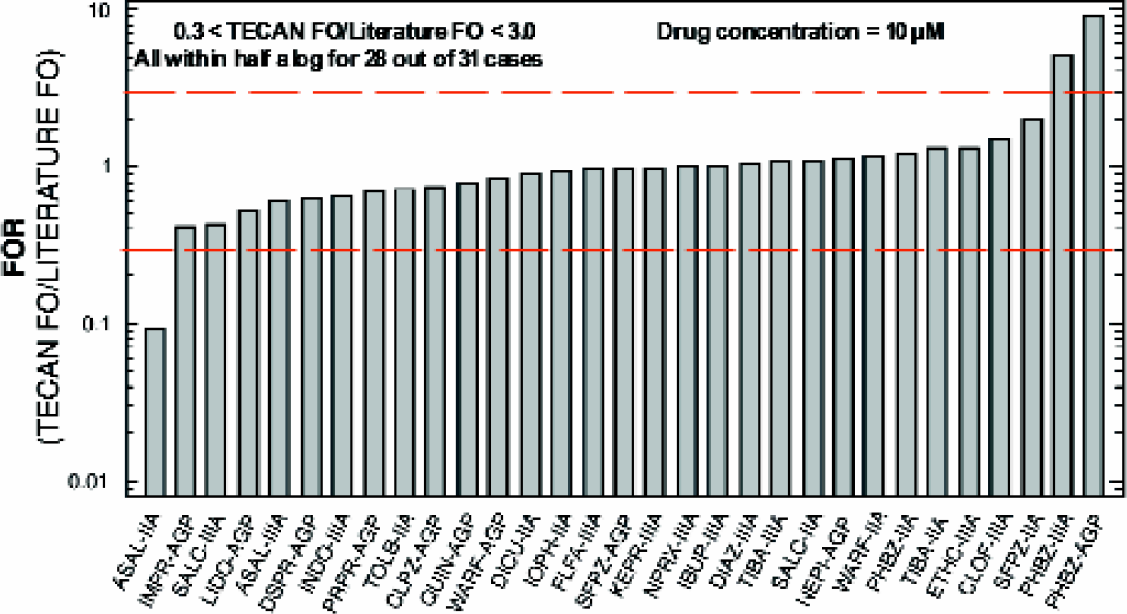
Fractional occupancy (FO) ratio analysis for LabCD System and literature values.
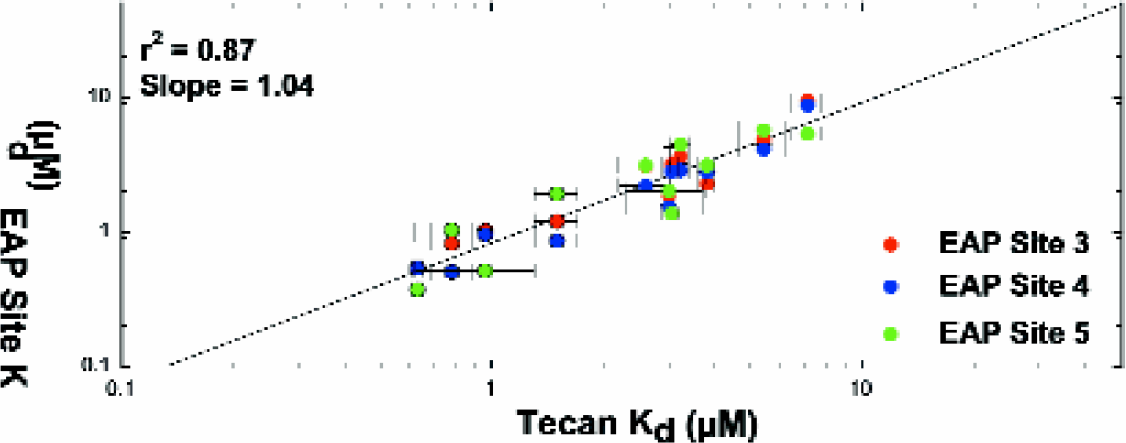
Tecan verses EAP Site apparent Kd's for compounds (as defined in Figure 4) plotted on a line of identity for LabCD System.
