Abstract
Green fluorescent protein (GFP) has many applications as a marker in living cells, and has become widely used as a reporter gene in microbial, plant and animal cells. Screening microbial colonies for GFP expression enables various types of assays (e.g. for secretory mutations). However, this is laborious, non-quantitative and potentially hazardous to the operator (due to UV illumination) when performed manually. In order to address this the GloPix robot was developed. The imaging system discriminates between colonies based on the level of fluorescence activity and the picking function automatically transfers cells to microplate wells. Measuring fluorescent activity allows quantitation of fluorescent tag concentration/expression.
Introduction
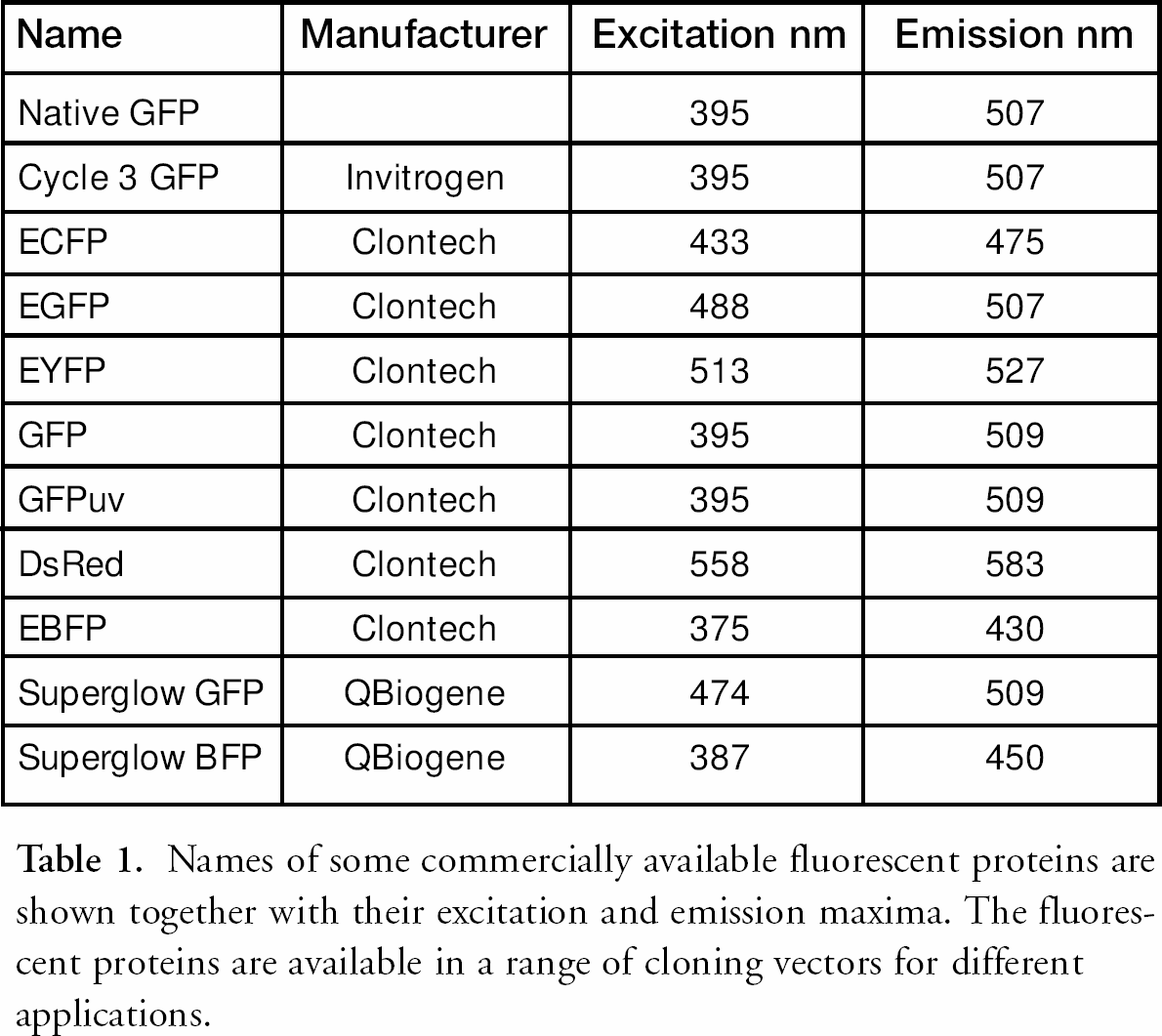
Green Fluorescent Protein (GFP) was originally isolated from the jellyfish Aequorea victoria in the 1960s 1,2 and has since become widely used in genomics and proteomics applications as a fluorescent tag in living cells. Benefits of the GFP system include remarkable stability of the chimeric protein fluorescence, low toxicity to the expressing cell, no requirement for any exogenous substrate and availability of a number of variants from the wild type protein with different fluorescence excitation and emission characteristics. Cloning vectors with different GFP variants and vectors constructed for different assays (e.g. mammalian/plant transfection, promoter probe etc.) are now commercially available (see Table 1). Some variants have been enhanced for optimal codon usage for mammalian and yeast expression. Other fluorescence proteins from different organisms are also used and the range available will probably continue to expand.
Names of some commercially available fluorescent proteins are shown together with their excitation and emission maxima. The fluorescent proteins are available in a range of cloning vectors for different applications.
The use of fluorescent proteins as markers has wide applications in genomics and proteomics. Biological processes to which GFP has been applied include monitoring gene promoter activity (in response to exogenous stimuli or mutations), protein folding, cell transfection, vesicle trafficking, exon cloning, secretory mutation screening and protein-protein interactions.
Colony picking robots have been available for approximately ten years and have had a huge impact on the speed and success of the human genome sequencing project. Such robots are able to image colonies of E. coli growing on the surface of agar, discriminate colonies based on user selectable criteria (e.g. diameter, roundness), and pick the colonies individually into microplates. These small-scale cultures in microplates are then ideal for downstream processing of the colonies (e.g. DNA sequencing in a high-throughput automated fashion). Each clone has a unique identifier (plate number and well coordinate) even when no other information has yet been obtained. Genetix, Ltd. has extended this picking technology to develop a high-throughput GFP picking robot, GloPix, which can detect and selectively pick GFP-expressing bacterial colonies
The Glopix Robot
The robot is based on the existing QPix and MegaPix colony picking systems (Genetix Ltd). These robots have XYZ drives that position the actuator head within the working area. The head is equipped with a camera to image colonies growing on agar. The colonies are imaged and assigned an XY coordinate, which enables them to be picked using a ‘picking head’ with 96 individually addressable pins. The head can be used to inoculate 96- or 384-well microplates. In order to develop a system for picking based on GFP expression, the imaging system was redesigned and equipped with a low noise, high dynamic range CCD camera.
The excitation illumination has been custom designed, incorporating a spectrally matched high-intensity discharge lamp system. This avoids any risk of UV induced cell damage and possible hazards to the operator. Illumination of the colonies is via a fibre-optic array optimised to produce the illumination field necessary for quantitative measurement of fluorescence within the picking area. Narrow bandwidth selectable excitation and emission wavelength filters enhance the optical contrast and reduce the impact of autofluorescence arising from intracellular and medium components. 3 The robot is fully enclosed and light-proofed, enabling it to be used in a normally illuminated laboratory. For high-throughput picking GloPix has stackers for 50 source plates and 210 destination microplates. A smaller version without the stackers is also available. Software, based on the established QSoft (Genetix, Ltd.) picking software, has been adapted to give fully quantitative picking criteria — on brightness (fluorescent activity), ratio or colour, as well as standard picking criteria including minimum and maximum diameter, roundness and proximity. Quantitative data can be output to other software such as spreadsheet applications. GloPix robots are also equipped to read barcodes on source and destination plates. This is particularly useful for data tracking through high-throughput experiments enabling the linking of the original imaged colony data to the microplate location after picking and through to further robotic applications, such as arraying.
E. Coli Expressing GFP
Colonies expressing GFP can be grown on and picked from the surface of agar plates. However, agar growth media often contain components, such as yeast extract, that contribute to fluorescence of the background. With GloPix, background fluorescence has been largely eliminated by a combination of growing the colonies on nylon membrane laid over the agar (Figures 1 and 2), illumination from above, and appropriate choice of filters. We have used 22 × 22 cm Genetix PerForma positively-charged nylon membrane laid on QTrays (Genetix, Ltd.) filled with LB medium. PerForma was designed for low background fluorescence for use with fluorescence detection systems, such as Attophos (Roche, Lewes, UK) or GFP. Colonies are spread homogeneously on top of the membrane by applying a cell suspension and rolling six mm glass beads (Genetix) around the surface to spread the liquid. Growing the colonies on a membrane has the further advantage that in cases where GFP expression has to be induced, such as by IPTG, the colonies can first be grown on medium without inducer and then transferred to medium with inducer by lifting and transferring the membrane. Since illumination and detection of the colonies is from above, the imaging of the colonies is not affected and the robot picking head can pick colonies from the membrane surface.