P-glycoprotein Interaction: Typical dose responses (Figure 12) obtained during the testing are shown. The results with Tecan's protocol as well as those of existing end-point detection methods (Sarkadi et al, 1992) were compared with other literature values. The results were internally consistent for both detection methods and reasonably consistent with the literature for both stimulatory and inhibitory effects (Figure 13).
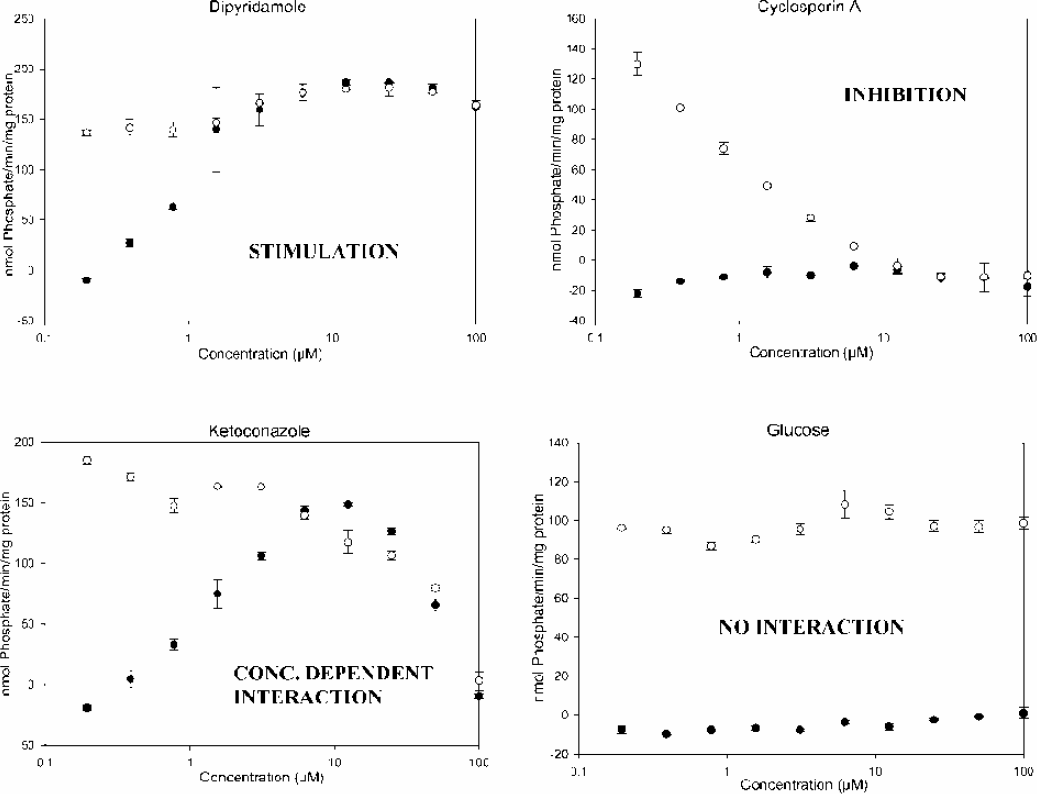
Drug Interactions with Pgp/ATPase system in the absence o and prescence • of verapamil, a known stimulant of system.
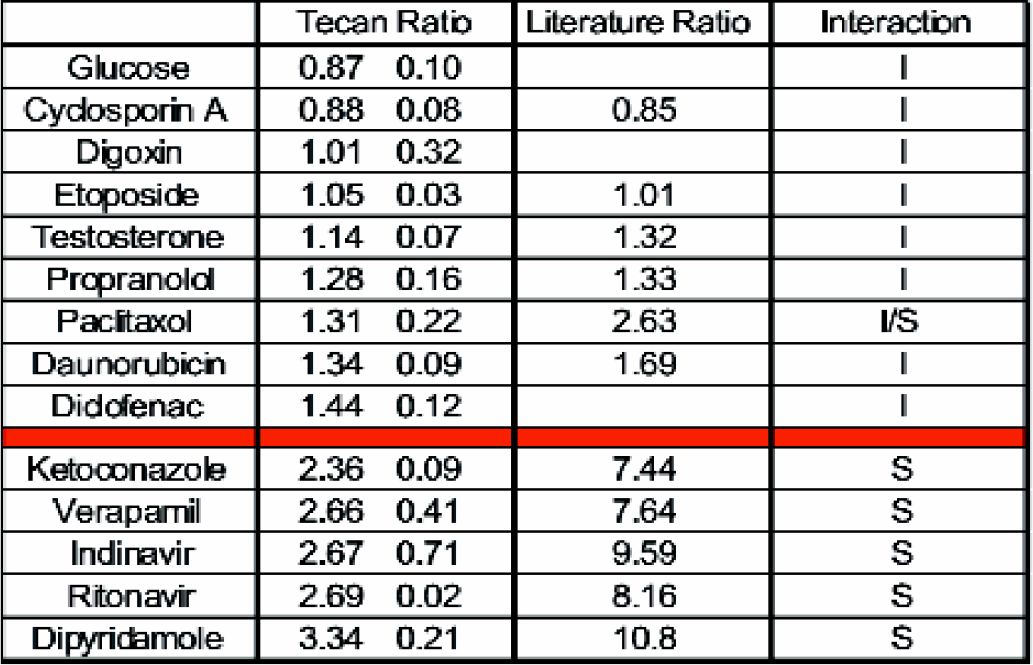
Tecan and literature values for ATPase assay ratios S = Stimulator of ATPase activity, I = Inhibitor or No interaction with ATPase activity.
Conclusions
The LabCD-ADMET System provides pre-validated and fully integrated assays for all major Cyp isoforms: Cyp 3A4, 2D6, 2C9, 2C19 and 1A2. In addition, the microfluidics approach enables a greater than 10-fold reduction in reagent usage while minimizing the non-specific binding of compounds commonly encountered.
The serum protein binding assays enable high throughput determinations of the binding affinities by eliminating the need for LC/MS detection or other complicated analytical methods. The system determines the apparent Kd of the compounds for the three major serum protein binding sites: Suβ-domains IIA and IIIA of HSA plus the major site for basic/acidic compounds on AGP.
These assays facilitate the rapid qualitative screening for high affinity drug interactions at the major sites of serum protein binding.
The miniaturized Pgp-ATPase assay enables a 50-fold reduction in enzyme usage over existing end-point detection methods. The assay permits simultaneous qualitative and quantitative detection of stimulation and inhibition of verapamil-induced activity at various concentrations. This integrated solution automatically calculates ATPase ratios that correlate well with literature values. Use of this method could also help elucidate drug-drug interactions at the ATP binding site.
The LabCD-ADMET System integrates the microfluidic LabCD disc with a suite of pre-validated assay protocols, application-specific software and hardware to deliver an automated, robust, and reproducible platform for ADMET assays. The miniaturization provided by the system enables significant reduction in reagent usage and associated assay cost. Built-in protocol specific automation scripts eliminate the need for upfront assay development, automation development and validation. In addition, the integrated data analysis package significantly reduces the data analysis, review and formatting time, with accompanying gains in throughput. In specific cases (serum protein binding and (MDR1)/ P-glycoprotein (Pgp)), the assays themselves are more amenable to higher throughput than conventional methods. The system thereby allows budgetary efficiencies to be realized by significantly reducing the FTE costs while providing reliable and high-quality data. Finally, the standardization and pre-validation of the assays allows for data comparisons across various sites of a corporation. Thus the LabCD-ADMET System represents an “industrial-strength” turnkey solution for today's ADMET screening needs.