GloPix high throughput GFP picking robot uses filtered excitation light and a CCD camera with emission filter to image fluorescent colonies.
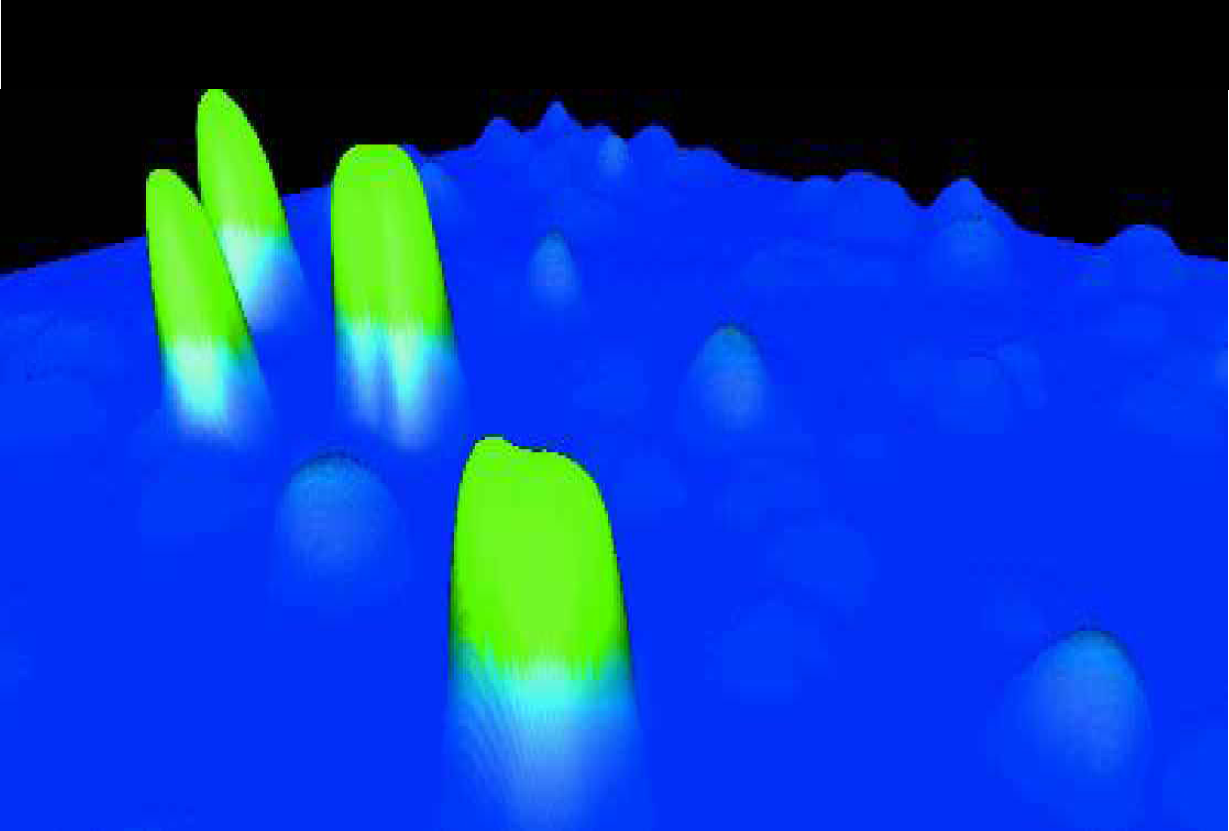

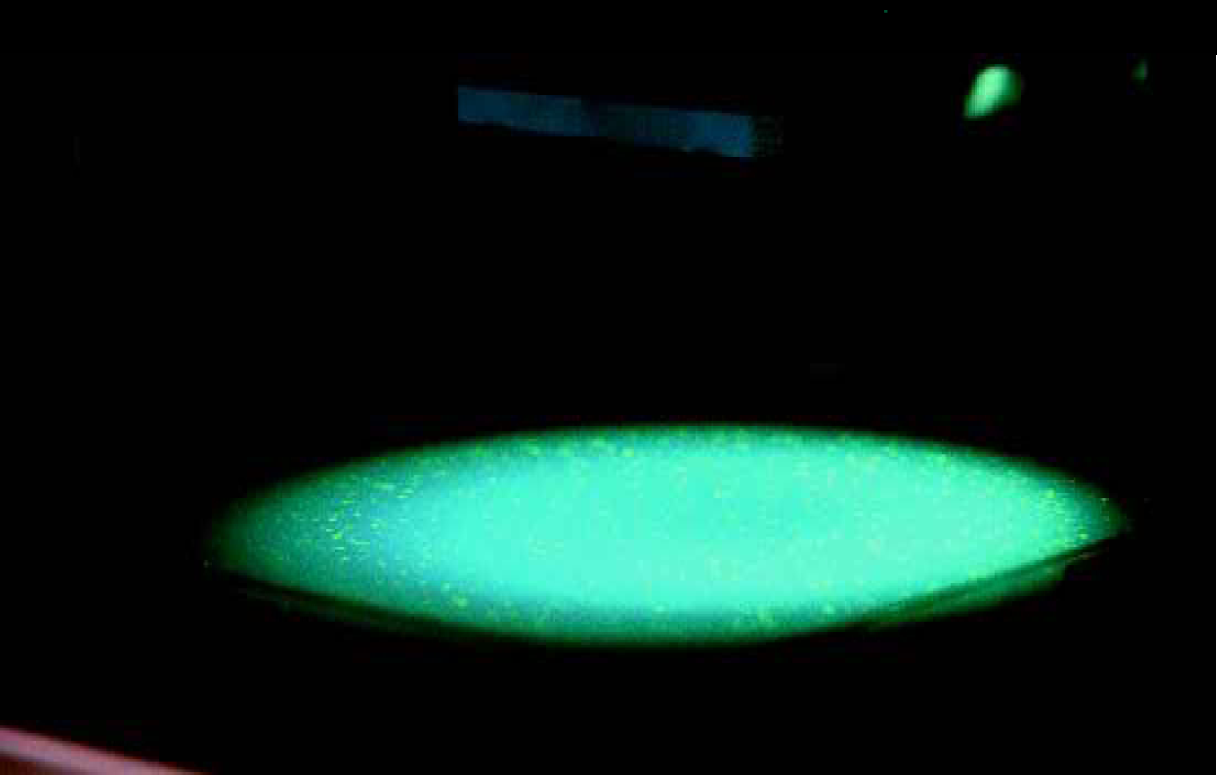
An image of colonies expressing different levels of GFP translated into a three-dimensional display of the data. The colonies were spread on PerForma membrane.
In addition to a random spread of the colonies (e.g. from a transformation mix), the arraying function on Genetix robots (QBot or QPix) can be used to array clones from microplates onto membranes for analysis of GFP expression. This also allows replicates of each clone to be introduced into the array for improved quantitative analysis (Figure 3).
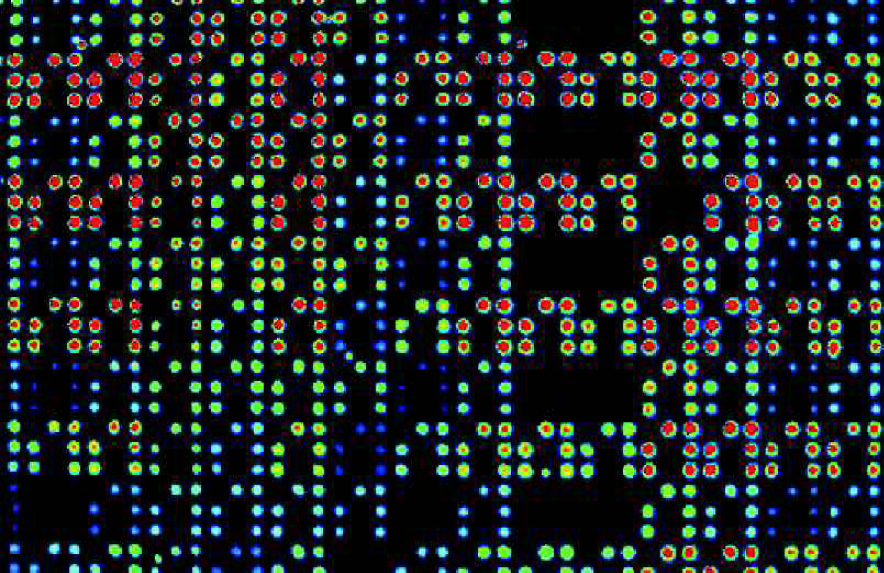

An array of GFP expressing E. coli produced using a Genetix QPix and imaged with GloPix. A false colour palette has been applied to indicate GFP intensity.
Conclusion
GloPix is a powerful tool for high-throughput analysis and manipulation of microbial colonies expressing fluorescent proteins. A broad range of biological applications can be envisaged such as gene expression, mutation screening, protein-protein interactions, protein folding and protein secretion studies.
